Submitted:
04 July 2025
Posted:
07 July 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Results
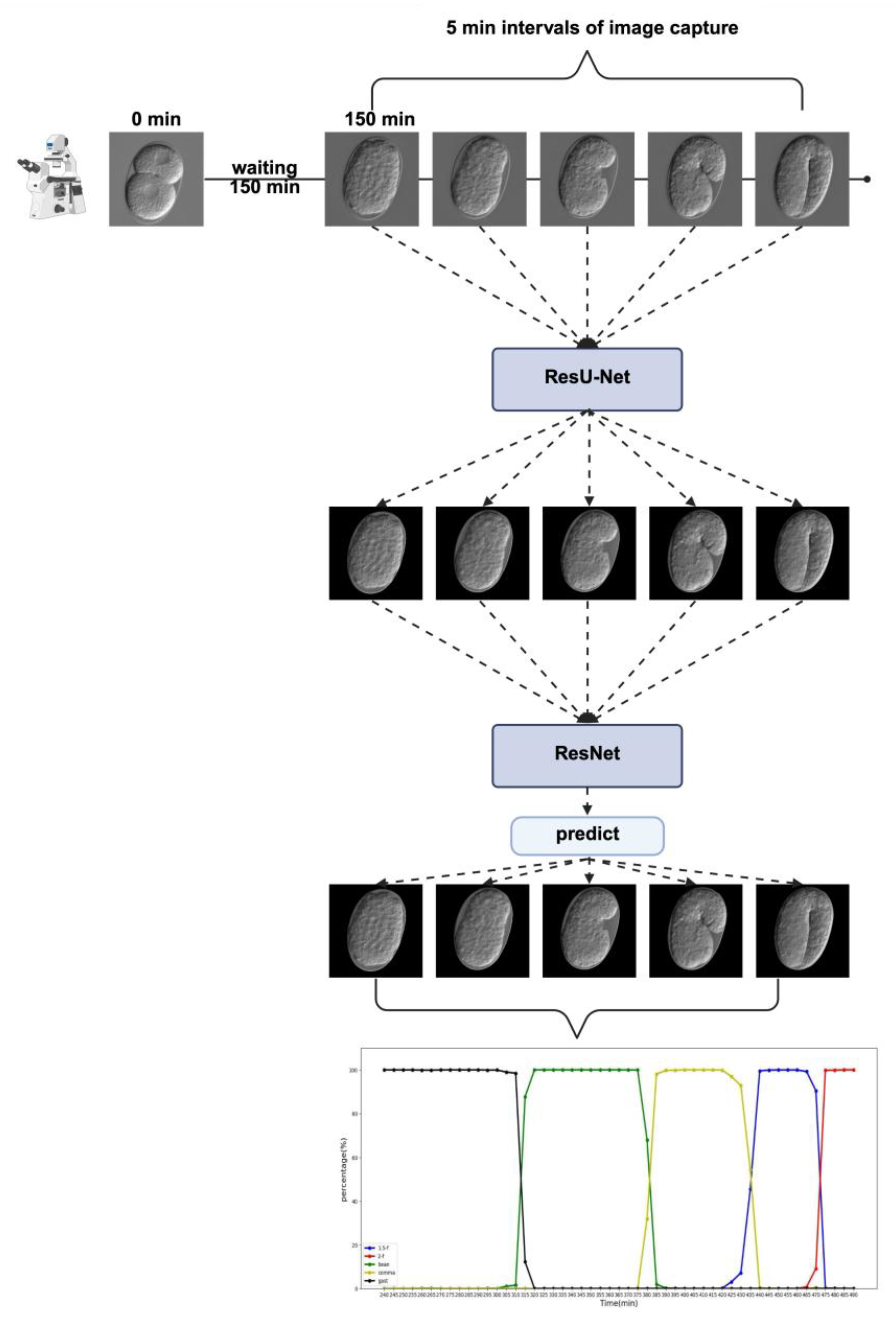
2.1. Embryo Segmentation and Classification Using ResU-Net and ResNet
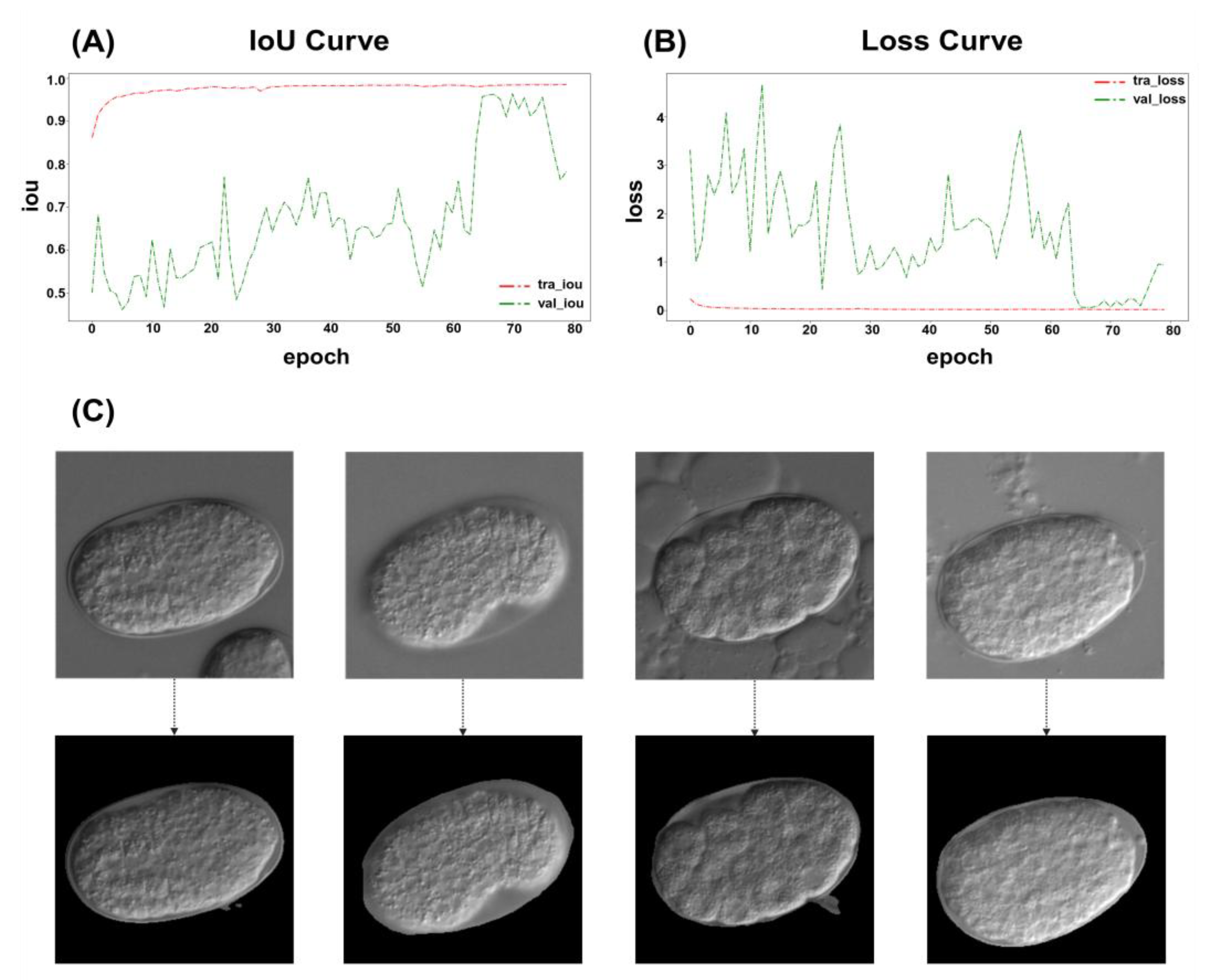
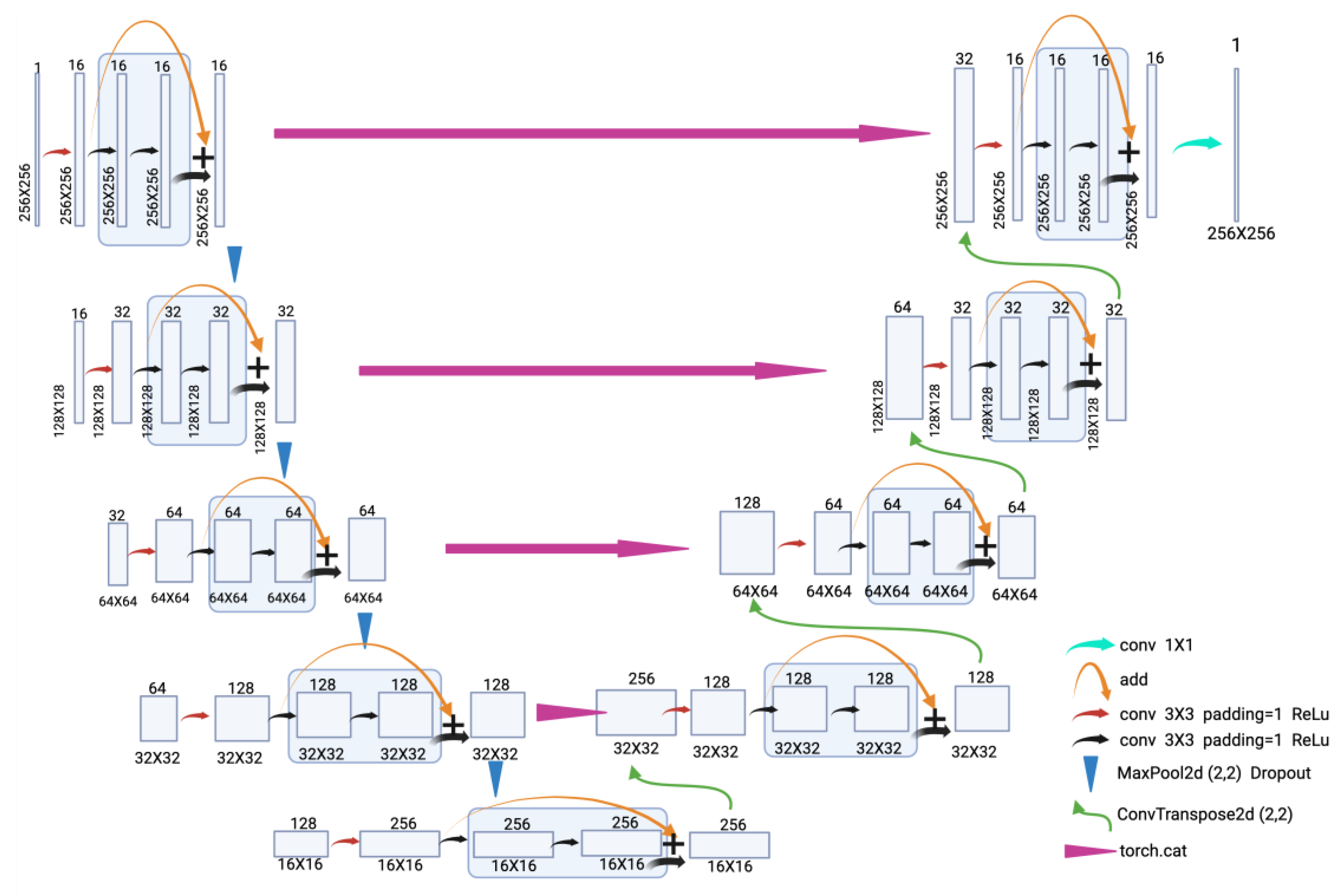
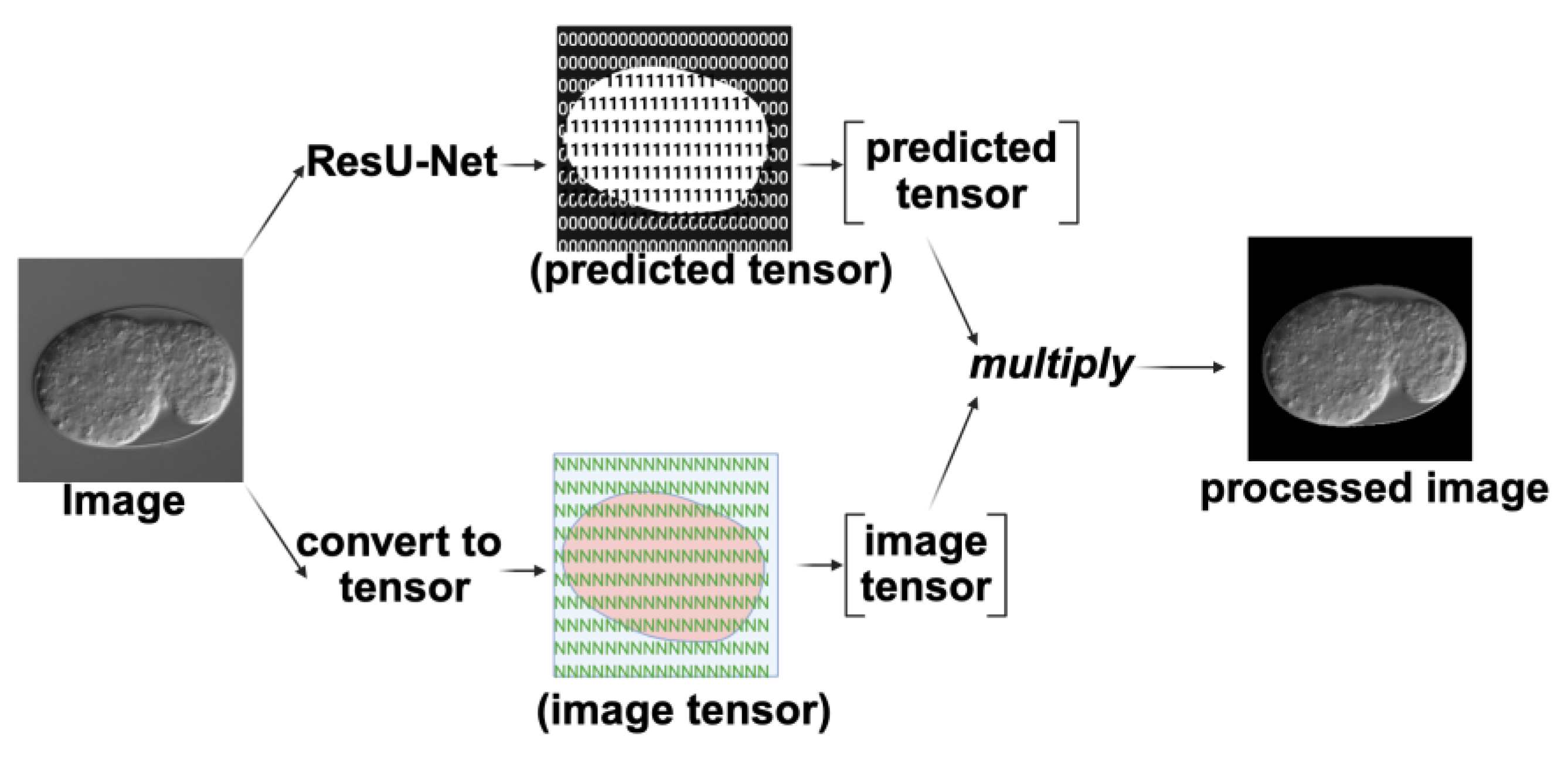
2.1.1. ResU-Net
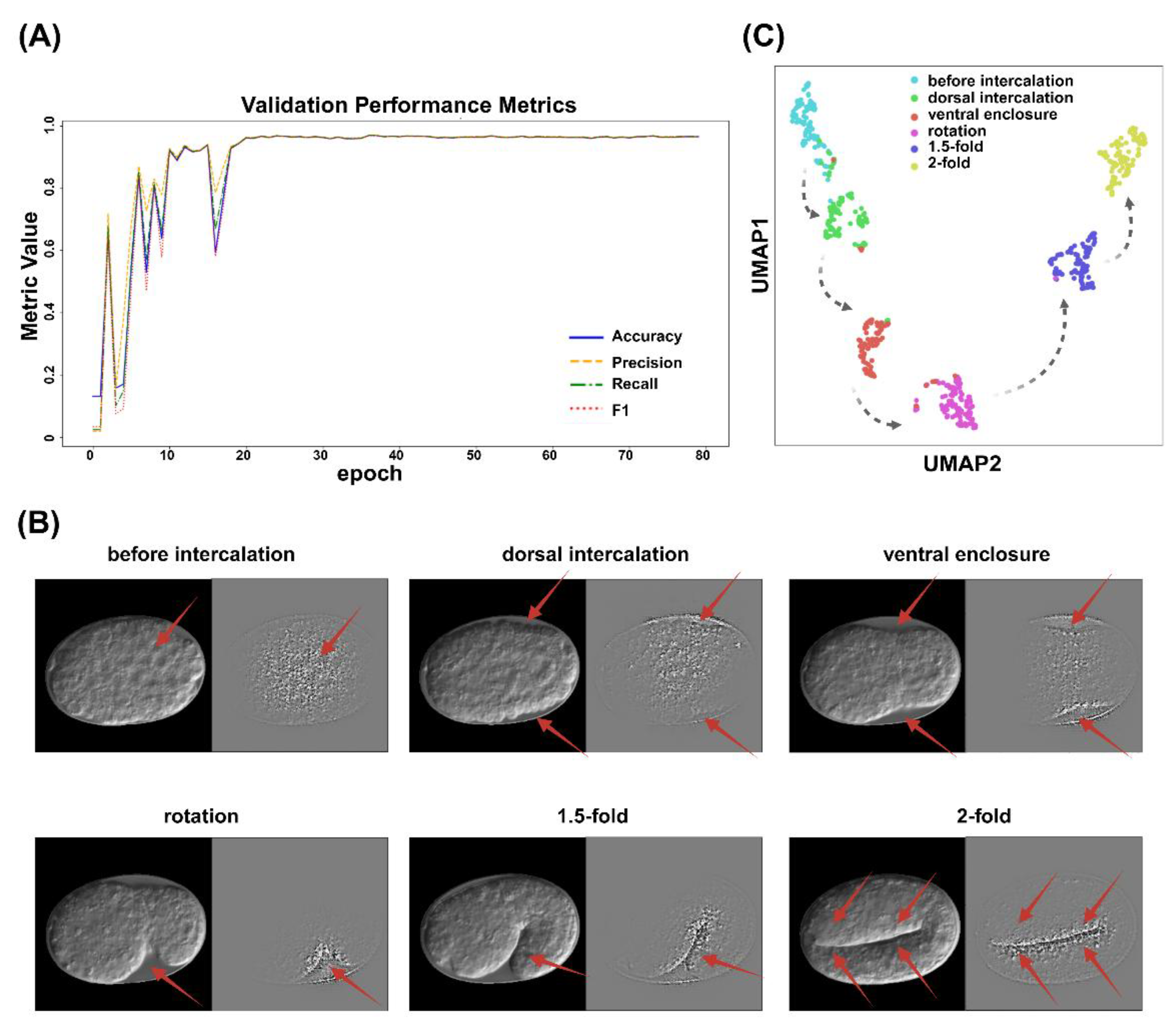
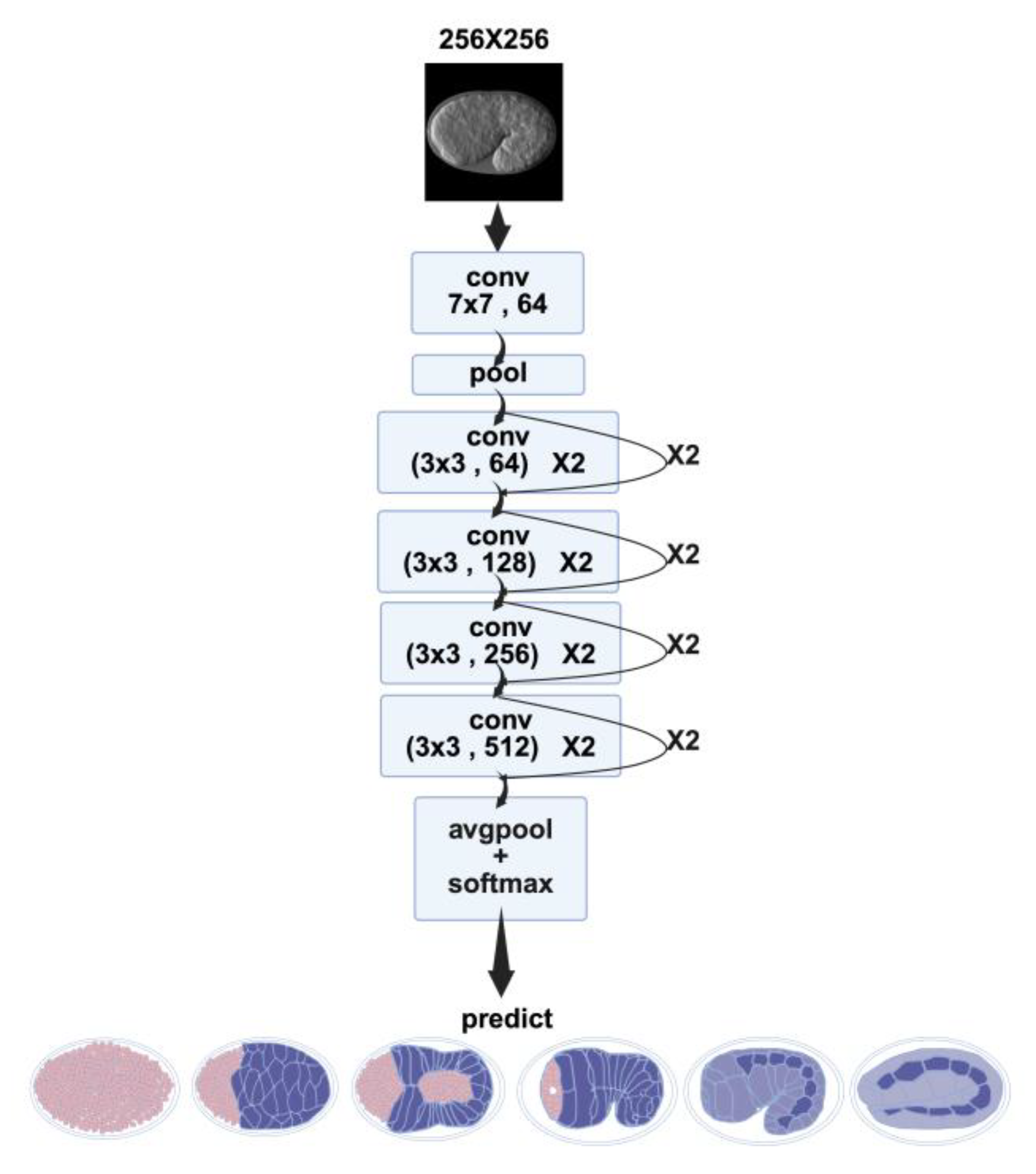
2.1.2. ResNet
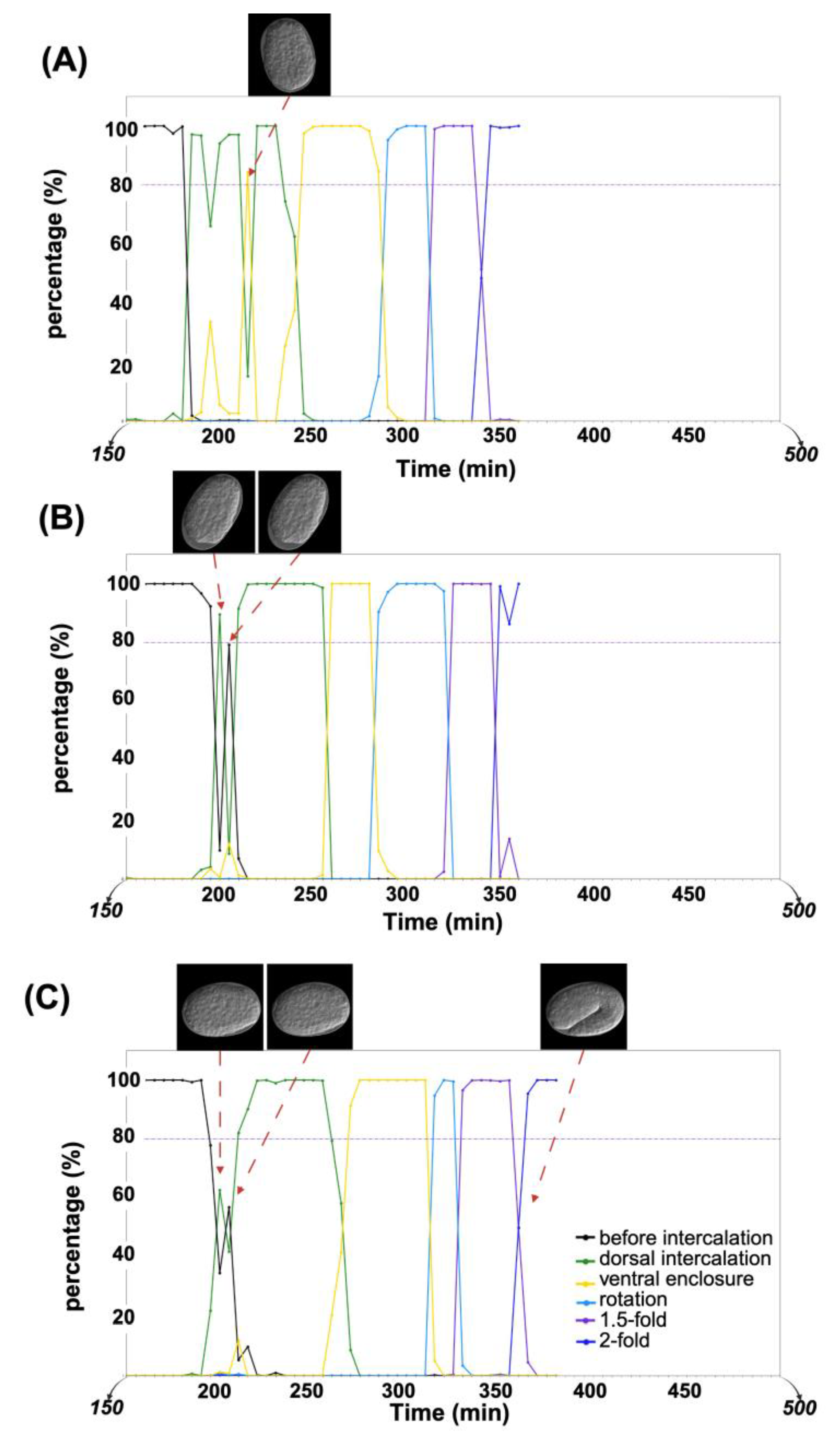
2.2. Analysis of Temporal Prediction Accuracy in Time-Lapse Data
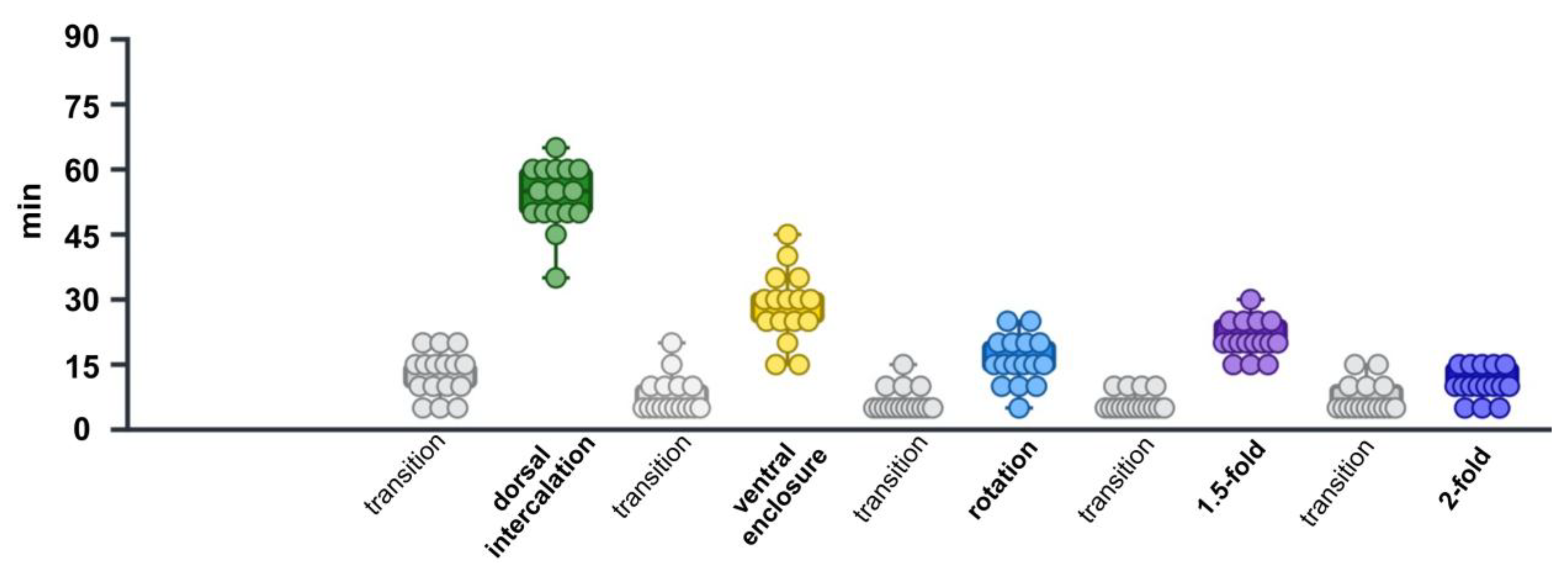
2.3. Analysis of Embryonic Stage Durations in Control (RNAi) Time-Lapse Data
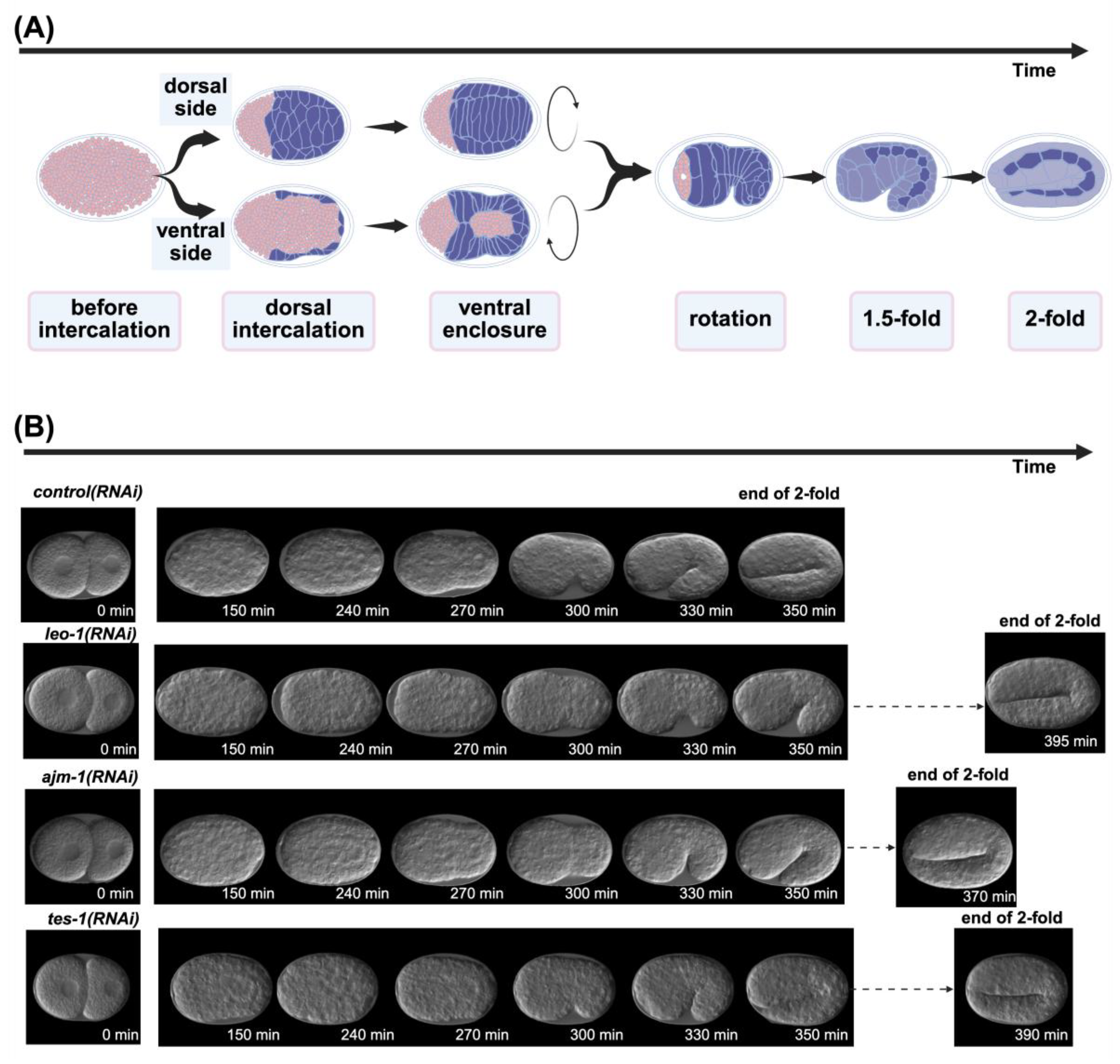
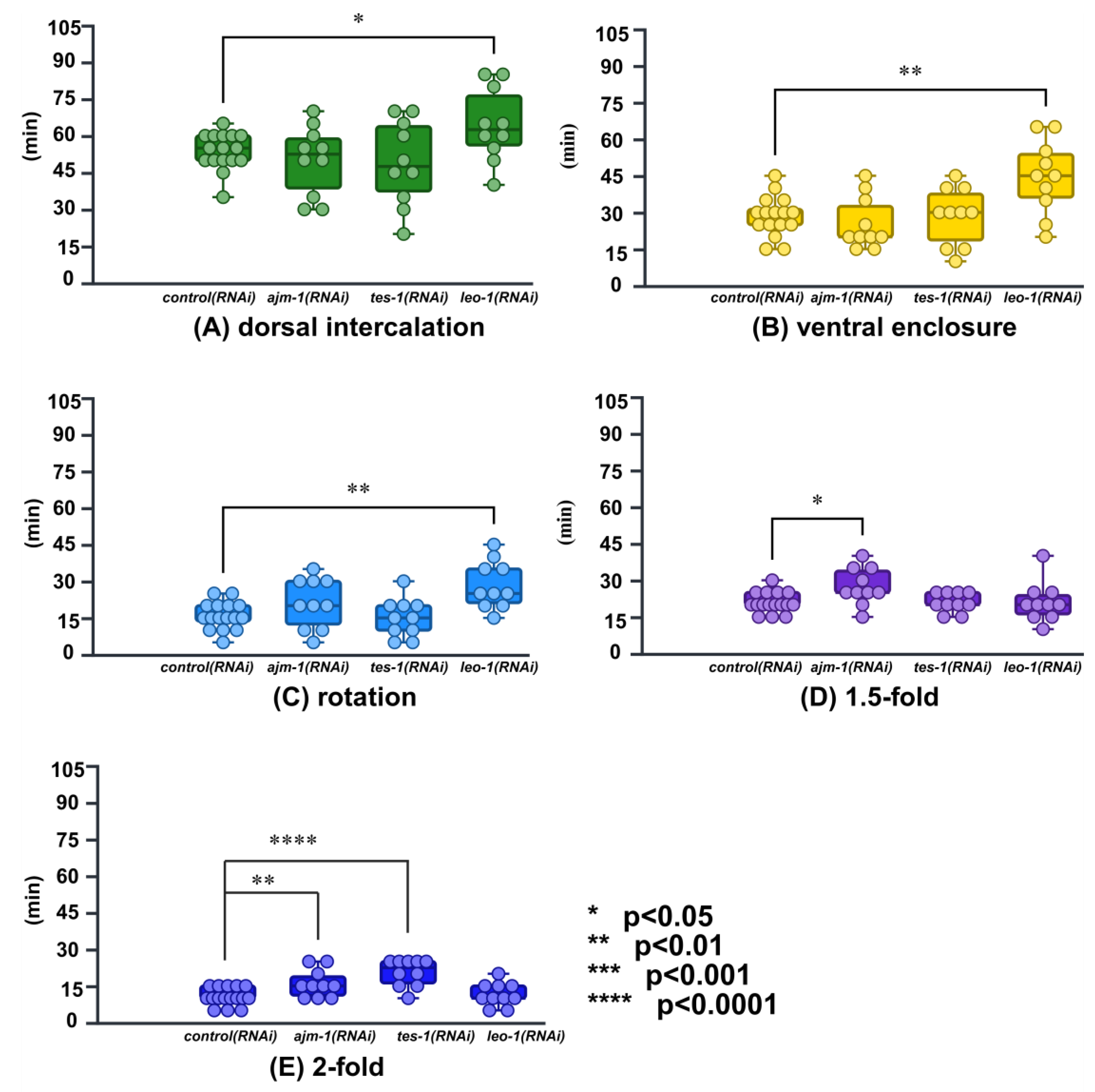
2.4. Application in RNAi Knockdown Time-Lapse Data
3. Discussion
3.1. Deep Learning-Based Diagnostic Tool for Temporal Analysis
3.2. Calculations of Time Required for Each Stage in Control(RNAi) Group
3.3. Trial to Temporal Analysis of the Gene Function in RNAi Knockdown Animals
3.4. Contributions and Limitations of the Current Approach
4. Materials and Methods
4.1. Caenorhabditis Elegans Strains
4.2. RNA Interference Assay
4.3. Epidermal Morphogenesis Stages
4.3.1. Before Intercalation
4.3.2. Dorsal Intercalation
4.3.3. Ventral Enclosure
4.3.4. Rotation
4.3.5. 1.5-Fold and 2-Fold
4.4. Microscope and Image Acquisition
4.5. Models Training and Evaluation
4.5.1. ResU-Net
4.5.2. ResNet
4.6. Timeline
4.7. Image Interpretability Analysis
4.7.1. Grad-CAM
4.7.2. UMAP
5. Conclusions
Supplementary Materials
Author Contributions
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Vuong-Brender, T.T.K.; Yang, X.; Labouesse, M.C. Elegans Embryonic Morphogenesis. In Current Topics in Developmental Biology; Wassarman, P.M., Ed.; Essays on Developmental Biology, Part A; Academic Press, 2016; Vol. 116, pp. 597–616.
- Chisholm, A.D.; Hardin, J. Epidermal morphogenesis. In WormBook: The Online Review of C. elegans Biology [Internet]; WormBook, 2005.
- Apfeld, J.; Alper, S. What Can We Learn About Human Disease from the Nematode C. Elegans. In Disease Gene Identification: Methods and Protocols; DiStefano, J.K., Ed.; Springer: New York, NY, 2018; ISBN 978-1-4939-7471-9. [Google Scholar]
- Sulston, J.E.; Schierenberg, E.; White, J.G.; Thomson, J.N. The Embryonic Cell Lineage of the Nematode Caenorhabditis Elegans. Developmental Biology 1983, 100, 64–119. [Google Scholar] [CrossRef] [PubMed]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and Specific Genetic Interference by Double-Stranded RNA in Caenorhabditis Elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef] [PubMed]
- Gönczy, P.; Echeverri, C.; Oegema, K.; Coulson, A.; Jones, S.J.; Copley, R.R.; Duperon, J.; Oegema, J.; Brehm, M.; Cassin, E.; et al. Functional Genomic Analysis of Cell Division in C. Elegans Using RNAi of Genes on Chromosome III. Nature 2000, 408, 331–336. [Google Scholar] [CrossRef]
- Fraser, A.G.; Kamath, R.S.; Zipperlen, P.; Martinez-Campos, M.; Sohrmann, M.; Ahringer, J. Functional Genomic Analysis of C. Elegans Chromosome I by Systematic RNA Interference. Nature 2000, 408, 325–330. [Google Scholar] [CrossRef] [PubMed]
- Williams-Masson, E.M.; Malik, A.N.; Hardin, J. An Actin-Mediated Two-Step Mechanism Is Required for Ventral Enclosure of the C. Elegans Hypodermis. Development 1997, 124, 2889–2901. [Google Scholar] [CrossRef]
- Quintin, S.; Michaux, G.; McMahon, L.; Gansmuller, A.; Labouesse, M. The Caenorhabditis Elegans Gene Lin-26 Can Trigger Epithelial Differentiation without Conferring Tissue Specificity. Dev Biol 2001, 235, 410–421. [Google Scholar] [CrossRef]
- Labouesse, M.; Sookhareea, S.; Horvitz, H.R. The Caenorhabditis Elegans Gene Lin-26 Is Required to Specify the Fates of Hypodermal Cells and Encodes a Presumptive Zinc-Finger Transcription Factor. Development 1994, 120, 2359–2368. [Google Scholar] [CrossRef]
- Pettitt, J.; Cox, E.A.; Broadbent, I.D.; Flett, A.; Hardin, J. The Caenorhabditis Elegans P120 Catenin Homologue, JAC-1, Modulates Cadherin-Catenin Function during Epidermal Morphogenesis. J Cell Biol 2003, 162, 15–22. [Google Scholar] [CrossRef]
- George, S.E.; Simokat, K.; Hardin, J.; Chisholm, A.D. The VAB-1 Eph Receptor Tyrosine Kinase Functions in Neural and Epithelial Morphogenesis in C. Elegans. Cell 1998, 92, 633–643. [Google Scholar] [CrossRef]
- Meeuse, M.W.; Hauser, Y.P.; Morales Moya, L.J.; Hendriks, G.; Eglinger, J.; Bogaarts, G.; Tsiairis, C.; Großhans, H. Developmental Function and State Transitions of a Gene Expression Oscillator in Caenorhabditis Elegans. Mol Syst Biol 2020, 16, e9975. [Google Scholar] [CrossRef]
- Green, R.A.; Khaliullin, R.N.; Zhao, Z.; Ochoa, S.D.; Hendel, J.M.; Chow, T.-L.; Moon, H.; Biggs, R.J.; Desai, A.; Oegema, K. Automated Profiling of Gene Function during Embryonic Development. Cell 2024, 187, 3141–3160.e23. [Google Scholar] [CrossRef] [PubMed]
- Shaikh, M.A.; Al-Rawashdeh, H.S.; Sait, A.R.W. A Review of Artificial Intelligence-Based Down Syndrome Detection Techniques. Life 2025, 15, 390. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.-H.; Lin, C.-H.; Lane, H.-Y. Machine Learning and Novel Biomarkers for the Diagnosis of Alzheimer’s Disease. Int J Mol Sci 2021, 22, 2761. [Google Scholar] [CrossRef]
- Lin, E.; Lin, C.-H.; Lane, H.-Y. Deep Learning with Neuroimaging and Genomics in Alzheimer’s Disease. Int J Mol Sci 2021, 22, 7911. [Google Scholar] [CrossRef]
- Song, Y.; Zheng, S.; Li, L.; Zhang, X.; Zhang, X.; Huang, Z.; Chen, J.; Wang, R.; Zhao, H.; Chong, Y.; et al. Deep Learning Enables Accurate Diagnosis of Novel Coronavirus (COVID-19) With CT Images. IEEE/ACM Trans Comput Biol Bioinform 2021, 18, 2775–2780. [Google Scholar] [CrossRef]
- Reddy, P.; J, A. Diagnosis of Autism in Children Using Deep Learning Techniques by Analyzing Facial Features. Engineering Proceedings 2024, 59, 198. [Google Scholar] [CrossRef]
- Azuma, Y.; Okada, H.; Onami, S. Systematic Analysis of Cell Morphodynamics in C. Elegans Early Embryogenesis. Front Bioinform 2023, 3, 1082531. [Google Scholar] [CrossRef]
- Zhang, J.; Liu, S.; Yuan, H.; Yong, R.; Duan, S.; Li, Y.; Spencer, J.; Lim, E.G.; Yu, L.; Song, P. Deep Learning for Microfluidic-Assisted Caenorhabditis Elegans Multi-Parameter Identification Using YOLOv7. Micromachines (Basel) 2023, 14, 1339. [Google Scholar] [CrossRef]
- Bates, K.; Le, K.N.; Lu, H. Deep Learning for Robust and Flexible Tracking in Behavioral Studies for C. Elegans. PLoS Comput Biol 2022, 18, e1009942. [Google Scholar] [CrossRef]
- Toulany, N.; Morales-Navarrete, H.; Čapek, D.; Grathwohl, J.; Ünalan, M.; Müller, P. Uncovering Developmental Time and Tempo Using Deep Learning. Nat Methods 2023, 20, 2000–2010. [Google Scholar] [CrossRef]
- Pan, Y.; Huang, Z.; Cai, H.; Li, Z.; Zhu, J.; Wu, D.; Xu, W.; Qiu, H.; Zhang, N.; Li, G.; et al. WormCNN-Assisted Establishment and Analysis of Glycation Stress Models in C. Elegans: Insights into Disease and Healthy Aging. Int J Mol Sci 2024, 25, 9675. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation 2015.
- Diakogiannis, F.I.; Waldner, F.; Caccetta, P.; Wu, C. ResUNet-a: A Deep Learning Framework for Semantic Segmentation of Remotely Sensed Data. ISPRS Journal of Photogrammetry and Remote Sensing 2020, 162, 94–114. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition 2015.
- Kubota, Y.; Tsuyama, K.; Takabayashi, Y.; Haruta, N.; Maruyama, R.; Iida, N.; Sugimoto, A. The PAF1 Complex Is Involved in Embryonic Epidermal Morphogenesis in Caenorhabditis Elegans. Dev Biol 2014, 391, 43–53. [Google Scholar] [CrossRef] [PubMed]
- Köppen, M.; Simske, J.S.; Sims, P.A.; Firestein, B.L.; Hall, D.H.; Radice, A.D.; Rongo, C.; Hardin, J.D. Cooperative Regulation of AJM-1 Controls Junctional Integrity in Caenorhabditis Elegans Epithelia. Nat Cell Biol 2001, 3, 983–991. [Google Scholar] [CrossRef]
- Lynch, A.M.; Zhu, Y.; Lucas, B.G.; Winkelman, J.D.; Bai, K.; Martin, S.C.T.; Block, S.; Slabodnick, M.M.; Audhya, A.; Goldstein, B.; et al. TES-1/Tes and ZYX-1/Zyxin Protect Junctional Actin Networks under Tension during Epidermal Morphogenesis in the C. Elegans Embryo. Curr Biol 2022, 32, 5189–5199.e6. [Google Scholar] [CrossRef]
- Reiss, A.L. Childhood Developmental Disorders: An Academic and Clinical Convergence Point for Psychiatry, Neurology, Psychology and Pediatrics. J Child Psychol Psychiatry 2009, 50, 87–98. [Google Scholar] [CrossRef]
- Brenner, S. The Genetics of Caenorhabditis Elegans. Genetics 1974, 77, 71–94. [Google Scholar] [CrossRef]
- Maeda, I.; Kohara, Y.; Yamamoto, M.; Sugimoto, A. Large-Scale Analysis of Gene Function in Caenorhabditis Elegans by High-Throughput RNAi. Curr Biol 2001, 11, 171–176. [Google Scholar] [CrossRef]
- Kubota, Y.; Ota, N.; Takatsuka, H.; Unno, T.; Onami, S.; Sugimoto, A.; Ito, M. The PAF1 Complex Cell Autonomously Promotes Oogenesis in Caenorhabditis Elegans. Genes Cells 2022, 27, 409–420. [Google Scholar] [CrossRef]
- Hardin, J.; Serre, J.; King, R.; Walck-Shannon, E.; Reiner, D. Imaging Epidermal Cell Rearrangement in the C. Elegans Embryo. Methods Mol Biol 2022, 2438, 345–376. [Google Scholar] [CrossRef]
- Oegema, K.; Hyman, T. Cell Division. In WormBook: The Online Review of C. elegans Biology [Internet]; WormBook, 2006.
| Metric | Value |
|---|---|
| True Positive (TP) | 519,293 |
| False Positive (FP) | 14,760 |
| True Negative (TN) | 714,840 |
| False Negative (FN) | 4,483 |
| Sensitivity (TPR) | 99.14% |
| Specificity (TNR) | 97.98% |
| Overall Accuracy | 98.47% |
| Precision (PPV) | 97.24% |
| F1-Score | 98.18% |
| Intersection over Union (IoU) | 96.43% |
| Metric | Value |
|---|---|
| Sensitivity (TPR) | 96.87% |
| Specificity (TNR) | 97.98% |
| Overall Accuracy | 96.86% |
| Precision (PPV) | 96.93% |
| F1-Score | 96.83% |
| RNAi | Number of timelines | Timeline number with misclassification | Total images | Images number with misclassification | Continuous misclassification |
|---|---|---|---|---|---|
| control(RNAi) | 16 | 1 | 681 | 0.15% (N=1) | 0 |
| leo-1(RNAi) | 10 | 2 | 541 | 0.37%(N=2) | 0 |
| ajm-1(RNAi) | 10 | 2 | 467 | 0.64%(N=3) | 0 |
| tes-1(RNAi) | 10 | 3 | 457 | 0.66%(N=3) | 0 |
| RNAi | Dorsal intercalation (min) | Ventral enclosure (min) | Rotation (min) |
1.5-fold (min) |
2-fold (min) |
|---|---|---|---|---|---|
| control(RNAi) | 53.75±1.85 | 28.43±2.02 | 15.93±1.38 | 20.93±1.04 | 10.62±0.89 |
| leo-1(RNAi) | 64.50 ± 4.74 * | 44.50 ± 4.80 ** | 28.50±3.08 ** | 21.00 ± 2.56 | 11.50 ± 1.50 |
| ajm-1(RNAi) | 50.50 ± 4.47 | 25.50 ± 3.37 | 21.00 ± 3.23 | 27.50 ± 2.39 * | 16.00 ± 1.80 ** |
| tes-1(RNAi) | 49.50 ± 5.46 | 28.50 ± 3.73 | 15.00 ± 2.47 | 21.00 ± 1.25 | 20.50 ± 1.74 **** |
| * P < 0.05; ** P < 0.01; *** P < 0.001; **** P < 0.0001 | |||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




