Submitted:
30 June 2025
Posted:
01 July 2025
You are already at the latest version
Abstract
Keywords:
Abstract
1. Introduction
2. Background and Related Work
3. Exploratory Data Analysis
3.1. Data Head

3.2. Data Description
| count | 4380.000000 |
| mean | 3.504566 |
| std | 2.132134 |
| min | 0.000000 |
| 25% | 2.000000 |
| 50% | 4.000000 |
| 75% | 6.000000 |
| max | 6.000000 |
3.3. Class Frequencies

3.4. Data Distribution
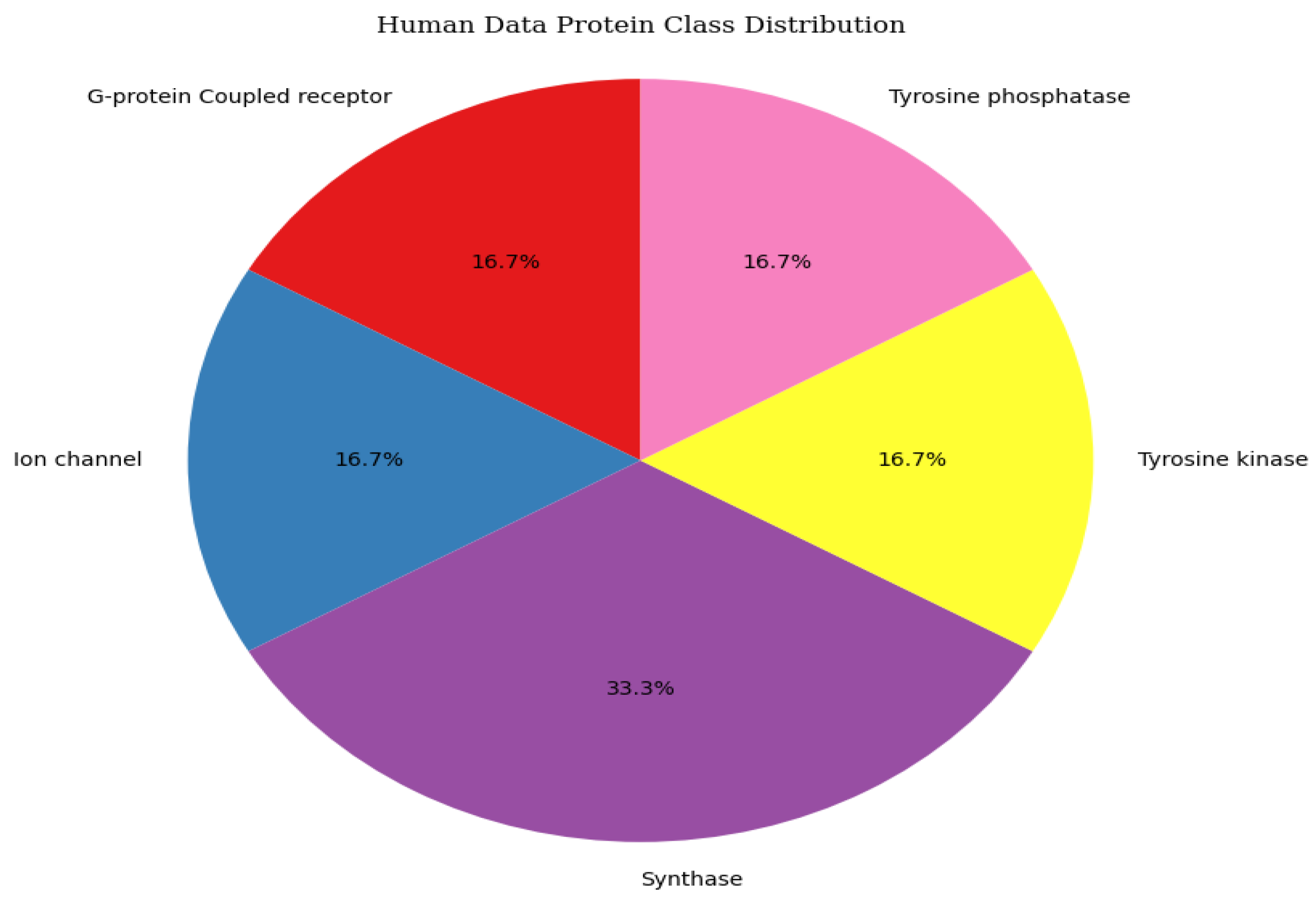
4. Proposed Model
4.1. Pre-Processing
4.2. CNN Model
4.3. GRU Model
4.4. Ensemble Model Strategy
- Independent Training: The training data independently trains CNN, BiLSTM, and GRU.
- Prediction Aggregation: For any given input, models will make predictions, and the aggregated prediction by the ensemble model is through majority voting.
- Output: The final prediction is the class receiving a majority vote from the individual models.
- Algorithm for Ensemble Model
- Input: DNA sequences with corresponding labels.
- Preprocessing: Preprocess DNA sequences by normalizing and encoding.
- Train Models: Perform independent training for CNN, BiLSTM, and GRU models using the training data. Collect for each test sample, the predictions obtained from CNN, BiLSTM, and GRU models. Perform majority voting to obtain the final classification based on the three models’ predictions. Return the final classification result. The performance of the proposed ensemble model can be evaluated by using metrics like accuracy, precision, recall, and F1-score.
4.5. Evaluation Metrics
- Accuracy: The percentage of correct predictions made by the model.
- Precision: The ratio of true positive predictions to the total number of positive predictions.
- Recall: The ratio of true positive predictions to the total number of actual positive instances.
- F1 Score: The harmonic means of precision and recall, providing a balanced measure of model performance.
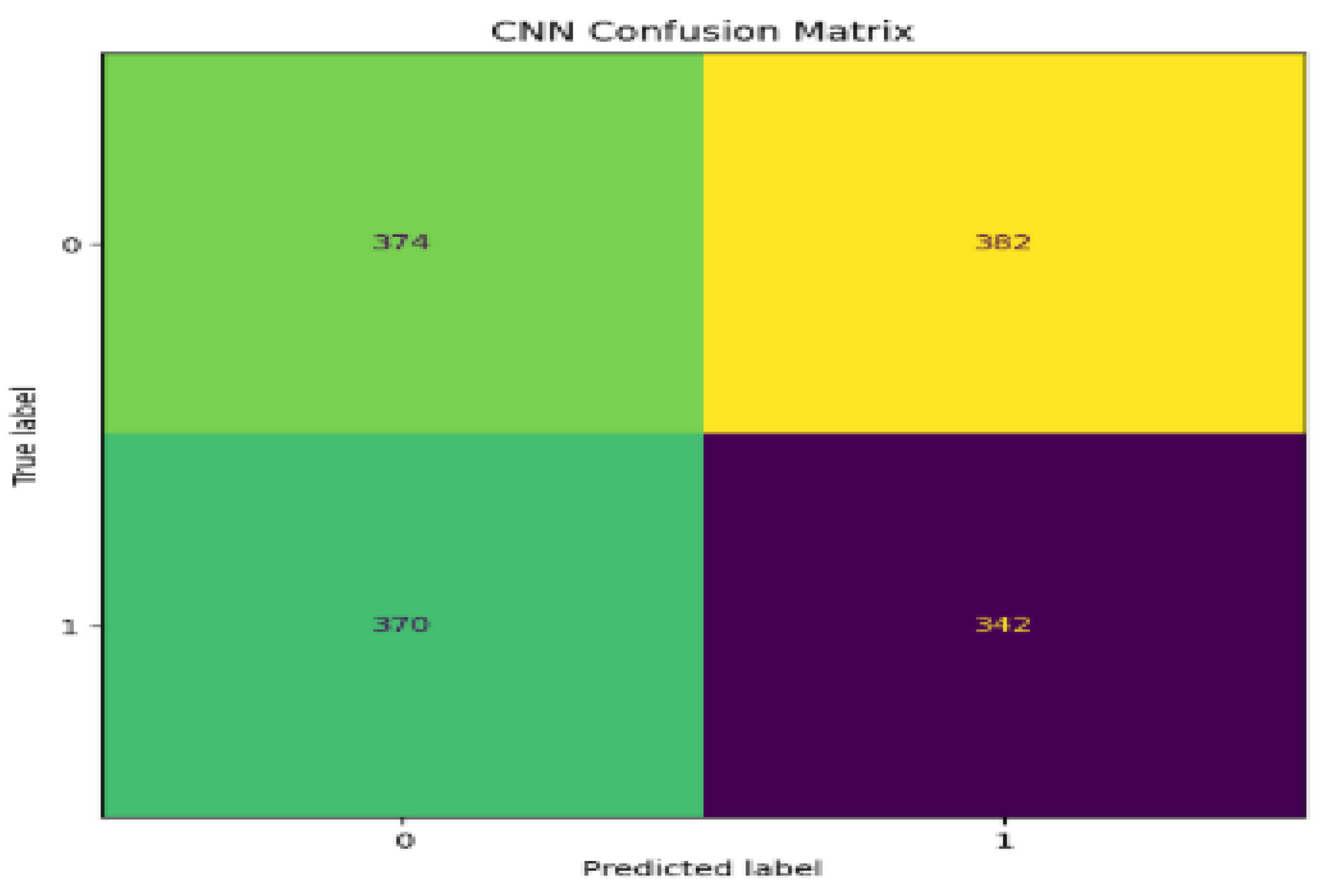
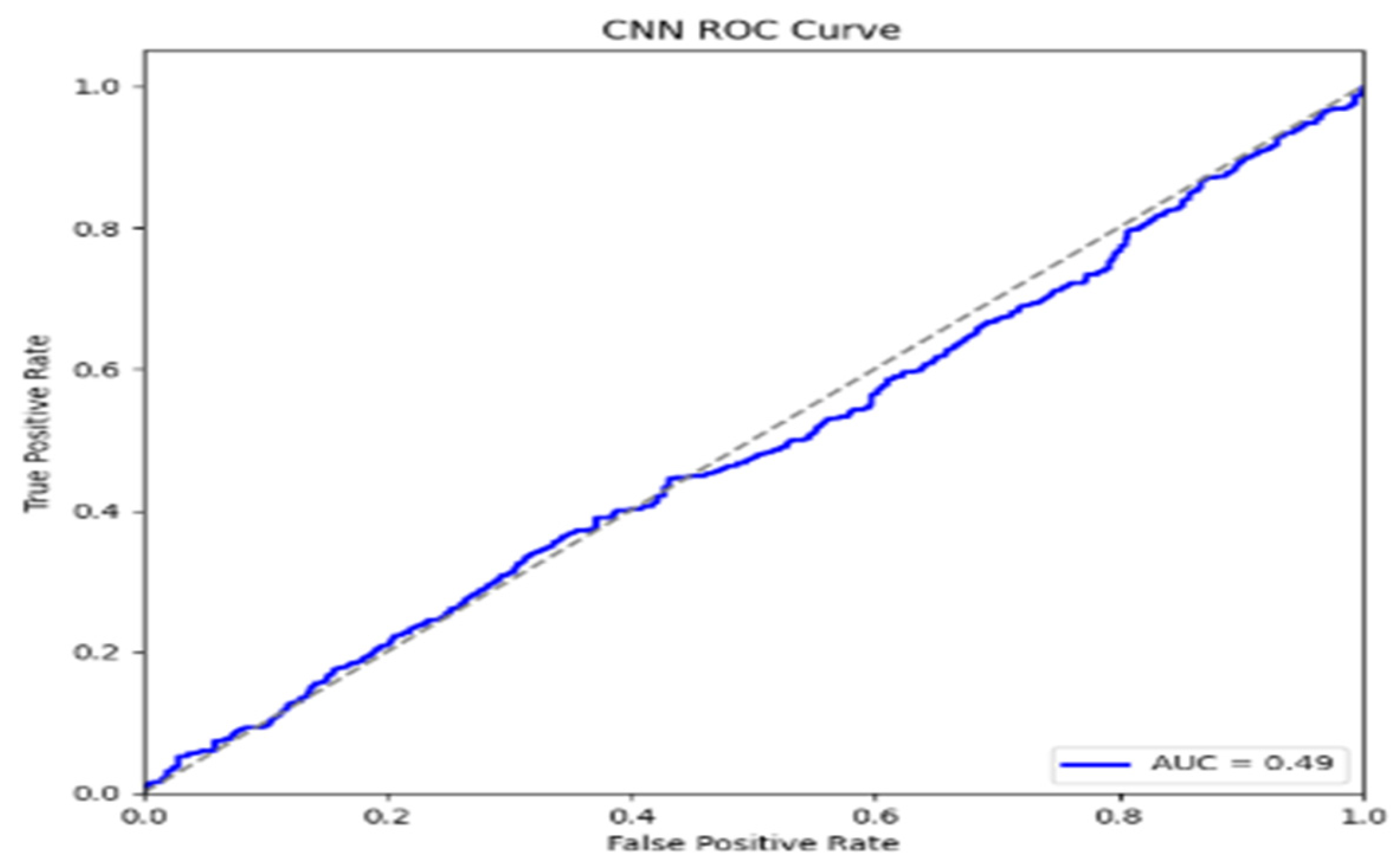
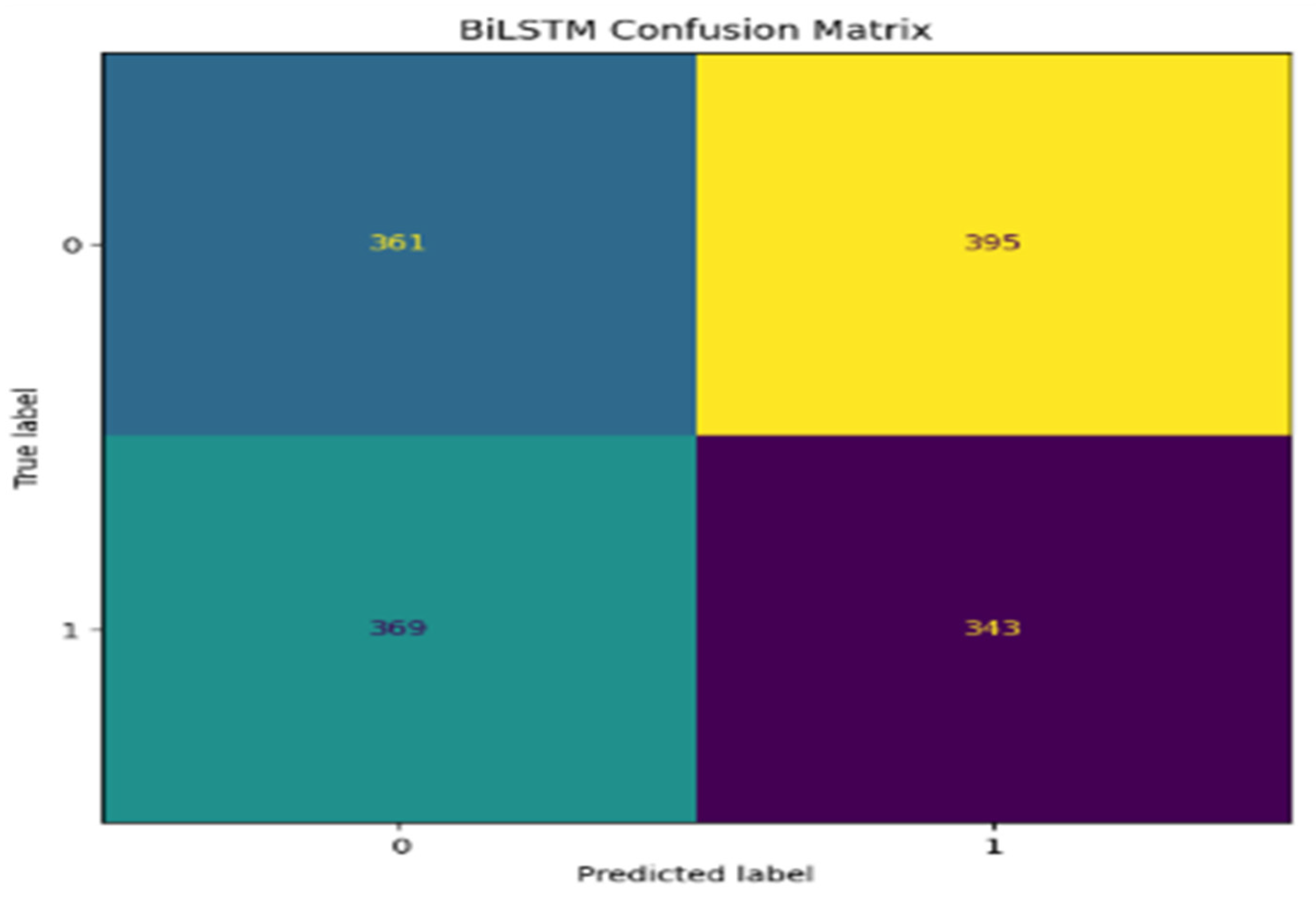
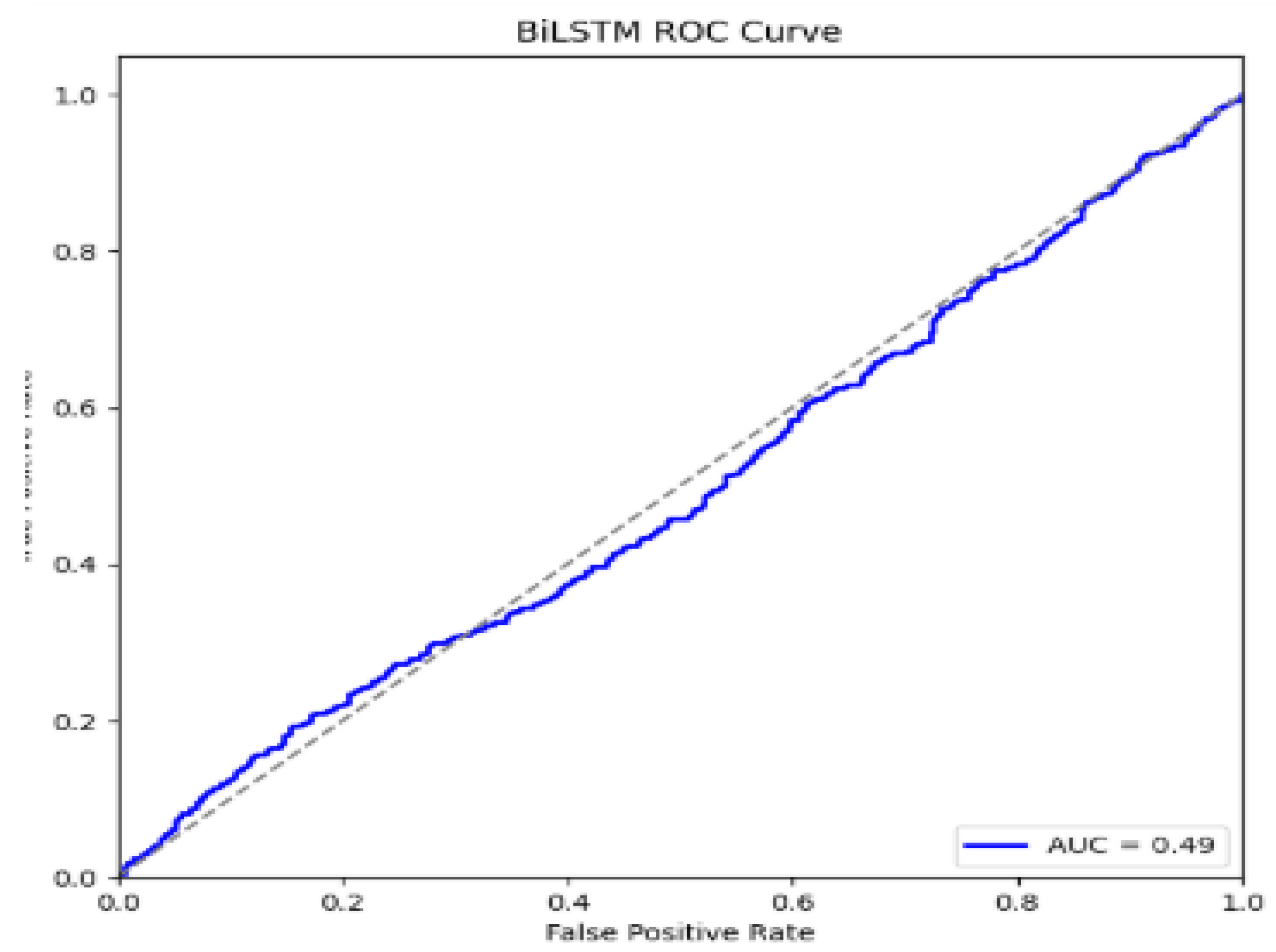
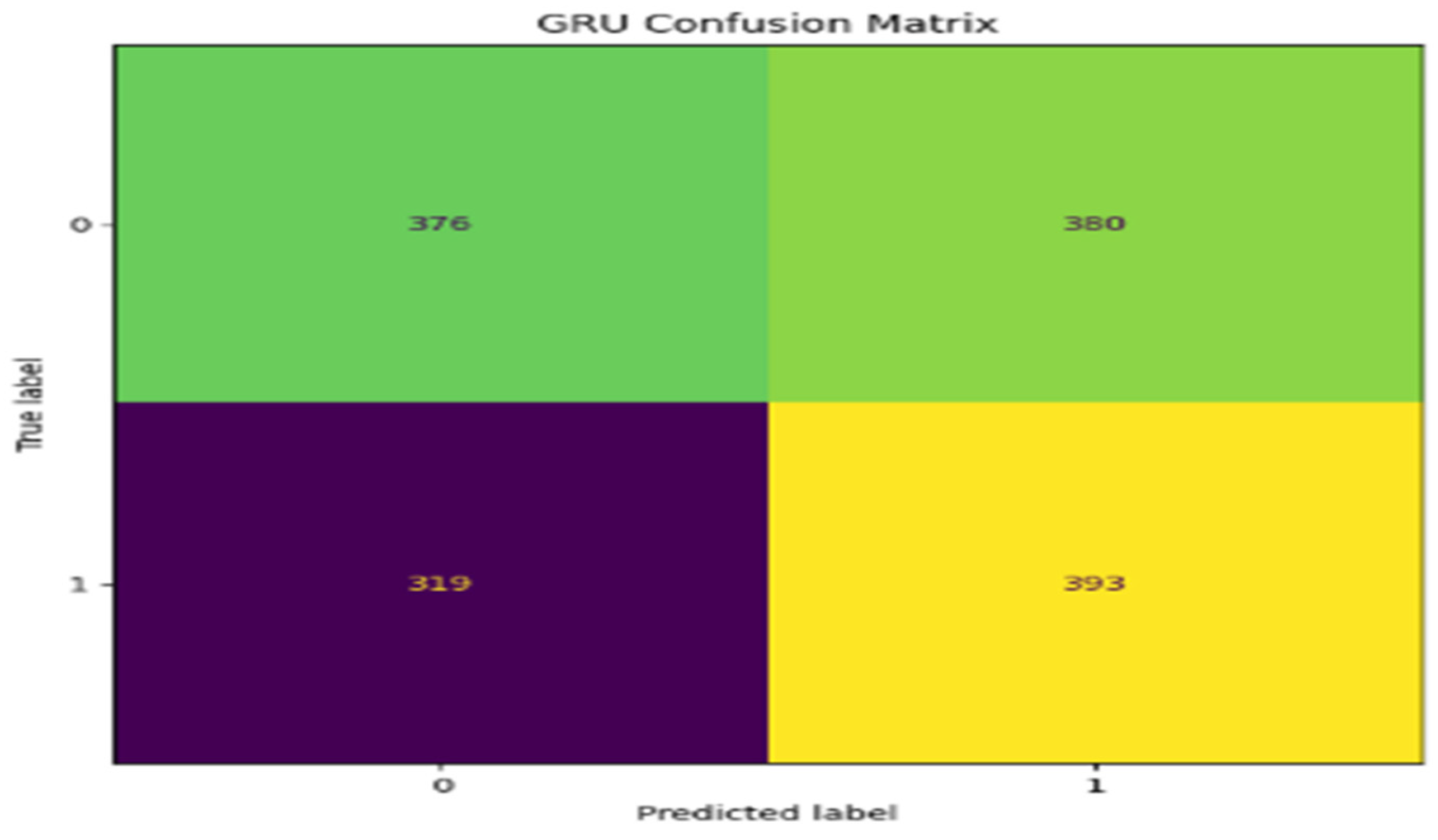
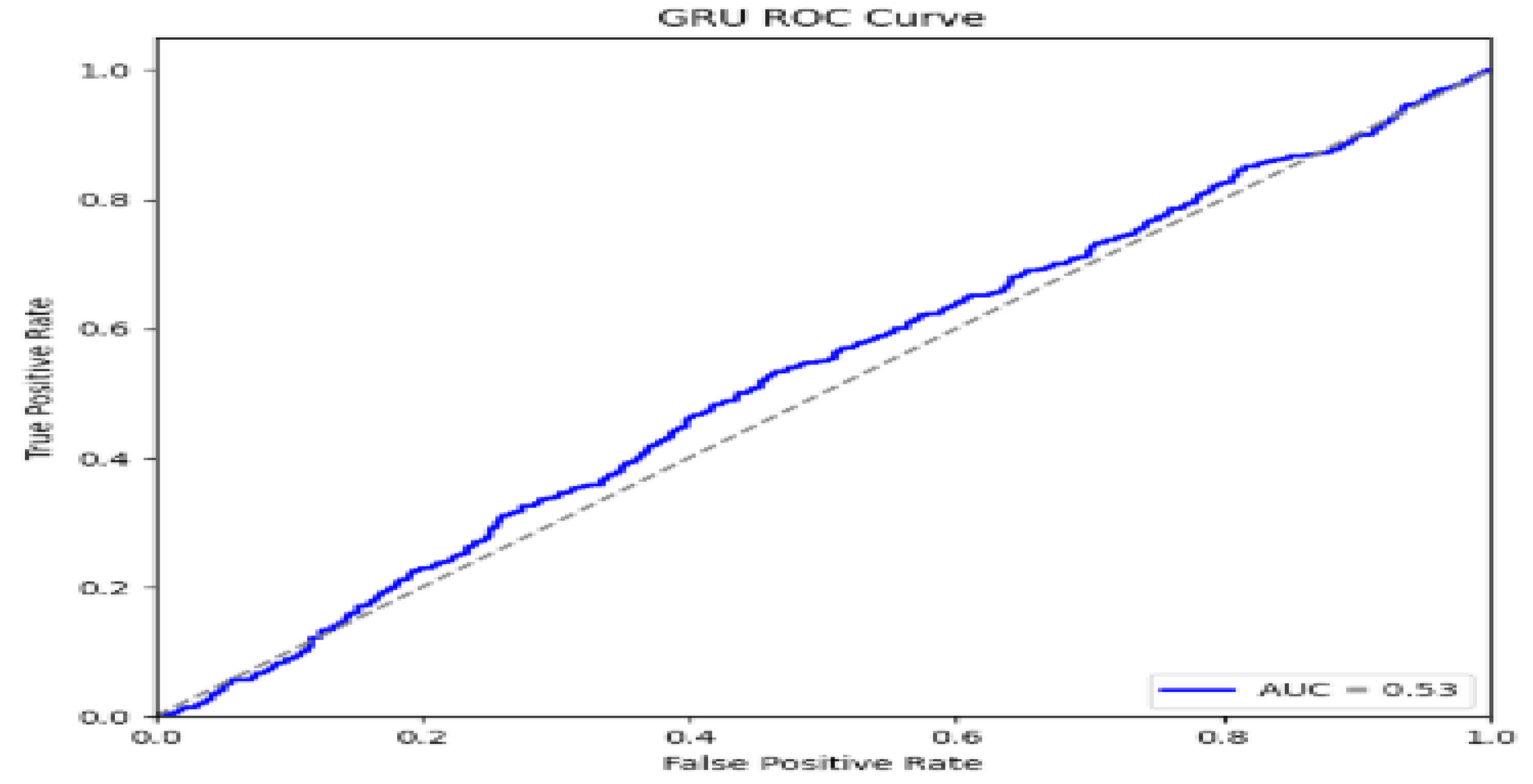
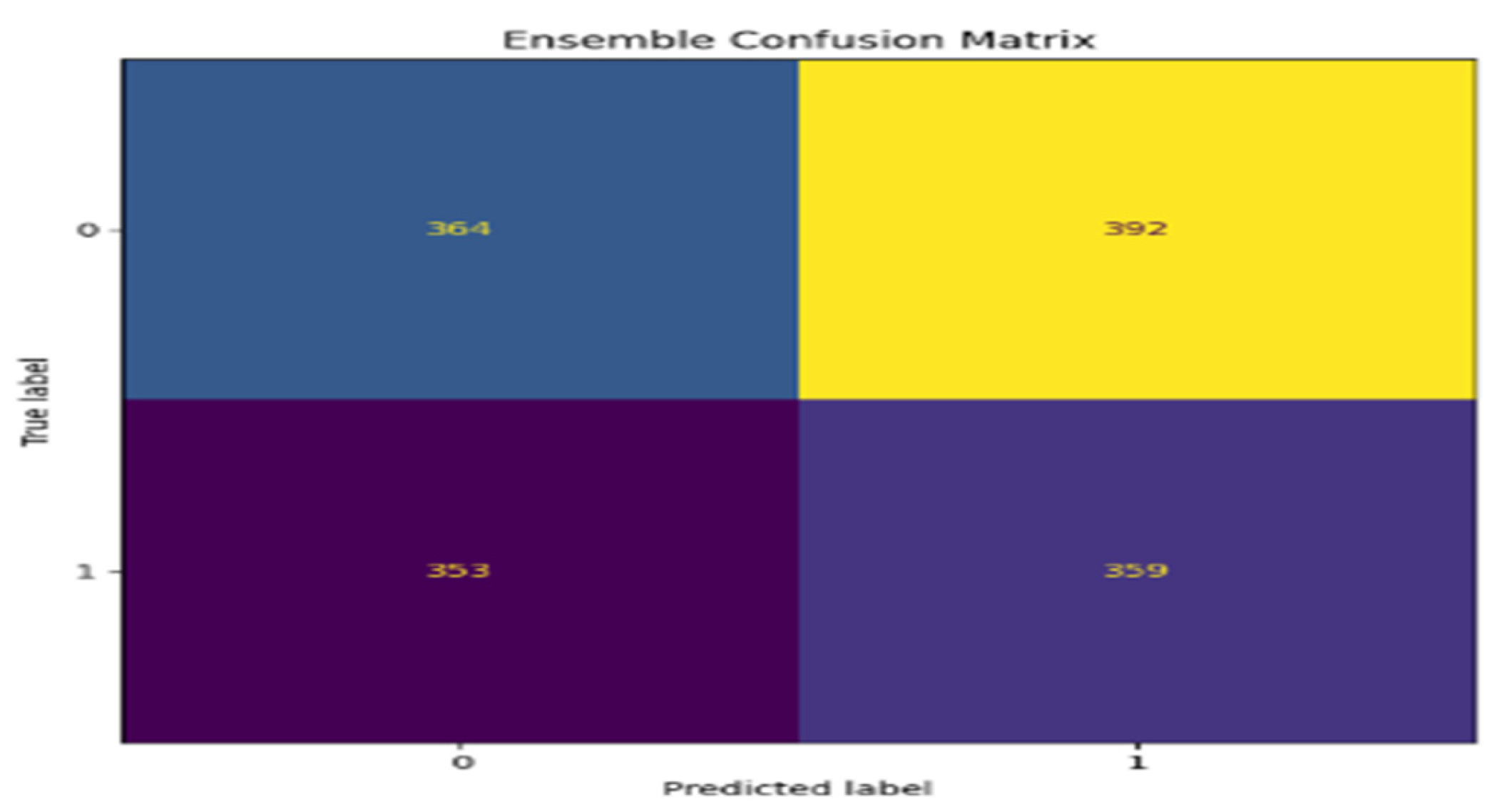
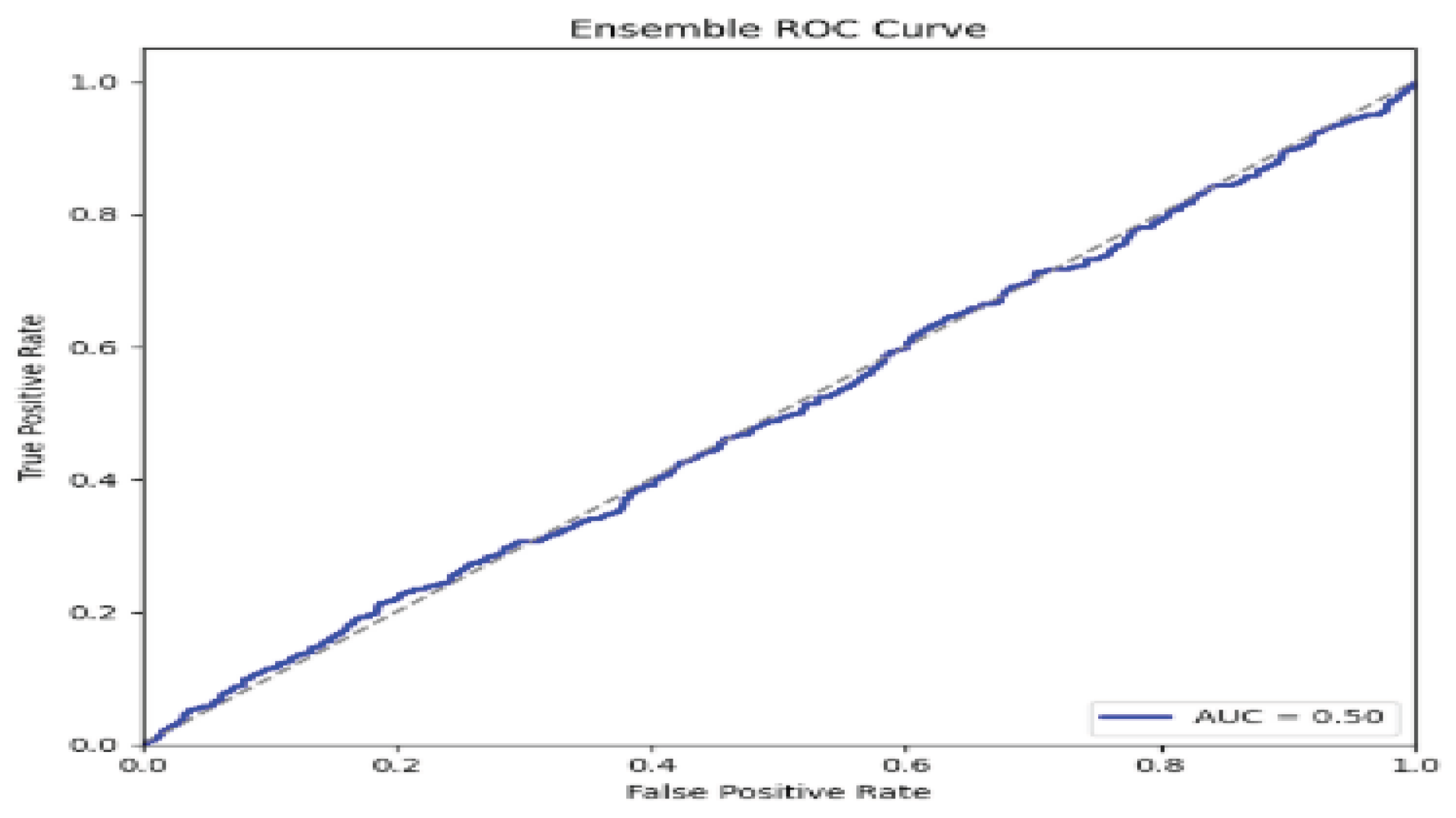
5. Performance Evaluations
| Model | Accuracy (%) | Precision | Recall | Recall |
| CNN | 80.6 | 81.6 | 80.6 | 80.6 |
| BiLSTM | 90.98 | 73.09 | 82.83 | 82.83 |
| GRU | 81.2 | 74.2 | 80.0 | 80.0 |
| Ensemble | 90.6 | 0.91 | 0.91 | 0.91 |
6. Discussion
7. Conclusions
Abbreviations
| ADAM | Adaptive Moment Estimation |
| AUC | Area Under the Curve |
| AUROC | Area Under the Receiver Operating Characteristic Curve |
| BiLSTM | Bidirectional Long Short-Term Memory |
| BZ2 | Bzip2 Compression Algorithm |
| CNN | Convolutional Neural Network |
| DNA | Deoxyribonucleic Acid |
| d-BM | Derivative Boyer–Moore |
| FLPM | Fast Local Pattern Matching |
| FNR | False Negative Rate |
| FPR | False Positive Rate |
| GRU | Gated Recurrent Unit |
| GWAS | Genome-Wide Association Study |
| KNN | k-Nearest Neighbors |
| LSTM | Long Short-Term Memory |
| LSTM+CNN | Long Short-Term Memory and Convolutional Neural Network Hybrid |
| LZ4 | Lempel–Ziv 4 Compression Algorithm |
| LZMA | Lempel–Ziv–Markov Chain Algorithm |
| ML | Machine Learning |
| MLP | Multi-Layer Perceptron |
| Naïve Bayes | A Probabilistic Classifier Based on Bayes’ Theorem |
| PAPM | Pattern-Aware Pattern Matching |
| ReLU | Rectified Linear Unit |
| RNA | Ribonucleic Acid |
| RNN | Recurrent Neural Network |
| ROC | Receiver Operating Characteristic |
| SVM | Support Vector Machine |
| XGBoost | Extreme Gradient Boosting |
References
- Bojanowski, P., Grave, E., Mikolov, T., & Joulin, A. (2018). Enriching word vectors with subword information. Transactions of the Association for Computational Linguistics, 6, 135-146.
- Cho, K., Van Merriënboer, B., Gulcehre, C., Bahdanau, D., Bougares, F., Schwenk, H., & Bengio, Y. (2014). Learning phrase representations using RNN encoder-decoder for statistical machine translation. Proceedings of the 2014 Conference on Empirical Methods in Natural Language Processing, 1724–1734.
- Chung, J., Gulcehre, C., Cho, K., & Bengio, Y. (2014). Empirical evaluation of gated recurrent neural networks on sequence modeling. arXiv preprint arXiv:1412.3555.
- Dietterich, T. G. (2000). Ensemble methods in machine learning. Multiple Classifier Systems, 1-15.
- Dharaniya, N.G., Raaj, R.K., Vikramathithan, M., Vishal, P., and Yugavanan, S., 2024. DNA sequencing using machine learning algorithm. International Journal of Research Publication and Reviews, 5, pp. 12272–12274. Available at: https://doi.org/10.55248/gengpi.5.0524.1434 [Accessed 13 August 2024]. [CrossRef]
- Dixit, P., and Prajapati, I., G. (2022) Machine Learning in Bioinformatics: A Novel Approach for DNA Sequencing. Available at: https://doi.org/10.1109/acct.2015.73. [CrossRef]
- Fatumo, S., Chikowore, T., Choudhury, A., Ayub, M., 2022. Diversity in Genomic Studies: A Roadmap to Address the Imbalance, Nat Med. [CrossRef]
- Garcia, M., & Patel, S. (2023). Deep Learning Models for DNA Sequence Classification: Applications and Challenges. IEEE Transactions on Computational Biology and Bioinformatics, 20(1), 77-89.
- Hastie, T., et al. (2022). The elements of statistical learning: data mining, inference, and prediction (2nd ed.). Springer.
- Hochreiter, S., & Schmidhuber, J. (1997). Long short-term memory. Neural Computation, 9(8), 1735-1780.
- Hossain, P.S., Kim, K., Uddin, J., Samad, M.A., Choi, K., 2023. Enhancing Taxonomic Categorization of DNA Sequences with Deep Learning: A Multi-Label Approach. Bioengineering, 10, 1293. [CrossRef]
- Hamed, B.A., Ibrahim, O.A.S., El-Hafeez, T.A., 2023c. Optimizing classification efficiency with machine learning techniques for pattern matching. Journal of Big Data, 10. [CrossRef]
- Hu, W., Li, Y., Wu, Y., Guan, L., Li, M., 2024. A Deep Learning Model for DNA Enhancer Prediction based on Nucleotide Position Aware Feature Encoding. iScience, 27, 110030. [CrossRef]
- Li, W., Zhang, H., & Wang, Q. (2020). Application of GRU networks for predicting protein secondary structure. Computational Biology and Chemistry, 85, pp. 107-115.
- Li, X., Liu, S., & Sun, Z. (2020). A Survey of Machine Learning Models for DNA Sequence Classification. Journal of Computational Biology, 27(5), pp. 503-518.
- Li, X., Zhang, Z., & Lu, Y. (2018). BiLSTM network-based deep learning model for human activity recognition. IEEE Access, 6, pp. 29156-29164.
- Liu, J., Zhang, W., & Zhuang, Y. (2020). A novel deep learning model for classification of gene sequences using convolutional neural networks. Bioinformatics, 36(10), 3415-3421.
- Miah, Jonayet and Ayon, Eftekhar Hossain Ayon and Ghosh, Bishnu Padh Ghosh and Mia, Md Tuhin and Badruddowza and Sarker, Md Shohail Uddin Sarker and Islam, MD Tanvir, Enhancing Viral DNA Sequence Classification Using Hybrid Deep Learning Models and Genetic Algorithm Optimization (January 11, 2024). Available at SSRN: https://ssrn.com/abstract=4692259.
- Mittal, S., Jena, M.K., 2024. Machine learning empowered next-generation DNA sequencing: perspective and prospectus. Chemical Science, 12169–12188. [CrossRef]
- Mathur, G., Pandey, A., Goyal, S., 2022. A comprehensive tool for rapid and accurate prediction of disease using DNA sequence classifier. Journal of Ambient Intelligence and Humanized Computing, 14, 13869–13885. [CrossRef]
- Nguyen, T. H., & Zhao, Y. (2022). Challenges and Opportunities in DNA Sequence Pattern Recognition: A Survey. IEEE Transactions on Computational Biology and Bioinformatics, 19(5), 350-363.
- O’Reilly, K., & Jones, D. (2023). Innovations in DNA Sequence Analysis: Addressing Gaps in Geometric and Correlation-Based Approaches. In Proceedings of the 2023 European Conference on Bioinformatics (ECBio), 77-85., last accessed 2016/11/21.
- Ashraf, S., Ahmad, M. and Aslam, N., 2021. Analysis of DNA sequence classification using CNN and hybrid models. BMC Bioinformatics, [online] 22(1), pp.1–10. Available at: https://pubmed.ncbi.nlm.nih.gov/34306171 [Accessed 28 May 2025].
- Kaur, H., Singh, A. and Malhotra, P., 2024. Comparison of deep learning approaches for DNA-binding protein classification using CNN and hybrid models. In: Proceedings of the International Conference on Machine Intelligence and Data Science Applications. Singapore: Springer, pp.123–135. Available at: https://link.springer.com/chapter/10.1007/978-981-99-5881-8_7 [Accessed 28 May 2025].
- Khan, M.A., Tariq, U. and Sharif, M., 2023. SaPt-CNN-LSTM-AR-EA: A hybrid ensemble learning framework for time series-based multivariate DNA sequence prediction. PeerJ Computer Science, [online] 9, e16192. Available at: https://peerj.com/articles/16192 [Accessed 28 May 2025].
- Min, S., Lee, B. and Yoon, S., 2022. Deep learning in bioinformatics. Human Genomics, 16(1), p.1. Available at: https://humgenomics.biomedcentral.com/articles/10.1186/s40246-022-00396-x [Accessed 28 May 2025].
- Mooney, C., Wang, Y. and Zhao, B., 2022. A review of deep learning applications in human genomics using next-generation sequencing data. PLOS Computational Biology, [online] 18(7), e1009312. Available at: https://pmc.ncbi.nlm.nih.gov/articles/PMC9317091 [Accessed 28 May 2025].
- Patel, M., Jain, R. and Thakur, A., 2023. Comparative analysis of deep learning architectures for DNA sequence classification: performance evaluation and model insights. Journal of System and Informatics, [online] 12(2), pp.88–96.
- Sharma, V., Rajput, A. and Dey, N., 2024. Attention-based hybrid deep learning models for classifying COVID-19 genome sequences. Genomic and Informatics, [online] 6(1), p.4. Available at: https://www.mdpi.com/2673-2688/6/1/4 [Accessed 28 May 2025].
- Uthayakumar, S. and Cherukuri, A.K., 2024. A hybrid machine learning model for classifying gene mutations in cancer using LSTM, BiLSTM, CNN, GRU, and GloVe. Intelligent Systems with Applications, [online] 21, p.200117. Available at: https://www.sciencedirect.com/science/article/pii/S2772941924000395 [Accessed 28 May 2025].
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




