Submitted:
09 June 2025
Posted:
11 June 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Patient Description
2.1.1. Patient 1
2.1.2. Patient 2
2.1.3. Patient 3:
2.2. Whole Genome Sequencing
2.3. Plasmids
2.4. Cell Culture
2.5. Enzyme Activity Measurements
2.6. Reporter Gene Assays
2.7. Western Blot
2.8. RNA Isolation and Quantitative Real-Time PCR
2.9. In Silico Mutagenesis and Analysis of Protein Stability
2.10. Statistical Analysis
3. Results
3.1. Pathogenic NGLY1 Variants in Two Families with NGLY1 Deficiency
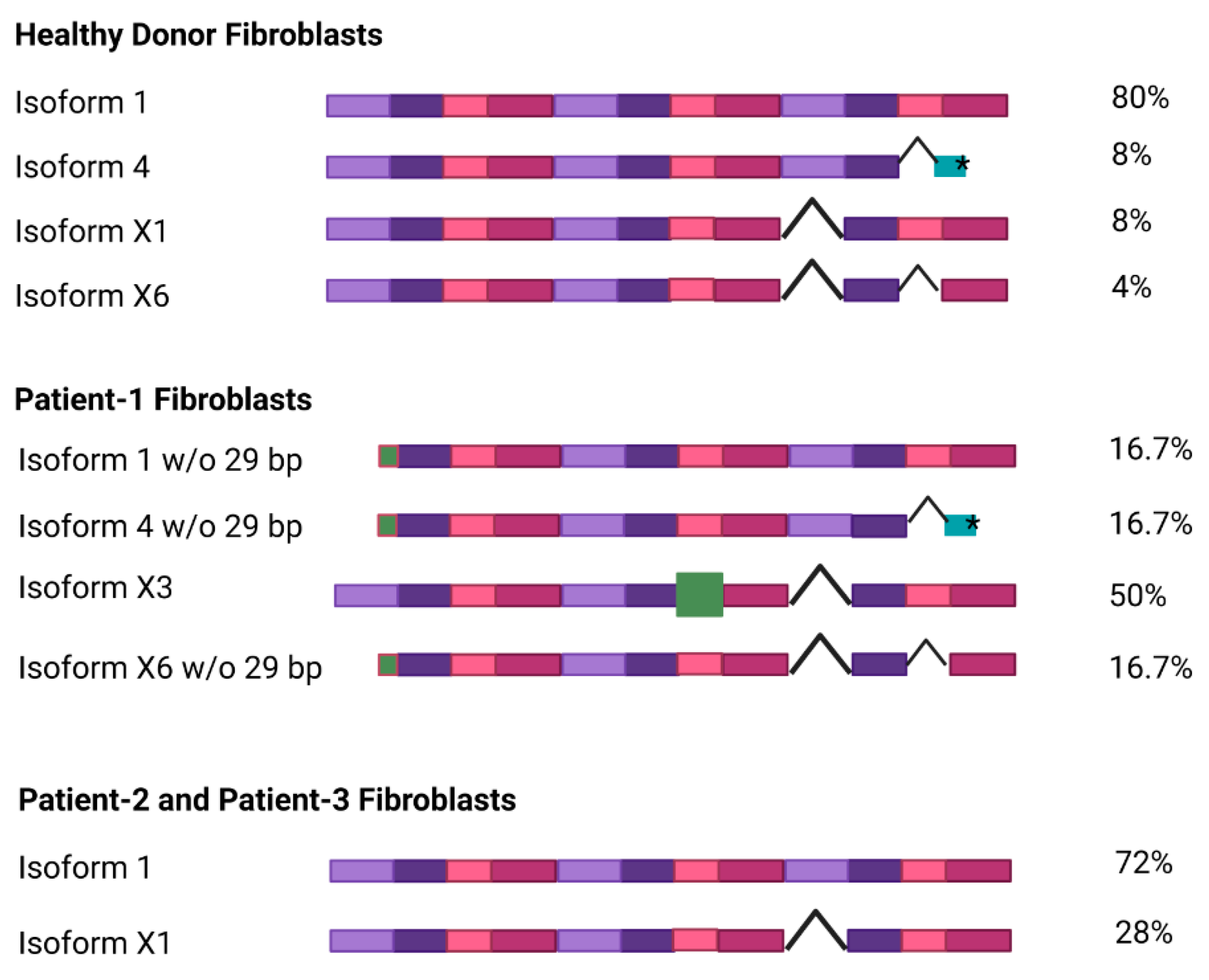
3.2. NGLY1 Isoforms in Patient Fibroblasts
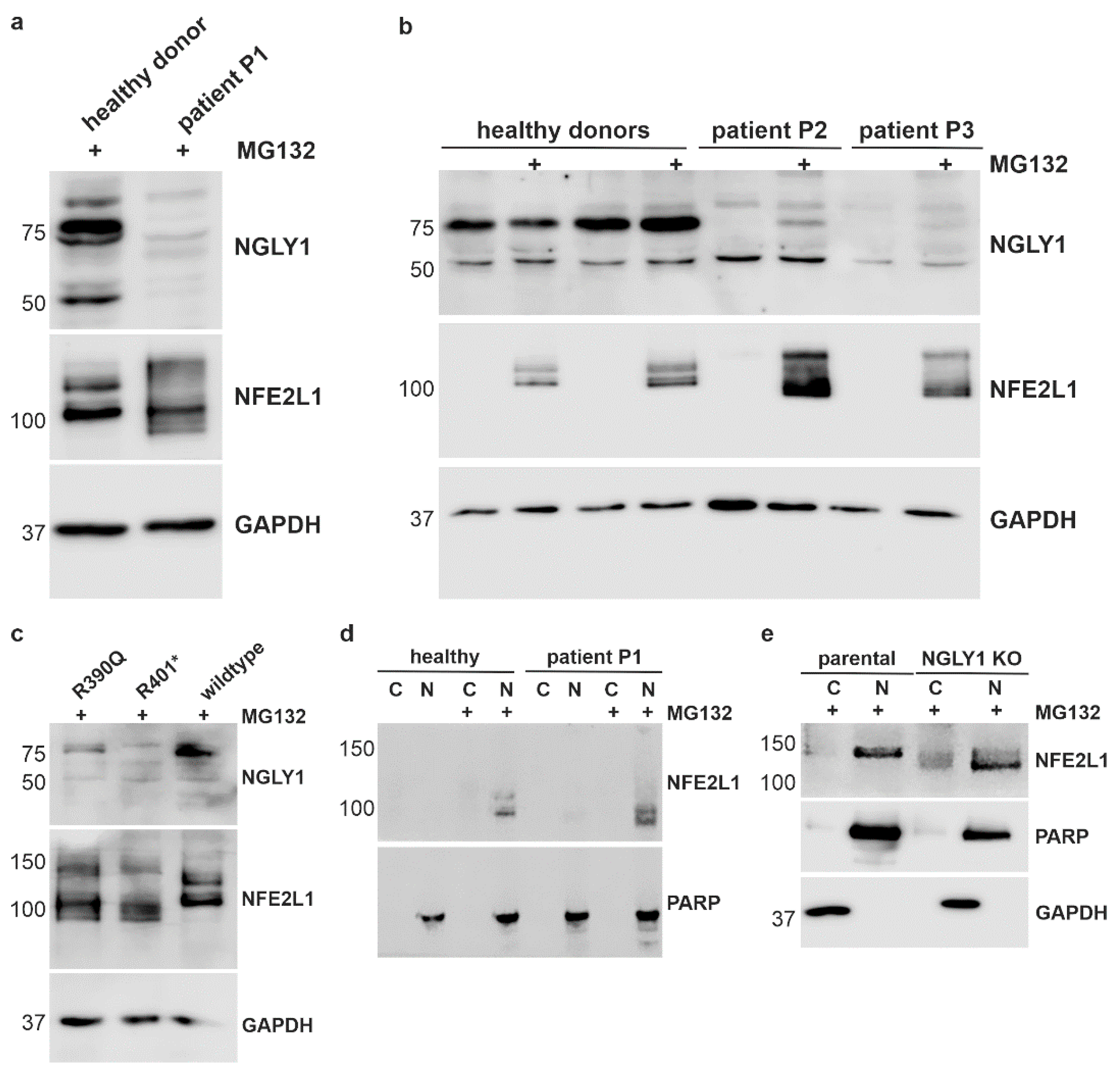
3.3. Misprocessing of the Transcription Factor NFE2L1 in Patient Cells and in NGLY1 Knockout HEK293 Cells
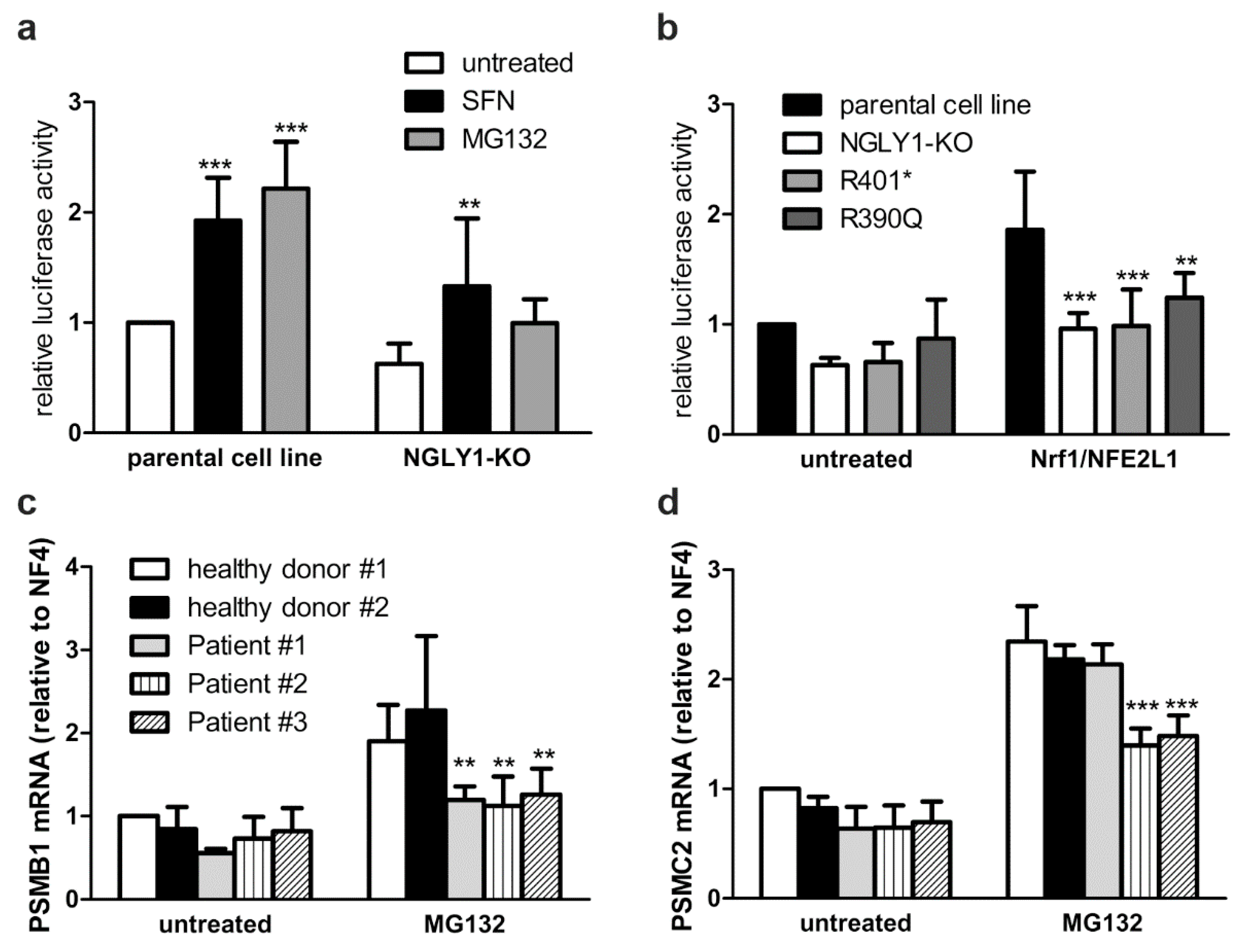
3.4. Reduced Activity of NFE2L1 and Reduced Expression of NFE2L1 Target Genes in NGLY1-Deficient Cells
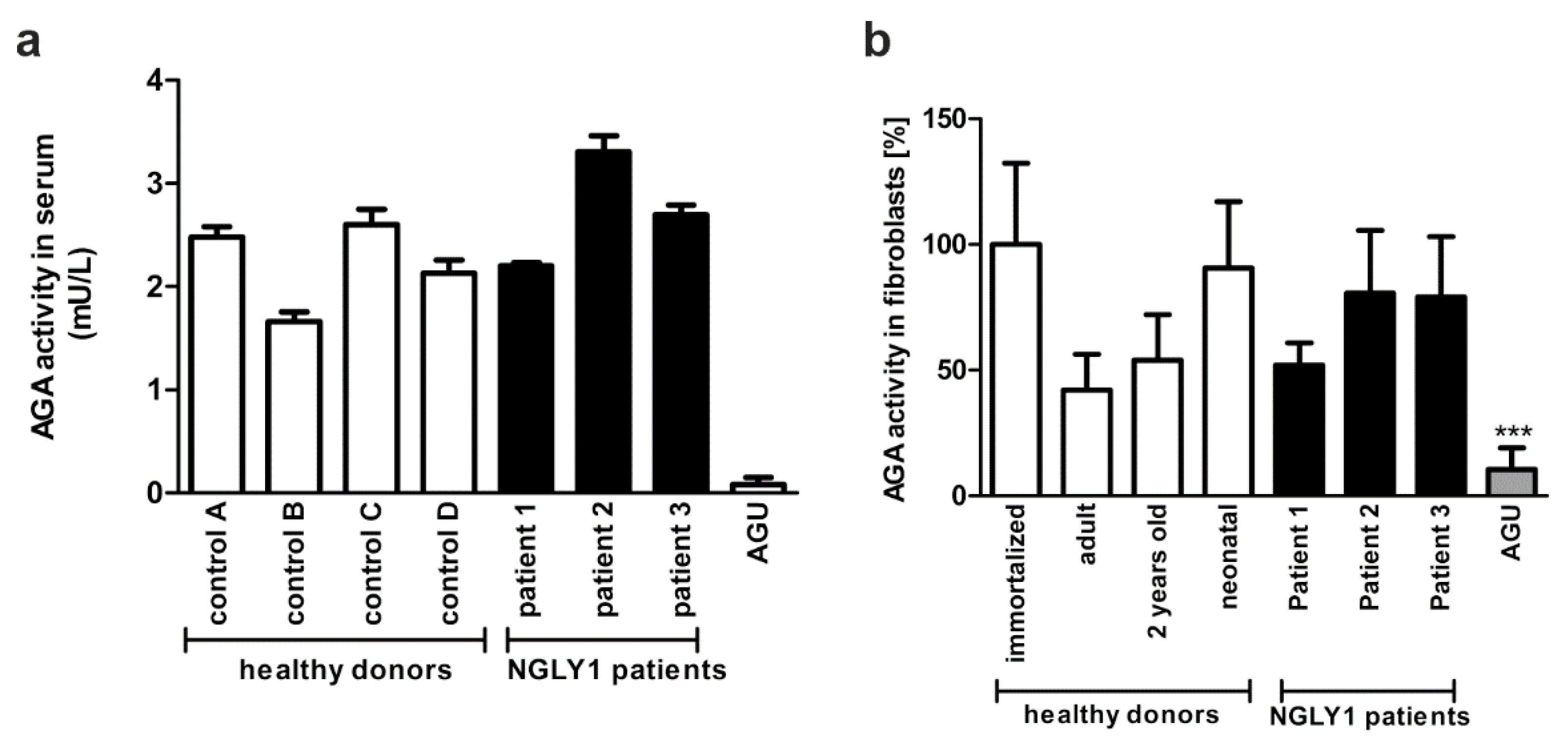
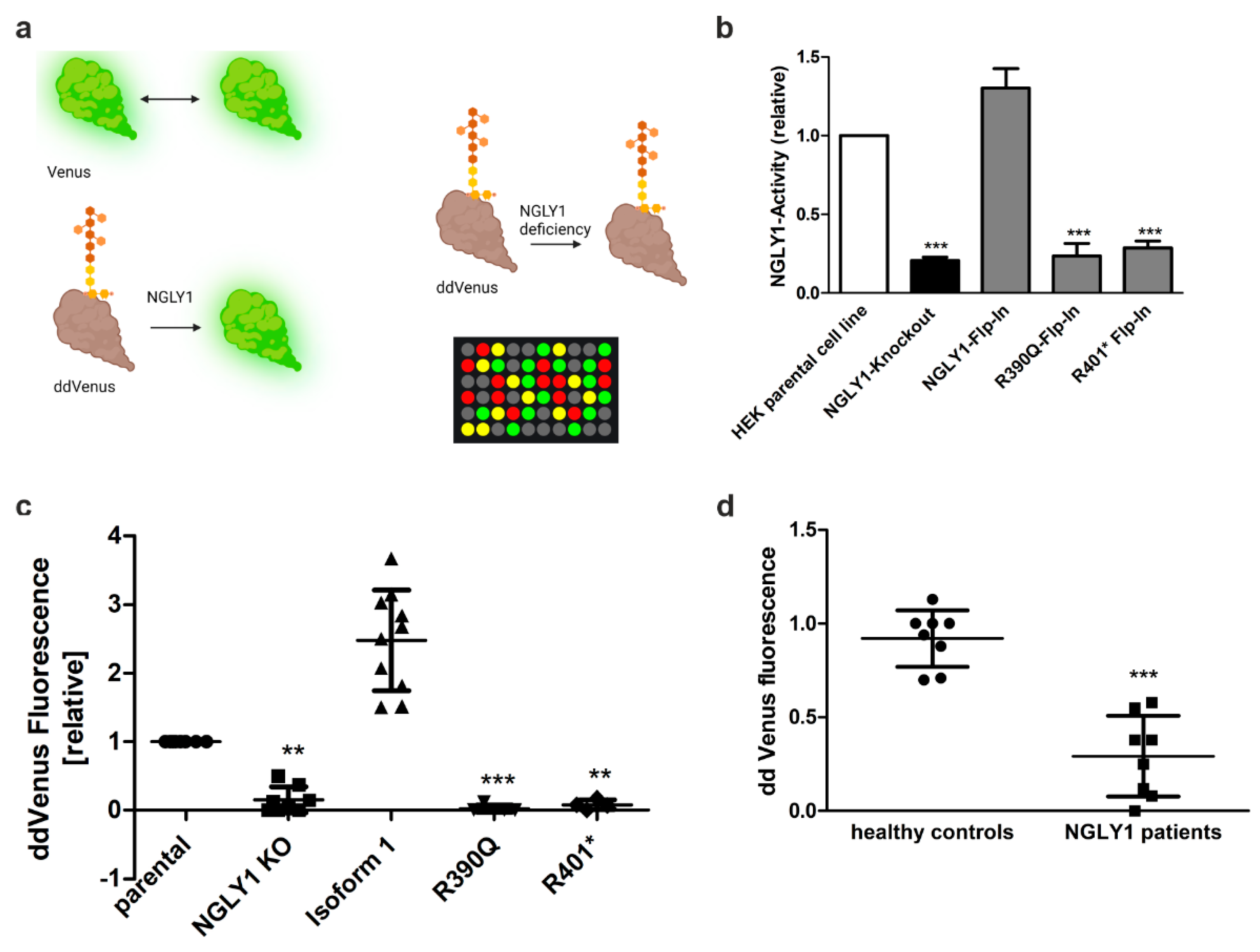
3.5. Establishment of an NGLY1 Activity Assay
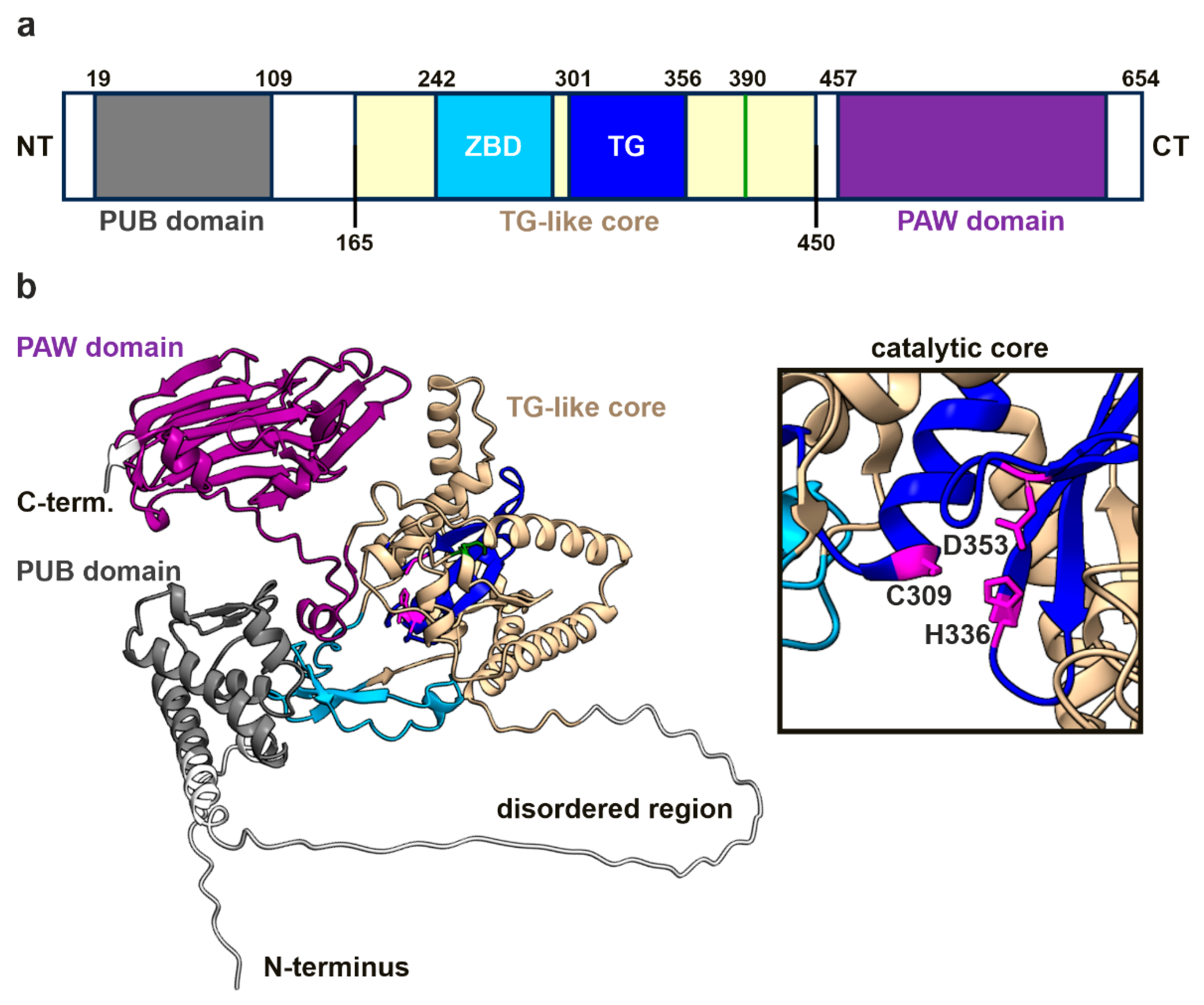
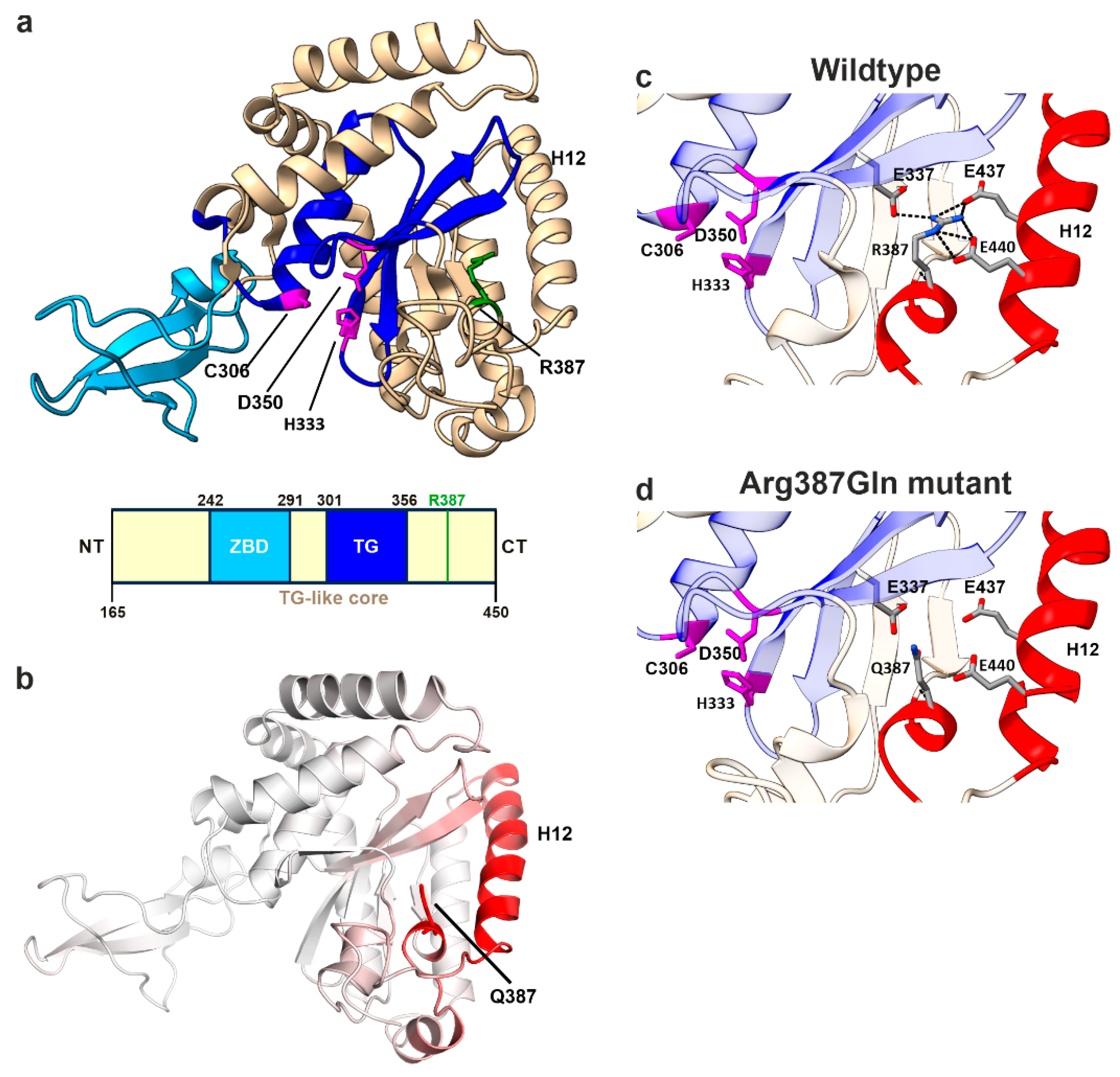
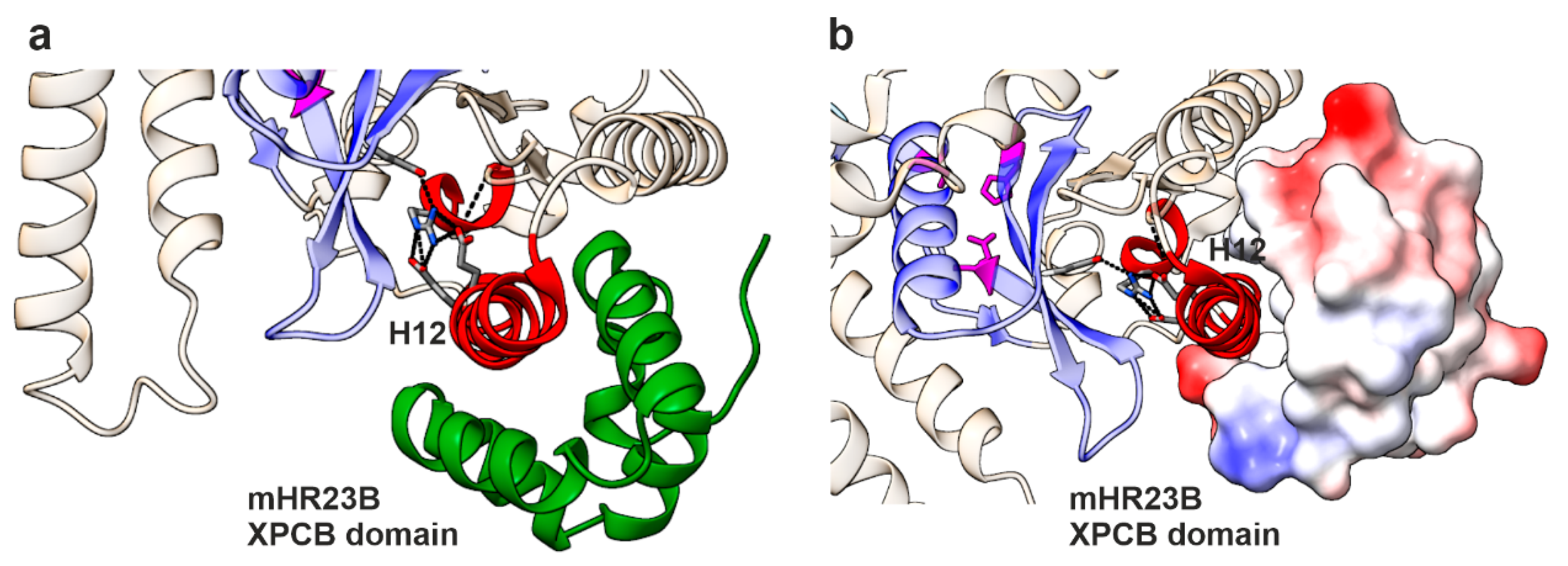
3.6. In-Silico Mutagenesis Reveals the Impact of the R390Q Substitution on NGLY1 Structure
4. Discussion
4.1. Urine Glycoasparagines in the Diagnosis of AGU and NGLY1 Deficiency
4.2. NGLY1 Activity Measurement
4.3. New NGLY1-Variants
4.4. Structural Insights into the Molecular Consequences of the Arg390Gln Variant
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Huang, C.; Harada, Y.; Hosomi, A.; Masahara-Negishi, Y.; Seino, J.; Fujihira, H.; Funakoshi, Y.; Suzuki, T.; Dohmae, N.; Suzuki, T. Endo-β-N-acetylglucosaminidase forms N-GlcNAc protein aggregates during ER-associated degradation in Ngly1-defective cells. Proceedings of the National Academy of Sciences of the United States of America 2015, 112, 1398–1403. [Google Scholar] [CrossRef]
- Suzuki, T.; Huang, C.; Fujihira, H. The cytoplasmic peptide:N-glycanase (NGLY1) - Structure, expression and cellular functions. Gene 2016, 577, 1–7. [Google Scholar] [CrossRef]
- Pandey, A.; Adams, J.M.; Han, S.Y.; Jafar-Nejad, H. NGLY1 Deficiency, a Congenital Disorder of Deglycosylation: From Disease Gene Function to Pathophysiology. Cells 2022, 11. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.; Zhou, X.; Wang, L.; Li, G.; Kisker, C.; Lennarz, W.J.; Schindelin, H. Structure of the mouse peptide N-glycanase-HR23 complex suggests co-evolution of the endoplasmic reticulum-associated degradation and DNA repair pathways. The Journal of biological chemistry 2006, 281, 13751–13761. [Google Scholar] [CrossRef]
- Miao, X.; Wu, J.; Chen, H.; Lu, G. Comprehensive Analysis of the Structure and Function of Peptide:N-Glycanase 1 and Relationship with Congenital Disorder of Deglycosylation. Nutrients 2022, 14. [Google Scholar] [CrossRef]
- Lehrbach, N.J.; Ruvkun, G. Proteasome dysfunction triggers activation of SKN-1A/Nrf1 by the aspartic protease DDI-1. eLife 2016, 5. [Google Scholar] [CrossRef]
- Tachida, Y.; Hirayama, H.; Suzuki, T. Amino acid editing of NFE2L1 by PNGase causes abnormal mobility on SDS-PAGE. Biochimica et biophysica acta. General subjects 2023, 1867, 130494. [Google Scholar] [CrossRef] [PubMed]
- Tomlin, F.M.; Gerling-Driessen, U.I.M.; Liu, Y.-C.; Flynn, R.A.; Vangala, J.R.; Lentz, C.S.; Clauder-Muenster, S.; Jakob, P.; Mueller, W.F.; Ordoñez-Rueda, D.; Paulsen, M.; Matsui, N.; Foley, D.; Rafalko, A.; Suzuki, T.; Bogyo, M.; Steinmetz, L.M.; Radhakrishnan, S.K.; Bertozzi, C.R. Inhibition of NGLY1 Inactivates the Transcription Factor Nrf1 and Potentiates Proteasome Inhibitor Cytotoxicity. ACS central science 2017, 3, 1143–1155. [Google Scholar] [CrossRef] [PubMed]
- Kong, J.; Peng, M.; Ostrovsky, J.; Kwon, Y.J.; Oretsky, O.; McCormick, E.M.; He, M.; Argon, Y.; Falk, M.J. Mitochondrial function requires NGLY1. Mitochondrion 2018, 38, 6–16. [Google Scholar] [CrossRef]
- Yang, K.; Huang, R.; Fujihira, H.; Suzuki, T.; Yan, N. N-glycanase NGLY1 regulates mitochondrial homeostasis and inflammation through NRF1. The Journal of experimental medicine 2018, 215, 2600–2616. [Google Scholar] [CrossRef]
- Need, A.C.; Shashi, V.; Hitomi, Y.; Schoch, K.; Shianna, K.V.; McDonald, M.T.; Meisler, M.H.; Goldstein, D.B. Clinical application of exome sequencing in undiagnosed genetic conditions. Journal of medical genetics 2012, 49, 353–361. [Google Scholar] [CrossRef] [PubMed]
- Ge, H.; Wu, Q.; Lu, H.; Huang, Y.; Zhou, T.; Tan, D. ; ZhongqinJin Two novel compound heterozygous mutations in NGLY1as a cause of congenital disorder of deglycosylation: a case presentation. BMC medical genetics 2020, 21, 135. [Google Scholar] [CrossRef] [PubMed]
- Lam, C.; Ferreira, C.; Krasnewich, D.; Toro, C.; Latham, L.; Zein, W.M.; Lehky, T.; Brewer, C.; Baker, E.H.; Thurm, A.; Farmer, C.A.; Rosenzweig, S.D.; Lyons, J.J.; Schreiber, J.M.; Gropman, A.; Lingala, S.; Ghany, M.G.; Solomon, B.; Macnamara, E.; Davids, M.; Stratakis, C.A.; Kimonis, V.; Gahl, W.A.; Wolfe, L. Prospective phenotyping of NGLY1-CDDG, the first congenital disorder of deglycosylation. Genetics in medicine : official journal of the American College of Medical Genetics 2017, 19, 160–168. [Google Scholar] [CrossRef] [PubMed]
- Levy, R.J.; Frater, C.H.; Gallentine, W.B.; Phillips, J.M.; Ruzhnikov, M.R. Delineating the epilepsy phenotype of NGLY1 deficiency. Journal of inherited metabolic disease 2022, 45, 571–583. [Google Scholar] [CrossRef]
- Enns, G.M.; Shashi, V.; Bainbridge, M.; Gambello, M.J.; Zahir, F.R.; Bast, T.; Crimian, R.; Schoch, K.; Platt, J.; Cox, R.; Bernstein, J.A.; Scavina, M.; Walter, R.S.; Bibb, A.; Jones, M.; Hegde, M.; Graham, B.H.; Need, A.C.; Oviedo, A.; Schaaf, C.P.; Boyle, S.; Butte, A.J.; Chen, R.; Chen, R.; Clark, M.J.; Haraksingh, R.; Consortium, F.C.; Cowan, T.M.; He, P.; Langlois, S.; Zoghbi, H.Y.; Snyder, M.; Gibbs, R.A.; Freeze, H.H.; Goldstein, D.B. Mutations in NGLY1 cause an inherited disorder of the endoplasmic reticulum-associated degradation pathway. Genet Med 2014, 16, 751–758. [Google Scholar] [CrossRef]
- Haijes, H.A.; de Sain-van der Velden, M.G.M.; Prinsen, H.; Willems, A.P.; van der Ham, M.; Gerrits, J.; Couse, M.H.; Friedman, J.M.; van Karnebeek, C.D.M.; Selby, K.A.; van Hasselt, P.M.; Verhoeven-Duif, N.M.; Jans, J.J.M. Aspartylglycosamine is a biomarker for NGLY1-CDDG, a congenital disorder of deglycosylation. Mol Genet Metab 2019, 127, 368–372. [Google Scholar] [CrossRef]
- Stanclift, C.R.; Dwight, S.S.; Lee, K.; Eijkenboom, Q.L.; Wilsey, M.; Wilsey, K.; Kobayashi, E.S.; Tong, S.; Bainbridge, M.N. NGLY1 deficiency: estimated incidence, clinical features, and genotypic spectrum from the NGLY1 Registry. Orphanet journal of rare diseases 2022, 17, 440. [Google Scholar] [CrossRef]
- ClinVar database. Available online: https://www.ncbi.nlm.nih.gov/clinvar/ (accessed on 22.05.2025).
- GnomAD database. Available online: https://gnomad.broadinstitute.org/gene/ENSG00000151092?dataset=gnomad_r4 (accessed on 22.05.2025).
- Genome Analysis Toolkit. Available online: https://gatk.broadinstitute.org (accessed on 10.05.2024).
- Plagnol, V.; Curtis, J.; Epstein, M.; Mok, K.Y.; Stebbings, E.; Grigoriadou, S.; Wood, N.W.; Hambleton, S.; Burns, S.O.; Thrasher, A.J.; Kumararatne, D.; Doffinger, R.; Nejentsev, S. A robust model for read count data in exome sequencing experiments and implications for copy number variant calling. Bioinformatics (Oxford, England) 2012, 28, 2747–2754. [Google Scholar] [CrossRef]
- Banning, A.; Deubel, S.; Kluth, D.; Zhou, Z.; Brigelius-Flohé, R. The GI-GPx gene is a target for Nrf2. Molecular and cellular biology 2005, 25, 4914–4923. [Google Scholar] [CrossRef]
- Mueller, W.F.; Jakob, P.; Sun, H.; Clauder-Münster, S.; Ghidelli-Disse, S.; Ordonez, D.; Boesche, M.; Bantscheff, M.; Collier, P.; Haase, B.; Benes, V.; Paulsen, M.; Sehr, P.; Lewis, J.; Drewes, G.; Steinmetz, L.M. Loss of N-Glycanase 1 Alters Transcriptional and Translational Regulation in K562 Cell Lines. G3 (Bethesda, Md.) 2020, 10, 1585–1597. [Google Scholar] [CrossRef]
- Banning, A.; Zakrzewicz, A.; Chen, X.; Gray, S.J.; Tikkanen, R. Knockout of the CMP-Sialic Acid Transporter SLC35A1 in Human Cell Lines Increases Transduction Efficiency of Adeno-Associated Virus 9: Implications for Gene Therapy Potency Assays. Cells 2021, 10. [Google Scholar] [CrossRef] [PubMed]
- Banning, A.; Gulec, C.; Rouvinen, J.; Gray, S.J.; Tikkanen, R. Identification of Small Molecule Compounds for Pharmacological Chaperone Therapy of Aspartylglucosaminuria. Sci Rep 2016, 6, 37583. [Google Scholar] [CrossRef]
- Banning, A.; Laine, M.; Tikkanen, R. Validation of Aspartylglucosaminidase Activity Assay for Human Serum Samples: Establishment of a Biomarker for Diagnostics and Clinical Studies. International journal of molecular sciences 2023, 24. [Google Scholar] [CrossRef] [PubMed]
- Dynamut Tool. Available online: https://biosig.unimelb.edu.au/dynamut (accessed on 15.10 2024).
- Laimer, J.; Hofer, H.; Fritz, M.; Wegenkittl, S.; Lackner, P. MAESTRO--multi agent stability prediction upon point mutations. BMC bioinformatics 2015, 16, 116. [Google Scholar] [CrossRef] [PubMed]
- ChimeraX tool. Available online: https://rbvi.ucsf.edu/chimerax (accessed on 15.10.2024).
- Serial Cloner (Version 2.6.1). Available online: http://www.serialbasics.free.fr/Serial_Cloner.html (accessed on 25.05.2024).
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A.J.; Bambrick, J.; Bodenstein, S.W.; Evans, D.A.; Hung, C.-C.; O'Neill, M.; Reiman, D.; Tunyasuvunakool, K.; Wu, Z.; Žemgulytė, A.; Arvaniti, E.; Beattie, C.; Bertolli, O.; Bridgland, A.; Cherepanov, A.; Congreve, M.; Cowen-Rivers, A.I.; Cowie, A.; Figurnov, M.; Fuchs, F.B.; Gladman, H.; Jain, R.; Khan, Y.A.; Low, C.M.R.; Perlin, K.; Potapenko, A.; Savy, P.; Singh, S.; Stecula, A.; Thillaisundaram, A.; Tong, C.; Yakneen, S.; Zhong, E.D.; Zielinski, M.; Žídek, A.; Bapst, V.; Kohli, P.; Jaderberg, M.; Hassabis, D.; Jumper, J.M. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024, 630, 493–500. [Google Scholar] [CrossRef]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; Voelkerding, K.; Rehm, H.L. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genetics in medicine : official journal of the American College of Medical Genetics 2015, 17, 405–424. [Google Scholar] [CrossRef]
- NGLY1 mRNA isoform 1. Available online: https://www.ncbi.nlm.nih.gov/gene/55768 (accessed on 10.05.2023).
- Hamazaki, J.; Murata, S. ER-Resident Transcription Factor Nrf1 Regulates Proteasome Expression and Beyond. International journal of molecular sciences 2020, 21. [Google Scholar] [CrossRef]
- Radhakrishnan, S.K.; Lee, C.S.; Young, P.; Beskow, A.; Chan, J.Y.; Deshaies, R.J. Transcription factor Nrf1 mediates the proteasome recovery pathway after proteasome inhibition in mammalian cells. Molecular cell 2010, 38, 17–28. [Google Scholar] [CrossRef]
- Grotzke, J.E.; Lu, Q.; Cresswell, P. Deglycosylation-dependent fluorescent proteins provide unique tools for the study of ER-associated degradation. Proceedings of the National Academy of Sciences of the United States of America 2013, 110, 3393–3398. [Google Scholar] [CrossRef]
- He, P.; Grotzke, J.E.; Ng, B.G.; Gunel, M.; Jafar-Nejad, H.; Cresswell, P.; Enns, G.M.; Freeze, H.H. A congenital disorder of deglycosylation: Biochemical characterization of N-glycanase 1 deficiency in patient fibroblasts. Glycobiology 2015, 25, 836–844. [Google Scholar] [CrossRef]
- Rodrigues, C.H.; Pires, D.E.; Ascher, D.B. DynaMut: predicting the impact of mutations on protein conformation, flexibility and stability. Nucleic acids research 2018, 46, W350–W355. [Google Scholar] [CrossRef]
- Mak, J.; Cowan, T.M. Detecting lysosomal storage disorders by glycomic profiling using liquid chromatography mass spectrometry. Molecular genetics and metabolism 2021, 134, 43–52. [Google Scholar] [CrossRef]
- Dabaj, I.; Sudrié-Arnaud, B.; Lecoquierre, F.; Raymond, K.; Ducatez, F.; Guerrot, A.-M.; Snanoudj, S.; Coutant, S.; Saugier-Veber, P.; Marret, S.; Nicolas, G.; Tebani, A.; Bekri, S. NGLY1 Deficiency: A Rare Newly Described Condition with a Typical Presentation. Life (Basel, Switzerland) 2021, 11. [Google Scholar] [CrossRef]
- Hall, P.L.; Lam, C.; Alexander, J.J.; Asif, G.; Berry, G.T.; Ferreira, C.; Freeze, H.H.; Gahl, W.A.; Nickander, K.K.; Sharer, J.D.; Watson, C.M.; Wolfe, L.; Raymond, K.M. Urine oligosaccharide screening by MALDI-TOF for the identification of NGLY1 deficiency. Molecular genetics and metabolism 2018, 124, 82–86. [Google Scholar] [CrossRef] [PubMed]
- Hagemeijer, M.C.; van den Bosch, J.C.; Bongaerts, M.; Jacobs, E.H.; van den Hout, H.; Oussoren, E.; Ruijter, G.J.G. Analysis of urinary oligosaccharide excretion patterns by UHPLC/HRAM mass spectrometry for screening of lysosomal storage disorders. J Inherit Metab Dis 2023. [Google Scholar] [CrossRef]
- Hirayama, H.; Tachida, Y.; Fujinawa, R.; Matsuda, Y.; Murase, T.; Nishiuchi, Y.; Suzuki, T. Development of a fluorescence and quencher-based FRET assay for detection of endogenous peptide:N-glycanase/NGLY1 activity. The Journal of biological chemistry 2024, 300, 107121. [Google Scholar] [CrossRef] [PubMed]
- Hirayama, H.; Tachida, Y.; Seino, J.; Suzuki, T. A method for assaying peptide: N-glycanase/N-glycanase 1 activities in crude extracts using an N-glycosylated cyclopeptide. Glycobiology 2022, 32, 110–122. [Google Scholar] [CrossRef] [PubMed]
- Fujihira, H.; Sato, K.; Nishiuchi, Y.; Murase, T.; Matsuda, Y.; Yoshida, Y.; Kamei, T.; Suzuki, T. ELISA-based highly sensitive assay system for the detection of endogenous NGLY1 activity. Biochemical and biophysical research communications 2024, 710, 149826. [Google Scholar] [CrossRef]
- Grønbæk-Thygesen, M.; Kampmeyer, C.; Hofmann, K.; Hartmann-Petersen, R. The moonlighting of RAD23 in DNA repair and protein degradation. Biochimica et biophysica acta. Gene regulatory mechanisms 2023, 1866, 194925. [Google Scholar] [CrossRef]
- Bi, Y.; Might, M.; Vankayalapati, H.; Kuberan, B. Repurposing of Proton Pump Inhibitors as first identified small molecule inhibitors of endo-β-N-acetylglucosaminidase (ENGase) for the treatment of NGLY1 deficiency, a rare genetic disease. Bioorganic & medicinal chemistry letters 2017, 27, 2962–2966. [Google Scholar] [CrossRef]
- Fujihira, H.; Masahara-Negishi, Y.; Tamura, M.; Huang, C.; Harada, Y.; Wakana, S.; Takakura, D.; Kawasaki, N.; Taniguchi, N.; Kondoh, G.; Yamashita, T.; Funakoshi, Y.; Suzuki, T. Lethality of mice bearing a knockout of the Ngly1-gene is partially rescued by the additional deletion of the Engase gene. PLoS genetics 2017, 13, e1006696. [Google Scholar] [CrossRef] [PubMed]
- Zolekar, A.; Lin, V.J.T.; Mishra, N.M.; Ho, Y.Y.; Hayatshahi, H.S.; Parab, A.; Sampat, R.; Liao, X.; Hoffmann, P.; Liu, J.; Emmitte, K.A.; Wang, Y.-C. Stress and interferon signalling-mediated apoptosis contributes to pleiotropic anticancer responses induced by targeting NGLY1. British journal of cancer 2018, 119, 1538–1551. [Google Scholar] [CrossRef] [PubMed]
| Primer Name | Sequence 5’–3’ |
|---|---|
| NGLY1-BamHI-UTR-fwd NGLY1-BamHI-P1-fwd NGLY1-XhoI-UTR-rev |
CTATAGGATCCGCTGGCGCTCAAGCATGG |
| CTATAGGATCCAGTGTGGGACGCGGAGAGCG | |
| CTATACTCGAGAACTGCCAACTAAGCATGCAC | |
| NGLY1-gRNA-fwd NGLY1-gRNA-rev |
CACCGGGACTGAAGAACTTCTAGAA |
| AAACTTCTAGAAGTTCTTCAGTCCC | |
| NGLY1-genomic-fwd NGLY1-genomic-rev |
AGGCTCTGACACAAATGTGGCT |
| TACAAGCCAACGCTTTCTCCTG | |
| NGLY1 c.1169G>A (p.Arg390Gln) fwd NGLY1 c.1169G>A (p.Arg390Gln) rev |
GTAGTTGATGTCACTTGGCAATATTCCTGCAAACATGAAG CTTCATGTTTGCAGGAATATTGCCAAGTGACATCAACTAC |
| NGLY1 c.1201A>T (p.R401*) fwd NGLY1 c.1201A>T (p.R401*) rev |
ACATGAAGAGGTGATTGCCTGAAGAACTAAGGTTAAAGAAG CTTCTTTAACCTTAGTTCTTCAGGCAATCACCTCTTCATGT |
| EcoRI-SS-C-venus-fwd Venus-NotI-stop rev |
CTATAGAATTCCCATGGTACCGTGCACGC CTATAGCGGCCGCTTACTTGTACAGCTCGTCCATG |
| mCherry EcoRI fwd NotI-stop-mCherry-rev |
CTATAGAATTCATGGTGAGCAAGGGCGAGGAG CTATAGCGGCCGCTTACTTATAAAGCTCGTCCATGCCG |
| NFE2L1/Nrf1 HindIII fwd NFE2L1/Nrf1 EcoRI rev |
CTATAAAGCTTATGCTTTCTCTGAAGAAATAC CTATAGAATTCTCACTTTCTCCGGTCCTTTG |
| PSMB1 fwd PSMB1 rev |
CCTGCTTGACAACCAGGTTGGT TATGCAGATCCGGAGTGCGTCC |
| PSMC2 fwd PSMC2 rev |
TTGCCCGATCTAGAGGGTCGGA CATACCAGCCTCTGTGCAGACG |
| B2M fwd B2M rev |
AGATGAGTATGCCTGCCGTGTG TGCGGCATCTTCAAACCTCCA |
| GAPDH fwd GAPDH rev |
CATCTTCCAGGAGCGAGATCCC CCAGCCTTCTCCATGGTGGT |
| RPL13a fwd RPL13a rev |
CCTGGAGGAGAAGAGGAAAGAGA TTGAGGACCTCTGTGTATTTGTCAA |
| YWHAZ fwd YWHAZ rev |
AGGTTGCCGCTGGTGATGAC GGCCAGACCCAGTCTGATAGGA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




