Submitted:
09 June 2025
Posted:
10 June 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Patients and Specimens
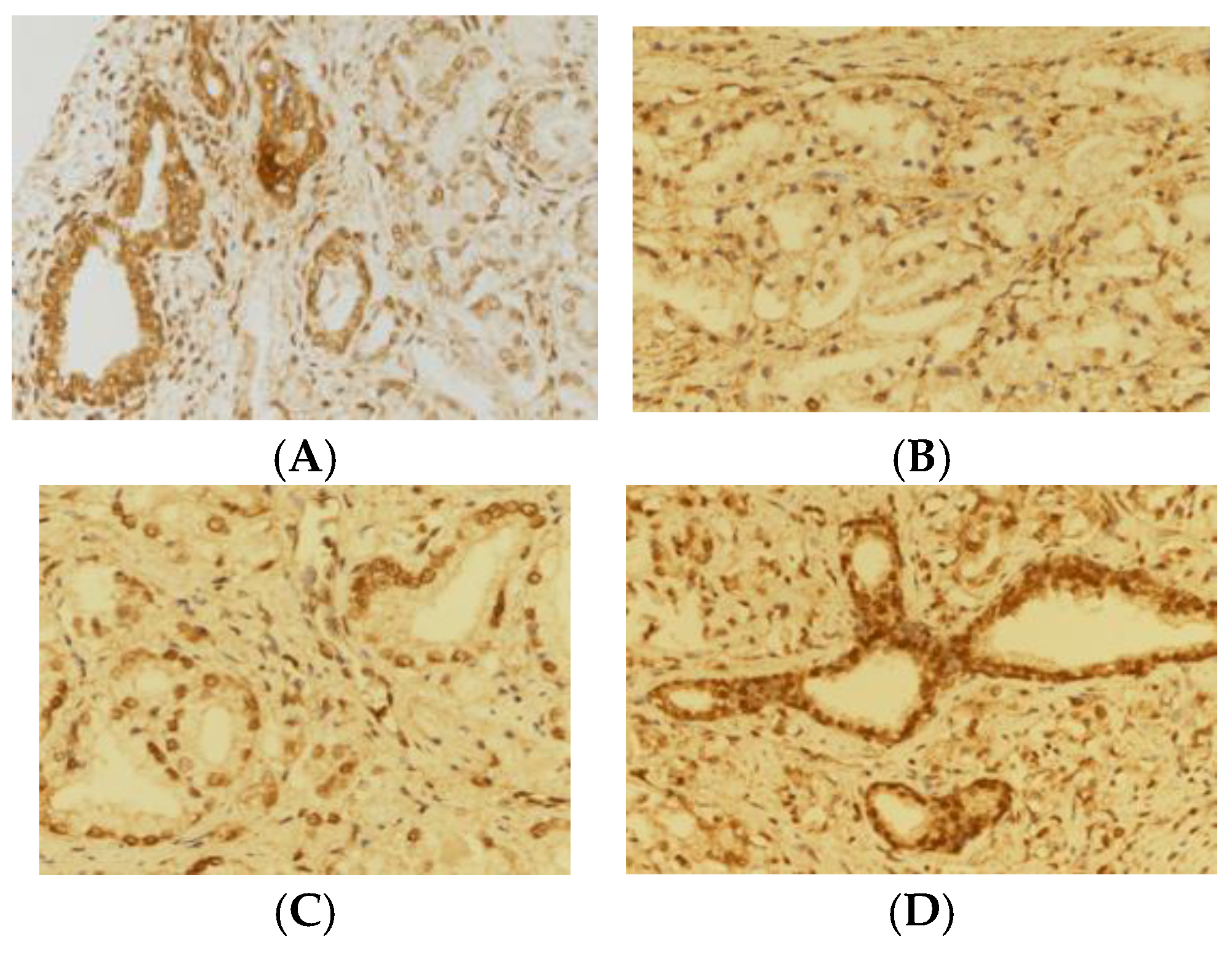
2.2. Immunohistochemical Staining of Tissue Microarrays
2.3. Survival Analysis
2.4. Statistical Analysis
3. Results
3.1. Clinicopathological Characteristics of Patients with Prostatic Cancer and USP13 Expression
3.2. Cox Proportional Hazard Model
Biochemical Recurrence
3.3. Overall Survival
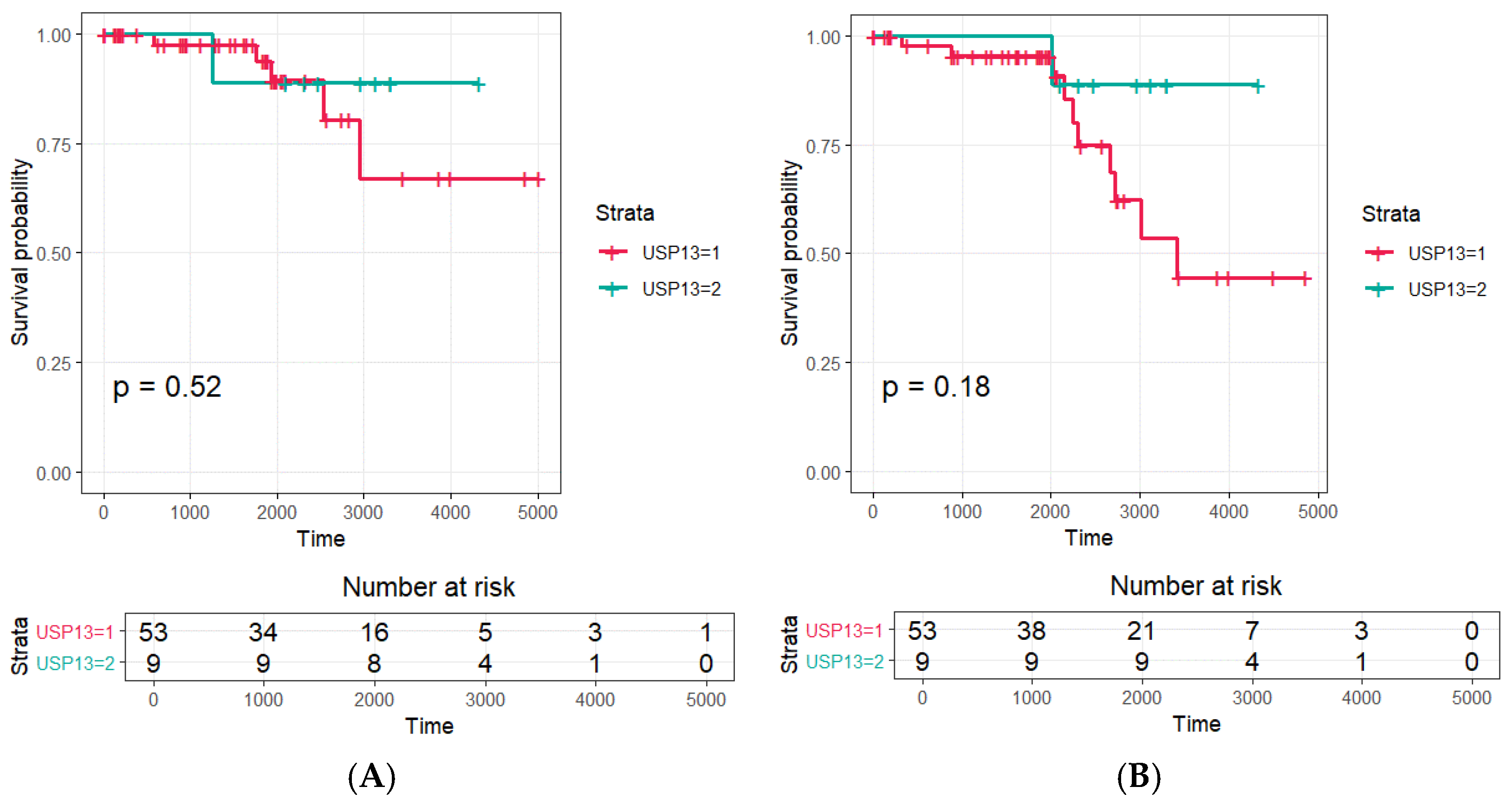
3.4. Kaplan-Meier Survival Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Consent to participate
Ethics Approval
Conflicts of Interest
References
- Salmena, L.; Carracedo, A.; Pandolfi, P.P. Tenets of PTEN tumor suppression. Cell 2008, 133, 403–414. [Google Scholar] [CrossRef] [PubMed]
- Maehama, T.; Dixon, J.E. The tumor suppressor, PTEN/MMAC1, dephosphorylates the lipid second messenger, phosphatidylinositol 3,4,5-trisphosphate. J Biol Chem 1998, 273, 13375–13378. [Google Scholar] [CrossRef]
- Stambolic, V.; Suzuki, A.; de la Pompa, J.L.; Brothers, G.M.; Mirtsos, C.; Sasaki, T.; Ruland, J.; Penninger, J.M.; Siderovski, D.P.; Mak, T.W. Negative regulation of PKB/Akt-dependent cell survival by the tumor suppressor PTEN. Cell 1998, 95, 29–39. [Google Scholar] [CrossRef]
- Song, M.S.; Salmena, L.; Pandolfi, P.P. The functions and regulation of the PTEN tumour suppressor. Nat Rev Mol Cell Biol 2012, 13, 283–296. [Google Scholar] [CrossRef]
- Hatano, K.; Nonomura, N. Systemic Therapies for Metastatic Castration-Resistant Prostate Cancer: An Updated Review. World J Mens Health 2023, 41, 769–784. [Google Scholar] [CrossRef]
- Leevers, S.J.; Vanhaesebroeck, B.; Waterfield, M.D. Signalling through phosphoinositide 3-kinases: the lipids take centre stage. Curr Opin Cell Biol 1999, 11, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Di Cristofano, A.; Pesce, B.; Cordon-Cardo, C.; Pandolfi, P.P. Pten is essential for embryonic development and tumour suppression. Nat Genet 1998, 19, 348–355. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Cao, I.; Song, M.S.; Hobbs, R.M.; Laurent, G.; Giorgi, C.; de Boer, V.C.; Anastasiou, D.; Ito, K.; Sasaki, A.T.; Rameh, L.; et al. Systemic elevation of PTEN induces a tumor-suppressive metabolic state. Cell 2012, 149, 49–62. [Google Scholar] [CrossRef]
- Liaw, D.; Marsh, D.J.; Li, J.; Dahia, P.L.; Wang, S.I.; Zheng, Z.; Bose, S.; Call, K.M.; Tsou, H.C.; Peacocke, M.; et al. Germline mutations of the PTEN gene in Cowden disease, an inherited breast and thyroid cancer syndrome. Nat Genet 1997, 16, 64–67. [Google Scholar] [CrossRef]
- Li, J.; Yen, C.; Liaw, D.; Podsypanina, K.; Bose, S.; Wang, S.I.; Puc, J.; Miliaresis, C.; Rodgers, L.; McCombie, R.; et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 1997, 275, 1943–1947. [Google Scholar] [CrossRef]
- Steck, P.A.; Pershouse, M.A.; Jasser, S.A.; Yung, W.K.; Lin, H.; Ligon, A.H.; Langford, L.A.; Baumgard, M.L.; Hattier, T.; Davis, T.; et al. Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced cancers. Nat Genet 1997, 15, 356–362. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Jiang, X. Post-translational regulation of PTEN. Oncogene 2008, 27, 5454–5463. [Google Scholar] [CrossRef] [PubMed]
- Fata, J.E.; Debnath, S.; Jenkins, E.C., Jr.; Fournier, M.V. Nongenomic Mechanisms of PTEN Regulation. Int J Cell Biol 2012, 2012, 379685. [Google Scholar] [CrossRef]
- Song, M.S.; Salmena, L.; Carracedo, A.; Egia, A.; Lo-Coco, F.; Teruya-Feldstein, J.; Pandolfi, P.P. The deubiquitinylation and localization of PTEN are regulated by a HAUSP-PML network. Nature 2008, 455, 813–817. [Google Scholar] [CrossRef]
- Ke, J.Y.; Dai, C.J.; Wu, W.L.; Gao, J.H.; Xia, A.J.; Liu, G.P.; Lv, K.S.; Wu, C.L. USP11 regulates p53 stability by deubiquitinating p53. J Zhejiang Univ Sci B 2014, 15, 1032–1038. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Tan, X.; Shi, Y.; Xu, G.; Mao, R.; Gu, X.; Fan, Y.; Yu, Y.; Burlingame, S.; Zhang, H.; et al. USP11 negatively regulates TNFalpha-induced NF-kappaB activation by targeting on IkappaBalpha. Cell Signal 2010, 22, 386–394. [Google Scholar] [CrossRef]
- Wu, H.C.; Lin, Y.C.; Liu, C.H.; Chung, H.C.; Wang, Y.T.; Lin, Y.W.; Ma, H.I.; Tu, P.H.; Lawler, S.E.; Chen, R.H. USP11 regulates PML stability to control Notch-induced malignancy in brain tumours. Nat Commun 2014, 5, 3214. [Google Scholar] [CrossRef]
- Wang, X.; Trotman, L.C.; Koppie, T.; Alimonti, A.; Chen, Z.; Gao, Z.; Wang, J.; Erdjument-Bromage, H.; Tempst, P.; Cordon-Cardo, C.; et al. NEDD4-1 is a proto-oncogenic ubiquitin ligase for PTEN. Cell 2007, 128, 129–139. [Google Scholar] [CrossRef]
- Maddika, S.; Kavela, S.; Rani, N.; Palicharla, V.R.; Pokorny, J.L.; Sarkaria, J.N.; Chen, J. WWP2 is an E3 ubiquitin ligase for PTEN. Nat Cell Biol 2011, 13, 728–733. [Google Scholar] [CrossRef]
- Ahmed, S.F.; Deb, S.; Paul, I.; Chatterjee, A.; Mandal, T.; Chatterjee, U.; Ghosh, M.K. The chaperone-assisted E3 ligase C terminus of Hsc70-interacting protein (CHIP) targets PTEN for proteasomal degradation. J Biol Chem 2012, 287, 15996–16006. [Google Scholar] [CrossRef]
- Van Themsche, C.; Leblanc, V.; Parent, S.; Asselin, E. X-linked inhibitor of apoptosis protein (XIAP) regulates PTEN ubiquitination, content, and compartmentalization. J Biol Chem 2009, 284, 20462–20466. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Lv, Y.; Li, H.; Gao, H.; Song, S.; Zhang, Y.; Xing, G.; Kong, X.; Wang, L.; Li, Y.; et al. Deubiquitylase OTUD3 regulates PTEN stability and suppresses tumorigenesis. Nat Cell Biol 2015, 17, 1169–1181. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhang, P.; Wei, Y.; Piao, H.L.; Wang, W.; Maddika, S.; Wang, M.; Chen, D.; Sun, Y.; Hung, M.C.; et al. Deubiquitylation and stabilization of PTEN by USP13. Nat Cell Biol 2013, 15, 1486–1494. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Paluch, B.E.; Wang, X.; Jiang, X. PTEN at a glance. J Cell Sci 2012, 125, 4687–4692. [Google Scholar] [CrossRef]
- Kanayama, M.; Nakao, K.; Horie, S.; Aiba, A. Generation of transgenic mouse line with prostate-specific expression of codon-improved Cre recombinase. Prostate International 2018, 6, 99–103. [Google Scholar] [CrossRef]
- Hatano, K.; Nonomura, N. Genomic Profiling of Prostate Cancer: An Updated Review. World J Mens Health 2022, 40, 368–379. [Google Scholar] [CrossRef]
| Total (n=264) |
Prostate cancer (n=242) |
Adjacent non-neoplastic tissue (n=22) |
p-value | |
| USP13 1 2 3 |
82 (31.06%) 136 (51.52%) 46 (17.42%) |
82 (33.88%) 132 (54.55%) 28 (11.57%) |
0 (0%) 4 (18.18%) 18 (81.82%) |
<0.001 |
| USP13_grade 1-2 3 |
218 (82.58%) 46 (17.42%) |
214 (88.43%) 28 (11.57%) |
4 (18.18%) 18 (81.82%) |
<0.001 |
| USP13_grade 1 2-3 |
82 (31.06%) 182 (68.94%) |
82 (33.88%) 160 (66.12%) |
0 (0%) 22 (100%) |
<0.001 |
| Total (n=242) |
USP13 | p-value | USP13 | p-value | |||
| 1-2 (n=214) | 3 (n=28) | 1 (n=82) | 2-3 (n=160) | ||||
| Age (years) | 65 (60, 70) | 65 (60, 70) | 66 (60, 72) | 0.392* | 66 (60,72) | 66 (62,72) | 0.982* |
| PSA (ng/mL) ≤10 10-20 >20 |
168 (69.14%) 40 (16.46%) 35 (14.4%) |
154 (71.96%) 33 (15.42%) 27 (12.62%) |
13 (46.43%) 7 (25.00%) 8 (28.57%) |
0.016** |
60 (73.17%) 14 (19.44%) 8 (9.76%) |
107 (66.88%) 26 (16.25%) 27 (16.88%) |
0.327** |
| Gleason score ≤ 6 7 8-10 |
24 (9.88%) 107 (44.03%) 112 (46.09%) |
22 (10.28%) 98 (45.79%) 94 (43.93%) |
1 (3.57%) 9 (32.14%) 18 (64.29%) |
0.136** |
8 (9.76%) 38 (46.34%) 36 (46.90%) |
15 (9.38%) 69 (43.13%) 76 (47.50%) |
0.866** |
| Pathological T stage ≤T2 ≥T3 |
48 (19.75%) 172 (80.37%) |
42 (19.63%) 172 (80.37%) |
5 (17.86%) 23 (82.14%) |
>0.99# |
13 (15.85%) 69 (84.15%) |
34 (21.25%) 126 (78.75%) |
0.391# |
| Seminal vesicle invasion Negative Positive |
197 (81.4%) 45 (18.6%) |
171 (80.28%) 42 (19.72%) |
25 (89.29%) 3 (10.71%) |
0.373# |
68 (82.93%) 14 (17.07%) |
128 (80.50%) 31 (19.50%) |
0.729# |
| Lymph node involvement Negative Positive |
241 (99.18%) 2 (0.47%) |
213 (99.53%) 1 (0.47%) |
27 (96.43%) 1 (3.57%) |
0.218** |
81 (98.78%) 1 (1.22%) |
159 (99.38%) 1 (0.62%) |
>0.99** |
| Total (n=115) |
USP13 | p-value | USP13 | p-value | |||
| 1-2 (n=98) | 3 (n=17) | 1 (n=41) | 2-3 (n=74) | ||||
| Age (years) | 69 (65, 75) | 69 (64, 76) | 69 (68, 75) | 0.695## | 71 (66, 75) | 69 (64, 75) | 0.637## |
| Height (cm) | 165.53±5.77 | 165.74±5.92 | 164.3±4.78 | 0.28## | 165.61±6.03 | 165.48±5.66 | 0.910## |
| Weight (kg) | 67.1 (60.9, 74.1) | 68.2 (61.2, 75) | 65.5 (55.8, 69.0) | 0.226* | 68.79±11.13 | 67.67±9.88 | 0.593* |
| PSA (ng/mL) ≤10 10-20 >20 |
53 (46.09%) 35 (30.43%) 27 (23.48%) |
50 (51.02%) 28 (28.57%) 20 (20.41%) |
3 (17.65%) 7 (41.18%) 7 (41.18%) |
0.026** |
23 11 7 |
30 24 20 |
0.250** |
| Gleason score ≤ 6 7 8-10 |
20 (17.39%) 64 (55.65%) 31 (26.96%) |
19 (19.39%) 56 (57.14%) 23 (23.47%) |
1 (5.88%) 8 (47.06%) 8 (47.06%) |
0.119** |
8 26 7 |
12 38 24 |
0.206** |
| Pathological T stage ≤T2 ≥T3 |
47 (40.87%) 68 (59.13%) |
42 (42.86%) 56 (57.14%) |
5 (29.41%) 12 (70.59%) |
0.439# |
13 28 |
34 40 |
0.167# |
| Seminal vesicle invasion Negative Positive |
101 (87.83%) 14 (18.6%) |
86 (57.76%) 12 (12.24%) |
15 (88.24%) 2 (11.76%) |
>0.99** |
35 6 |
66 8 |
0.563** |
| Lymph node involvement Negative Positive |
113 (98.26%) 2 (1.74%) |
97 (98.98%) 1 (1.02%) |
16 (94.12%) 1 (5.88%) |
0.275** |
40 (97.56%) 1 (2.44%) |
73 (98.65%) 1 (1.35%) |
>0.999** |
| Surgical margin Negative Positive |
64 (55.65%) 51 (44.35%) |
54 (55.10%) 44 (44.90%) |
10 (58.82%) 7 (41.18%) |
0.984# |
19 (46.34%) 22 (53.66%) |
45 (60.81%) 29 (39.19%) |
0.171# |
| Biochemical recurrence Negative Positive |
101 (87.83%) 14 (12.17%) |
86 (87.76%) 12 (12.24%) |
15 (88.24%) 2 (11.76%) |
>0.99** |
35 (85.37%) 6 (14.63%) |
66 (89.19%) 8 (10.81%) |
0.563** |
| Clinical recurrence Negative Positive |
112 (97.39%) 3 (2.61%) |
95 (96.94%) 3 (3.06%) |
17 (100%) 0 (0%) |
>0.99** |
39 (95.12%) 2 (4.88%) |
73 (98.65%) 1 (1.35%) |
0.289** |
| Mortality No Yes |
97 (84.35%) 18 (15.65%) |
84 (85.71%) 14 (14.29%) |
13 (76.47%) 4 (23.53%) |
0.303** |
36 (87.80%) 5 (12.20%) |
61 (82.43%) 13 (17.57%) |
0.594** |
| Univariable analysis | Multivariable analysis | |||||||||||
| Model 1 | Model 2 | |||||||||||
| HR | 95% CI | p-value | HR | 95% CI | p-value | HR | 95% CI | p-value | ||||
| lower | upper | lower | upper | lower | upper | |||||||
| Age (years) | 1.052 | 0.974 | 1.136 | 0.197 | ||||||||
| Height (cm) | 0.929 | 0.844 | 1.023 | 0.134 | 0.872 | 0.777 | 0.979 | 0.020 | ||||
| Weight (kg) | 1.028 | 0.984 | 1.074 | 0.223 | 1.061 | 1.009 | 1.117 | 0.021 | ||||
| USP13 1 2 3 |
1.970 1.720 1.000 |
0.396 0.341 |
9.805 8.680 |
0.408 0.512 |
1.744 2.110 1.000 |
0.336 0.400 |
9.038 11.133 |
0.508 0.379 |
||||
| USP13 grade 1-2 3 |
1.841 1.000 |
0.408 |
8.311 |
0.428 |
3.724 |
0.779 |
17.795 |
0.100 |
||||
| PSA (ng/mL) ≤10 10-20 >20 |
1.000 1.728 3.635 |
0.384 0.938 |
7.778 14.082 |
0.476 0.062 |
1.000 1.040 3.887 |
0.210 0.919 |
5.144 16.437 |
0.962 0.065 |
||||
| Gleason score ≤ 6 7 8-10 |
||||||||||||
| Pathological T stage ≤T2 ≥T3 |
1.000 11.465 |
1.495 |
87.924 |
0.019 |
1.000 10.521 |
1.335 |
82.902 |
0.026 |
1.000 9.182 |
1.172 |
71.933 |
0.035 |
| Seminal vesicle invasion Negative Positive |
1.000 2.610 |
0.807 |
8.445 |
0.109 |
1.000 3.624 |
0.960 |
13.683 |
0.058 |
||||
| Lymph node involvement Negative Positive |
1.000 2.984 |
0.384 |
23.191 |
0.298 |
||||||||
| Surgical margin Negative Positive |
1.000 5.349 |
1.480 |
19.330 |
0.011 |
||||||||
| Univariable analysis | Multivariable analysis | |||||||||||
| Model 1 | Model 2 | |||||||||||
| HR | 95% CI | p-value | HR | 95% CI | p-value | HR | 95% CI | p-value | ||||
| lower | upper | lower | upper | lower | upper | |||||||
| Age (years) | 1.064 | 0.981 | 1.154 | 0.135 | ||||||||
| Height (cm) | 0.955 | 0.885 | 1.032 | 0.246 | ||||||||
| Weight (kg) | 0.955 | 0.906 | 1.005 | 0.078 | 0.946 | 0.896 | 0.999 | 0.047 | 0.952 | 0.904 | 1.004 | 0.068 |
| USP13 1 2 3 |
1.120 1.286 1.000 |
0.297 0.392 |
4.218 4.217 |
0.868 0.679 |
1.080 1.765 1.000 |
0.287 0.515 |
4.070 6.052 |
0.910 0.366 |
||||
| USP13 grade 1-2 3 |
1.463 1.000 |
0.371 |
5.771 |
0.587 |
1.411 1.000 |
0.456 |
4.367 |
0.550 |
||||
| PSA (ng/mL) ≤10 10-20 >20 |
1.000 1.402 1.367 |
0.438 0.413 |
4.491 4.523 |
0.570 0.608 |
||||||||
| Gleason score ≤ 6 7 8-10 |
1.000 0.726 0.753 |
0.218 0.210 |
2.422 2.700 |
0.602 0.663 |
||||||||
| Pathological T stage ≤T2 ≥T3 |
1.000 1.347 |
0.521 |
3.484 |
0.539 |
||||||||
| Seminal vesicle invasion Negative Positive |
1.000 1.769 |
0.68 |
5.060 |
0.288 |
||||||||
| Lymph node involvement Negative Positive |
1.000 1.884 |
0.246 |
14.422 |
0.542 |
||||||||
| Surgical margin Negative Positive |
1.000 1.504 |
0.577 |
3.919 |
0.404 |
||||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




