Submitted:
11 April 2025
Posted:
11 April 2025
You are already at the latest version
Abstract
Keywords:
Introduction
Materials and Methods
Study Cohort
Identification of Candidate CCMs Using the EpiSwitch Platform
Prioritization of Candidate CCMs Based on Estimated Interactions with TMB and OS
Assessment of Potential CCM Gene Expression
Visium Data Processing
Results
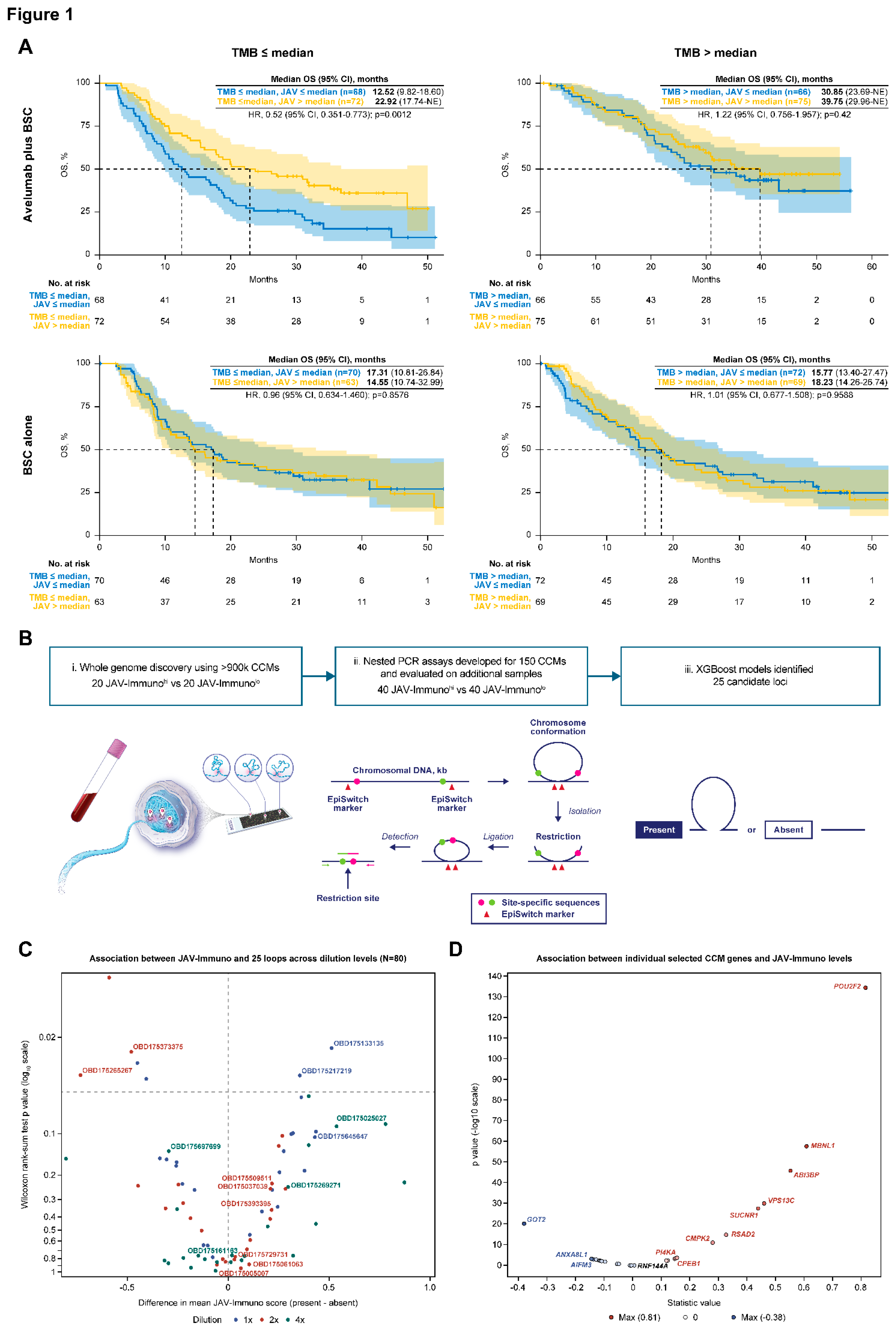
Study Cohort and Methodology
Identification of Candidate CCMs
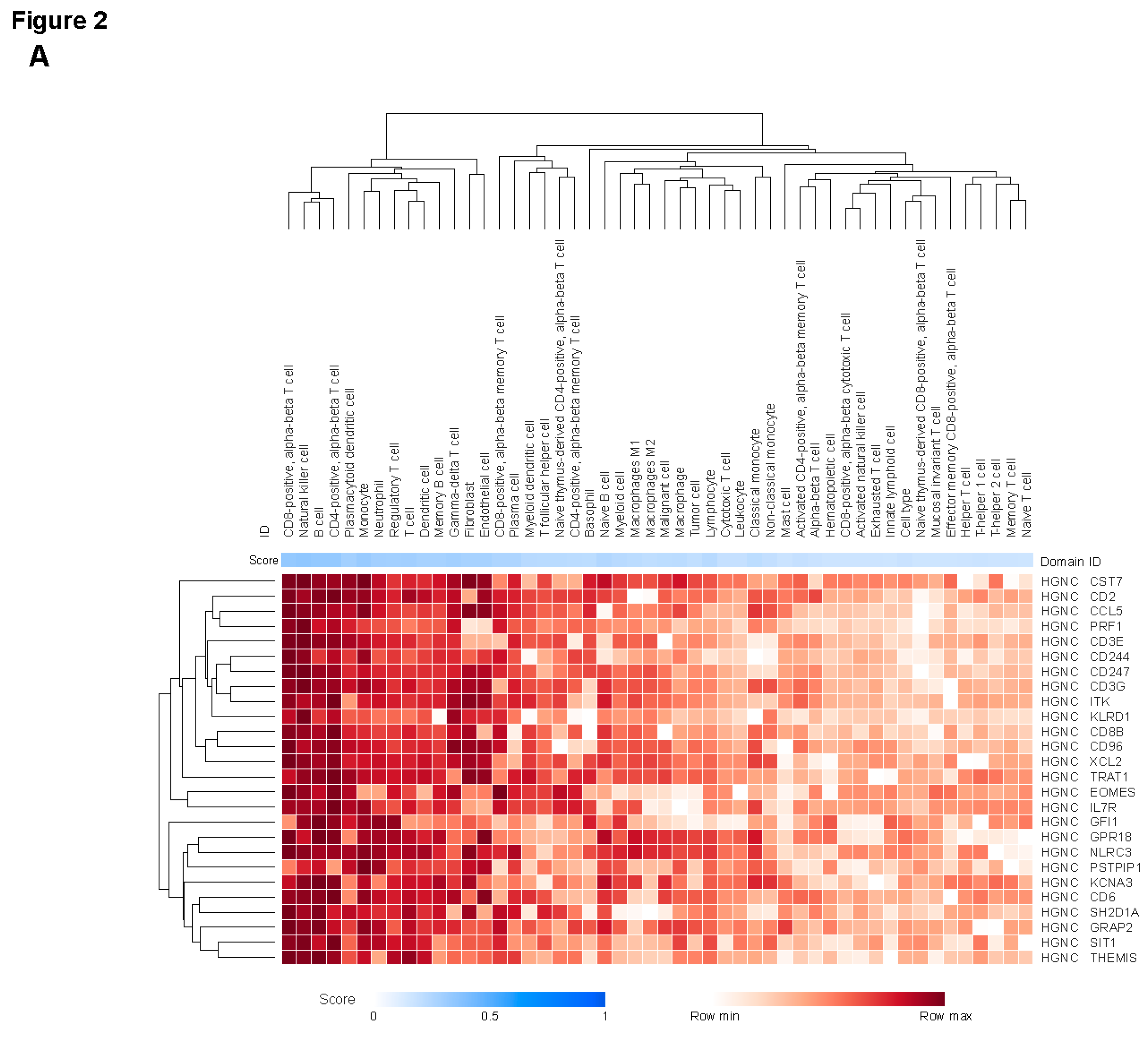
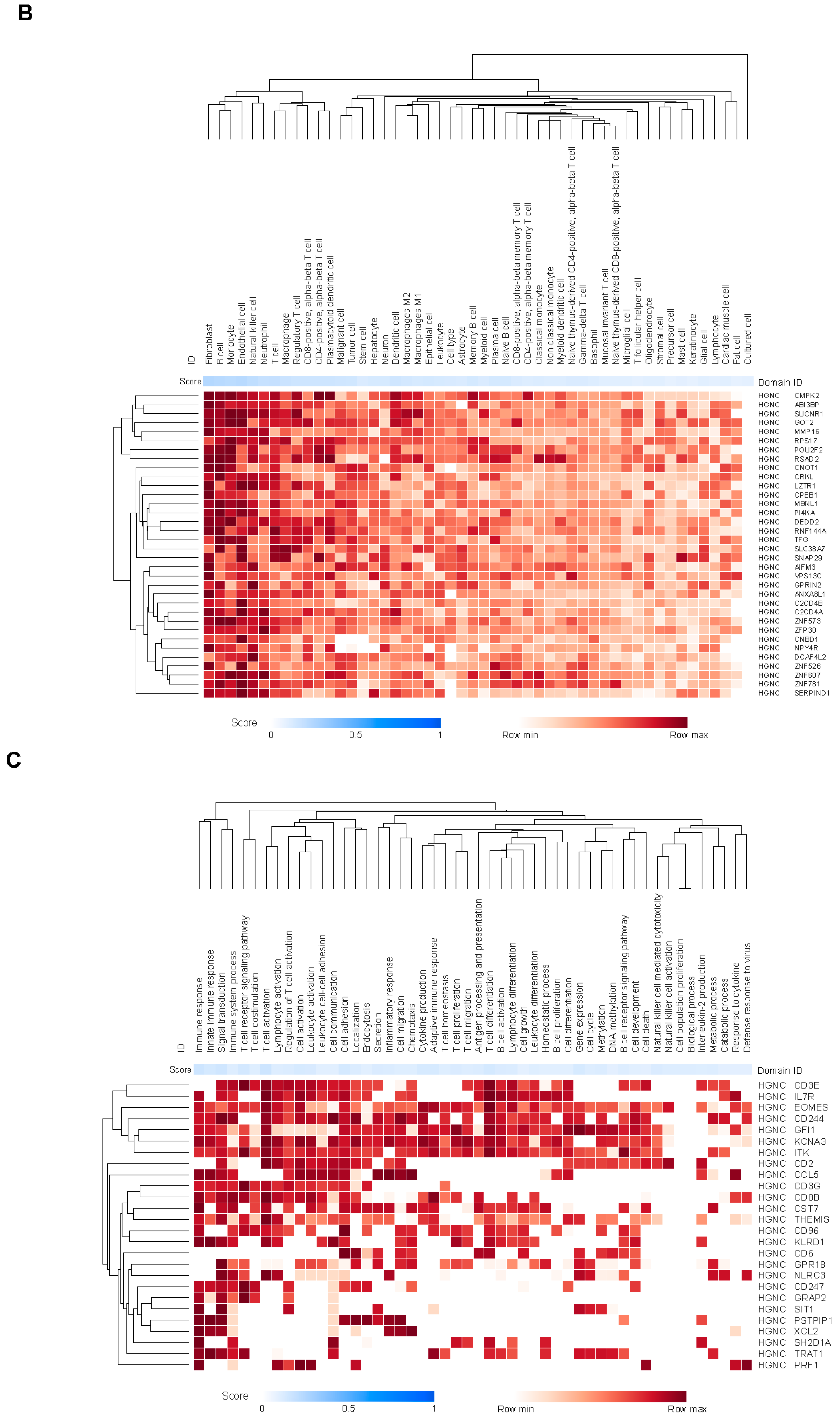
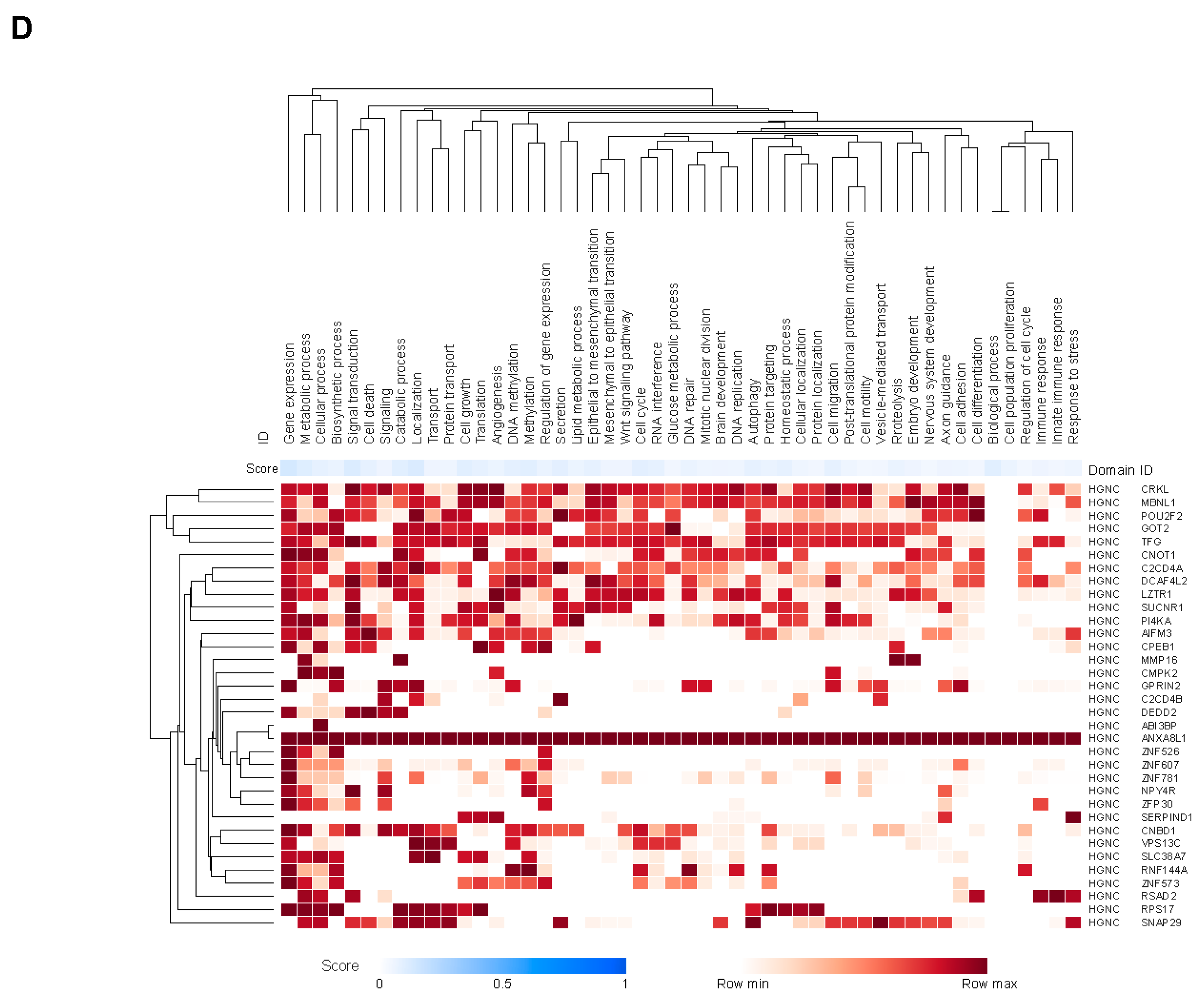
Biological Annotation of Genes Covered by the CCMs
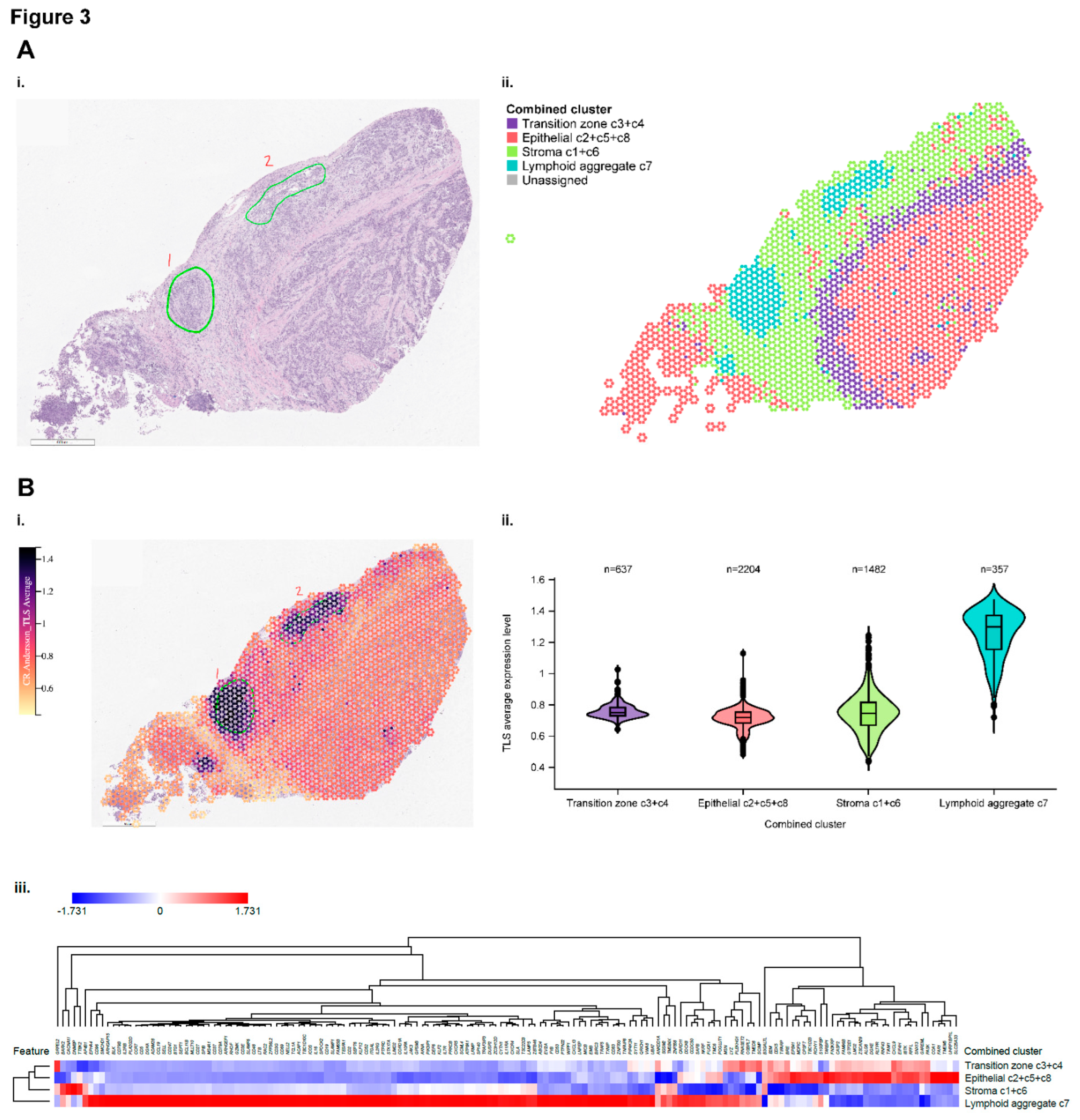
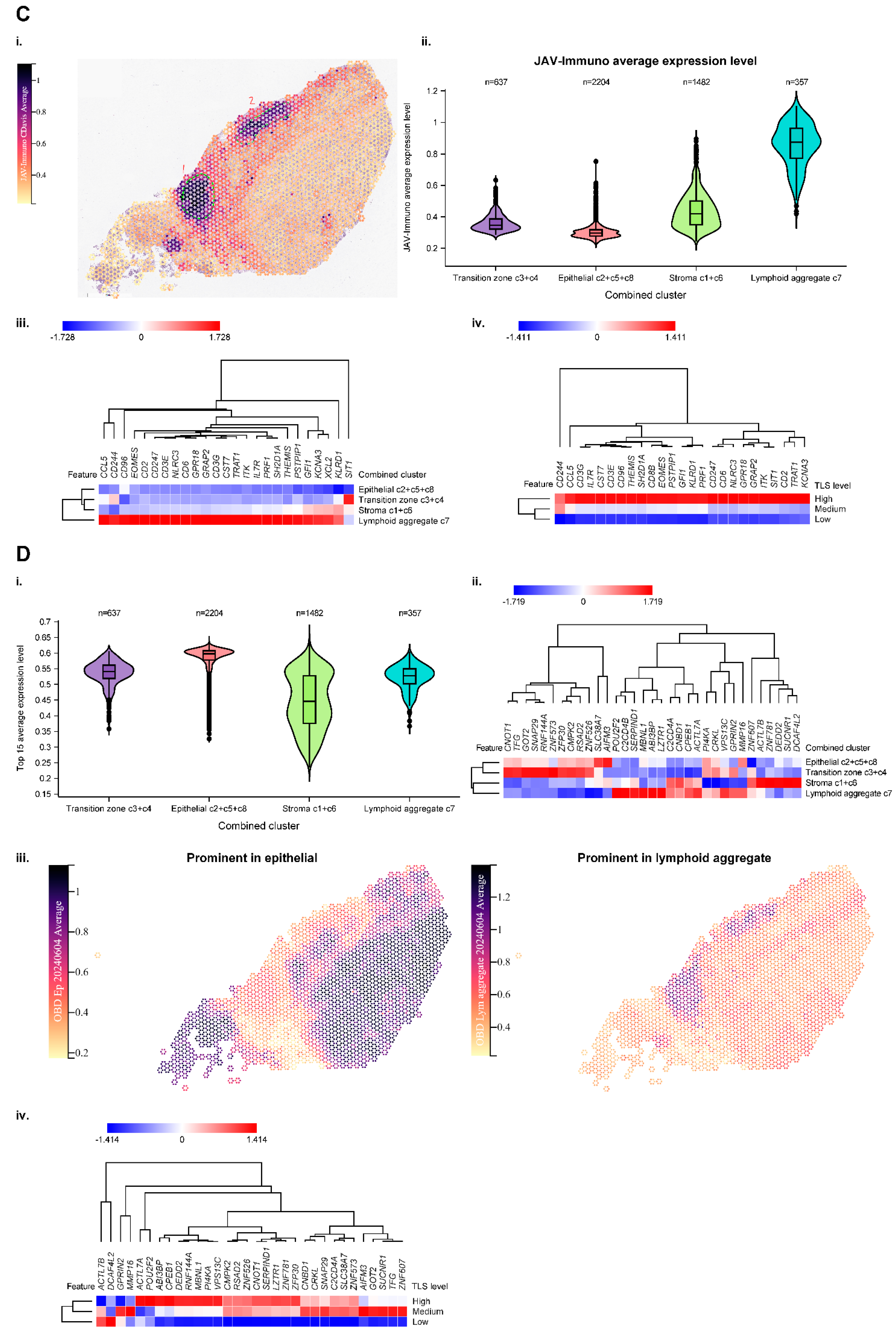
Spatial Profiling
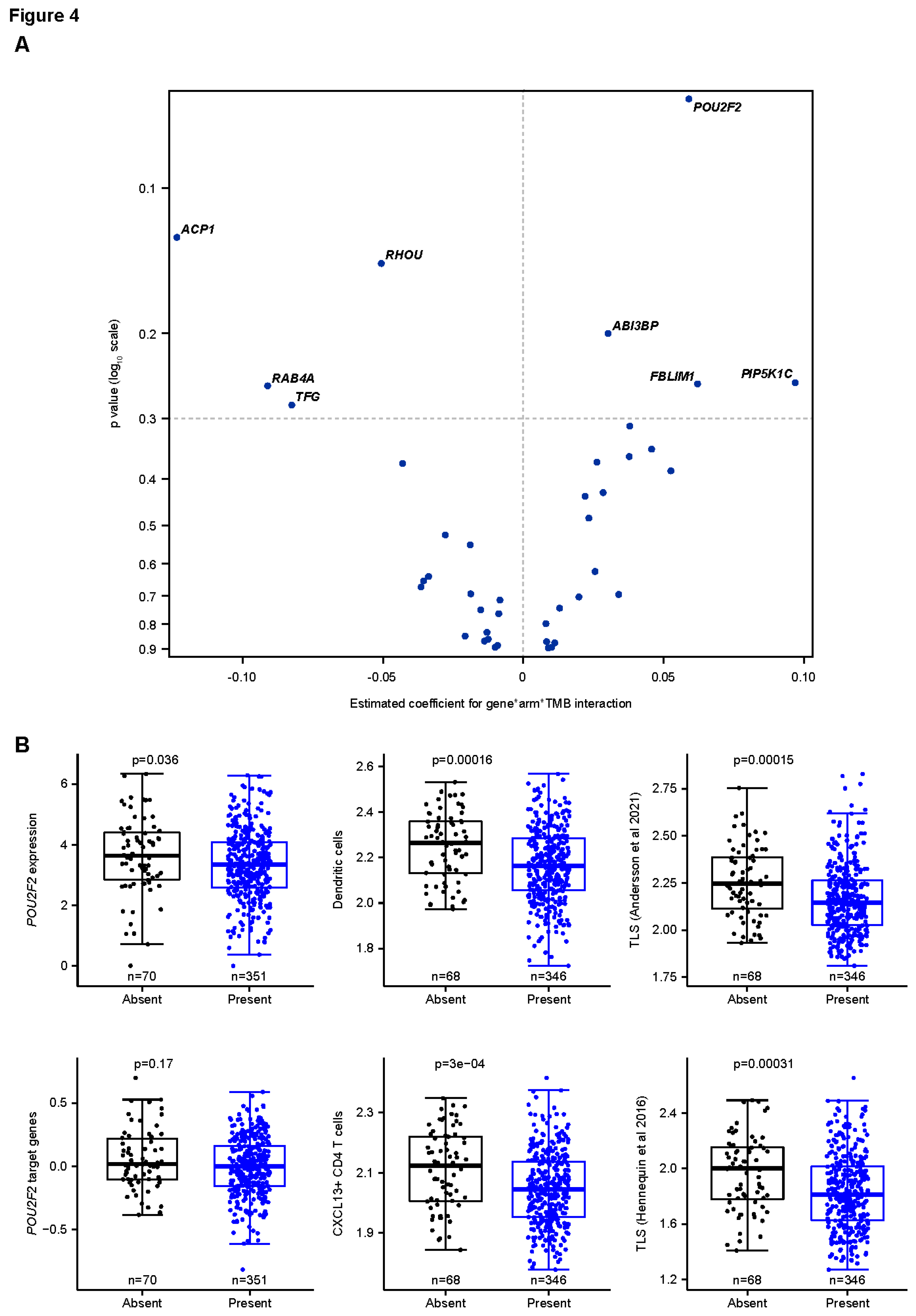
Associations Between POU2F2, TMB, and OS
Discussion
Supplementary Materials
Author Contributions
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Banchereau, R.; et al. Molecular determinants of response to PD-L1 blockade across tumor types. Nat Commun 2021, 12, 3969. [Google Scholar] [CrossRef] [PubMed]
- Powles, T.; et al. Avelumab maintenance in advanced urothelial carcinoma: biomarker analysis of the phase 3 JAVELIN Bladder 100 trial. Nat Med 2021, 27, 2200–2211. [Google Scholar] [CrossRef]
- Powles, T.; et al. Avelumab Maintenance Therapy for Advanced or Metastatic Urothelial Carcinoma. N Engl J Med 2020, 383, 1218–1230. [Google Scholar] [CrossRef] [PubMed]
- Powles, T.; et al. ESMO Clinical Practice Guideline interim update on first-line therapy in advanced urothelial carcinoma. Ann Oncol 2024, 35, 485–490. [Google Scholar] [CrossRef]
- Grivas, P.; et al. Avelumab first-line maintenance treatment for advanced urothelial carcinoma: review of evidence to guide clinical practice. ESMO Open 2023, 8, 102050. [Google Scholar] [CrossRef]
- Mellman, I.; et al. The cancer-immunity cycle: Indication, genotype, and immunotype. Immunity 2023, 56, 2188–2205. [Google Scholar] [CrossRef]
- Khan, Z.; et al. Polygenic risk for skin autoimmunity impacts immune checkpoint blockade in bladder cancer. Proc Natl Acad Sci U S A 2020, 117, 12288–12294. [Google Scholar] [CrossRef] [PubMed]
- Dizman, N.; et al. Nivolumab plus ipilimumab with or without live bacterial supplementation in metastatic renal cell carcinoma: a randomized phase 1 trial. Nat Med 2022, 28, 704–712. [Google Scholar] [CrossRef]
- Cuartero, S.; Stik, D.; Stadhouders, R. Three-dimensional genome organization in immune cell fate and function. Nat Rev Immunol 2023, 23, 206–221. [Google Scholar] [CrossRef]
- Johanson, T.M.; et al. Genome organization in immune cells: unique challenges. Nat Rev Immunol 2019, 19, 448–456. [Google Scholar] [CrossRef]
- Tordini, F.; et al. The genome conformation as an integrator of multi-omic data: the example of damage spreading in cancer. Front Genet 2016, 7, 194. [Google Scholar] [CrossRef] [PubMed]
- Bonev, B.; Cavalli, G. Organization and function of the 3D genome. Nat Rev Genet 2016, 17, 661–678. [Google Scholar] [CrossRef]
- Tien, F.-M.; et al. Epigenetic remodeling of the immune landscape in cancer: therapeutic hurdles and opportunities. J Biomed Sci 2023, 30, 3. [Google Scholar] [CrossRef] [PubMed]
- Kempfer, R.; Pombo, A. Methods for mapping 3D chromosome architecture. Nat Rev Genet 2020, 21, 207–226. [Google Scholar] [CrossRef]
- Dekker, J.; et al. Capturing chromosome conformation. Science 2002, 295, 1306–11. [Google Scholar] [CrossRef] [PubMed]
- Jakub, J.W.; et al. A pilot study of chromosomal aberrations and epigenetic changes in peripheral blood samples to identify patients with melanoma. Melanoma Res 2015, 25, 406–11. [Google Scholar] [CrossRef]
- Carini, C.; et al. Chromosome conformation signatures define predictive markers of inadequate response to methotrexate in early rheumatoid arthritis. J Transl Med 2018, 16, 18. [Google Scholar] [CrossRef]
- Alshaker, H.; et al. Chromatin conformation changes in peripheral blood can detect prostate cancer and stratify disease risk groups. J Transl Med 2021, 19, 46. [Google Scholar] [CrossRef]
- Pchejetski, D.; et al. Circulating chromosome conformation signatures significantly enhance PSA positive predicting value and overall accuracy for prostate cancer detection. Cancers 2023, 15, 821. [Google Scholar] [CrossRef]
- Hunter, E.; et al. Development and validation of blood-based predictive biomarkers for response to PD-1/PD-L1 checkpoint inhibitors: evidence of a universal systemic core of 3D immunogenetic profiling across multiple oncological indications. Cancers 2023, 15, 2696. [Google Scholar] [CrossRef]
- Quinlan, A.R. BEDTools: the Swiss-army tool for genome feature analysis. Curr Protoc Bioinformatics 2014, 47, 11.12–1. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Guestrin, C. XGBoost: a scalable tree boosting system, in Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. 2016, Association for Computing Machinery: San Francisco, California, USA. pp. 785–794.
- Biecek, P.; Burzykowski, T. Shapley Additive Explanations (SHAP) for Average Attributions; 2021; pp. 95–106. [Google Scholar]
- Salter, M.; et al. Initial identification of a blood-based chromosome conformation signature for aiding in the diagnosis of amyotrophic lateral sclerosis. EBioMedicine 2018, 33, 169–184. [Google Scholar] [CrossRef] [PubMed]
- Zou, H.; Hastie, T. Regularization and variable selection via the elastic net. J R Stat Soc Series B Stat Methodol 2005, 67, 301–320. [Google Scholar] [CrossRef]
- Tibshirani, R. Regression shrinkage and selection via the Lasso. J R Stat Soc Series B Stat Methodol 1996, 58, 267–288. [Google Scholar] [CrossRef]
- Hoerl, A.E.; Kennard, R.W. Ridge regression: applications to nonorthogonal problems. Technometrics 1970, 12, 69–82. [Google Scholar] [CrossRef]
- Tiddi, I.; Schlobach, S. Knowledge graphs as tools for explainable machine learning: A survey. Artificial Intelligence 2022, 302, 103627. [Google Scholar] [CrossRef]
- Zhao, E.; et al. Spatial transcriptomics at subspot resolution with BayesSpace. Nat Biotechnol 2021, 39, 1375–1384. [Google Scholar] [CrossRef]
- Hu-Lieskovan, S.; et al. Association of tumor mutational burden and immune gene expression with response to PD-1 blockade by sasanlimab across tumor types and routes of administration. Targeted Onco 2021, 16, 773–787. [Google Scholar] [CrossRef]
- Motzer, R.J.; et al. Avelumab plus axitinib versus sunitinib in advanced renal cell carcinoma: biomarker analysis of the phase 3 JAVELIN Renal 101 trial. Nat Med 2020, 26, 1733–1741. [Google Scholar] [CrossRef]
- Yan, H.; et al. Epigenetic chromatin conformation changes in peripheral blood can detect thyroid cancer. Surgery 2019, 165, 44–49. [Google Scholar] [CrossRef]
- Karlsson, M.; et al. A single-cell type transcriptomics map of human tissues. Sci Adv 2021, 7. [Google Scholar] [CrossRef] [PubMed]
- Tabula Sapiens Consortium, et al. The Tabula Sapiens: a multiple-organ, single-cell transcriptomic atlas of humans. Science 2022, 376, eabl4896. [CrossRef]
- Ashburner, M.; et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 2000, 25, 25–9. [Google Scholar] [CrossRef]
- Helmink, B.A.; et al. B cells and tertiary lymphoid structures promote immunotherapy response. Nature 2020, 577, 549–555. [Google Scholar] [CrossRef] [PubMed]
- Higgs, B.W.; et al. Interferon gamma messenger RNA signature in tumor biopsies predicts outcomes in patients with non-small cell lung carcinoma or urothelial cancer treated with durvalumab. Clin Cancer Res 2018, 24, 3857–3866. [Google Scholar] [CrossRef]
- Fehrenbacher, L.; et al. Atezolizumab versus docetaxel for patients with previously treated non-small-cell lung cancer (POPLAR): a multicentre, open-label, phase 2 randomised controlled trial. Lancet 2016, 387, 1837–46. [Google Scholar] [CrossRef] [PubMed]
- Eckstein, M.; et al. Cytotoxic T-cell-related gene expression signature predicts improved survival in muscle-invasive urothelial bladder cancer patients after radical cystectomy and adjuvant chemotherapy. J Immunother Cancer 2020, 8, e000162. [Google Scholar] [CrossRef]
- Ayers, M.; et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J Clin Invest 2017, 127, 2930–2940. [Google Scholar] [CrossRef]
- Wiedemann, G.M.; et al. Cutting edge: STAT1-mediated epigenetic control of Rsad2 promotes clonal expansion of antiviral NK cells. J Immunol 2020, 205, 21–25. [Google Scholar] [CrossRef]
- Arnold, N.; et al. Genomic and functional analysis of the host response to acute simian varicella infection in the lung. Sci Rep 2016, 6, 34164. [Google Scholar] [CrossRef]
- Larsson, I.; et al. Genome-scale metabolic modeling of glioblastoma reveals promising targets for drug development. Front Genet 2020, 11, 381. [Google Scholar] [CrossRef] [PubMed]
- Ho, S.R.; et al. RNF144A deficiency promotes PD-L1 protein stabilization and carcinogen-induced bladder tumorigenesis. Cancer Lett 2021, 520, 344–360. [Google Scholar] [CrossRef]
- Yang, Y.L.; et al. RNF144A functions as a tumor suppressor in breast cancer through ubiquitin ligase activity-dependent regulation of stability and oncogenic functions of HSPA2. Cell Death Differ 2020, 27, 1105–1118. [Google Scholar] [CrossRef]
- Zhang, J.; et al. Association between succinate receptor SUCNR1 expression and immune infiltrates in ovarian cancer. Front Mol Biosci 2020, 7, 150. [Google Scholar] [CrossRef] [PubMed]
- Najm, R.; Hachim, M.Y.; Kandasamy, R.K. Divulging a pleiotropic role of succinate receptor SUCNR1 in renal cell carcinoma microenvironment. Cancers (Basel) 2022, 14, 6064. [Google Scholar] [CrossRef]
- Hodson, D.J.; et al. Regulation of normal B-cell differentiation and malignant B-cell survival by OCT2. Proc Natl Acad Sci U S A 2016, 113, E2039–E2046. [Google Scholar] [CrossRef] [PubMed]
- Corcoran, L.; et al. Oct2 and Obf1 as facilitators of B:T cell collaboration during a humoral immune response. Front Immunol 2014, 5, 108. [Google Scholar] [CrossRef]
- Powles, T.; et al. Clinical efficacy and biomarker analysis of neoadjuvant atezolizumab in operable urothelial carcinoma in the ABACUS trial. Nat Med 2019, 25, 1706–1714. [Google Scholar] [CrossRef]
| Association between tumor gene expression and JAV-Immuno score | Cell type | Immune compartment | Immune process | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Gene | Gene description | Statistic value | p value | q value | Endothelial cell | Fibroblast | B cell | Monocyte | Natural killer cell | Dendritic cell | Cytotoxic T cell | Lymph node | Germinal center | MHC class I antigen presentation | MHC class II antigen presentation | Response to stress |
| POU2F2 | POU class 2 homeobox 2 | 0.8141921 | 2.75E−135 | 4.96E−134 | 0.09011 | 0.2449 | 0.2558 | 0.2563 | 0.228 | 0.2125 | 0.02845 | 0.09261 | 0.09931 | 0 | 0.04072 | 0.004806 |
| MBNL1 | Muscle blindlike splicing regulator 1 | 0.6073972 | 2.31E−58 | 2.77E−57 | 0.2185 | 0.222 | 0.2186 | 0.2325 | 0.07344 | 0.04999 | 0.02845 | 0 | 0 | 0.008677 | 0.008505 | 0.1479 |
| ABI3BP | ABI family member 3 binding protein | 0.5517624 | 2.09E−46 | 1.88E−45 | 0.212 | 0.2435 | 0.2236 | 0.1964 | 0.1889 | 0.1974 | 0.02832 | 0.07382 | 0 | 0 | 0 | 0 |
| VPS13C | Vacuolar protein sorting 13 homolog C | 0.4595461 | 6.43E−31 | 4.63E−30 | 0.07422 | 0.1806 | 0.05065 | 0.06594 | 0.06424 | 0.05733 | 0.02213 | 0 | 0 | 0 | 0 | 0 |
| SUCNR1 | Succinate receptor 1 | 0.4390713 | 4.48E−28 | 2.69E−27 | 0.2147 | 0.2118 | 0.1985 | 0.2108 | 0.2092 | 0.208 | 0.15 | 0.09239 | 0 | 0.003451 | 0.00712 | 0.0858 |
| RSAD2 | Radical S-adenosyl methionine domain containing 2 | 0.3267927 | 1.49E−15 | 6.72E−15 | 0.2266 | 0.2148 | 0.234 | 0.2137 | 0.2234 | 0.229 | 0.1652 | 0.09274 | 0.07864 | 0 | 0.06117 | 0.1613 |
| CMPK2 | Cytidine/uridine monophosphate kinase 2 | 0.2787244 | 1.47E−11 | 5.89E−11 | 0.2066 | 0.2194 | 0.2099 | 0.2243 | 0.1865 | 0.2063 | 0.1437 | 0.07898 | 0.06812 | 0 | 0.0499 | 0.08855 |
| PI4KA | Phosphatidylinositol 4-kinase alpha | 0.152742 | 0.000265 | 0.000954 | 0.2101 | 0.2087 | 0.2186 | 0.1967 | 0.197 | 0.04704 | 0.02591 | 0 | 0 | 0.01603 | 0 | 0 |
| RNF144A | Ring finger protein 144A | 0.1471635 | 0.000444 | 0.00145 | 0.09449 | 0.2008 | 0.1847 | 0.1624 | 0.06401 | 0.1495 | 0.08942 | 0.07085 | 0.01967 | 0 | 0 | 0 |
| CPEB1 | Cytoplasmic polyadenylation element binding protein 1 | 0.1458114 | 0.000502 | 0.0015 | 0.1894 | 0.2084 | 0.1893 | 0.1863 | 0.07279 | 0.05963 | 0 | 0 | 0 | 0 | 0 | 0.01358 |
| NPY4R | Neuropeptide Y receptor Y4 | 0.1201768 | 0.00419 | 0.00944 | 0.04041 | 0.1401 | 0.1273 | 0.1208 | 0.1318 | 0.03498 | 0 | 0 | 0 | 0.003384 | 0 | 0 |
| ZNF573 | Zinc finger protein 573 | 0.1156841 | 0.00586 | 0.0113 | 0.06559 | 0.1173 | 0.1437 | 0.1506 | 0.1212 | 0.09401 | 0.04908 | 0.02246 | 0 | 0.005744 | 0 | 0 |
| SNAP29 | Synaptosome associated protein 29 | 0.1154207 | 0.00598 | 0.0113 | 0.2049 | 0.1919 | 0.06076 | 0.06829 | 0.06407 | 0.04174 | 0.02845 | 0 | 0 | 0.006851 | 0.005367 | 0.1535 |
| ZNF781 | Zinc finger protein 781 | 0.1124934 | 0.00739 | 0.0121 | 0.1024 | 0.0865 | 0.1235 | 0.08621 | 0.09041 | 0.04321 | 0.005367 | 0.04858 | 0 | 0.003986 | 0.002624 | 0.000162 |
| SLC38A7 | Solute carrier family 38 member 7 | 0.0978701 | 0.0199 | 0.0286 | 0.1976 | 0.05722 | 0.192 | 0.06142 | 0.05257 | 0.03522 | 0.02277 | 0 | 0 | 0 | 0 | 0.08463 |
| DEDD2 | Death effector domain containing 2 | 0.0966533 | 0.0215 | 0.0297 | 0.08156 | 0.1777 | 0.1756 | 0.1643 | 0.1515 | 0.146 | 0.1052 | 0.06802 | 0 | 0 | 0.02047 | 0.06925 |
| C2CD4B | C2 calcium dependent domain containing 4B | 0.0826085 | 0.0495 | 0.066 | 0.2162 | 0.1775 | 0.154 | 0.1431 | 0.1367 | 0.05737 | 0.01532 | 0.0178 | 0 | 0 | 0 | 0 |
| TMEM14E | transmembrane protein 14E, pseudogene | 0.0678954 | 0.107 | 0.137 | 0.03418 | 0.0232 | 0.0338 | 0.03111 | 0.04544 | 0.02271 | 0.004312 | 0 | 0 | 0 | 0 | 0 |
| MMP16 | Matrix metallopeptidase 16 | 0.0639516 | 0.129 | 0.16 | 0.2134 | 0.2523 | 0.2056 | 0.2038 | 0.2085 | 0.06526 | 0.007903 | 0.09234 | 0 | 0 | 0 | 0 |
| LZTR1 | Leucine zipper like transcription regulator 1 | 0.0550683 | 0.191 | 0.222 | 0.2021 | 0.1965 | 0.06215 | 0.1714 | 0.2034 | 0.03179 | 0.01318 | 0.08179 | 0 | 0.007257 | 0.008901 | 0.006873 |
| RPS17 | Ribosomal protein S17 | 0.0165515 | 0.694 | 0.757 | 0.2079 | 0.183 | 0.2076 | 0.2269 | 0.2022 | 0.179 | 0.1641 | 0 | 0 | 0.03428 | 0.07347 | 0.1058 |
| CNBD1 | Cyclic nucleotide binding domain containing 1 | 0.0040582 | 0.923 | 0.923 | 0.1355 | 0.1114 | 0.1151 | 0.1164 | 0.1073 | 0.09581 | 0.01741 | 0.04782 | 0 | 0.003208 | 0.003264 | 0.00432 |
| C2CD4A | C2 calcium dependent domain containing 4A | −0.004789 | 0.909 | 0.923 | 0.171 | 0.1587 | 0.1331 | 0.1213 | 0.1156 | 0.05684 | 0.004867 | 0.04788 | 0 | 0 | 0.003982 | 0.08443 |
| DCAF4L2 | DDB1 and CUL4 associated factor 4 like 2 | −0.008794 | 0.835 | 0.884 | 0.06324 | 0.03736 | 0.06659 | 0.09815 | 0.1282 | 0.05735 | 0 | 0.06016 | 0 | 0.00528 | 0.01013 | 0.00442 |
| ZNF526 | Zinc finger protein 526 | −0.048678 | 0.248 | 0.279 | 0.1291 | 0.1437 | 0.07571 | 0.06573 | 0.04032 | 0.03047 | 0.004803 | 0 | 0 | 0 | 0 | 0 |
| CNOT1 | CCR4-NOT transcription complex subunit 1 | −0.055119 | 0.19 | 0.222 | 0.0678 | 0.21 | 0.2023 | 0.2026 | 0.03454 | 0.1721 | 0.01716 | 0 | 0 | 0.01848 | 0.003042 | 0.09112 |
| ZFP30 | ZFP30 zinc finger protein | −0.098922 | 0.0186 | 0.0279 | 0.1219 | 0.1227 | 0.1167 | 0.1308 | 0.1257 | 0.07336 | 0.02253 | 0 | 0 | 0 | 0 | 0 |
| ZNF607 | Zinc finger protein 607 | −0.110554 | 0.00848 | 0.0133 | 0.04603 | 0.0471 | 0.06989 | 0.08244 | 0.1002 | 0.03778 | 0.0391 | 0.02276 | 0 | 0.004409 | 0.003307 | 0.01889 |
| GPRIN2 | G protein regulated inducer of neurite outgrowth 2 | −0.112585 | 0.00734 | 0.0121 | 0.06067 | 0.1465 | 0.1516 | 0.03462 | 0.1517 | 0.04801 | 0.00213 | 0.05896 | 0.02013 | 0 | 0.01973 | 0.00166 |
| TFG | Trafficking from ER to golgi regulator | −0.114673 | 0.00631 | 0.0114 | 0.2094 | 0.1879 | 0.2205 | 0.2067 | 0.06115 | 0.02826 | 0.02845 | 0.08952 | 0 | 0.003158 | 0.00423 | 0.005551 |
| SERPIND1 | Serpin family D member 1 | −0.116083 | 0.00569 | 0.0113 | 0.2145 | 0.1994 | 0.189 | 0.1905 | 0.05945 | 0.1672 | 0 | 0.09033 | 0 | 0 | 0.004157 | 0.1498 |
| CRKL | CRK like proto-oncogene, adaptor protein | −0.130329 | 0.00189 | 0.00454 | 0.2009 | 0.2036 | 0.2249 | 0.1969 | 0.2029 | 0.1667 | 0.0242 | 0.09401 | 0 | 0.02339 | 0.02136 | 0.02151 |
| AIFM3 | Apoptosis inducing factor mitochondria associated 3 | −0.138936 | 0.000919 | 0.00236 | 0.19 | 0.1729 | 0.05 | 0.06931 | 0.0611 | 0.05562 | 0.006126 | 0.08887 | 0 | 0.0107 | 0.004403 | 0.1262 |
| ANXA8L1 | Annexin A8 like 1 | −0.14415 | 0.000582 | 0.00161 | 0.1466 | 0.1922 | 0.1392 | 0.1212 | 0.1354 | 0.1276 | 0.02692 | 0.0607 | 0 | 0 | 0 | 0 |
| GOT2 | Glutamic-oxaloacetic transaminase 2 | −0.381291 | 5.01E−21 | 2.58E−20 | 0.2346 | 0.2296 | 0.2188 | 0.2172 | 0.07141 | 0.1993 | 0.1728 | 0.09115 | 0.07472 | 0 | 0.006362 | 0.09641 |
| Treatment | TMB | POU2F2 marker | No. of patients (N=457) | No. of events | OS, median (95% CI), months | HR (POU2F2 marker absent vs present) (95% CI) | p value |
|---|---|---|---|---|---|---|---|
| Avelumab plus BSC | ≤Median | Absent | 22 | 10 | 36.99 (18.17-NE) | 0.46 (0.240-0.894) | 0.0218 |
| Avelumab plus BSC | ≤Median | Present | 112 | 82 | 17.77 (13.34-22.34) | ||
| Avelumab plus BSC | >Median | Absent | 11 | 7 | 19.25 (17.81-NE) | 1.38 (0.623-3.048) | 0.4281 |
| Avelumab plus BSC | >Median | Present | 98 | 48 | 35.12 (26.05-NE) | ||
| BSC alone | ≤Median | Absent | 18 | 13 | 13.68 (8.8-NE) | 1.14 (0.622-2.071) | 0.6788 |
| BSC alone | ≤Median | Present | 89 | 60 | 16.07 (10.25-24.18) | ||
| BSC alone | >Median | Absent | 21 | 14 | 14.78 (11.5-NE) | 1.14 (0.635-2.044) | 0.6608 |
| BSC alone | >Median | Present | 86 | 58 | 17.81 (13.54-26.64) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




