Submitted:
19 March 2025
Posted:
20 March 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
Data Source
Sequence Alignment
Sequences Comparison
Recombination Analysis
Phylogenetic Analysis
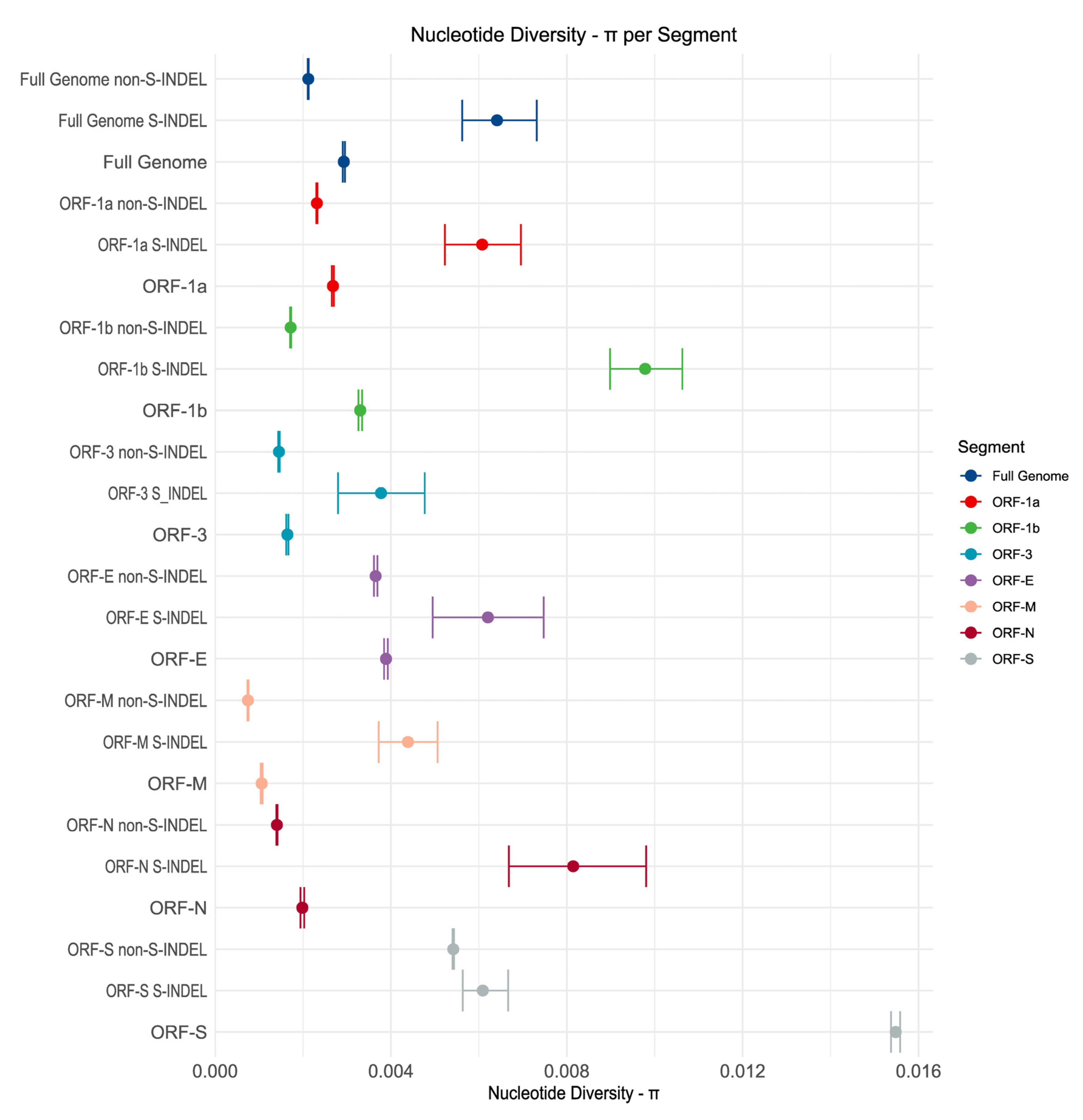
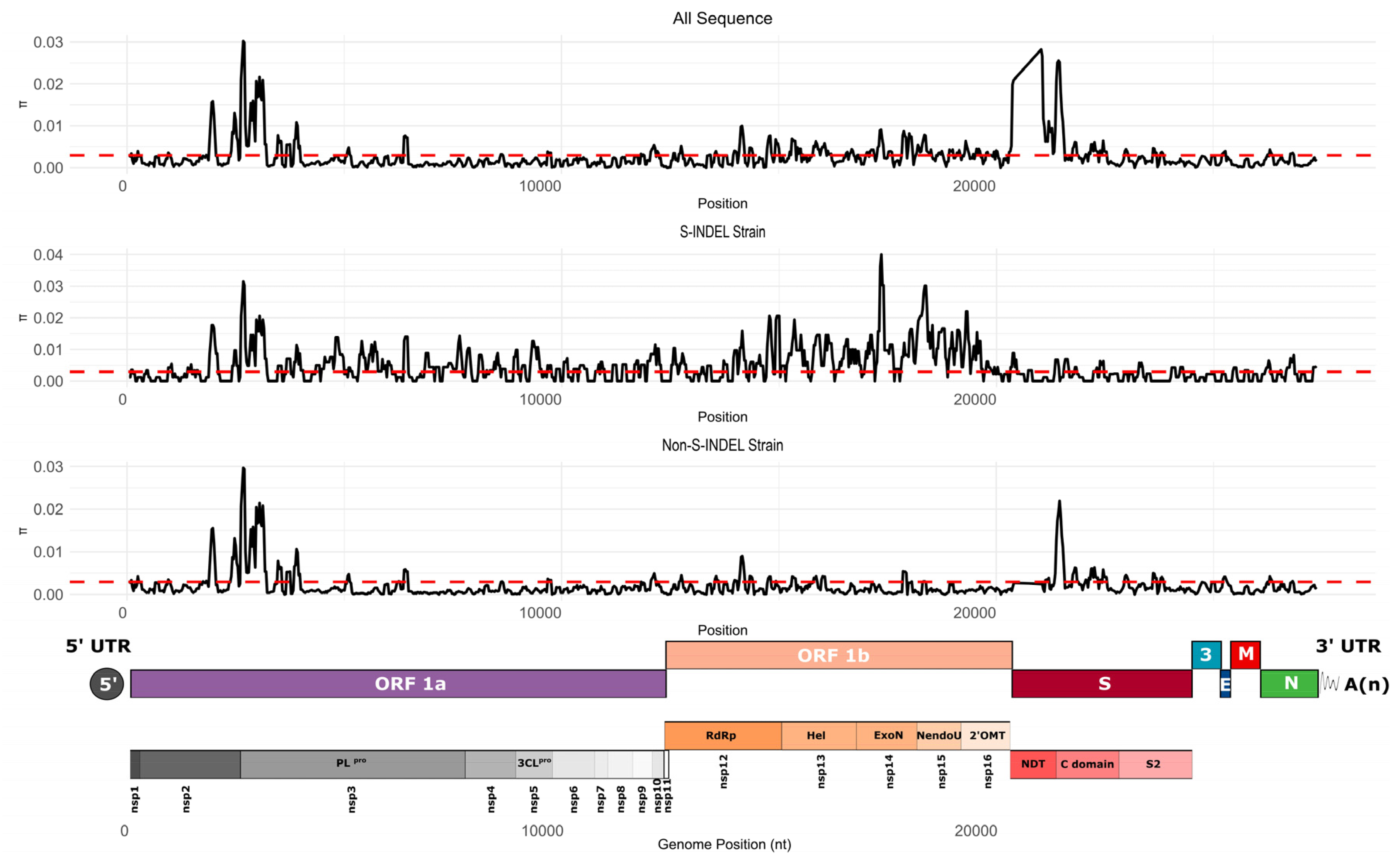
Genetic Diversity
Selections Analysis
3. Results
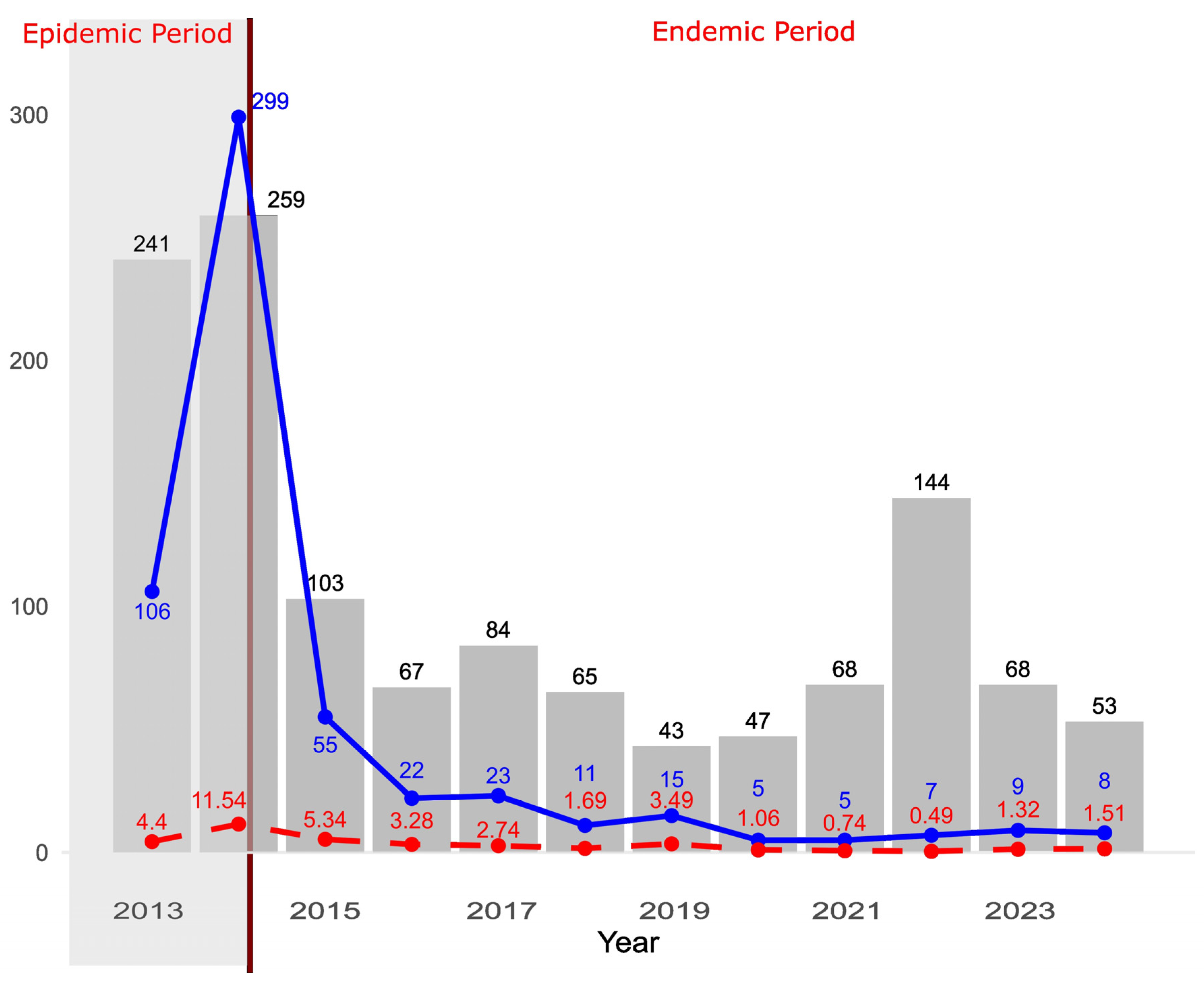
Stabilization of PEDV Case Numbers After the Epidemic Period
Recombination Analysis
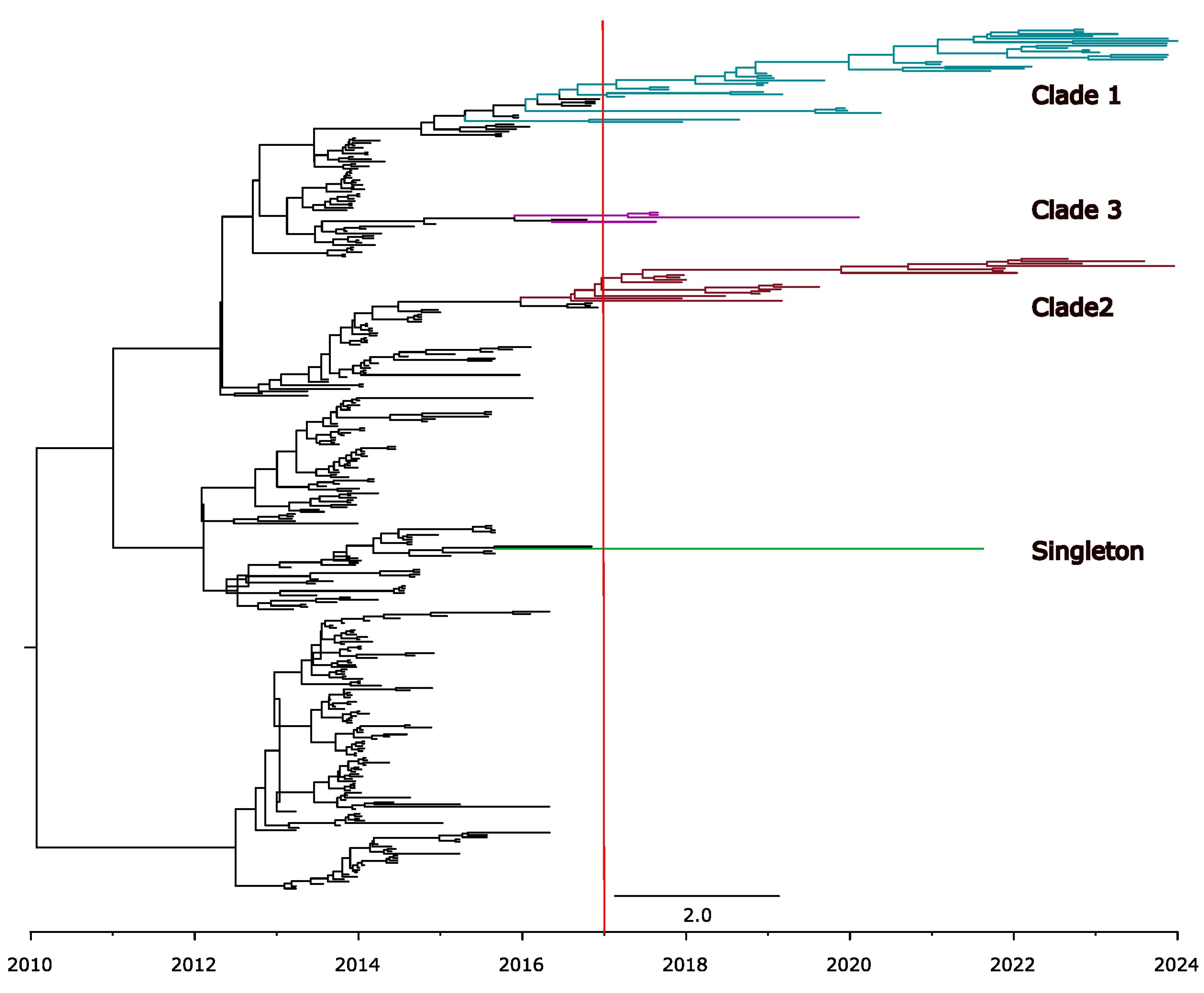
Temporal Signal and Nucleotide Substitution Rates
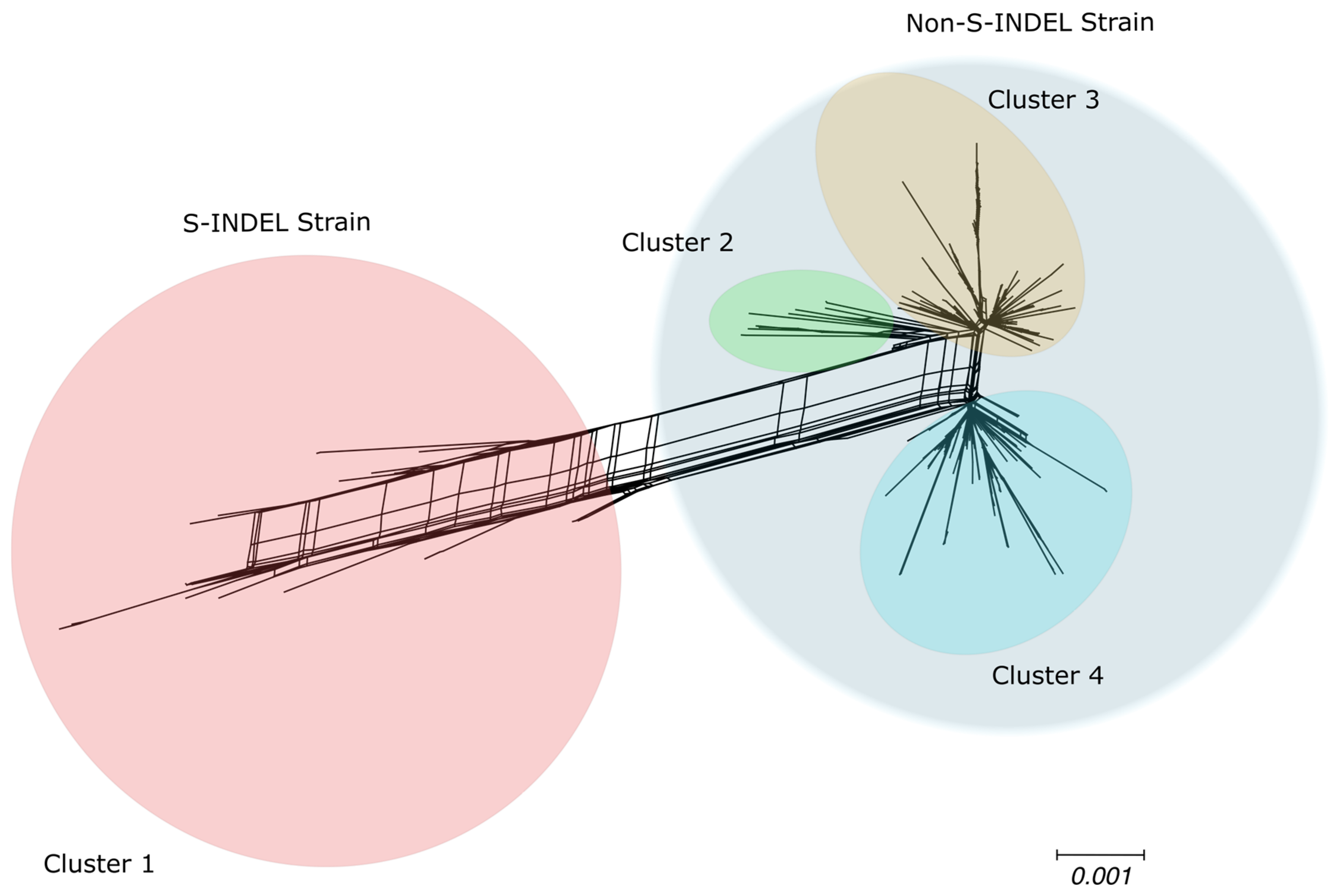
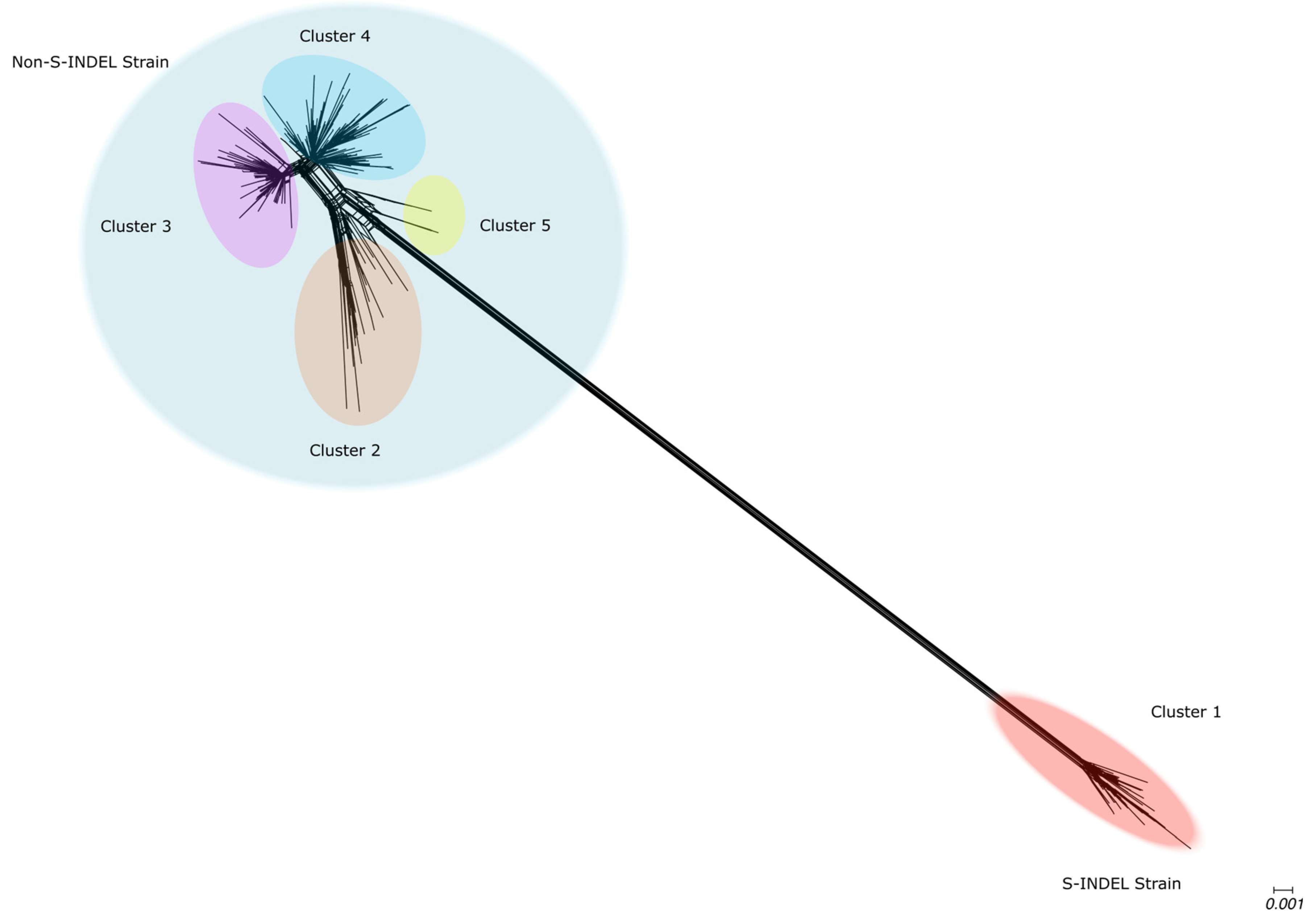
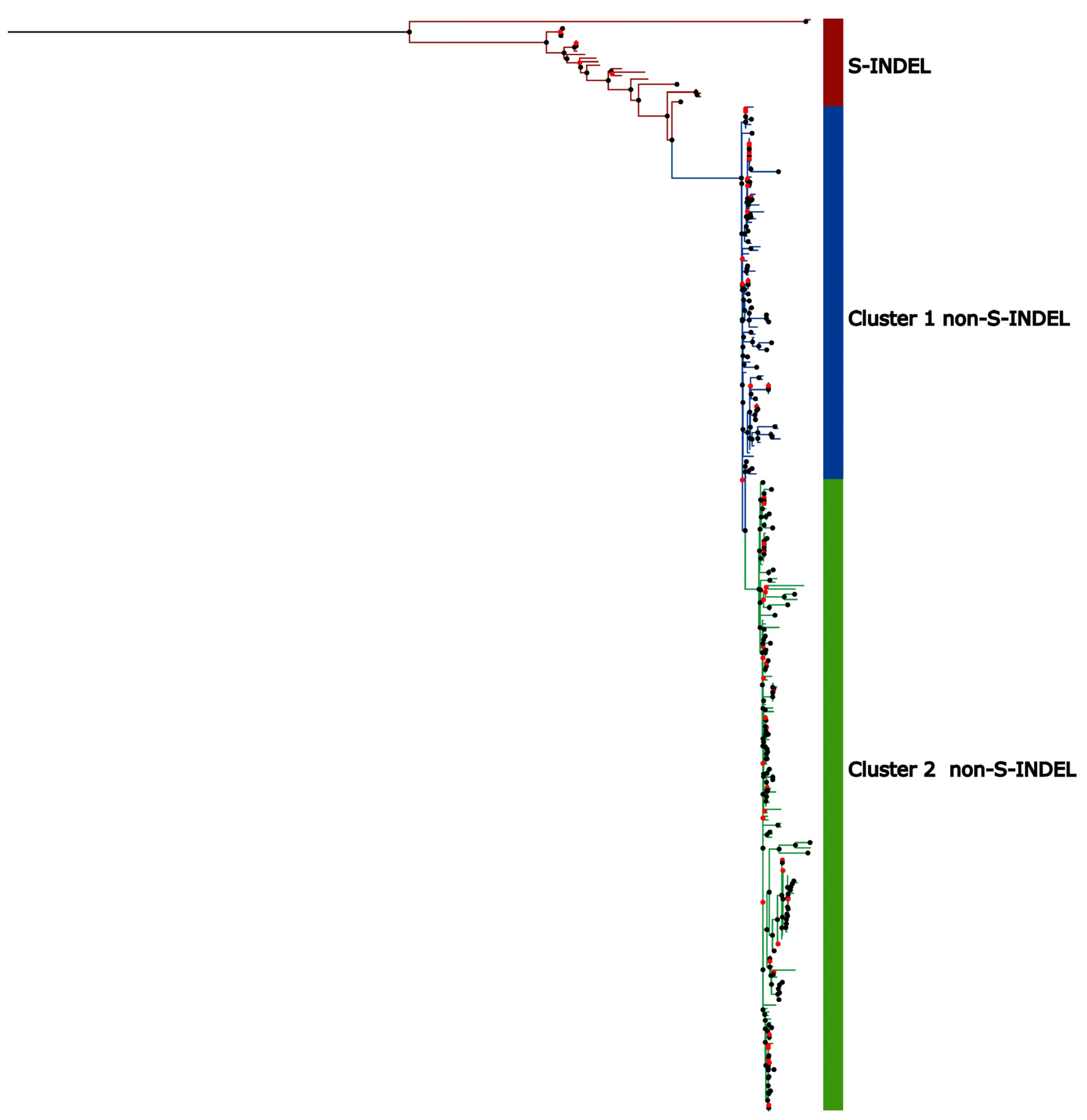
Genetic Variability in U.S. Population
High Sequence Similarity Between PEDV Strains and Among the Current Clades
Positive Selection Evidence
4. Discussion
5. Conclusions
6. Acknowledgements
References
- Wang, X.-m.; Niu, B.-b.; Yan, H.; Gao, D.-s.; Yang, X.; Chen, L.; Chang, H.-t.; Zhao, J.; Wang, C.-q. Archives of virology 2013. Genetic properties of endemic Chinese porcine epidemic diarrhea virus strains isolated since 2010, 158, 2487–2494. [Google Scholar]
- Stadler, J.; Zoels, S.; Fux, R.; Hanke, D.; Pohlmann, A.; Blome, S.; Weissenböck, H.; Weissenbacher-Lang, C.; Ritzmann, M.; Ladinig, A. Emergence of porcine epidemic diarrhea virus in southern Germany. BMC veterinary research 2015, 11, 1–8. [Google Scholar] [CrossRef]
- Stevenson, G.W.; Hoang, H.; Schwartz, K.J.; Burrough, E.R.; Sun, D.; Madson, D.; Cooper, V.L.; Pillatzki, A.; Gauger, P.; Schmitt, B.J. Emergence of Porcine epidemic diarrhea virus in the United States: Clinical signs, lesions, and viral genomic sequences. Journal of veterinary diagnostic investigation 2013, 25, 649–654. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zou, C.; Peng, O.; Ashraf, U.; Xu, Q.; Gong, L.; Fan, B.; Zhang, Y.; Xu, Z.; Xue, C. Global dynamics of porcine enteric coronavirus PEDV epidemiology, evolution, and transmission. Molecular Biology and Evolution 2023, 40, msad052. [Google Scholar] [CrossRef] [PubMed]
- Ojkic, D.; Hazlett, M.; Fairles, J.; Marom, A.; Slavic, D.; Maxie, G.; Alexandersen, S.; Pasick, J.; Alsop, J.; Burlatschenko, S. The first case of porcine epidemic diarrhea in Canada. The Canadian Veterinary Journal 2015, 56, 149. [Google Scholar]
- Vlasova, A.N.; Marthaler, D.; Wang, Q.; Culhane, M.R.; Rossow, K.D.; Rovira, A.; Collins, J.; Saif, L.J. Distinct characteristics and complex evolution of PEDV strains, North America, May 2013–February 2014. Emerging infectious diseases 2014, 20, 1620. [Google Scholar] [CrossRef]
- Song, D.; Park, B. Porcine epidemic diarrhoea virus: A comprehensive review of molecular epidemiology, diagnosis, and vaccines. Virus genes 2012, 44, 167–175. [Google Scholar] [CrossRef]
- Wang, L.; Byrum, B.; Zhang, Y. New variant of porcine epidemic diarrhea virus, United States, 2014. Emerging infectious diseases 2014, 20, 917. [Google Scholar] [CrossRef]
- Makau, D.N.; Pamornchainavakul, N.; VanderWaal, K.; Kikuti, M.; Picasso-Risso, C.; Geary, E.; Corzo, C.A. Postepidemic Epidemiology of Porcine Epidemic Diarrhea Virus in the United States. Transboundary and Emerging Diseases 2024, 2024, 5531899. [Google Scholar] [CrossRef]
- Yue, X.; Kikuti, M.; Melini, C.M.; Corzo, C.A. 2024. PEDV: Sow herds are eliminating the virus faster. National Hog Farmer.
- Project MSHM. 2024. PED aggregate prevalence of sow herd status (last updated 24). Report | Morrison Swine Health Monitoring Project (2024). Available online: https://mshmpumnedu/reports#Charts (accessed on 21 June 2024).
- Jung, K.; Saif, L.J. Porcine epidemic diarrhea virus infection: Etiology, epidemiology, pathogenesis and immunoprophylaxis. The Veterinary Journal 2015, 204, 134–143. [Google Scholar] [CrossRef]
- O’Dea, E.B.; Snelson, H.; Bansal, S. Using heterogeneity in the population structure of US swine farms to compare transmission models for porcine epidemic diarrhoea. Scientific Reports 2016, 6, 22248. [Google Scholar] [CrossRef]
- Schumacher, L.L.; Huss, A.R.; Cochrane, R.A.; Stark, C.R.; Woodworth, J.C.; Bai, J.; Poulsen, E.G.; Chen, Q.; Main, R.G.; Zhang, J. Characterizing the rapid spread of porcine epidemic diarrhea virus (PEDV) through an animal food manufacturing facility. PLoS ONE 2017, 12, e0187309. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Yang, M.; Goyal, S.M.; Cheeran, M.C.; Torremorell, M. Evaluation of biosecurity measures to prevent indirect transmission of porcine epidemic diarrhea virus. BMC veterinary research 2017, 13, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Dee, S.; Clement, T.; Schelkopf, A.; Nerem, J.; Knudsen, D.; Christopher-Hennings, J.; Nelson, E. An evaluation of contaminated complete feed as a vehicle for porcine epidemic diarrhea virus infection of naive pigs following consumption via natural feeding behavior: Proof of concept. BMC veterinary research 2014, 10, 1–9. [Google Scholar] [CrossRef]
- Woo, P.C.; de Groot, R.J.; Haagmans, B.; Lau, S.K.; Neuman, B.W.; Perlman, S.; Sola, I.; van der Hoek, L.; Wong, A.C.; Yeh, S.-H. ICTV Virus Taxonomy Profile: Coronaviridae 2023: This article is part of the ICTV Virus Taxonomy Profiles collection. Journal of general virology 2023, 104, 001843. [Google Scholar] [CrossRef]
- Elena, S.F.; Sanjuán, R. Adaptive value of high mutation rates of RNA viruses: Separating causes from consequences. Journal of virology 2005, 79, 11555–11558. [Google Scholar] [CrossRef]
- Makau, D.N.; Lycett, S.; Michalska-Smith, M.; Paploski, I.A.; Cheeran, M.C.-J.; Craft, M.E.; Kao, R.R.; Schroeder, D.C.; Doeschl-Wilson, A.; VanderWaal, K. Ecological and evolutionary dynamics of multi-strain RNA viruses. Nature Ecology & Evolution 2022, 6, 1414–1422. [Google Scholar]
- Duffy, S.; Shackelton, L.A.; Holmes, E.C. Rates of evolutionary change in viruses: Patterns and determinants. Nature Reviews Genetics 2008, 9, 267–276. [Google Scholar] [CrossRef]
- Dolan, P.T.; Whitfield, Z.J.; Andino, R. Mechanisms and concepts in RNA virus population dynamics and evolution. Annual Review of Virology 2018, 5, 69–92. [Google Scholar] [CrossRef]
- Minskaia, E.; Hertzig, T.; Gorbalenya, A.E.; Campanacci, V.; Cambillau, C.; Canard, B.; Ziebuhr, J. Discovery of an RNA virus 3′→ 5′ exoribonuclease that is critically involved in coronavirus RNA synthesis. Proceedings of the National Academy of Sciences 2006, 103, 5108–5113. [Google Scholar] [CrossRef]
- Eckerle, L.D.; Lu, X.; Sperry, S.M.; Choi, L.; Denison, M.R. High fidelity of murine hepatitis virus replication is decreased in nsp14 exoribonuclease mutants. Journal of virology 2007, 81, 12135–12144. [Google Scholar] [CrossRef] [PubMed]
- Brown, D.W. Threat to humans from virus infections of non-human primates. Reviews in medical virology 1997, 7, 239–246. [Google Scholar] [CrossRef]
- Gibbs, M.J.; Weiller, G.F. Evidence that a plant virus switched hosts to infect a vertebrate and then recombined with a vertebrate-infecting virus. Proceedings of the National Academy of Sciences 1999, 96, 8022–8027. [Google Scholar] [CrossRef] [PubMed]
- Simon-Loriere, E.; Holmes, E.C. Why do RNA viruses recombine? Nature Reviews Microbiology 2011, 9, 617–626. [Google Scholar] [CrossRef]
- Wells, H.L.; Bonavita, C.M.; Navarrete-Macias, I.; Vilchez, B.; Rasmussen, A.L.; Anthony, S.J. The coronavirus recombination pathway. Cell host & microbe 2023, 31, 874–889. [Google Scholar]
- Müller, N.F.; Kistler, K.E.; Bedford, T. A Bayesian approach to infer recombination patterns in coronaviruses. Nature communications 2022, 13, 4186. [Google Scholar] [CrossRef] [PubMed]
- Mei, X.; Guo, J.; Fang, P.; Ma, J.; Li, M.; Fang, L. The characterization and pathogenicity of a recombinant porcine epidemic diarrhea virus variant ECQ1. Viruses 2023, 15, 1492. [Google Scholar] [CrossRef]
- Jarvis, M.C.; Lam, H.C.; Zhang, Y.; Wang, L.; Hesse, R.A.; Hause, B.M.; Vlasova, A.; Wang, Q.; Zhang, J.; Nelson, M.I. Genomic and evolutionary inferences between American and global strains of porcine epidemic diarrhea virus. Preventive Veterinary Medicine 2016, 123, 175–184. [Google Scholar] [CrossRef]
- Huang, Y.-W.; Dickerman, A.W.; Piñeyro, P.; Li, L.; Fang, L.; Kiehne, R.; Opriessnig, T.; Meng, X.-J. Origin, evolution, and genotyping of emergent porcine epidemic diarrhea virus strains in the United States. MBio 2013, 4. [Google Scholar] [CrossRef]
- Lee, C. Porcine epidemic diarrhea virus: An emerging and re-emerging epizootic swine virus. Virology journal 2015, 12, 1–16. [Google Scholar] [CrossRef]
- Wang, Q.; Vlasova, A.N.; Kenney, S.P.; Saif, L.J. Emerging and re-emerging coronaviruses in pigs. Current opinion in virology 2019, 34, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Jung, K.; Saif, L.J.; Wang, Q. Porcine epidemic diarrhea virus (PEDV): An update on etiology, transmission, pathogenesis, and prevention and control. Virus research 2020, 286, 198045. [Google Scholar] [CrossRef]
- Lee, S.; Lee, C. Outbreak-related porcine epidemic diarrhea virus strains similar to US strains, South Korea, 2013. Emerging Infectious Diseases 2014, 20, 1223. [Google Scholar] [CrossRef]
- Lin, C.-N.; Chung, W.-B.; Chang, S.-W.; Wen, C.-C.; Liu, H.; Chien, C.-H.; Chiou, M.-T. US-like strain of porcine epidemic diarrhea virus outbreaks in Taiwan, 2013–2014. Journal of Veterinary Medical Science 2014, 76, 1297–1299. [Google Scholar] [CrossRef] [PubMed]
- He, W.-T.; Bollen, N.; Xu, Y.; Zhao, J.; Dellicour, S.; Yan, Z.; Gong, W.; Zhang, C.; Zhang, L.; Lu, M. Phylogeography reveals association between swine trade and the spread of porcine epidemic diarrhea virus in China and across the world. Molecular Biology and Evolution 2022, 39, msab364. [Google Scholar] [CrossRef] [PubMed]
- Sayers, E.W.; Cavanaugh, M.; Clark, K.; Ostell, J.; Pruitt, K.D.; Karsch-Mizrachi, I. GenBank. Nucleic acids research 2019, 47, D94–D99. [Google Scholar] [CrossRef]
- Perez, A.M.; Linhares, D.C.; Arruda, A.G.; VanderWaal, K.; Machado, G.; Vilalta, C.; Sanhueza, J.M.; Torrison, J.; Torremorell, M.; Corzo, C.A. Individual or common good? Voluntary data sharing to inform disease surveillance systems in food animals. Frontiers in Veterinary Science 2019, 6, 194. [Google Scholar]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Molecular biology and evolution 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Muhire, B.M.; Varsani, A.; Martin, D.P. SDT: A virus classification tool based on pairwise sequence alignment and identity calculation. PLoS ONE 2014, 9, e108277. [Google Scholar] [CrossRef]
- Martin, D.; Rybicki, E. RDP: Detection of recombination amongst aligned sequences. Bioinformatics 2000, 16, 562–563. [Google Scholar] [CrossRef]
- Padidam, M.; Sawyer, S.; Fauquet, C.M. Possible emergence of new geminiviruses by frequent recombination. Virology 1999, 265, 218–225. [Google Scholar] [CrossRef] [PubMed]
- Martin, D.P.; Posada, D.; Crandall, K.; Williamson, C. A modified bootscan algorithm for automated identification of recombinant sequences and recombination breakpoints. AIDS Research & Human Retroviruses 2005, 21, 98–102. [Google Scholar]
- Smith, J.M. Analyzing the mosaic structure of genes. Journal of molecular evolution 1992, 34, 126–129. [Google Scholar] [CrossRef]
- Posada, D.; Crandall, K.A. Evaluation of methods for detecting recombination from DNA sequences: Computer simulations. Proceedings of the National Academy of Sciences 2001, 98, 13757–13762. [Google Scholar] [CrossRef]
- Gibbs, M.J.; Armstrong, J.S.; Gibbs, A.J. Sister-scanning: A Monte Carlo procedure for assessing signals in recombinant sequences. Bioinformatics 2000, 16, 573–582. [Google Scholar] [CrossRef] [PubMed]
- Lam, H.M.; Ratmann, O.; Boni, M.F. Improved algorithmic complexity for the 3SEQ recombination detection algorithm. Molecular biology and evolution 2018, 35, 247–251. [Google Scholar] [CrossRef]
- Martin, D.P.; Varsani, A.; Roumagnac, P.; Botha, G.; Maslamoney, S.; Schwab, T.; Kelz, Z.; Kumar, V.; Murrell, B. RDP5: A computer program for analyzing recombination in, and removing signals of recombination from, nucleotide sequence datasets. Virus Evolution 2021, 7, veaa087. [Google Scholar] [CrossRef] [PubMed]
- Huson, D.H.; Bryant, D. Application of phylogenetic networks in evolutionary studies. Molecular biology and evolution 2006, 23, 254–267. [Google Scholar] [CrossRef]
- Kozlov, A.M.; Darriba, D.; Flouri, T.; Morel, B.; Stamatakis, A. RAxML-NG: A fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 2019, 35, 4453–4455. [Google Scholar] [CrossRef]
- Darriba, D.; Posada, D.; Kozlov, A.M.; Stamatakis, A.; Morel, B.; Flouri, T. ModelTest-NG: A new and scalable tool for the selection of DNA and protein evolutionary models. Molecular biology and evolution 2020, 37, 291–294. [Google Scholar] [CrossRef]
- Rambaut, A.; Lam, T.; Max Carvalho, L.; Pybus, O. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol 2016, 2, vew007. [Google Scholar] [CrossRef] [PubMed]
- Duchene, S.; Lemey, P.; Stadler, T.; Ho, S.Y.; Duchene, D.A.; Dhanasekaran, V.; Baele, G. Bayesian evaluation of temporal signal in measurably evolving populations. Molecular Biology and Evolution 2020, 37, 3363–3379. [Google Scholar] [CrossRef] [PubMed]
- Duchene, S.; Featherstone, L.; Haritopoulou-Sinanidou, M.; Rambaut, A.; Lemey, P.; Baele, G. Temporal signal and the phylodynamic threshold of SARS-CoV-2. Virus evolution 2020, 6, veaa061. [Google Scholar] [CrossRef]
- Suchard, M.A.; Lemey, P.; Baele, G.; Ayres, D.L.; Drummond, A.J.; Rambaut, A. Bayesian phylogenetic and phylodynamic data integration using BEAST 1. 10. Virus Evol 2018, 4, vey016. [Google Scholar] [CrossRef] [PubMed]
- Kass, R.E.; Raftery, A.E. Bayes factors. Journal of the american statistical association 1995, 90, 773–795. [Google Scholar] [CrossRef]
- Baele, G.; Lemey, P.; Suchard, M.A. Genealogical working distributions for Bayesian model testing with phylogenetic uncertainty. Systematic Biology 2016, 65, 250–264. [Google Scholar] [CrossRef]
- Rambaut, A.; Drummond, A.J.; Xie, D.; Baele, G.; Suchard, M.A. Posterior summarization in Bayesian phylogenetics using Tracer 1. 7. Systematic biology 2018, 67, 901–904. [Google Scholar] [CrossRef]
- Nei, M. Molecular evolutionary genetics. Columbia university press.
- Lima, A.T.; Silva, J.C.; Silva, F.N.; Castillo-Urquiza, G.P.; Silva, F.F.; Seah, Y.M.; Mizubuti, E.S.; Duffy, S.; Zerbini, F.M. 2017. The diversification of begomovirus populations is predominantly driven by mutational dynamics. Virus evolution 1987, 3, vex005. [Google Scholar]
- Peng, R.D. 2008. simpleboot: Simple Bootstrap Routines.
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Molecular Biology and Evolution 2017, 34, 3299–3302. [Google Scholar] [CrossRef]
- Kosakovsky Pond, S.L.; Posada, D.; Gravenor, M.B.; Woelk, C.H.; Frost, S.D. GARD: A genetic algorithm for recombination detection. Bioinformatics 2006, 22, 3096–3098. [Google Scholar] [CrossRef]
- Kosakovsky Pond, S.L.; Frost, S.D. Not so different after all: A comparison of methods for detecting amino acid sites under selection. Molecular biology and evolution 2005, 22, 1208–1222. [Google Scholar] [CrossRef]
- Murrell, B.; Wertheim, J.O.; Moola, S.; Weighill, T.; Scheffler, K.; Kosakovsky Pond, S.L. Detecting individual sites subject to episodic diversifying selection. PLoS genetics 2012, 8, e1002764. [Google Scholar] [CrossRef] [PubMed]
- Murrell, B.; Moola, S.; Mabona, A.; Weighill, T.; Sheward, D.; Kosakovsky Pond, S.L.; Scheffler, K. FUBAR: A fast, unconstrained bayesian approximation for inferring selection. Molecular biology and evolution 2013, 30, 1196–1205. [Google Scholar] [CrossRef] [PubMed]
- Weaver, S.; Shank, S.D.; Spielman, S.J.; Li, M.; Muse, S.V.; Kosakovsky Pond, S.L. Datamonkey 2. 0: A modern web application for characterizing selective and other evolutionary processes. Molecular Biology and Evolution 2018, 35, 773–777. [Google Scholar]
- Li, X.; Wu, Y.; Yan, Z.; Li, G.; Luo, J.; Huang, S.; Guo, X. A Comprehensive View on the Protein Functions of Porcine Epidemic Diarrhea Virus. Genes 2024, 15, 165. [Google Scholar] [CrossRef] [PubMed]
- Weng, L.; Weersink, A.; Poljak, Z.; de Lange, K.; von Massow, M. An economic evaluation of intervention strategies for Porcine Epidemic Diarrhea (PED). Preventive Veterinary Medicine 2016, 134, 58–68. [Google Scholar] [CrossRef]
- Kimberly, V.; Andres, P.; Montse, T.; Meggan, C. 2018. Role of animal movement and indirect contact among farms in transmission of porcine epidemic diarrhea virus.
- Gallien, S.; Andraud, M.; Moro, A.; Lediguerher, G.; Morin, N.; Gauger, P.C.; Bigault, L.; Paboeuf, F.; Berri, M.; Rose, N. Better horizontal transmission of a US non-InDel strain compared with a French InDel strain of porcine epidemic diarrhoea virus. Transboundary and emerging diseases 2018, 65, 1720–1732. [Google Scholar] [CrossRef]
- Kikuti, M.; Drebes, D.; Robbins, R.; Dufresne, L.; Sanhueza, J.M.; Corzo, C.A. Growing pig incidence rate, control and prevention of porcine epidemic diarrhea virus in a large pig production system in the United States. Porcine Health Management 2022, 8, 23. [Google Scholar] [CrossRef]
- Lowe, J.; Gauger, P.; Harmon, K.; Zhang, J.; Connor, J.; Yeske, P.; Loula, T.; Levis, I.; Dufresne, L.; Main, R. Role of transportation in spread of porcine epidemic diarrhea virus infection, United States. Emerging infectious diseases 2014, 20, 872. [Google Scholar] [CrossRef]
- Bowman, A.S.; Krogwold, R.A.; Price, T.; Davis, M.; Moeller, S.J. Investigating the introduction of porcine epidemic diarrhea virus into an Ohio swine operation. BMC veterinary research 2015, 11, 1–7. [Google Scholar] [CrossRef]
- Narechania, A.; Bobo, D.; Deitz, K.; DeSalle, R.; Planet, P.J.; Mathema, B. Rapid SARS-CoV-2 surveillance using clinical, pooled, or wastewater sequence as a sensor for population change. Genome Research 2024, 34, 1651–1660. [Google Scholar] [CrossRef] [PubMed]
- Korber, B.; Fischer, W.M.; Gnanakaran, S.; Yoon, H.; Theiler, J.; Abfalterer, W.; Hengartner, N.; Giorgi, E.E.; Bhattacharya, T.; Foley, B. Tracking changes in SARS-CoV-2 spike: Evidence that D614G increases infectivity of the COVID-19 virus. Cell 2020, 182, 812–827. [Google Scholar] [CrossRef] [PubMed]
- Cardenas, N.C.; Valencio, A.; Sanchez, F.; O’Hara, K.C.; Machado, G. Analyzing the intrastate and interstate swine movement network in the United States. Preventive Veterinary Medicine 2024, 230, 106264. [Google Scholar] [CrossRef] [PubMed]
- Niederwerder, M.; Hesse, R. Swine enteric coronavirus disease: A review of 4 years with porcine epidemic diarrhoea virus and porcine deltacoronavirus in the United States and Canada. Transboundary and emerging diseases 2018, 65, 660–675. [Google Scholar] [CrossRef]
- Sun, M.; Ma, J.; Wang, Y.; Wang, M.; Song, W.; Zhang, W.; Lu, C.; Yao, H. Genomic and epidemiological characteristics provide new insights into the phylogeographical and spatiotemporal spread of porcine epidemic diarrhea virus in Asia. Journal of clinical microbiology 2015, 53, 1484–1492. [Google Scholar] [CrossRef]
- Tian, Y.; Yu, Z.; Cheng, K.; Liu, Y.; Huang, J.; Xin, Y.; Li, Y.; Fan, S.; Wang, T.; Huang, G. Molecular characterization and phylogenetic analysis of new variants of the porcine epidemic diarrhea virus in Gansu, China in 2012. Viruses 2013, 5, 1991–2004. [Google Scholar] [CrossRef]
- Sun, R.; Leng, Z.; Zhai, S.-L.; Chen, D.; Song, C. Genetic variability and phylogeny of current Chinese porcine epidemic diarrhea virus strains based on spike, ORF3, and membrane genes. The scientific world Journal 2014, 2014, 208439. [Google Scholar]
- Peng, Q.; Fan, B.; Song, X.; He, W.; Wang, C.; Zhao, Y.; Guo, W.; Zhang, X.; Liu, S.; Gao, J. Genetic signatures associated with the virulence of porcine epidemic diarrhea virus AH2012/12. Journal of Virology 2023, 97, e01063–e23. [Google Scholar] [CrossRef]
- Li, D.; Li, Y.; Liu, Y.; Chen, Y.; Jiao, W.; Feng, H.; Wei, Q.; Wang, J.; Zhang, Y.; Zhang, G. Isolation and identification of a recombinant porcine epidemic diarrhea virus with a novel insertion in S1 domain. Frontiers in microbiology 2021, 12, 667084. [Google Scholar] [CrossRef]
- Wang, H.; Zhang, L.; Shang, Y.; Tan, R.; Ji, M.; Yue, X.; Wang, N.; Liu, J.; Wang, C.; Li, Y. Emergence and evolution of highly pathogenic porcine epidemic diarrhea virus by natural recombination of a low pathogenic vaccine isolate and a highly pathogenic strain in the spike gene. Virus evolution 2020, 6, veaa049. [Google Scholar] [CrossRef]
- Kong, N.; Meng, Q.; Jiao, Y.; Wu, Y.; Zuo, Y.; Wang, H.; Sun, D.; Dong, S.; Zhai, H.; Tong, W. Identification of a novel B-cell epitope in the spike protein of porcine epidemic diarrhea virus. Virology journal 2020, 17, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Servín-Blanco, R.; Zamora-Alvarado, R.; Gevorkian, G.; Manoutcharian, K. Antigenic variability: Obstacles on the road to vaccines against traditionally difficult targets. Human vaccines & immunotherapeutics 2016, 12, 2640–2648. [Google Scholar]
| Breakpoint Position | ||||||||
| A | ||||||||
| Full genome | In Recombinant sequence | |||||||
| event | Found in | Begin | End | Recombinationsequencies | Minor Parental | Major Parental | Detection Methods # | P-value* |
| 1 | 6 | 16816 | 22432 | KR265760 | KM975738 | KF452322 | RGBMCSP3 | 2.64 e-76 |
| 2 | 5 | 3910 | 20375 | KR265761 | KR265763 | KM975738 | RGMCS3 | 6.01 e-75 |
| 3 | 4 | 13244 | 21566 | KM975740 | KM975738 | KF452322 | RGMS3 | 3.03 e-53 |
| 4 | 2 | 13244 | 17879 | KR265786 | KJ645704 | KF468753 | RGMC3 | 3.02 e-46 |
| 5 | 2 | 11075 | 27425 | KM975738 | KR265759 | Unknown | RGBMCS3 | 5.96 e-37 |
| 6 | 106 | 16816 | 19902 | KU558702 | KJ645704 | 248 | RGMC3 | 3.48 e-32 |
| 7 | 12 | 20376 | 21478 | KR265761 | Unknown | KJ645643 | RGBMS3 | 1.48 e-33 |
| 8 | 7 | 4545 | 16815 | KR265759 | Unknown | 205 | RGBMS3 | 7.98 e-28 |
| 9 | 6 | 4377 | 16815 | 264 | Unknown | KJ645641 | MCS3 | 1.16 e-22 |
| 10 | 116 | 9921 | 14874 | KU893873 | KF452322 | KJ645635 | RGMC3 | 3.91 e-19 |
| 11 | 223 | 7680 | 13243 | KJ645704 | KJ645635 | KR265844 | RGM3 | 1.23 e-14 |
| 12 | 152 | 19912 | 21587 | KJ645641 | MG837058 | Unknown | RGMC3 | 1.20 e-6 |
| B | ||||||||
| Spike Protein | Breakpoint Position | |||||||
| event | Found in | Begin | End | Recombinationsequencies | Minor Parental | Major Parental | Detection Methods # | P-value* |
| 1 | 3 | 8 | 915 | KU982979 | 265 | 340 | RMSP3 | 1.75 e-23 |
| 2 | 34 | 8 | 955 | 340 | Unknown | 296 | RBMSP3 | 4.98 e-24 |
| 3 | 24 | 8 | 1056 | 262 | Unknown | KU982968 | MP3 | 3.88 e-23 |
| Clade | Max | Min | Average |
|---|---|---|---|
| Spike Protein | |||
| Non-S-INDEL | 99.8 % | 97.6 % | 99.1% |
| Clade1non-S-INDEL | 99.8 % | 98.2 % | 99.3% |
| Clade1non-S-INDEL vs older cladesnon-S-INDEL | 99.6 % | 97.7 % | 98.6% |
| Clade2non-S-INDEL | 99.9 % | 99.3 % | 99.2% |
| Clade2non-S-INDEL vs older cladesnon-S-INDEL | 99.7 % | 97.8 % | 99.3% |
| Clade1non-S-INDELvsc Clade 2non-S-INDEL | 99.0 % | 97.9% | 98.6% |
| S-INDEL strain | 100.0 % | 94.5 % | 99.1% |
| S-INDEL vs non-S-INDEL strains | 94.25% | 91.14% | 93,61% |
| Full Genome | |||
| Non-S-INDEL strain | 100% | 98.75% | 99.78% |
| INDEL strain | 100% | 99.21% | 99.55% |
| INDEL vs Non-INDEL strains | 99.85% | 98.52% | 98.99% |
| Gene | dN/dS | SLAC | MEME | FUBAR | |||
| diversifying | purifying | diversifying | purifying | diversifying | purifying | ||
| S-INDEL strain | |||||||
| Spike | 0.331 | - | 368, 659 | 27, 83, 240, 351, 429, 500, 632 | - | 83, 196, 310, 351, 719 | 21, 41, 44, 48, 73, 76, 92, 93, 125, 141, 149, 199, 226, 237, 238, 269, 312 |
| Envelop | 0.491 | - | - | - | - | 66 | - |
| Matrix | 0.0967 | - | - | - | - | - | 27, 41, 71, 78, 116, 121, 122, 188, 201 |
| Nucleo capside | 0.240 | - | - | 27, 54, 415 | - | 27 | 28, 43, 51, 140, 147, 207, 211, 249, 267, 271, 298, 327, 360, 364, 414 |
| Non-S-INDEL | |||||||
| Spike | 0.887 | 144, 380, 488, 525, 526, 568, 610, 614, 724 | 80, 94, 100, 101, 114, 139, 154, 209, 276, 359, 374, 462, 558, 582, 588, 621, 654, 664, 697, 729 |
24, 58, 144, 277, 355, 380, 412, 417, 433, 488, 495, 525, 526, 568, 610, 614, 724 |
- | 24, 55, 58, 70, 144, 157, 196, 277, 355, 380, 412, 433, 488, 494, 501, 525, 526, 568, 610, 614, 637, 653, 676, 695, 724 |
53, 94, 100, 101, 109, 139, 154, 244, 276, 359, 374, 394, 462, 468, 558, 582, 621, 625, 654, 697, 729 |
| Envelop | 0.743 | - | - | - | - | - | - |
| Matrix | 0.195 | - | - | - | 200 | - | - |
| Nucleo capside | 0.585 | - | 240 | - | - | 193 | 55, 87, 100, 190, 240, 244, 252, 386 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




