Submitted:
11 March 2025
Posted:
13 March 2025
You are already at the latest version
Abstract
Liver cancer poses a significant global health challenge, necessitating precise tumor segmentation in CT scans for diagnosis and treatment. While deep learning models like U-Net and Vision Transformers show promise, their computational demands hinder edge deployment. To address this gap, we propose an optimized Swin-UNet framework enhanced by the Search and Rescue (SAR) algorithm, enabling real-time edge computing without compromising the model’s performance. This work proposes a hybrid objective function with quadratic penalties for model compression and area under the curve(AUC). The models are trained using focal AUC loss to mitigate class imbalance. Evaluations on 3DIRCADb, LiTS, and MSD datasets show state-of-the-art performance, with Dice scores of 94.78%, 89.06%, and 88.95%, respectively, and an 80.3% parameter reduction versus baselines. The solution achieves efficient segmentation on edge devices (e.g., Jetson Nano), with a Volume Overlap Error of 1.73% (MSD) and Relative Volume Difference of 0.23% (3DIRCADb), outperforming existing methods. This work advances memory-efficient deep learning for clinical deployment, enabling AI-driven diagnostics in low-resource settings.
Keywords:
1. Introduction
- Model Size Optimization: The discrete design space of Swin-UNet achieves a balance between model size and accuracy, enabling deployment on memory-constrained devices like Jetson Nano.
- Quadratic Penalty Objective Function: A quadratic penalty-based objective function balances model size and AUC, encouraging compact, accurate models.
- Search and Rescue Algorithm: The Search and Rescue algorithm identifies optimal configurations, yielding an optimized model termed SAR-Swin-UNet.
- Focal AUC Loss Function: The Focal AUC loss function addresses class imbalance during training, enhancing the model’s ability to segment minority class pixels.
2. Preliminary Knowledge
2.1. The Transformer Model
2.2. The Vision Transformer (ViT)
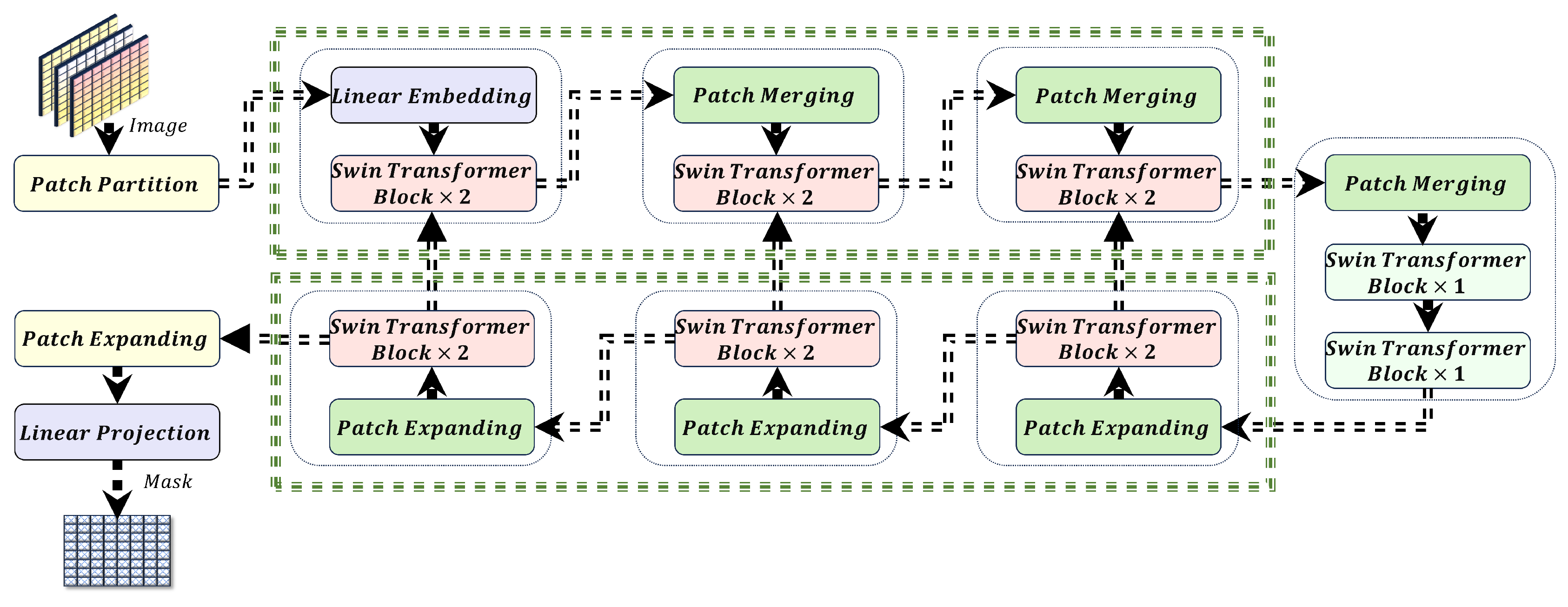
2.3. The Swin Transformer
2.4. U-Net for Segmentation
3. Proposed Methodology
3.1. Swin-UNet and Hyperparameter Optimization
3.1.1. Multi-Head Self-Attention (MHSA) in Swin-UNet
3.1.2. Hyperparameter Space and Constraints
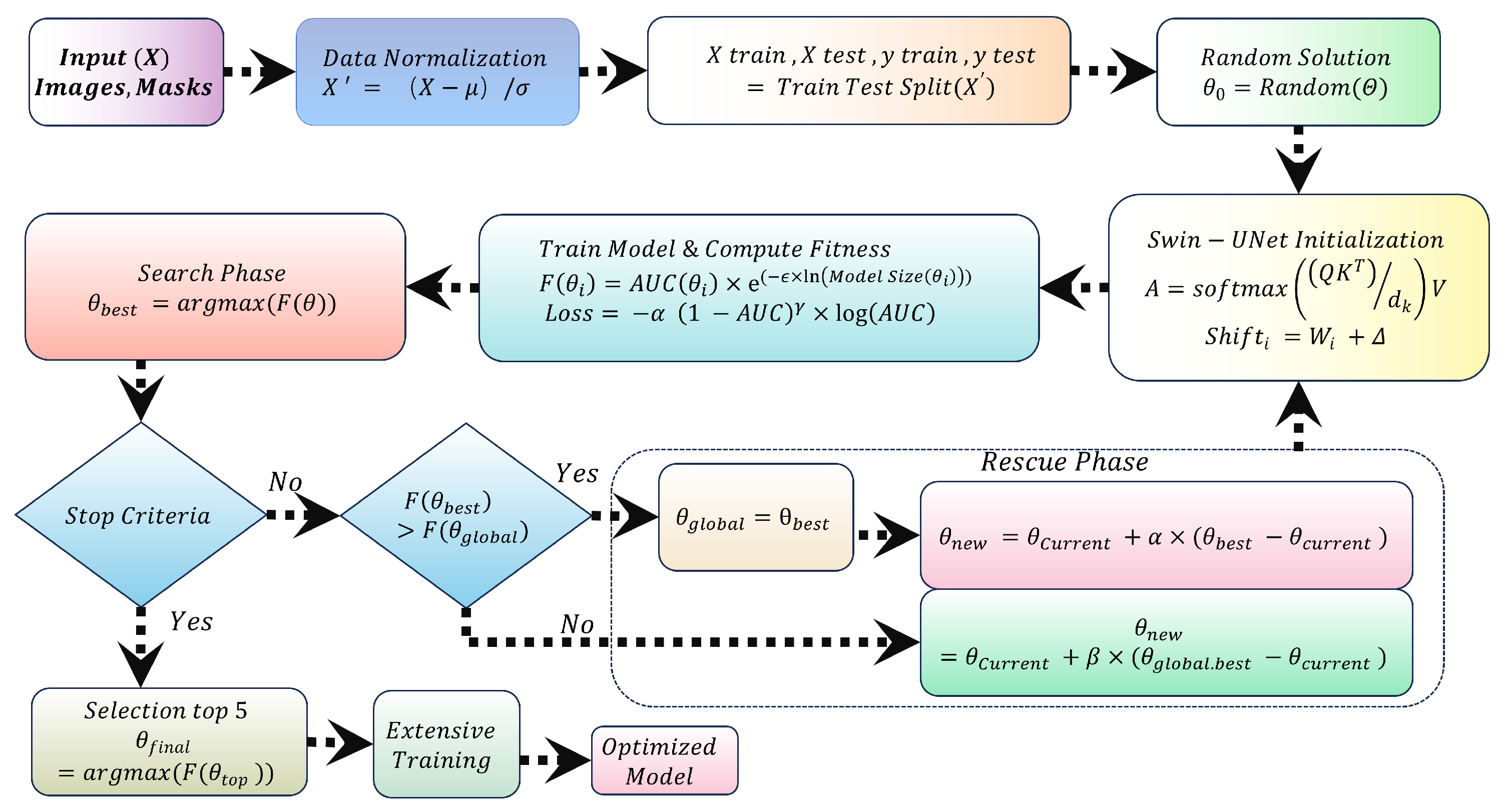
3.2. Search and Rescue (SAR) Algorithm
3.2.1. SAR Optimization Phases
3.3. AUC Focal Loss for Class Imbalance
| Algorithm 1 Search and Rescue (SAR) Algorithm for Swin-UNet Optimization |
|
3.4. Optimality Analysis of the Objective Function with SAR
- Local Optimality in Continuous Subspace Each continuous optimization step finds a locally optimal solution within the convex region.
- Global Exploration in Discrete Space By systematically exploring different discrete configurations, SAR ensures broad coverage of the search space.
4. Results and Discussion
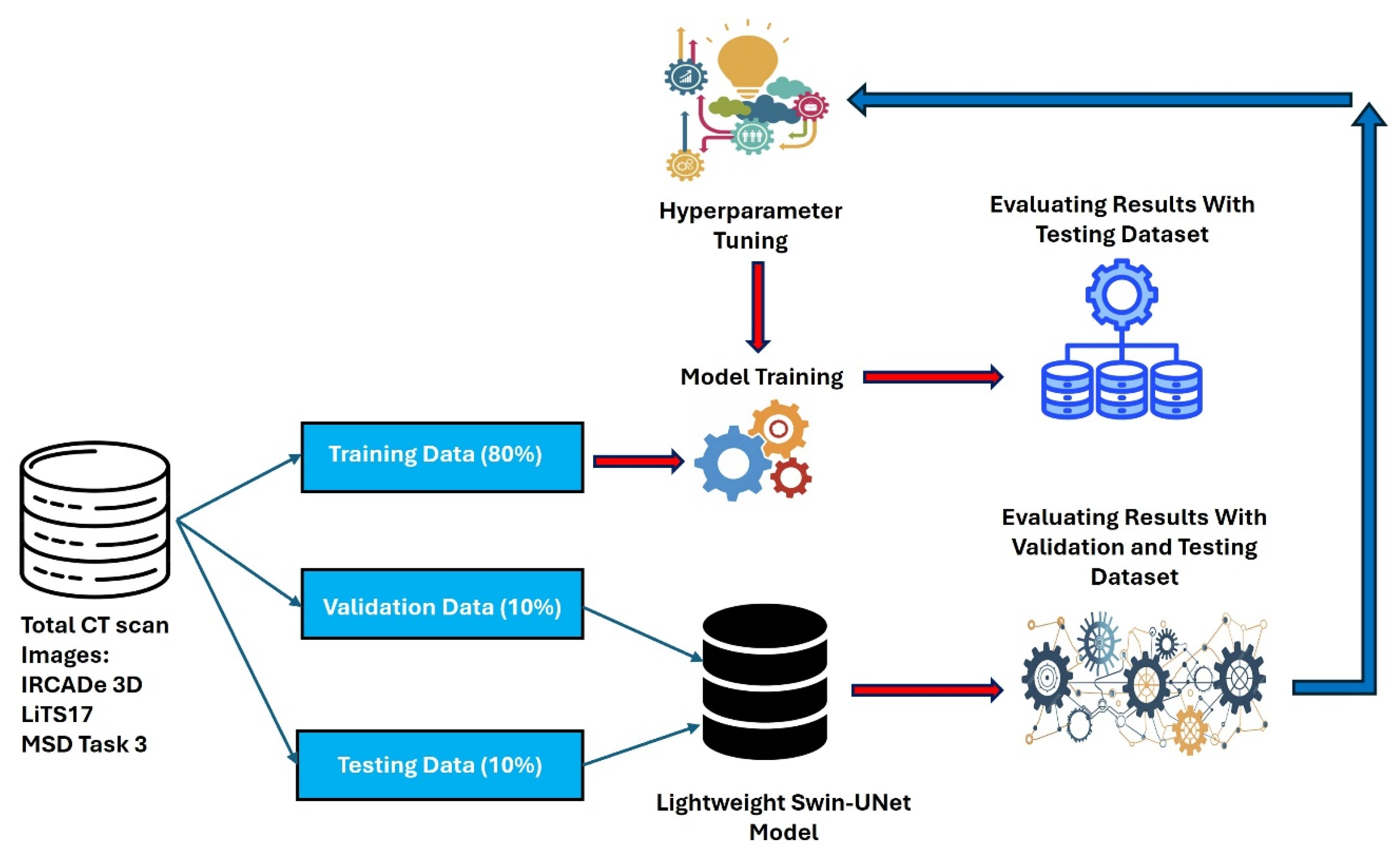
4.1. Datasets and Preprocessing
4.2. Experimental Setup
| Hyperparameter | Value |
|---|---|
| Learning Rate | 0.0001 |
| Epochs | 2000 |
| Batch Size | 64 |
| Optimizer | Adam |
| Patience (Early Stopping) | 10 epochs |
| Device for Optimization | Nvidia 3090 Ti |
| Device for Deployment | Jetson Nano |
4.3. Comparison Metrics
4.4. Hyperparameter Optimization and Model Performance
4.5. Model Performance on Liver Tumor Segmentation
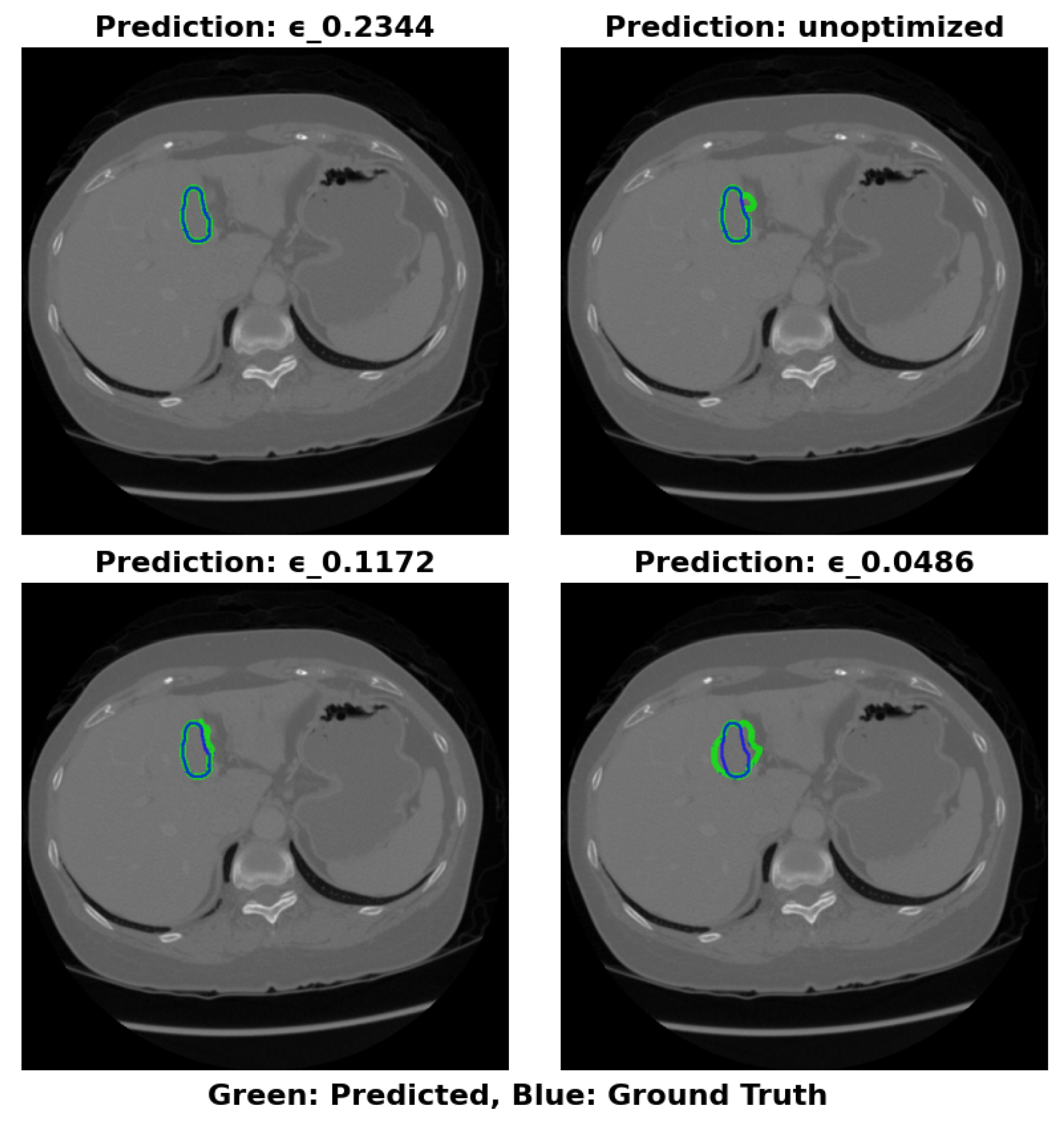
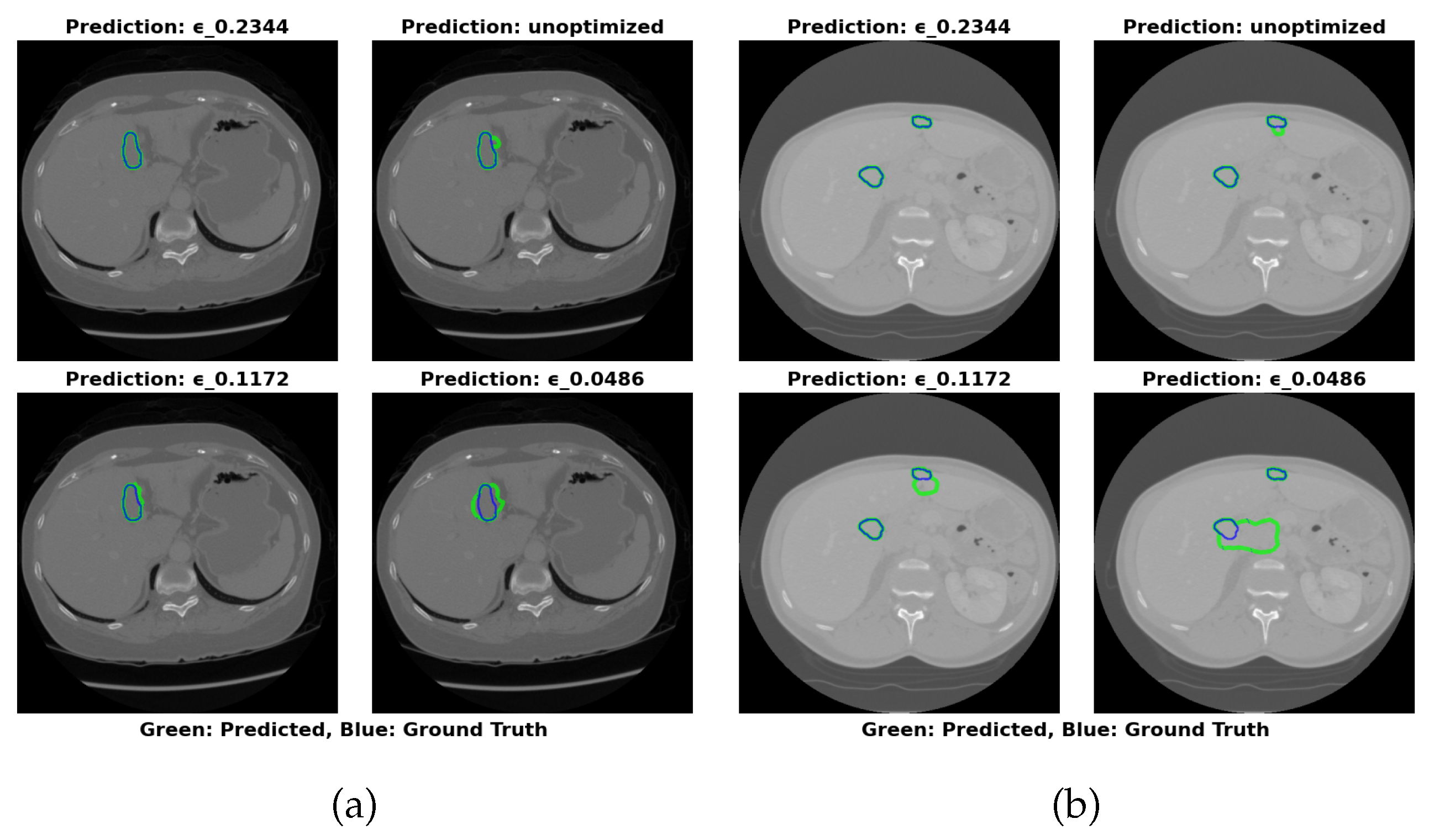
4.5.1. 3DIRCADb Dataset Analysis
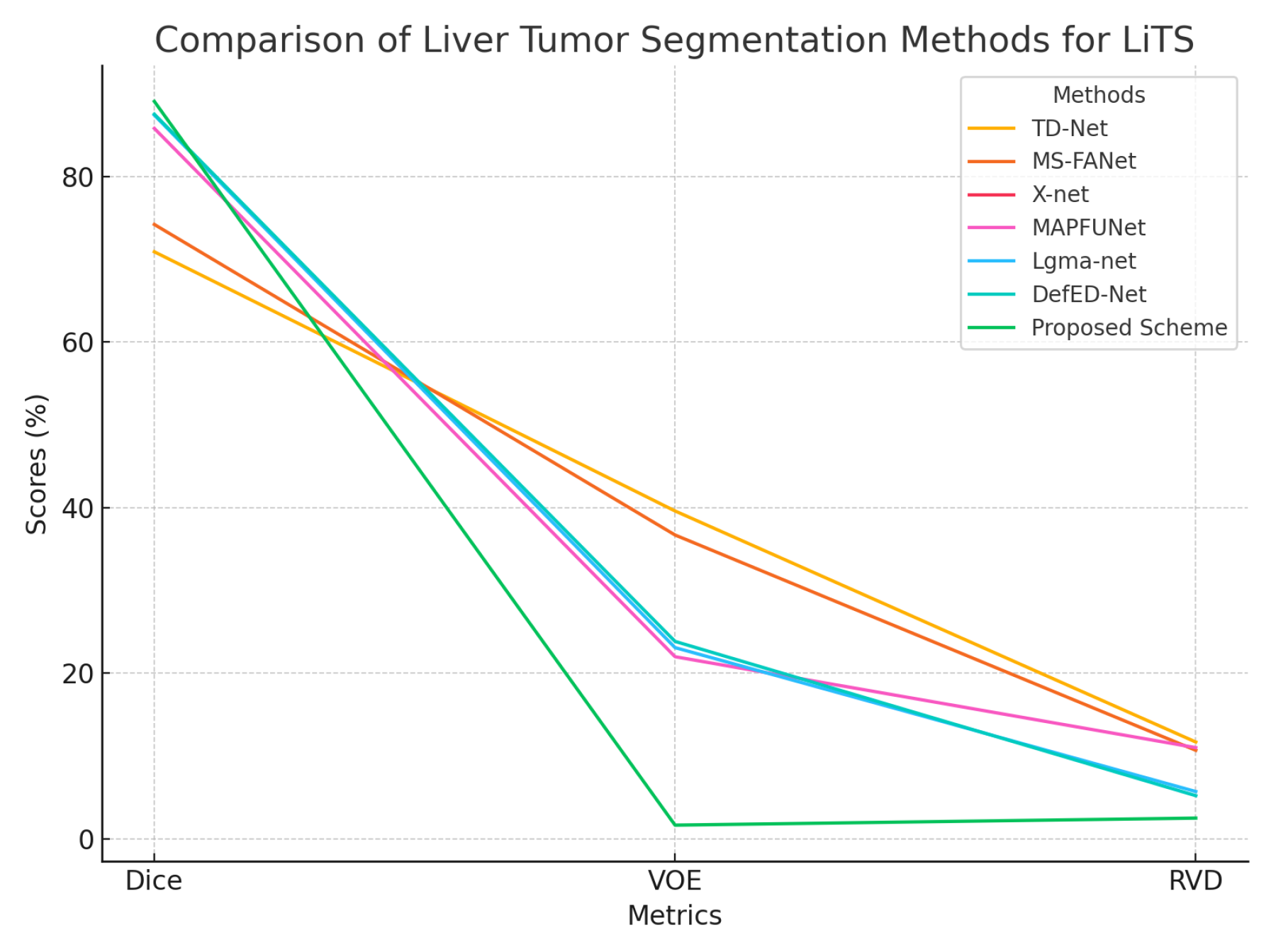
4.5.2. LiTS Dataset Analysis
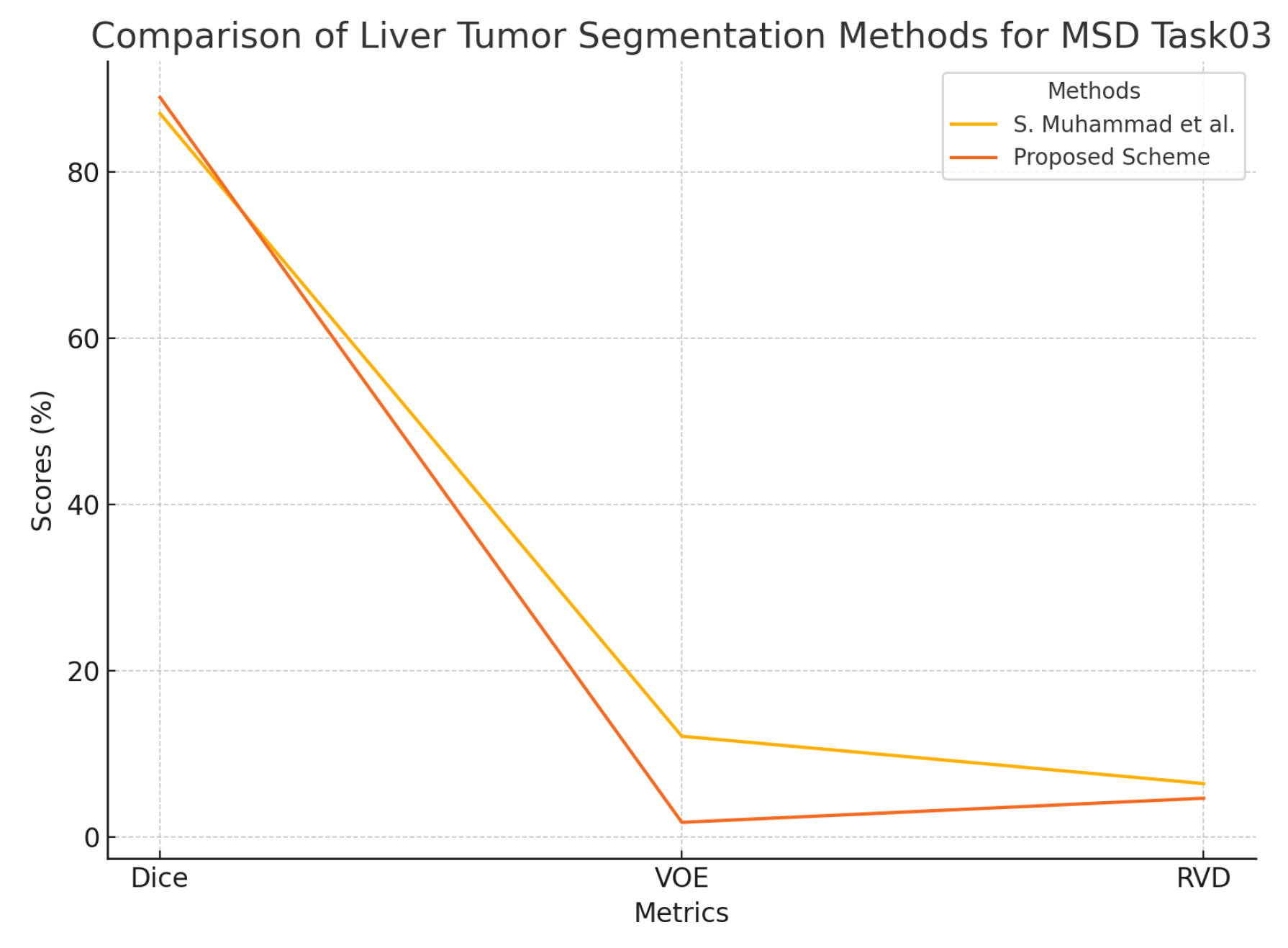
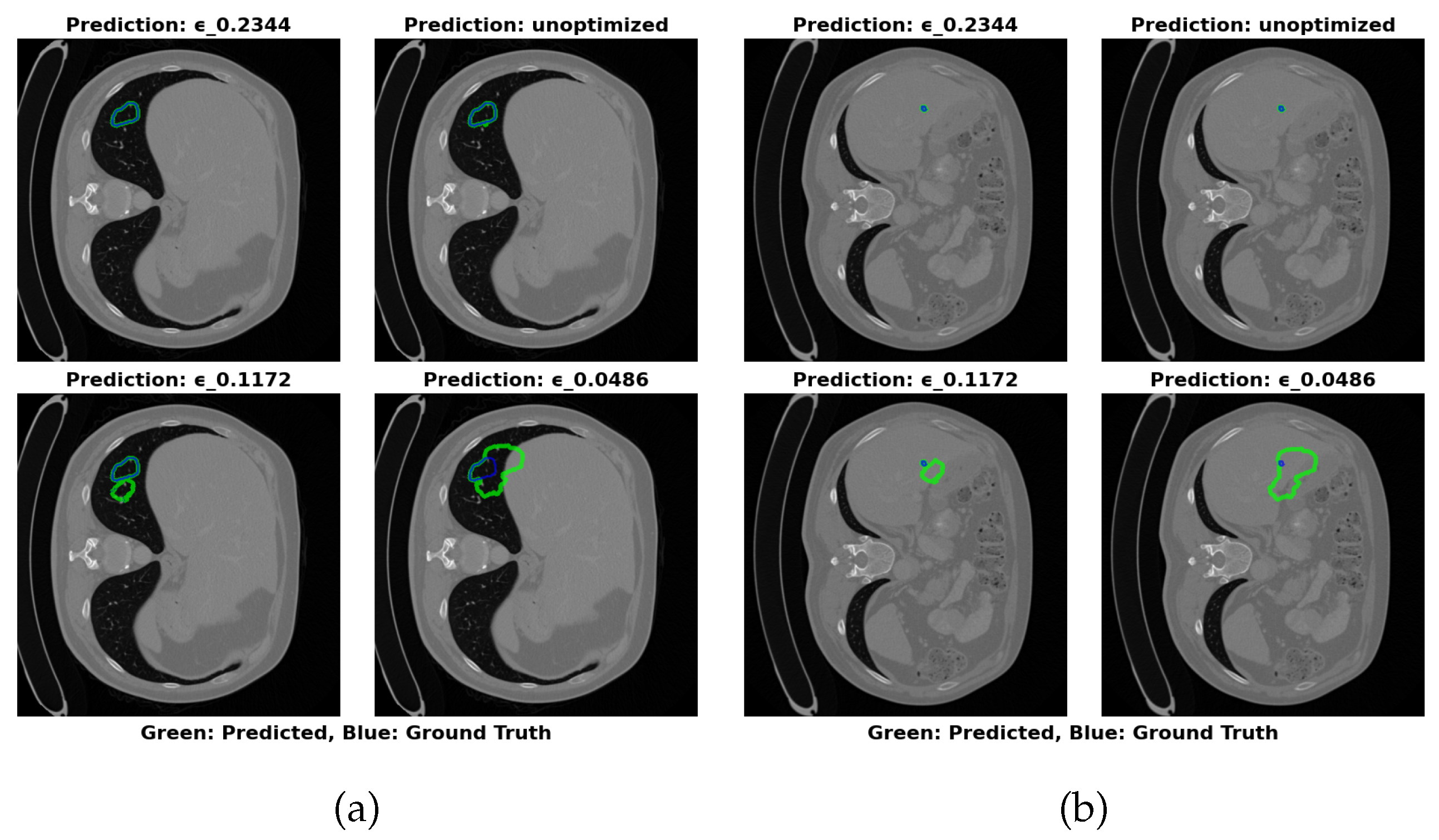
4.5.3. MSD Task03 Dataset Analysis
4.5.4. Optimal Configuration Selection
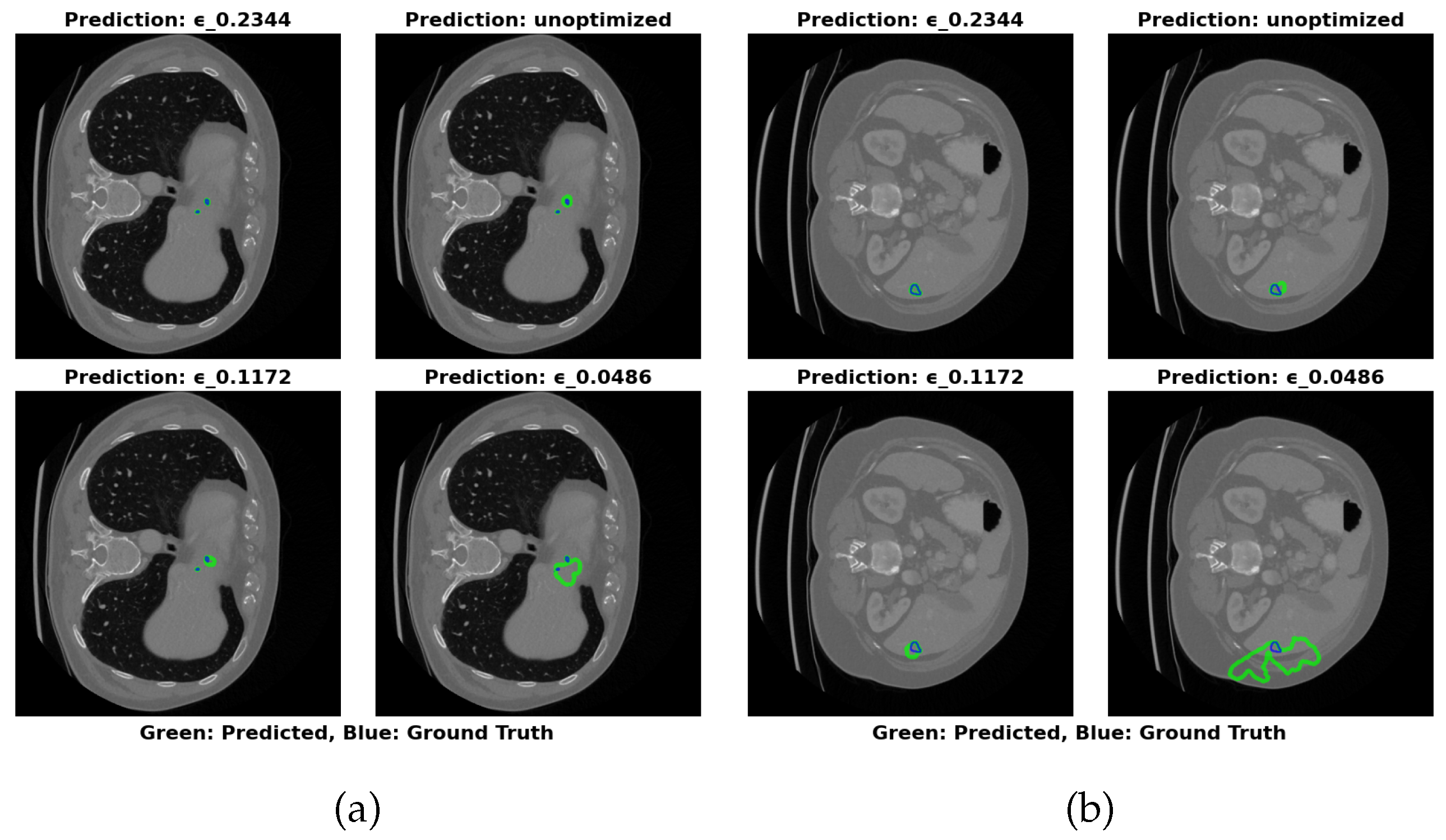
- Maximal Compression: =0.0486 (17.22 MB) for storage-constrained deployments.
- Optimal Balance: =0.2344 (64.16 MB) maintains greater than 83% Dice with RVD less than 25% across datasets.
- Maximal Accuracy: Unoptimized (324.91 MB) for non-constrained environments.
4.6. Optimized Model Performance Comparison with SOTA
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; others. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians 2021, 71, 209–249. [CrossRef]
- Kazi, I.A.; Jahagirdar, V.; Kabir, B.W.; Syed, A.K.; Kabir, A.W.; Perisetti, A. Role of Imaging in Screening for Hepatocellular Carcinoma. Cancers 2024, 16, 3400. [CrossRef]
- Dharaneswar, S.; Kumar, B.S. Elucidating the novel framework of liver tumour segmentation and classification using improved Optimization-assisted EfficientNet B7 learning model. Biomedical Signal Processing and Control 2025, 100, 107045. [CrossRef]
- Ghobadi, V.; Ismail, L.I.; Hasan, W.Z.W.; Ahmad, H.; Ramli, H.R.; Norsahperi, N.M.H.; Tharek, A.; Hanapiah, F.A. Challenges and solutions of deep learning-based automated liver segmentation: A systematic review. Computers in Biology and Medicine 2025, 185, 109459. [CrossRef]
- Rahman, H.; Aoun, N.B.; Bukht, T.F.N.; Ahmad, S.; Tadeusiewicz, R.; Pławiak, P.; Hammad, M. Automatic Liver Tumor Segmentation of CT and MRI Volumes Using Ensemble ResUNet-InceptionV4 Model. Information Sciences 2025, p. 121966. [CrossRef]
- Hammad, M.; ElAffendi, M.; Asim, M.; Abd El-Latif, A.A.; Hashiesh, R. Automated lung cancer detection using novel genetic TPOT feature optimization with deep learning techniques. Results in Engineering 2024, 24, 103448. [CrossRef]
- Rehman, A.; Mujahid, M.; Damasevicius, R.; Alamri, F.S.; Saba, T. Densely convolutional BU-NET framework for breast multi-organ cancer nuclei segmentation through histopathological slides and classification using optimized features. CMES-Computer modeling In engineering and sciences. 2024, 141, 2375–2397. [CrossRef]
- Hussain, S.S.; Degang, X.; Shah, P.M.; Islam, S.U.; Alam, M.; Khan, I.A.; Awwad, F.A.; Ismail, E.A. Classification of Parkinson’s disease in patch-based MRI of substantia nigra. Diagnostics 2023, 13, 2827. [CrossRef]
- Javed, R.; Saba, T.; Alahmadi, T.J.; Al-Otaibi, S.; AlGhofaily, B.; Rehman, A. EfficientNetB1 Deep Learning Model for Microscopic Lung Cancer Lesion Detection and Classification Using Histopathological Images. Computers, Materials & Continua 2024, 81. [CrossRef]
- Gul, S.; Khan, M.S.; Bibi, A.; Khandakar, A.; Ayari, M.A.; Chowdhury, M.E. Deep learning techniques for liver and liver tumour segmentation: A review. Computers in Biology and Medicine 2022, 147, 105620. [CrossRef]
- Moghe, A.A.; Singhai, J.; Shrivastava, S. Automatic threshold based liver lesion segmentation in abdominal 2D-CT images. International Journal of Image Processing (IJIP) 2011, 5, 166.
- Peng, W.; Zhao, Y. Liver CT image segmentation based on modified Canny algorithm. 2019 12th International Congress on Image and Signal Processing, BioMedical Engineering and Informatics (CISP-BMEI). IEEE, 2019, pp. 1–5. [CrossRef]
- Anter, A.; Hassenian, A. CT liver tumor segmentation hybrid approach using neutrosophic sets, fast fuzzy c-means and adaptive watershed algorithm. Artif. Intell. Med. 2019, 97, 105–117. [CrossRef]
- Xu, Y.; Quan, R.; Xu, W.; Huang, Y.; Chen, X.; Liu, F. Advances in medical image segmentation: A comprehensive review of traditional, deep learning and hybrid approaches. Bioengineering 2024, 11, 1034. [CrossRef]
- Al-Kofahi, Y.; Lassoued, W.; Lee, W.; Roysam, B. Improved automatic detection and segmentation of cell nuclei in histopathology images. IEEE Trans Biomed Eng 2010, 57, 841–852. [CrossRef]
- Kong, H.; Akakin, H.; Sarma, S. A generalized Laplacian of Gaussian filter for blob detection and its applications. IEEE Trans Cybern 2013, 43, 1719–1733. [CrossRef]
- Basu, M. Gaussian-based edge-detection methods-a survey. IEEE Trans Syst Man Cybern Part C (appl Rev) 2002, 32, 252–260. [CrossRef]
- Chan, T.; Vese, L. Active contours without edges. IEEE Trans Image Process 2001, 10, 266–277. [CrossRef]
- Moga, A.; Gabbouj, M. Parallel marker-based image segmentation with watershed transformation. J Parallel Distrib Comput 1998, 51, 27–45. [CrossRef]
- Saha Roy, S.; Roy, S.; Mukherjee, P.; Roy, A. An automated liver tumour segmentation and classification model by deep learning based approaches. Comput. Methods Biomech. Biomed. Eng. Imaging Vis. 2022, pp. 1–13. [CrossRef]
- Chen, Y.; others. MS-FANet: multi-scale feature attention network for liver tumor segmentation. Computers in Biology and Medicine 2023, 163, 107208. [CrossRef]
- Lakshmi, P.; Sampurna, P.; others. Deploying the model of improved heuristic-assisted adaptive SegUnet++ and multi-scale deep learning network for liver tumor segmentation and classification. J. Real-Time Image Process. 2025, 22, 8. [CrossRef]
- Reyad, M.; others. Architecture optimization for hybrid deep residual networks in liver tumor segmentation using a GA. Int. J. Comput. Intell. Syst. 2024, 17, 209. [CrossRef]
- Di, S.; others. TD-Net: A hybrid end-to-end network for automatic liver tumor segmentation from CT images. IEEE Journal of Biomedical and Health Informatics 2022, 27, 1163–1172. [CrossRef]
- Liu, Z.; others. PA-Net: A phase attention network fusing venous and arterial phase features of CT images for liver tumor segmentation. Comput. Methods Programs Biomed. 2024, 244, 107997. [CrossRef]
- Valanarasu, J.; Oza, P.; Hacihaliloglu, I.; Patel, V. Medical transformer: Gated axial-attention for medical image segmentation. Proc. Med. Image Comput. Comput. Assist. Interv., 2021, pp. 36–46. [CrossRef]
- Dosovitskiy, A.; others. An image is worth 16×16 words: Transformers for image recognition at scale. Proc. 9th Int. Conf. Learn. Representations, 2021, pp. 1–22.
- Zhu, X.; Su, W.; Lu, L.; Li, B.; Wang, X.; Dai, J. Deformable DETR: Deformable transformers for end-to-end object detection. Proc. Int. Conf. Learn. Representations, 2021, pp. 1–16. [CrossRef]
- Liang, J.; Homayounfar, N.; Ma, W.C.; Xiong, Y.; Hu, R.; Urtasun, R. PolyTransform: Deep polygon transformer for instance segmentation. Proc. IEEE/CVF Conf. Comput. Vis. Pattern Recognit., 2020, pp. 9128–9137.
- Balasubramanian, P.; Lai, W.C.; Seng, G.; Selvaraj, J. APESTNet with Mask R-CNN for liver tumour segmentation and classification. Cancers 2023, 15, 330. [CrossRef]
- Chen, J.; others. TransUNet: Transformers make strong encoders for medical image segmentation. arXiv:2102.04306 2021. [CrossRef]
- Ni, Y.; others. DA-Tran: Multiphase liver tumor segmentation with a domain-adaptive transformer network. Pattern Recognition 2024, 149, 110233. [CrossRef]
- Aslam, L.; Zou, R.; Awan, E.S.; Hussain, S.S.; Shaki, K.A.; Wani, M.A.; Asim, M. Hardware-Centric Exploration of the Discrete Design Space in Transformer-LSTM Models for Wind Speed Prediction on Memory-Constrained Devices 2025. [CrossRef]
- Aslam, L.; Zou, R.; Awan, E.; Butt, S.A. Integrating Physics-Informed Vectors for Improved Wind Speed Forecasting with Neural Networks. 2024 14th Asian Control Conference (ASCC). IEEE, 2024, pp. 1902–1907.
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser. Attention is all you need. Advances in Neural Information Processing Systems 2017, 30.
- Alexey, D. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv preprint arXiv: 2010.11929 2020. [CrossRef]
- Liu, Z.; Lin, Y.; Cao, Y.; Hu, H.; Wei, Y.; Zhang, Z.; Xie, Z.; Lin, S.; Li, H. Swin Transformer: Hierarchical Vision Transformer using Shifted Windows. Proceedings of the IEEE/CVF International Conference on Computer Vision (ICCV) 2021, pp. 10012–10022.
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI) 2015, pp. 234–241. [CrossRef]
- Shabani, A.; Asgarian, B.; Salido, M.; Gharebaghi, S.A. Search and rescue optimization algorithm: A new optimization method for solving constrained engineering optimization problems. Expert Systems with Applications 2020, 161, 113698. [CrossRef]
- Bilic, P.; Christ, P.; Li, H.B.; Vorontsov, E.; Ben-Cohen, A.; Kaissis, G.; Menze, B.H. The liver tumor segmentation benchmark (LiTS). Medical Image Analysis 2023, 84, 10268. [CrossRef]
- Soler, L.; Hostettler, A.; Agnus, V.; Charnoz, A.; Fasquel, J.B.; Moreau, J.; Marescaux, J. 3D image reconstruction for comparison of algorithm database, 2010.
- Antonelli, M.; Reinke, A.; Bakas, S.; Farahani, K.; Kopp-Schneider, A.; Landman, B.A.; Cardoso, M.J. The medical segmentation decathlon. Nature Communications 2022, 13, 4128. [CrossRef]
- Lei, T.; others. DefED-Net: Deformable encoder-decoder network for liver and liver tumor segmentation. IEEE Transactions on Radiation and Plasma Medical Sciences 2021, 6, 68–78. [CrossRef]
- Chi, J.; others. X-Net: Multi-branch UNet-like network for liver and tumor segmentation from 3D abdominal CT scans. Neurocomputing 2021, 459, 81–96. [CrossRef]
- Ren, W.; others. Lgma-net: liver and tumor segmentation methods based on local–global feature mergence and attention mechanisms. Signal, Image and Video Processing 2025, 19, 1–11. [CrossRef]
- Kushnure, D.T.; Talbar, S.N. MS-UNet: A multi-scale UNet with feature recalibration approach for automatic liver and tumor segmentation in CT images. Computerized Medical Imaging and Graphics 2021, 89, 101885. [CrossRef]
- Sun, J.; others. MAPFUNet: Multi-attention Perception-Fusion U-Net for Liver Tumor Segmentation. Journal of Bionic Engineering 2024, pp. 1–25. [CrossRef]
- Muhammad, S.; Zhang, J. Segmentation of Liver Tumors by Monai and PyTorch in CT Images with Deep Learning Techniques. Applied Sciences 2024, 14, 5144. [CrossRef]
| Hyperparameter | Symbol | Range |
|---|---|---|
| Depth | D | [2, 8] |
| Initial Filter Number | [32, 256] | |
| Patch Size | P | [2, 16] |
| Number of Attention Heads | H | [1, 16] |
| Window Size | W | [1, 8] |
| MLP Size | M | [32, 512] |
| AUC Focal Loss Alpha | [0, 5] | |
| AUC Focal Loss Gamma | [0, 5] |
| Dataset | Total Images | Training (80%) | Validation (10%) | Test (10%) |
|---|---|---|---|---|
| LiTS17[40] | 131 | 104 | 13 | 13 |
| IRCADe 3D [41] | 20 | 16 | 4 | 4 |
| MSD Task-3 [42] | 131 | 104 | 13 | 13 |
| Total | 282 | 224 | 30 | 30 |
| Hyperparameter | Unoptimized | |||
|---|---|---|---|---|
| Filter Number Begin | 128 | 32 | 32 | 64 |
| Depth | 4 | 4 | 4 | 4 |
| Stack Num Down | 2 | 2 | 2 | 2 |
| Stack Num Up | 2 | 2 | 2 | 2 |
| Patch Size | 4 | 16 | 16 | 16 |
| Number of Heads | 4, 8, 8, 8 | 4, 2, 8, 2 | 4, 1, 4, 2 | 8, 4, 2, 4 |
| Window Size | 4, 2, 2, 2 | 1, 1, 2, 2 | 8, 1, 2, 2 | 8, 1, 4, 1 |
| Num MLP | 512 | 46 | 158 | 46 |
| Gamma | 2 | 2.6326 | 1.7471 | 1.4890 |
| Alpha | 0.5 | 4.9448 | 4.9407 | 3.7244 |
| Dataset | Size (MB) | Acc. | Prec. | Rec. | Spec. | Dice | VOE | RVD(%) | |
|---|---|---|---|---|---|---|---|---|---|
| 3DIRCADb | unoptimized | 324.91 | 0.9985 | 0.8272 | 0.8488 | 0.9999 | 0.8340 | 0.1801 | 0.30 |
| 3DIRCADb | 0.2344 | 64.16 | 0.9998 | 0.9297 | 0.9915 | 1.0000 | 0.9478 | 0.0783 | 0.23 |
| 3DIRCADb | 0.1172 | 30.88 | 0.9976 | 0.7423 | 0.8401 | 0.9994 | 0.7891 | 0.2661 | 0.89 |
| 3DIRCADb | 0.0486 | 17.22 | 0.9962 | 0.7400 | 0.8329 | 0.9962 | 0.7644 | 0.2634 | 4.82 |
| LiTS | unoptimized | 324.91 | 0.9998 | 0.9797 | 0.9709 | 1.0000 | 0.8753 | 0.0291 | 2.89 |
| LiTS | 0.2344 | 64.16 | 0.9999 | 0.9923 | 0.9910 | 1.0000 | 0.8906 | 0.0166 | 2.51 |
| LiTS | 0.1172 | 30.88 | 0.9996 | 0.9288 | 0.9817 | 0.9998 | 0.8405 | 0.0791 | 3.17 |
| LiTS | 0.0486 | 17.22 | 0.9987 | 0.8724 | 0.9925 | 0.9988 | 0.7998 | 0.1345 | 16.8 |
| MSD Task03 | unoptimized | 324.91 | 0.9999 | 0.9712 | 0.9921 | 0.9999 | 0.8758 | 0.0366 | 19.33 |
| MSD Task03 | 0.2344 | 64.16 | 0.9999 | 0.9906 | 0.9920 | 1.0000 | 0.8895 | 0.0173 | 4.63 |
| MSD Task03 | 0.1172 | 30.88 | 0.9999 | 0.9569 | 0.9921 | 0.9999 | 0.8654 | 0.0508 | 27.39 |
| MSD Task03 | 0.0486 | 17.22 | 0.9964 | 0.8124 | 0.9933 | 0.9965 | 0.8363 | 0.1938 | 48.04 |
| Dataset | Method/Scheme | Dice (%) | VOE (%) | RVD (%) |
|---|---|---|---|---|
| 3DIRCADb | DefED-Net [43] | 66.2 | 34.3 | 0.8 |
| X-net [44] | 69.1 | 36.1 | 0.7 | |
| TD-Net [24] | 68.2 | 40.8 | 8.4 | |
| MS-FANet [21] | 78.0 | 31.3 | 15.5 | |
| Lgma-net [45] | 83.2 | 24.3 | 0.76 | |
| MS-UNet [46] | 84.1 | 27.3 | 0.22 | |
| MAPFUNet [47] | 85.9 | 23.7 | 0.22 | |
| Proposed Scheme | 94.78 | 7.83 | 0.23 | |
| LiTS | TD-Net [24] | 70.9 | 39.6 | 11.7 |
| MS-FANet [21] | 74.2 | 36.7 | 10.7 | |
| X-net [44] | 76.4 | - | - | |
| MAPFUNet [47] | 85.8 | 22.0 | 11.02 | |
| Lgma-net [45] | 87.4 | 23.1 | 5.72 | |
| DefED-Net [43] | 87.52 | 23.85 | 5.22 | |
| Proposed Scheme | 89.06 | 1.66 | 2.51 | |
| MSD Task03 | S. Muhammad et al. [48] | 87.0 | 12.09 | 6.39 |
| Proposed Scheme | 88.95 | 1.73 | 4.63 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




