1. Introduction
Liver cancer (LC) is a significant global health concern, ranking as one of the most commonly diagnosed cancers and the leading cause of cancer-related deaths. Recent global statistics position liver cancer as the sixth most prevalent cancer, following breast, lung, colorectal, prostate and gastric cancers and as the third leading cause of cancer mortality. In 2020, approximately 905,700 new cases and 830,200 deaths from LC occurred worldwide [
1]. Hepatocellular carcinoma (HCC), the most common form of LC, accounts for nearly 90% of all cases [
2]. Various imaging techniques such as ultrasound, elastography, MRI and CT scans detect LC, with CT scans providing detailed images of internal structures [
3]. However, CT imaging encounters challenges in detecting and diagnosing liver abnormalities accurately. The liver’s varying sizes and shapes cause incorrect segmentation due to the similarity in intensity between tumors and surrounding tissues and unclear lesion boundaries. These factors make manual annotation by radiologists time-intensive and error-prone, leading to inconsistencies in diagnosis.
Medical image segmentation achieves progress in improving cancer diagnosis accuracy and computer-aided diagnosis (CAD) systems assist in the detection, classification, and segmentation of tumours on medical images, reduces radiologists’ workload and increases diagnostic consistency [
4,
5,
6,
7,
8,
9]. Deep learning models, particularly Convolutional Neural Networks (CNNs), automate this segmentation process. Fully Convolutional Networks (FCNs) and U-Net architectures also segment liver tumors by performing pixel-wise classification [
10]. Despite their high accuracy compared to manual segmentation, such models require significant computational resources, complicating deployment on edge devices with limited processing power and memory. Large servers are necessary to run these models, causing issues like bandwidth usage, data security, high server costs and substantial carbon footprints due to increased energy consumption compared to resource-constrained embedded systems. Thus, resource-efficient models are needed for accurate liver tumor segmentation on constrained devices.
Liver tumor segmentation techniques include traditional image processing, supervised learning and unsupervised learning methods. Traditional image processing techniques, such as thresholding [
11], Canny edge detection [
12] and watershed segmentation [
13], rely on edge detection and intensity thresholding to differentiate tumor regions from normal tissue. Thresholding separates objects based on intensity, while watershed segmentation uses gradients to define boundaries. These methods struggle with the complexity of medical images, where tumors exhibit irregular shapes, variable sizes and similar intensity values to surrounding tissues, leading to errors [
14]. Additionally, traditional methods often require manual or semi-automatic intervention, increasing dependency on expert input for accuracy.
Unsupervised learning methods address limitations in traditional techniques by segmenting tumors without labeled data. Notable methods include clustering-based techniques and edge-based algorithms. For instance, Al-Kofahi et al. [
15] introduced a multi-scale Laplacian of Gaussian (LoG) filter for histopathology images to detect nuclei of varying sizes. Kong et al. [
16] developed a generalized LoG filter (gLoG) to detect elliptical nuclei in histopathology images, which can apply to LC segmentation in CT scans. Despite their potential, unsupervised methods require careful parameter tuning, are sensitive to noise and struggle to define tumor boundaries with low contrast [
17]. Combining unsupervised techniques with region-based methods, such as Active Contour Models (ACM) [
18] and marker-based watershed transforms [
19], improves segmentation accuracy but remains computationally expensive and less generalizable across datasets.
CNNs, FCNs and U-Net variants dominate the field of medical image analysis due to their ability to learn hierarchical features from complex medical images. For instance, Saha Roy et al. [
20] proposed an automated model that utilizes Mask R-CNN followed by Maximally Stable Extremal Regions (MSER) for tumor identification, enabling multi-class tumor classification. Chen et al. [
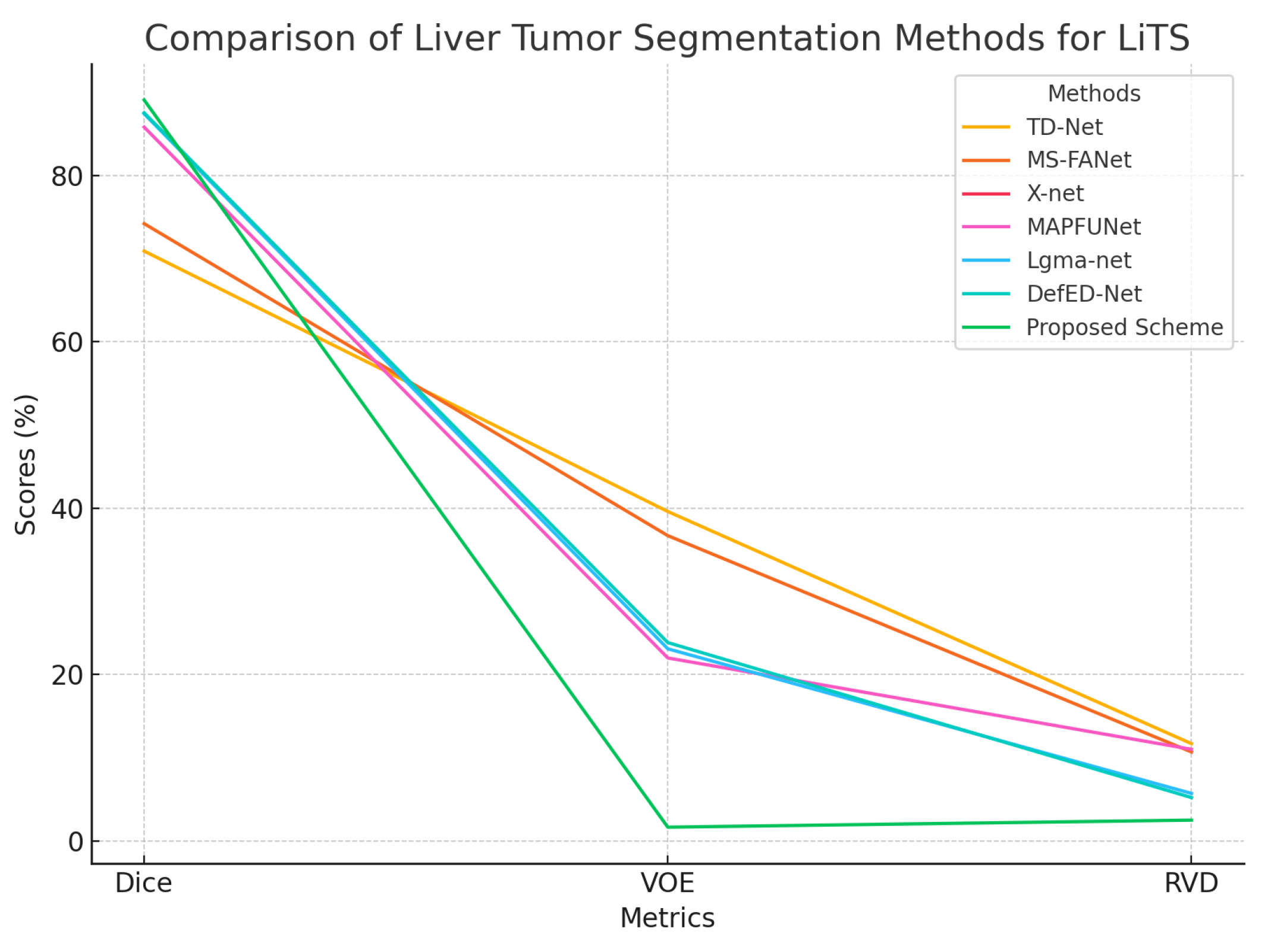
21] proposed MS-FANet, a multi-scale feature attention network that performs liver tumor segmentation through multi-scale attention mechanisms which boost segmentation capabilities while capturing both global and local context. Lakshmi et al. [
22] designed the Adaptive SegUnet++ (ASUnet++) framework and optimized it with the Enhanced Lemurs Optimizer (ELO) for tumor segmentation and classification. The authors’ model tackles traditional machine learning hurdles including slow training times and gradient explosion issues as well as overfitting using both residual connections and multiscale approaches. Reyad et al. [
23] proposed an architecture optimization framework for hybrid deep residual networks in liver tumor segmentation, utilizing a Genetic Algorithm (GA) to improve segmentation accuracy and model efficiency. Di et al. [
24] developed a framework for automatic liver tumor segmentation which integrates 3D U-Net architecture with hierarchical superpixels and SVM-based classification, achieving robust performance on noisy and low-contrast CT images. Liu et al. [
25] introduced PA-Net, a phase attention network that fuses venous and arterial phase features of CT images for liver tumor segmentation, effectively leveraging phase-specific information to enhance segmentation performance. CNN-based approaches have shown powerful representation abilities together with resilience to different image appearances. However, CNNs are inherently limited in modeling long-range dependencies, which can lead to suboptimal segmentation outcomes. Specifically, the localized receptive fields of convolutional operations restrict the network’s focus to local context rather than global context [
26].
Transformers which were initially created for sequence-to-sequence prediction tasks now play a primary role in computer vision tasks. Transformers demonstrate outstanding performance across multiple computer vision tasks including image classification [
27], object detection [
28], semantic segmentation [
29] and generative tasks like text-to-image synthesis [
25]. Transformers achieve success because their self-attention mechanism provides large receptive fields and long-range dependency capturing abilities. Medical image segmentation tasks have seen multiple proposals for hybrid methods that integrate both CNNs and Transformers. For instance, Balasubramanian et al. [
30] proposed APESTNet, a Mask R-CNN-based Enhanced Swin Transformer Network for tumor segmentation and classification. This method combines the strengths of Mask R-CNN with the attention mechanisms of the Swin Transformer to improve segmentation accuracy. Chen et al. [
31] introduced TransUNet, a cascaded architecture that integrates CNN and Transformer modules to enhance segmentation performance. Ni et al. [
32] presented DA-Tran, a domain-adaptive transformer network for multiphase liver tumor segmentation. DA-Tran leverages domain adaptation techniques to effectively integrate multiphase CT images, improving segmentation accuracy and robustness across varying imaging conditions.
Despite their accuracy, existing models require substantial memory and computational power, which are unsuitable for edge devices like Jetson Nano. These models typically operate on server systems, demanding sensitive patient data transfer over the Internet, raising privacy and security concerns. High server energy consumption limits feasibility in resource-constrained settings. Optimizing segmentation models to reduce size and power consumption while maintaining accuracy enables deployment on edge devices for real-time, secure and energy-efficient tumor segmentation.
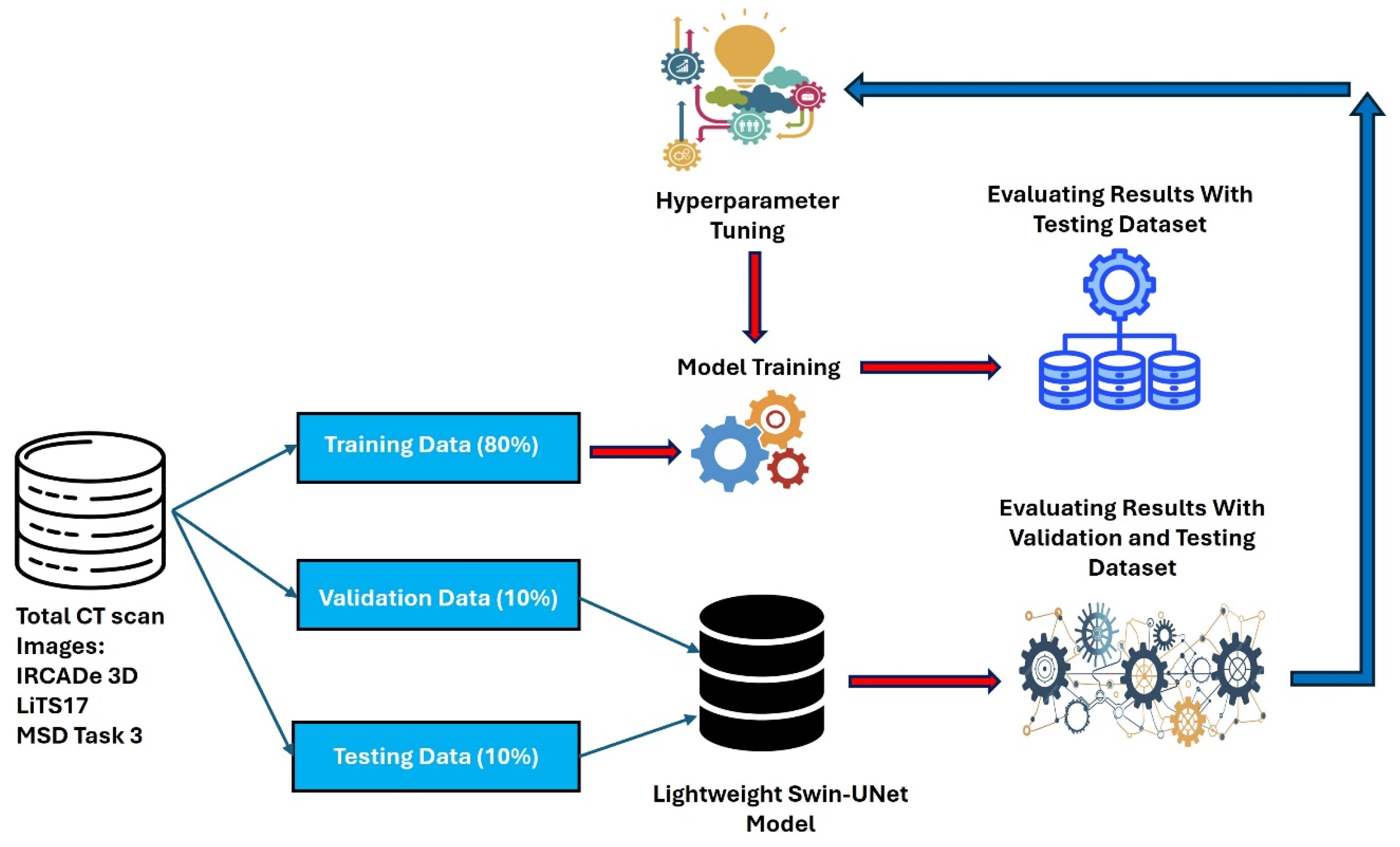
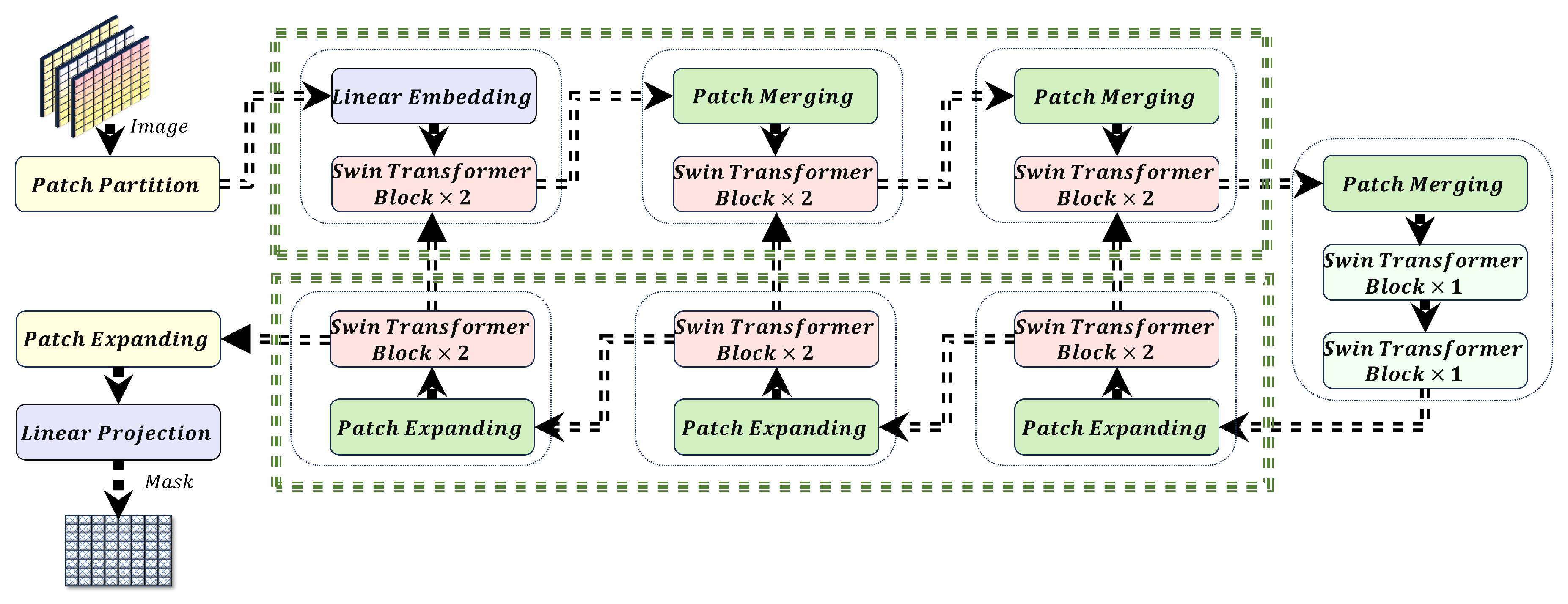
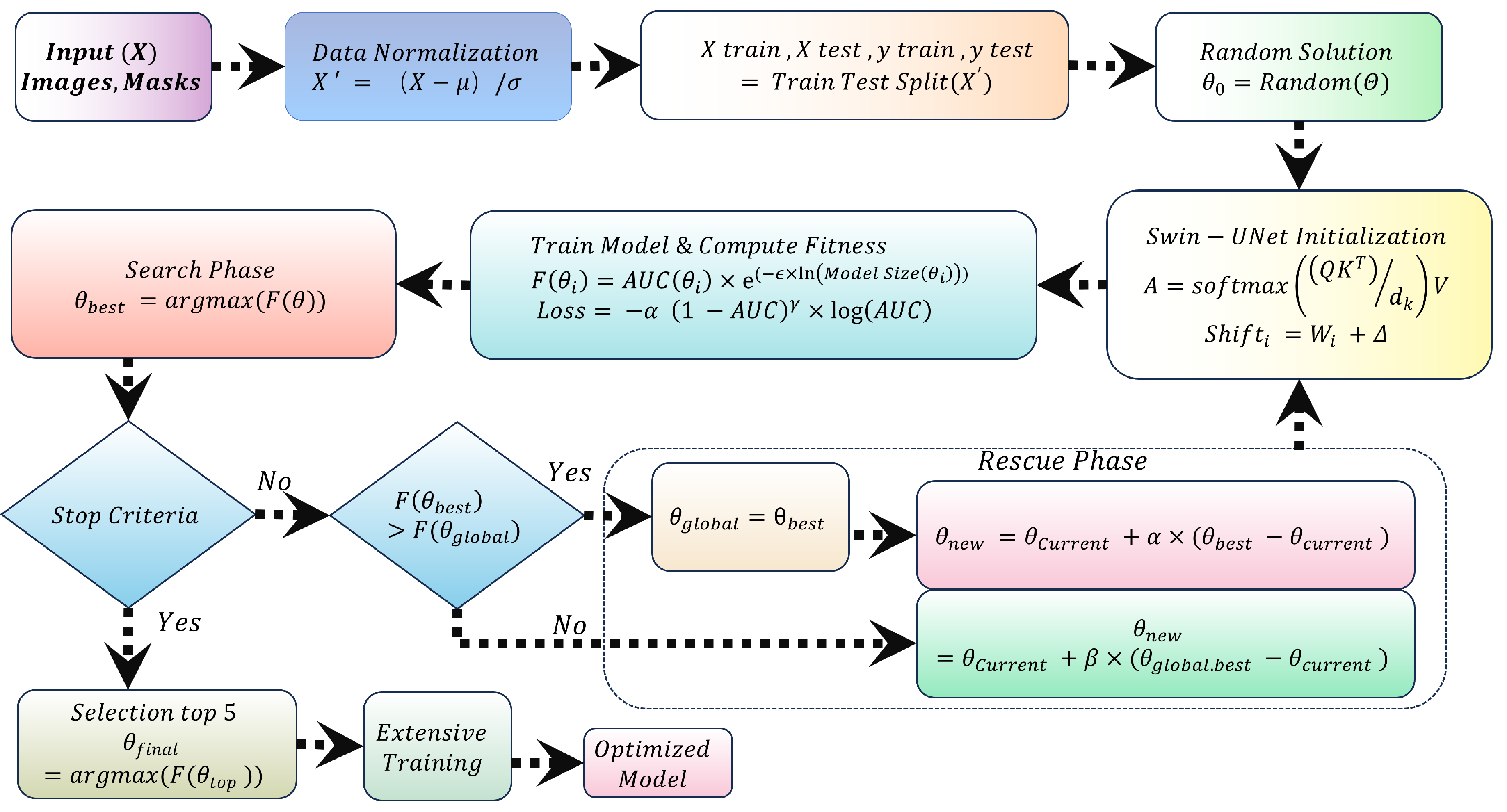
This study proposes a novel approach to optimize the Swin-UNet model for efficient liver cancer segmentation on edge devices, balancing model size and Area Under the Curve (AUC). Contributions include:
Model Size Optimization: The discrete design space of Swin-UNet achieves a balance between model size and accuracy, enabling deployment on memory-constrained devices like Jetson Nano.
Quadratic Penalty Objective Function: A quadratic penalty-based objective function balances model size and AUC, encouraging compact, accurate models.
Search and Rescue Algorithm: The Search and Rescue algorithm identifies optimal configurations, yielding an optimized model termed SAR-Swin-UNet.
Focal AUC Loss Function: The Focal AUC loss function addresses class imbalance during training, enhancing the model’s ability to segment minority class pixels.
This approach facilitates accurate tumor segmentation on edge devices, ensuring real-time analysis with data security and energy efficiency.