Submitted:
28 February 2025
Posted:
28 February 2025
You are already at the latest version
Abstract
Keywords:
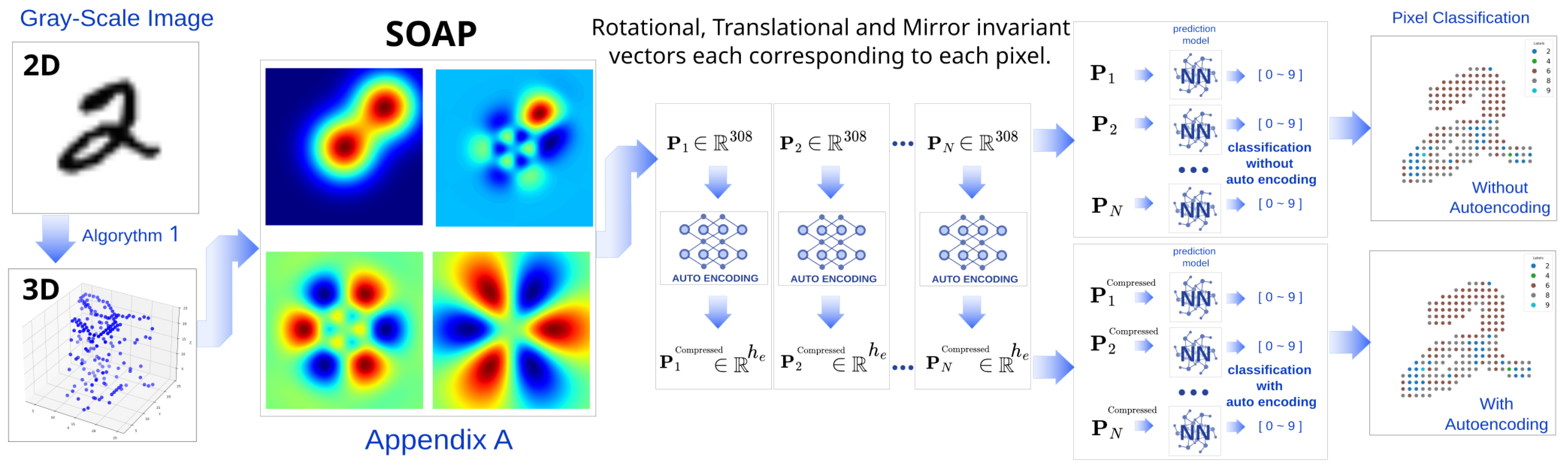
1. Introduction
- Translation invariance: Images may undergo shifts in spatial position, causing pixel values to move across the image grid. Training models to handle translation typically involves augmenting the dataset with translated versions of the original images.
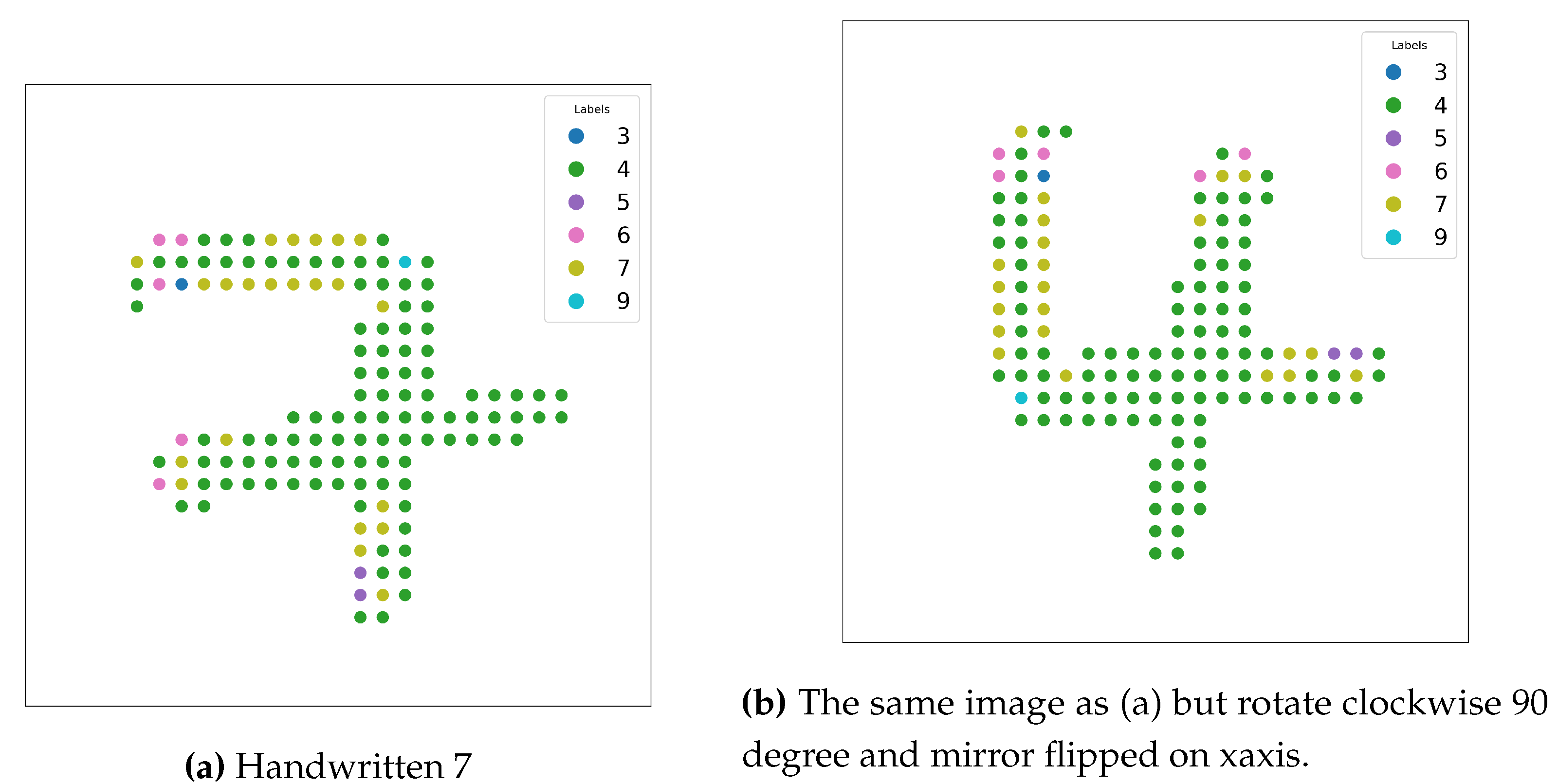
- Rotation invariance: Images can appear in different orientations. Achieving robustness to rotations requires augmenting the dataset with rotated images, increasing computational cost and memory requirements.
- Mirror symmetry: Certain images may appear as mirror reflections. Training models to handle such transformations often involves flipping the images horizontally or vertically, further expanding the dataset.
2. Related Work
3. Methodology
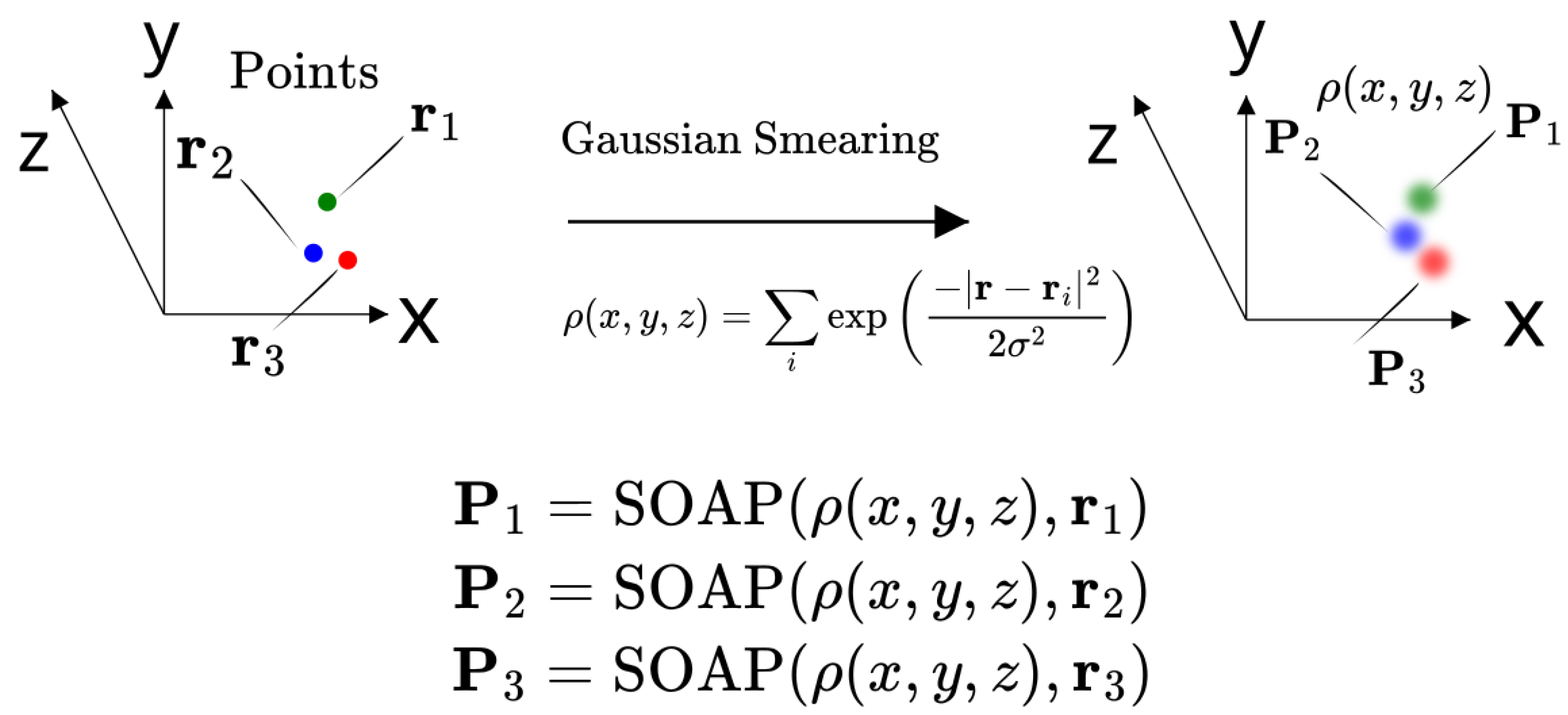
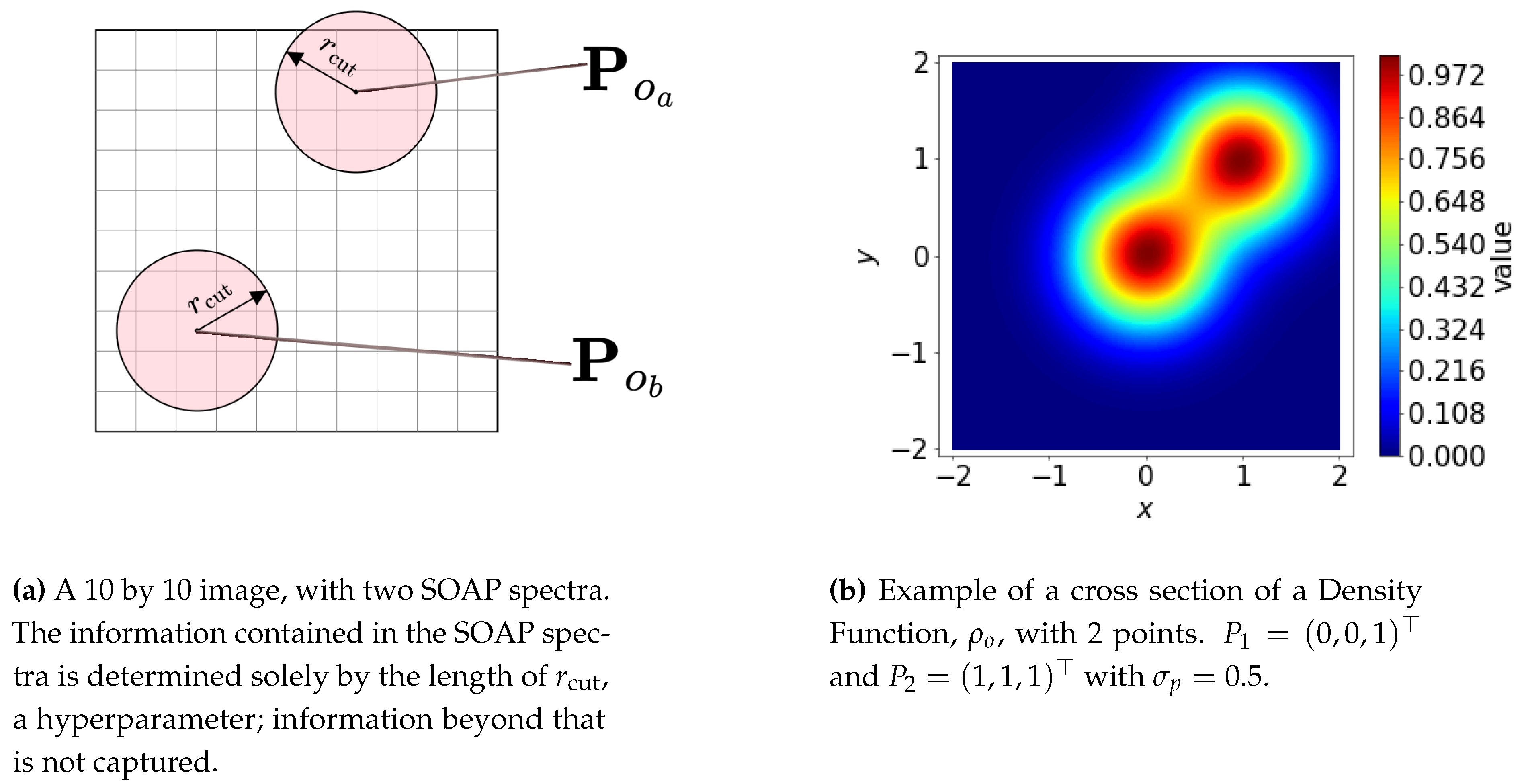
3.1. SOAP Formulation
3.1.1. Density Function
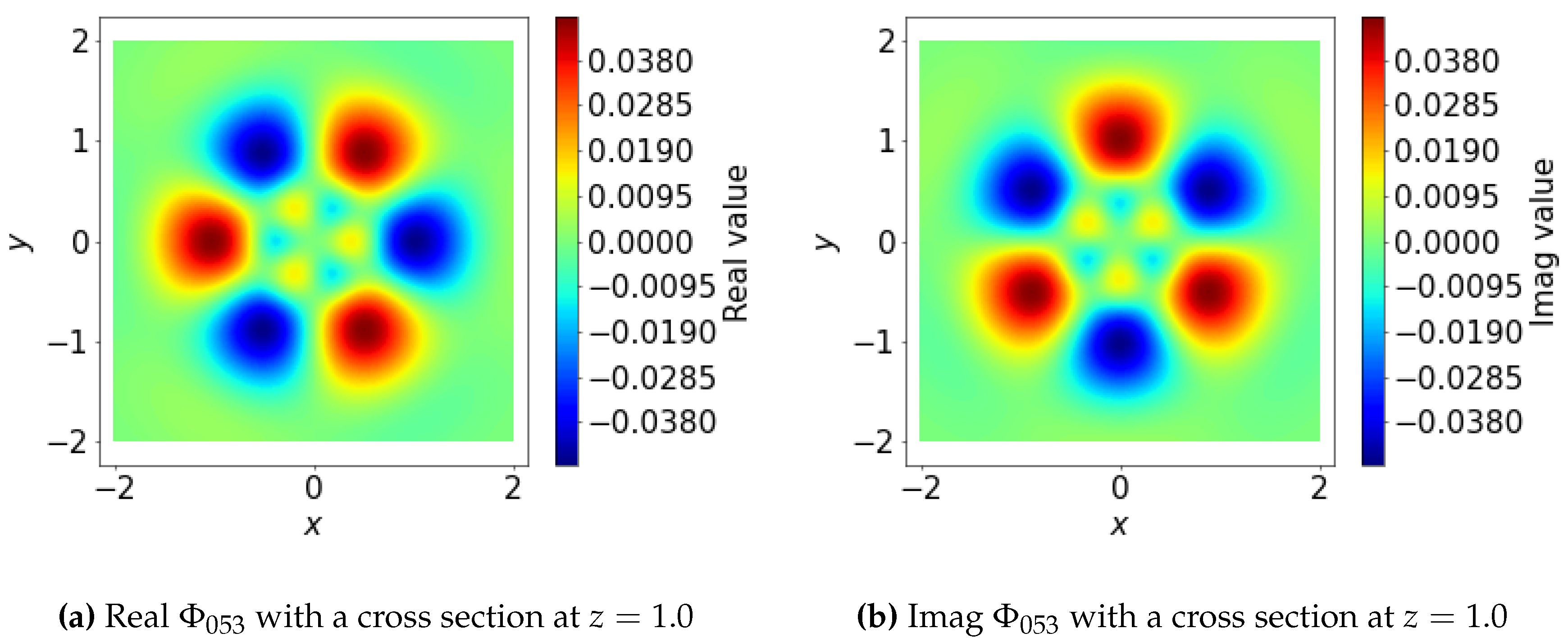
3.1.2. Spatial Basis Function
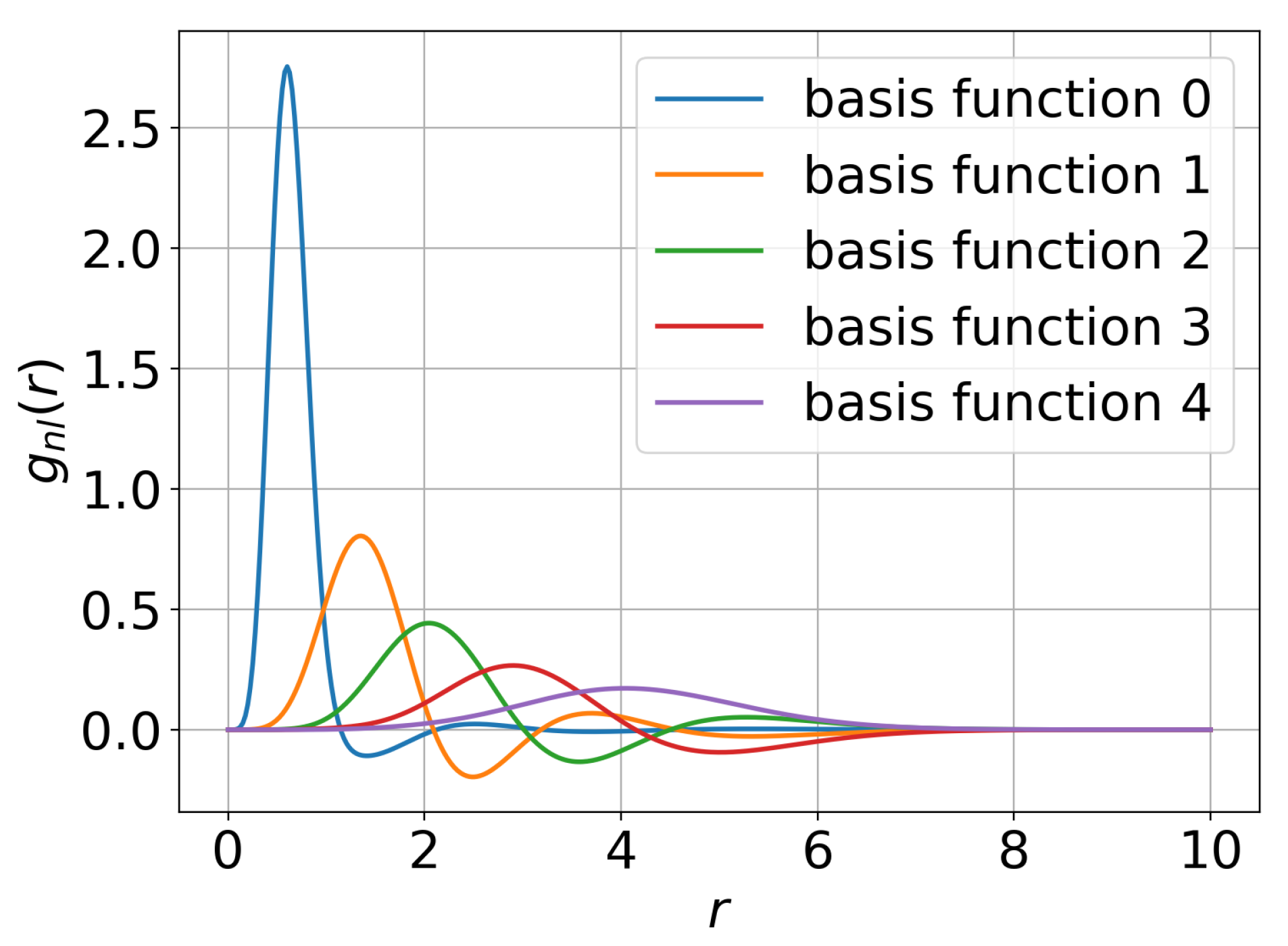
3.1.3. Radial Basis Functions
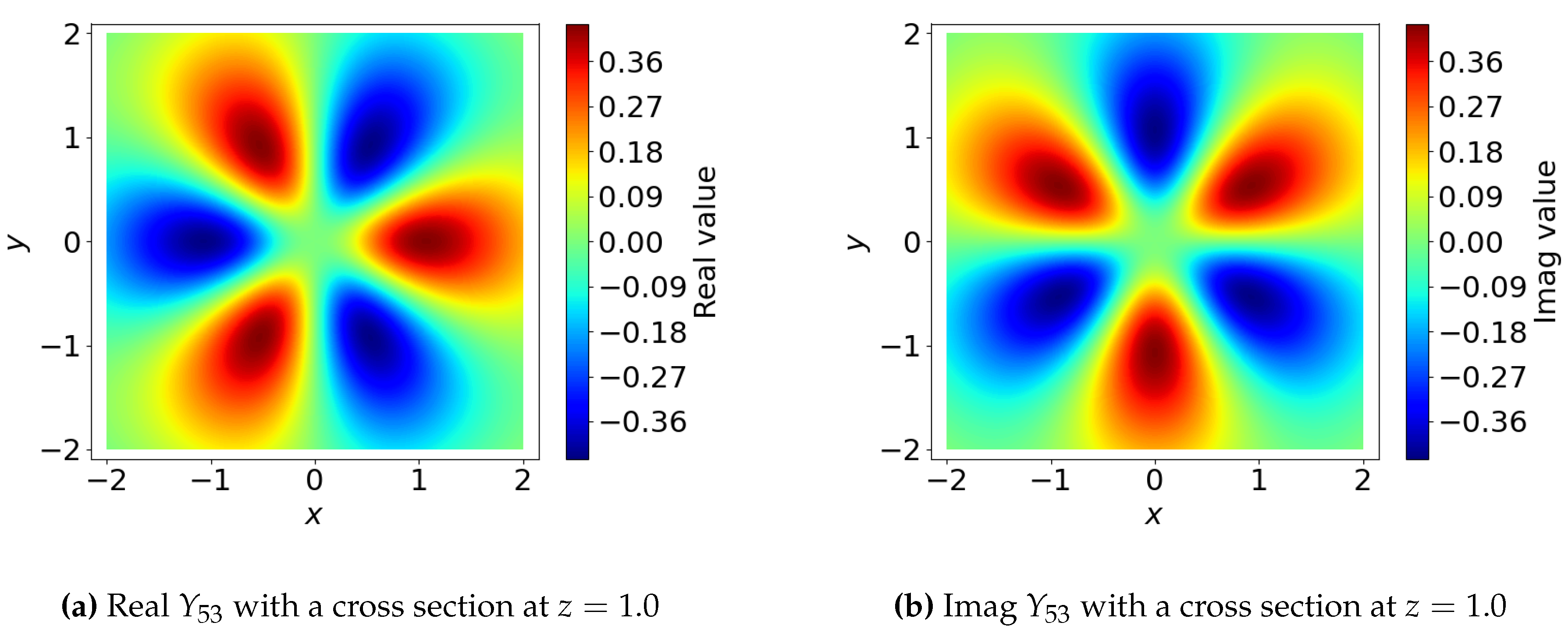
3.1.4. Angular Basis Functions
3.1.5. SOAP Expansion Coefficients
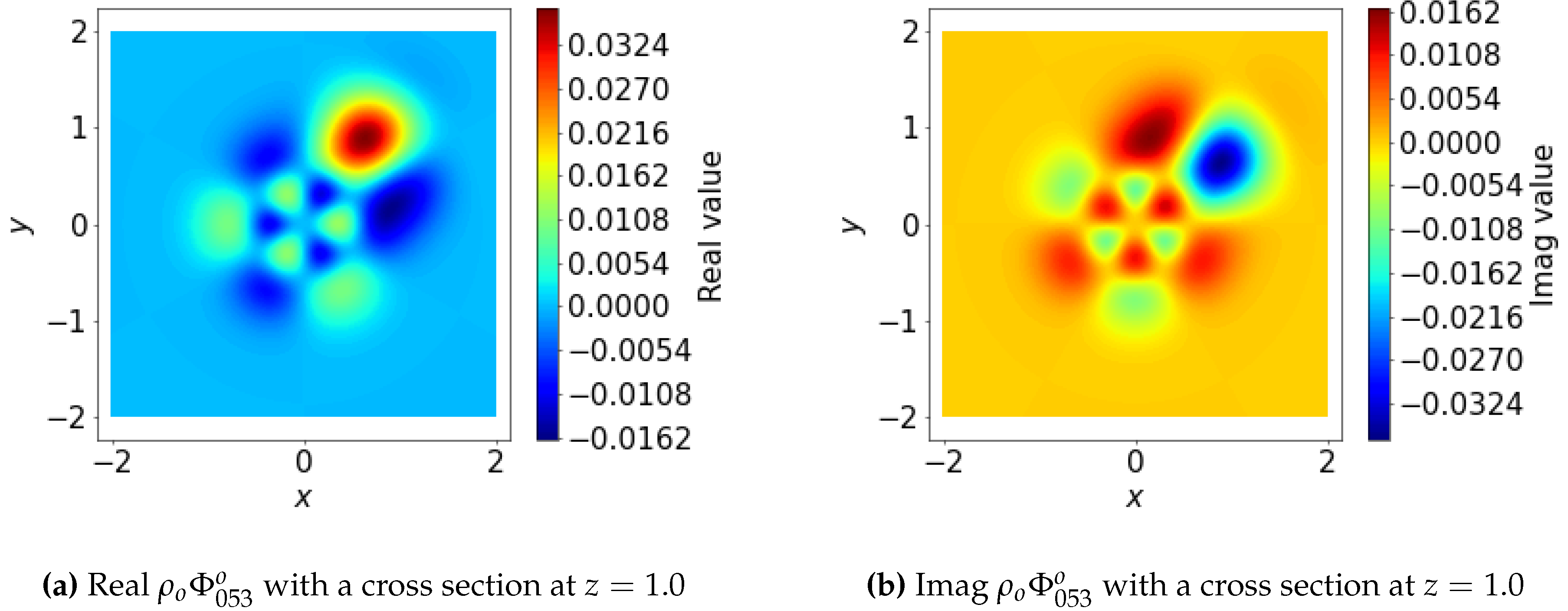
3.1.6. SOAP Power Spectrum
3.2. Converting Images to 3D Points and Computing SOAP Descriptors
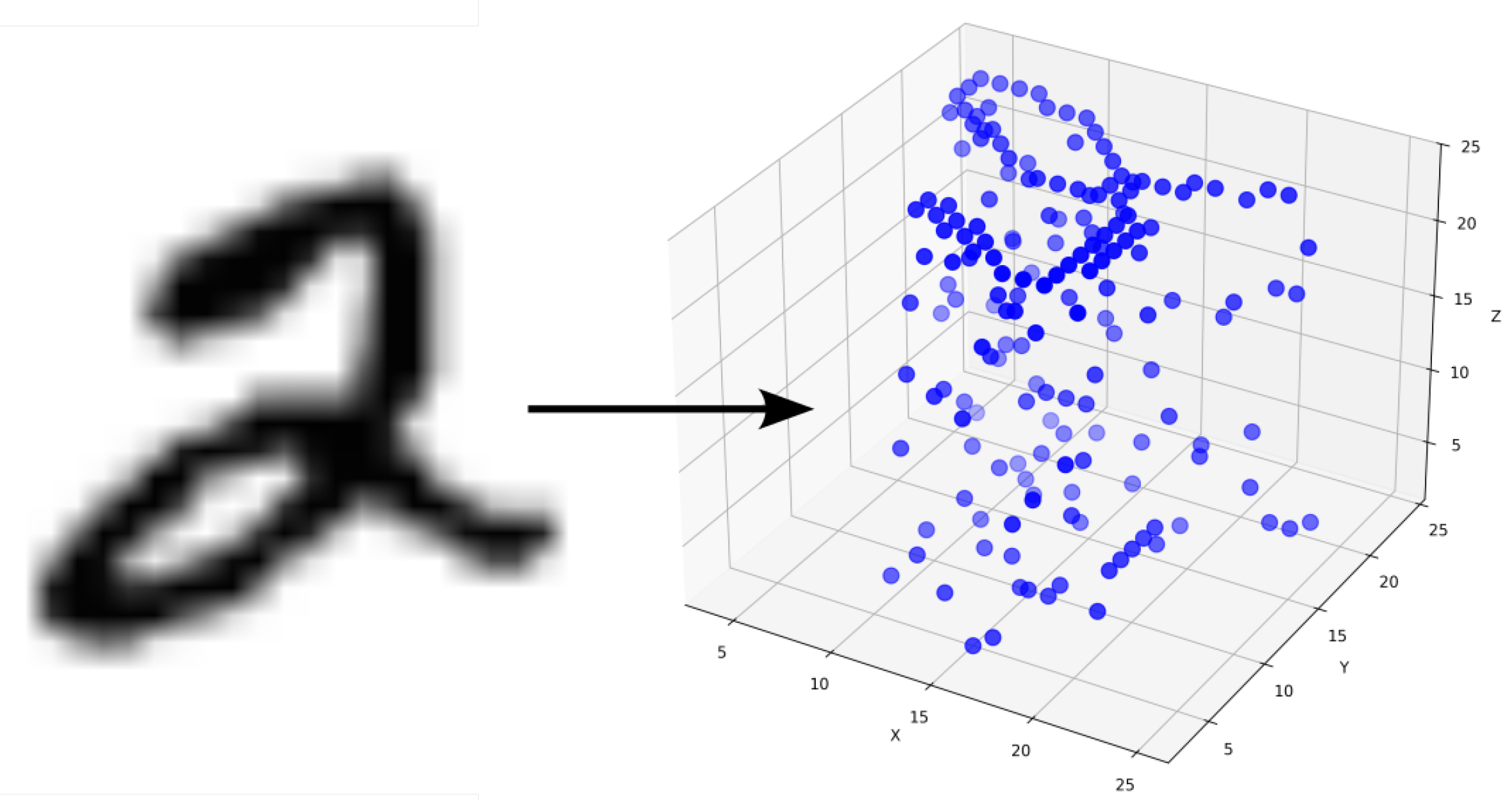
3.2.1. Converting Gray-Scale Images to 3D Points
| Algorithm 1: ConvertImagesToXYZ (3D structure) |
|
Require: ● : A collection of n images, each of size .
● : Standard deviation for Gaussian displacement. Ensure: ● Points derived from the intensity values of each pixel in each image, with optional random displacement.
|
3.2.2. Computing SOAP Descriptors for Image 3D Structures
| Algorithm 2:Compute SOAP Matrices for a Set of 3D Structures |
|
Require: ● A collection of 3D structures , where each structure is an matrix containing coordinates in 3D space: ● Parameters for the SOAP descriptor: . Ensure: ● A set of SOAP matrices , where each is an matrix holding the per-point SOAP vectors for . In other words, ▹Algorithm Steps
|
4. Experiments and Results
4.1. Training Data Preparation
| Algorithm 3: Random Extraction of SOAP Spectra with Rescaling |
|
Require:
Collection of SOAP vectors from 3D structures, . Ensure: Extracted and rescaled SOAP descriptors , and corresponding labels .
|
4.2. Experiment 1: Hyperparameter Optimization for SOAP and Predictions
4.2.1. Objective
4.2.2. Methods
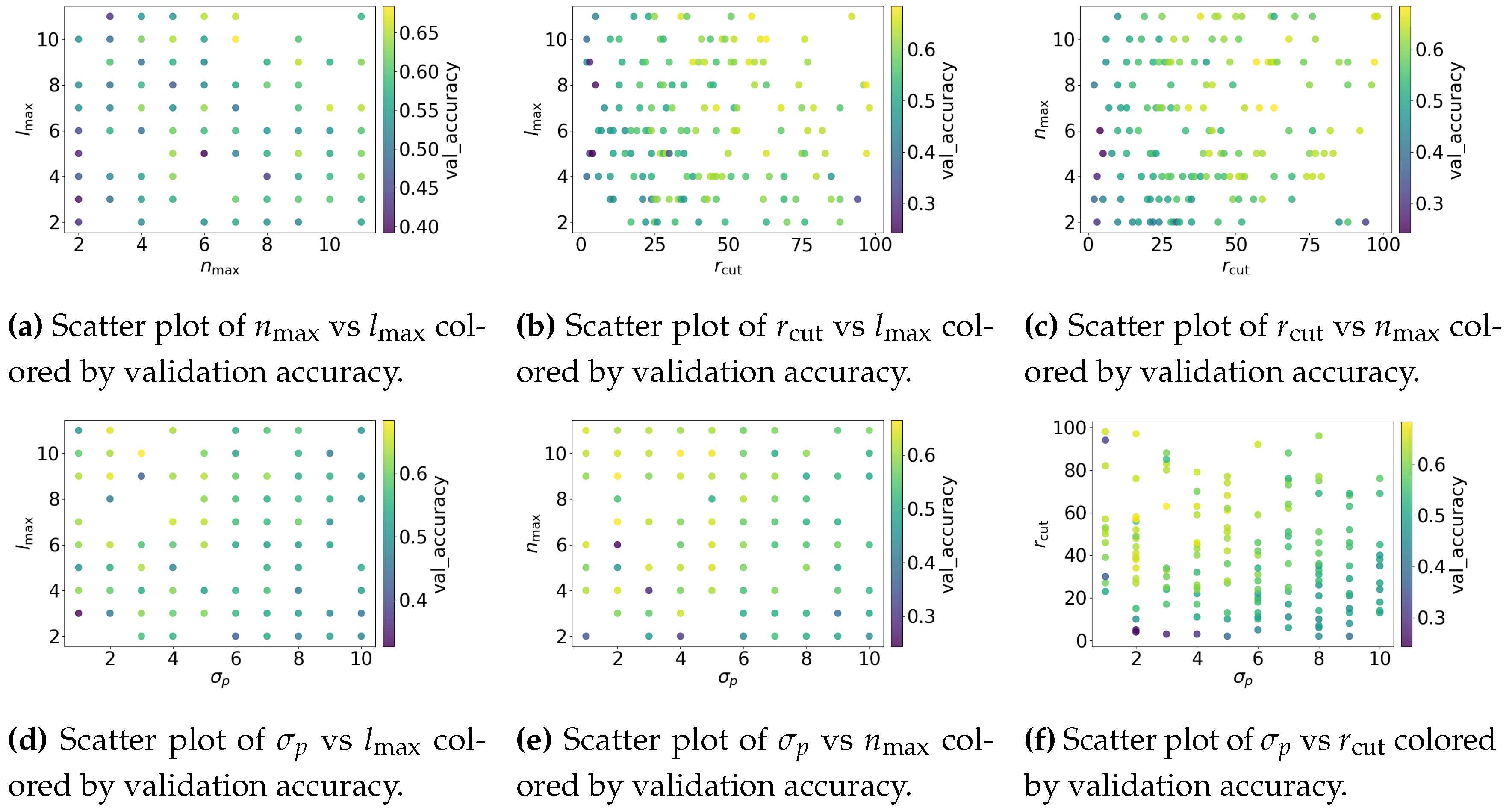
4.2.3. Results
4.2.4. Discussion
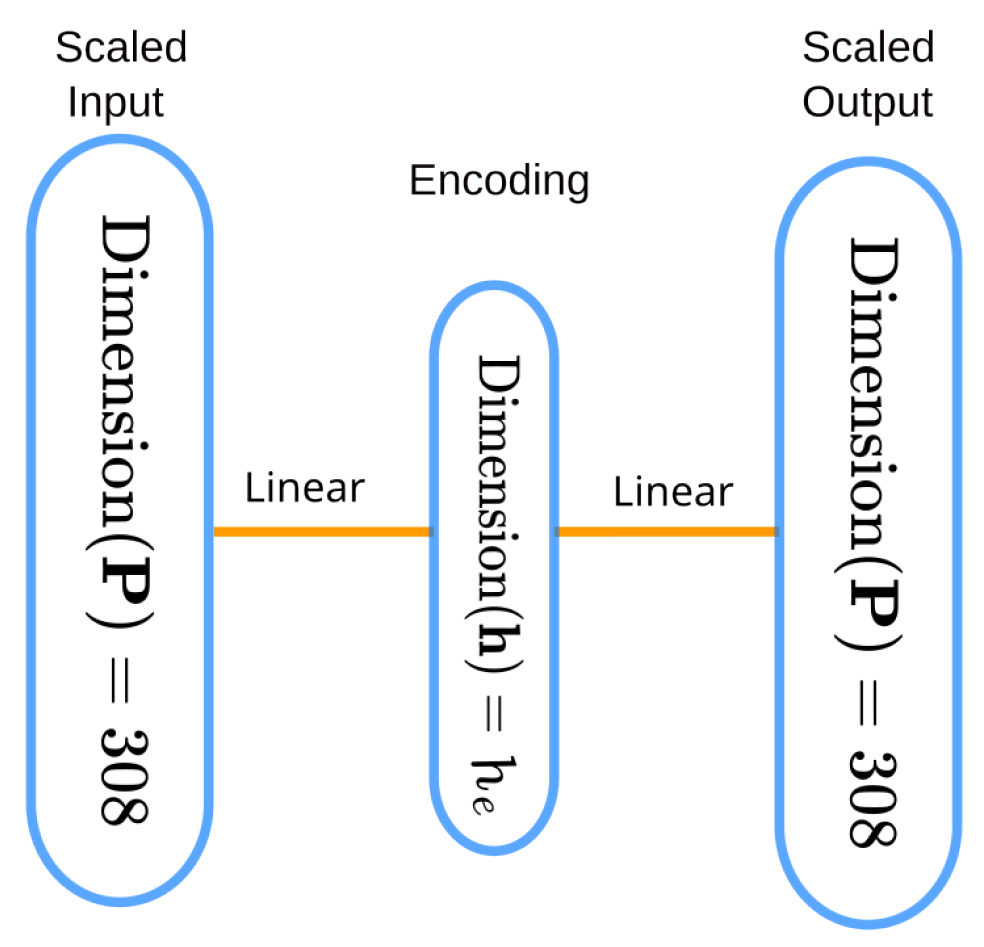
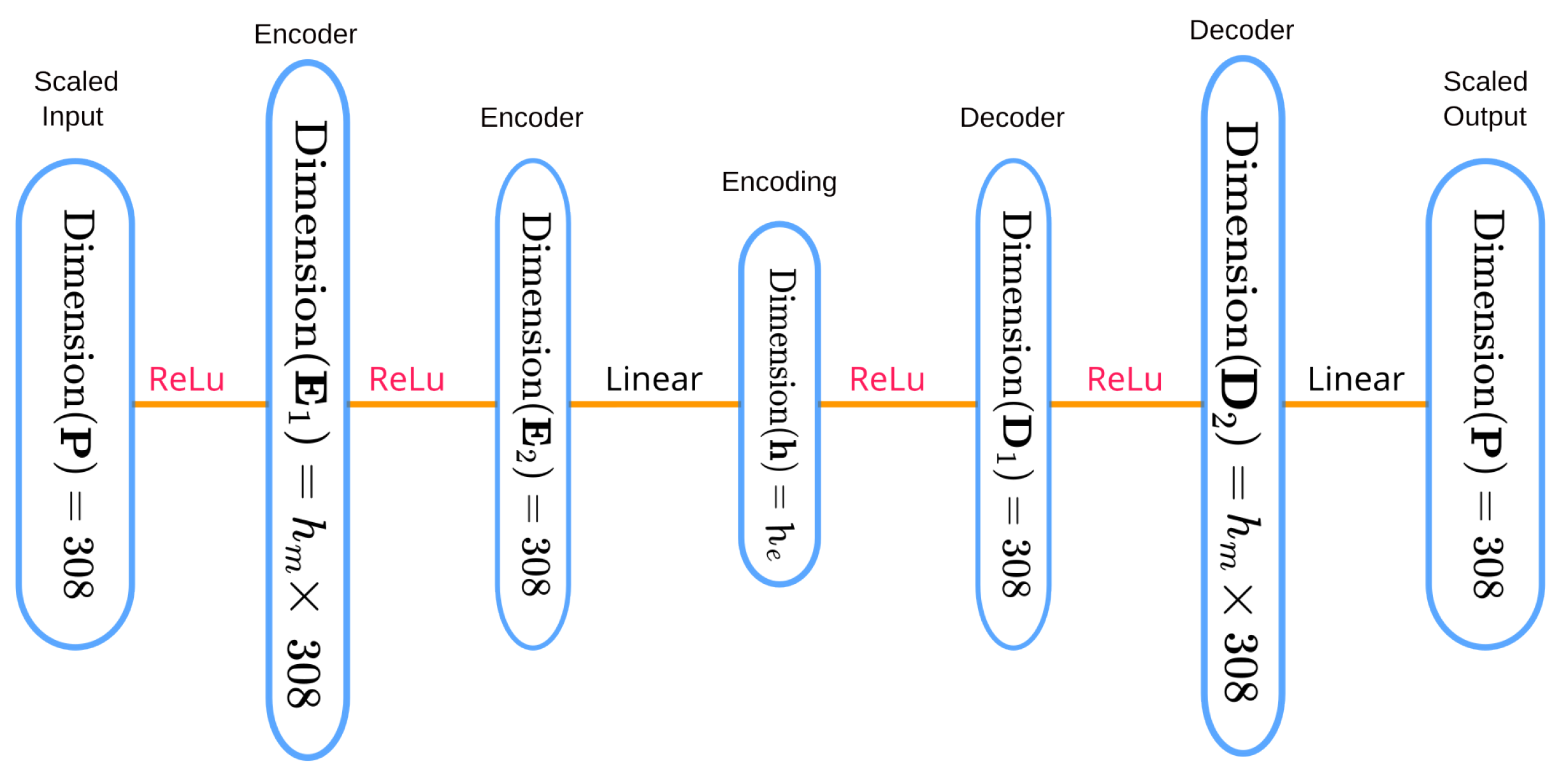
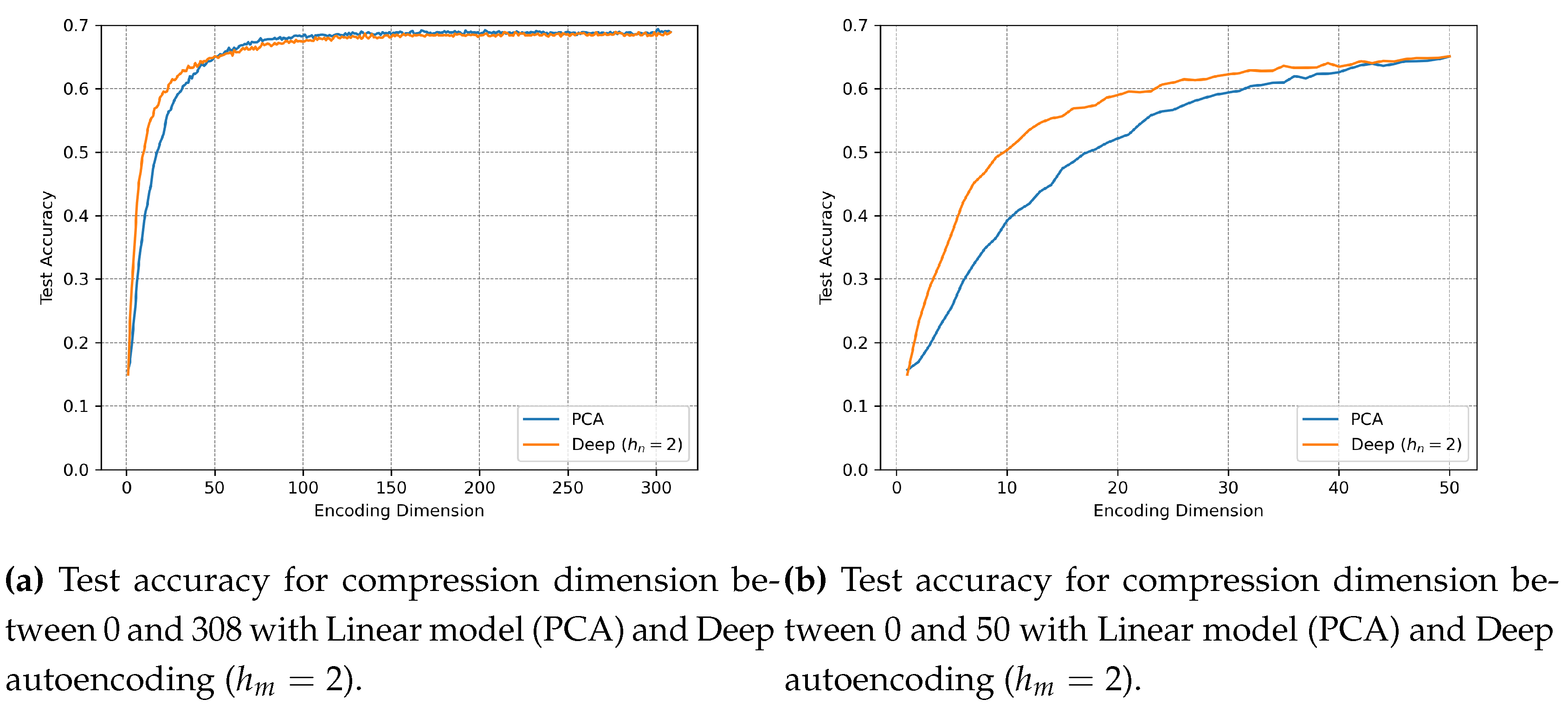
4.3. Experiment 2: SOAP Vector Compression and Impact on Prediction Accuracy
4.3.1. Objective
4.3.2. Methods
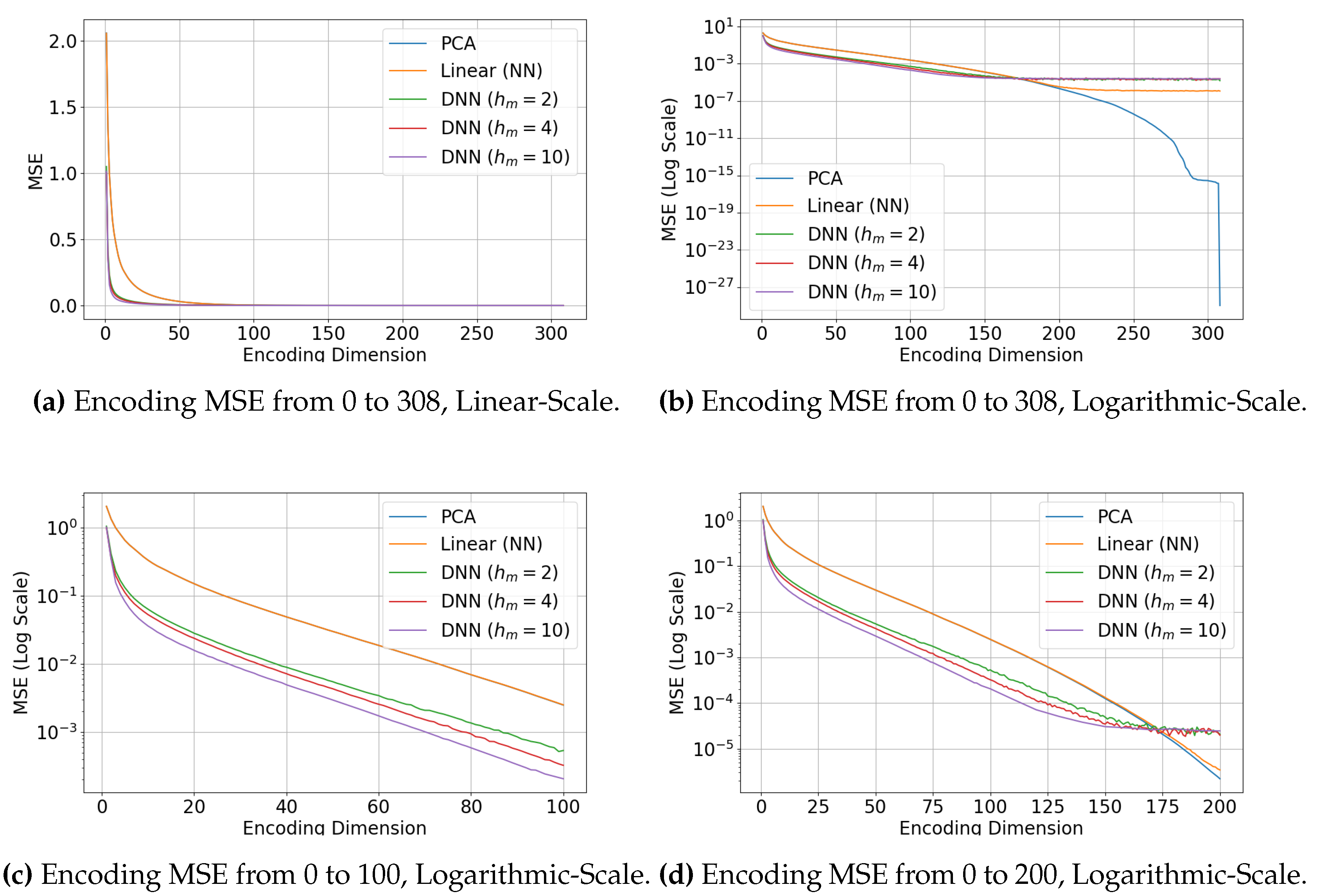
4.3.3. Results
4.3.4. Discussion
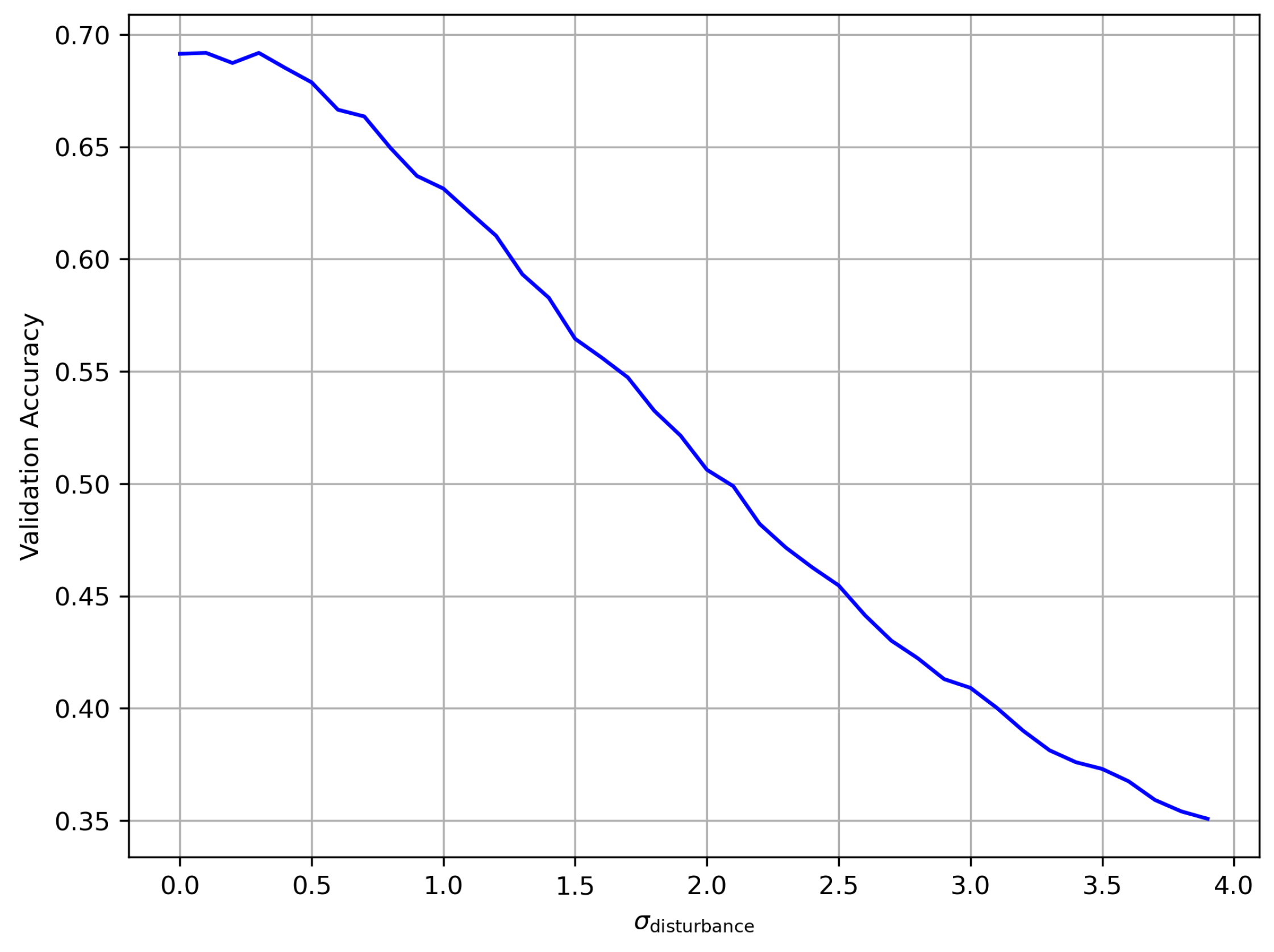
4.4. Experiment 3: Robustness to Pixel Position Perturbations
4.4.1. Objective
4.4.2. Methods
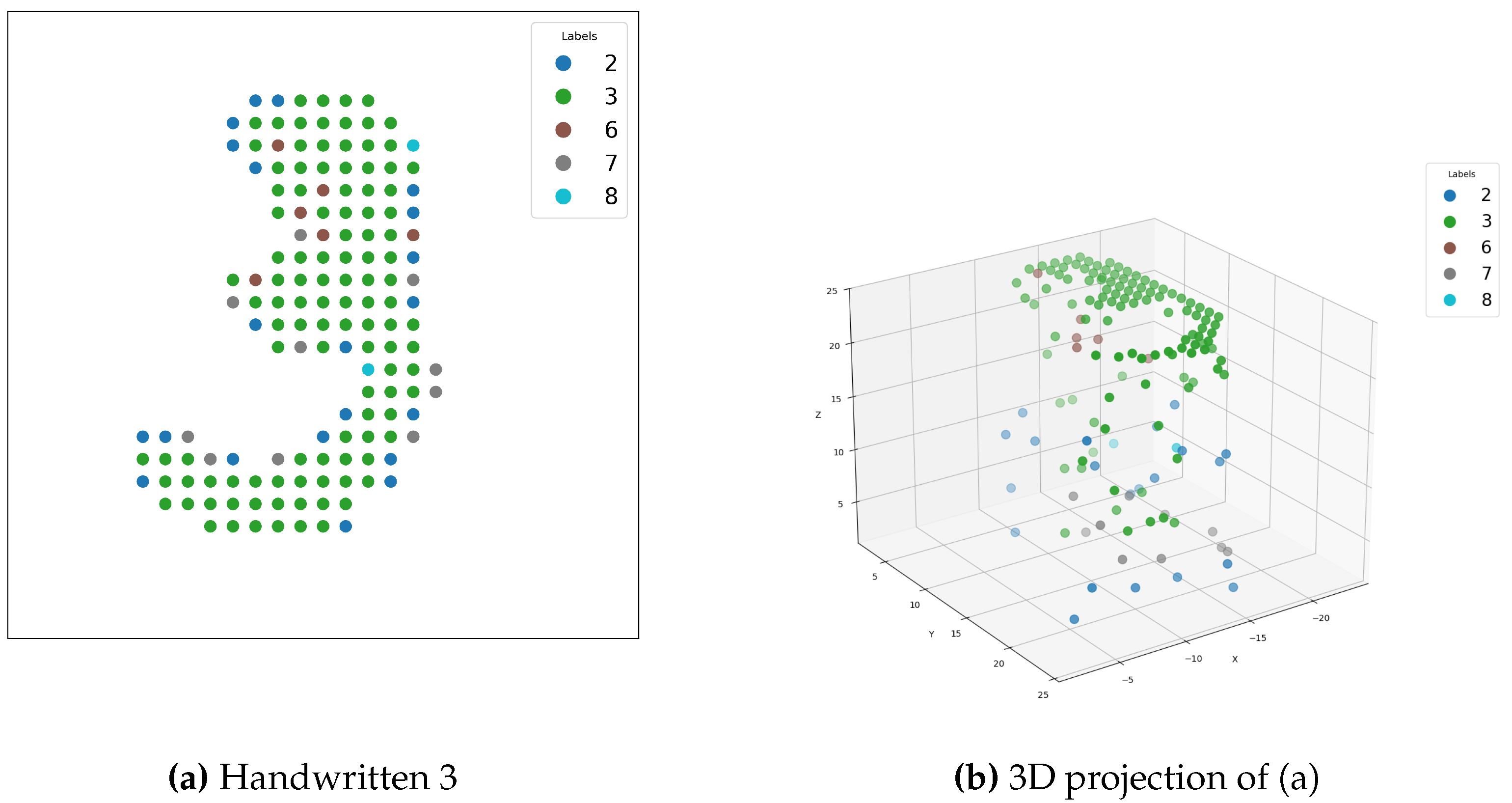
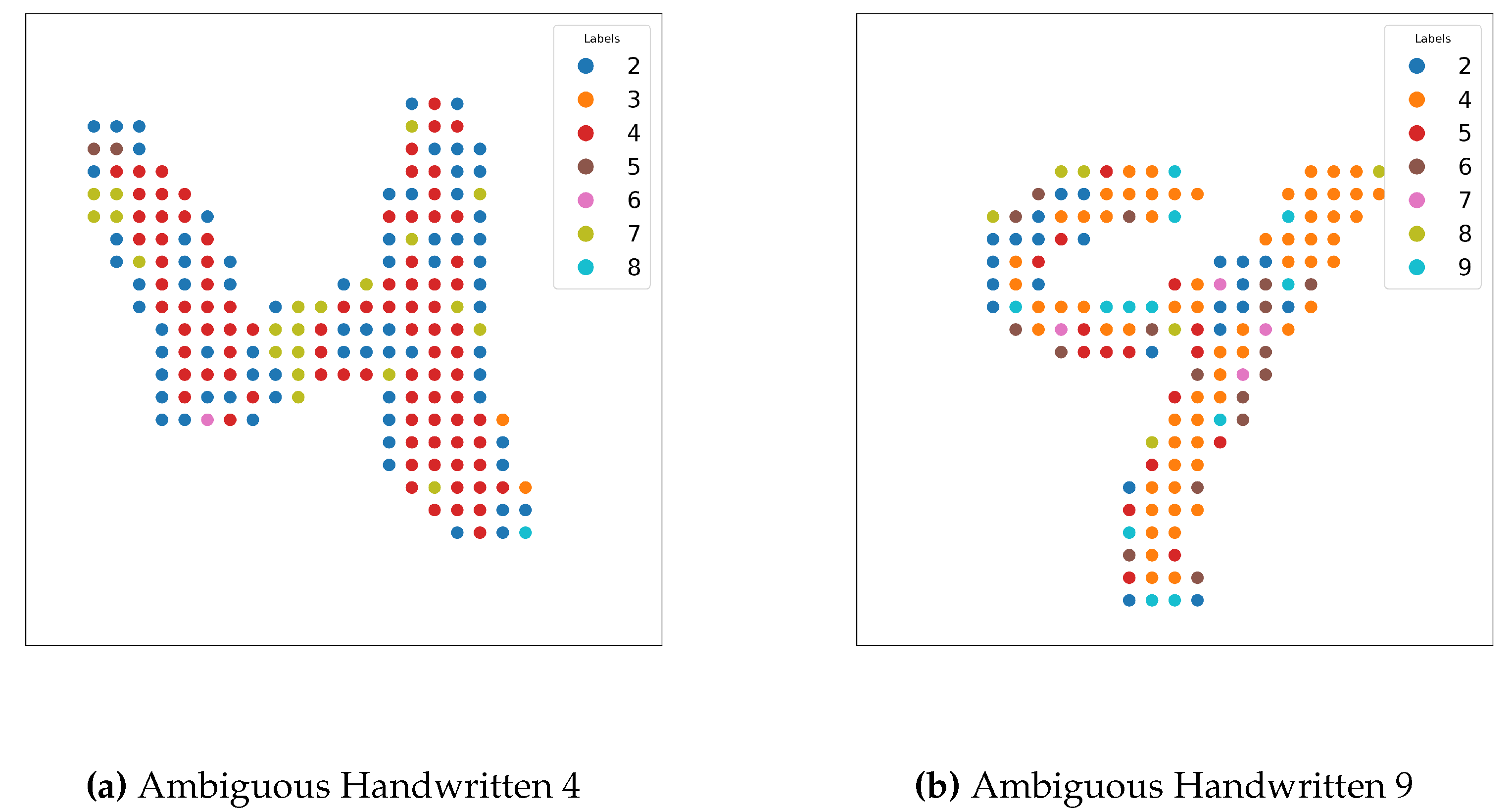
4.4.3. Results
4.4.4. Discussion
5. Future Work
6. Conclusions
Data Availability Statement
Appendix A. Example of C’s

| Algorithm 4: GetBasisGTO () |
|
Require: ● : The radial cutoff distance.
● : The number of GTO radial basis functions. ● : The maximum angular momentum quantum number. Ensure: ● : A array of radial decay exponents. ● : A array of Löwdin-orthonormalization factors.
|
Appendix B. Table of Variables
| Variable | Type/Dimension | Description |
|---|---|---|
| Collection of matrices | 3D structure. | |
| matrix | The k-th 3D structure containing coordinates of each 3D-pixel. | |
| Scalars | Parameters defining the SOAP descriptor computation. | |
| matrix | SOAP descriptors for each 3D-pixel in the k-th structure. | |
| vector | SOAP descriptor for the o-th 3D-pixel in structure . | |
| k | Integer | Index for iterating over each structure (). |
| o | Integer | Index for iterating over each 3D-pixel within a structure (). |
| Collection of matrix | Output set of SOAP descriptors for all structures and their 3D structure. |
| Variable | Type/Dim | Description |
|---|---|---|
| Final collection of extracted and rescaled SOAP descriptors, where in our case for the training data and validation data, and for the test data. | ||
| T | Labels for each row of . | |
| A single descriptor randomly chosen from . | ||
| p | Robust Rescale Parameters for later use. |
References
- Quiroga, F.; Ronchetti, F.; Lanzarini, L.; Bariviera, A. F. Revisiting data augmentation for rotational invariance in convolutional neural networks. In Modelling and Simulation in Management Sciences: Proceedings of the International Conference on Modelling and Simulation in Management Sciences (MS-18); Springer, 2020; pp. 127–141. [Google Scholar]
- Maharana, K.; Mondal, S.; Nemade, B. A review: Data pre-processing and data augmentation techniques. Global Transitions Proceedings 2022, 3, 91–99. [Google Scholar] [CrossRef]
- Omae, Y.; Saito, Y.; Fukamachi, D.; Nagashima, K.; Okumura, Y.; Toyotani, J. Impact of chest radiograph image size and augmentation on estimating pulmonary artery wedge pressure by regression convolutional neural network. In AIP Conference Proceedings; AIP Publishing, 2023; Volume 2872, p. 1. [Google Scholar]
- Yoo, J.; Kang, S. Class-adaptive data augmentation for image classification. IEEE Access 2023, 11, 26393–26402. [Google Scholar] [CrossRef]
- Bartók, A. P.; Kondor, R.; Csányi, G. On representing chemical environments. Physical Review B—Condensed Matter and Materials Physics 2013, 87, 184115. [Google Scholar] [CrossRef]
- Himanen, L.; Jäger, M. O. J.; Morooka, E. V.; Canova, F. F.; Ranawat, Y. S.; Gao, D. Z.; Rinke, P.; Foster, A. S. DScribe: Library of descriptors for machine learning in materials science. Computer Physics Communications 2020, 247, 106949. [Google Scholar] [CrossRef]
- Caro, M. A. Optimizing many-body atomic descriptors for enhanced computational performance of machine learning based interatomic potentials. Physical Review B 2019, 100, 024112. [Google Scholar] [CrossRef]
- Jäger, M. O. J.; Morooka, E. V.; Federici Canova, F.; Himanen, L.; Foster, A. S. Machine learning hydrogen adsorption on nanoclusters through structural descriptors. npj Computational Materials 2018, 4, 37. [Google Scholar] [CrossRef]
- De, S.; Bartók, A. P.; Csányi, G.; Ceriotti, M. Comparing molecules and solids across structural and alchemical space. Physical Chemistry Chemical Physics 2016, 18, 13754–13769. [Google Scholar] [CrossRef]
- Caruso, C.; Cardellini, A.; Crippa, M.; Rapetti, D.; Pavan, G. M. TimeSOAP: Tracking high-dimensional fluctuations in complex molecular systems via time variations of SOAP spectra. The Journal of Chemical Physics 2023, 158, 21. [Google Scholar] [CrossRef]
- LeCun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proceedings of the IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef]
- Gewers, F. L.; Ferreira, G. R.; Arruda, H. F. D.; Silva, F. N.; Comin, C. H.; Amancio, D. R.; Costa, L. F. Principal component analysis: A natural approach to data exploration. ACM Computing Surveys (CSUR) 2021, 54, 1–34. [Google Scholar] [CrossRef]
- Berahmand, K.; Daneshfar, F.; Salehi, E. S.; Li, Y.; Xu, Y. Autoencoders and their applications in machine learning: A survey. Artificial Intelligence Review 2024, 57, 28. [Google Scholar] [CrossRef]
- Malik, J. S.; Hemani, A. Gaussian random number generation: A survey on hardware architectures. ACM Computing Surveys (CSUR) 2016, 49, 1–37. [Google Scholar]
- Tanner, M. A.; Wong, W. H. The calculation of posterior distributions by data augmentation. Journal of the American Statistical Association 1987, 82, 528–540. [Google Scholar]
- Wei, L. Empirical Bayes test of regression coefficient in a multiple linear regression model. Acta Mathematicae Applicatae Sinica 1990, 6, 251–262. [Google Scholar]
- Simard, P. Y.; Steinkraus, D.; Platt, J. C. Best practices for convolutional neural networks applied to visual document analysis. Proceedings of ICDAR; Edinburgh: 2003; 3; 2003. [Google Scholar]
- Chawla, N. V.; Bowyer, K. W.; Hall, L. O.; Kegelmeyer, W. P. SMOTE: Synthetic minority over-sampling technique. Journal of Artificial Intelligence Research 2002, 16, 321–357. [Google Scholar]
- Elreedy, D.; Atiya, A. F. A comprehensive analysis of synthetic minority oversampling technique (SMOTE) for handling class imbalance. Information Sciences 2019, 505, 32–64. [Google Scholar] [CrossRef]
- Goodfellow, I.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative adversarial nets. Advances in Neural Information Processing Systems 2014, 27. [Google Scholar]
- Bao, G.; Yan, B.; Tong, L.; Shu, J.; Wang, L.; Yang, K.; Zeng, Y. Data augmentation for EEG-based emotion recognition using generative adversarial networks. Frontiers in Computational Neuroscience 2021, 15, 723843. [Google Scholar]
- Chen, L.; Li, Y.; Deng, X.; Liu, Z.; Lv, M.; Zhang, H. Dual auto-encoder GAN-based anomaly detection for industrial control system. Applied Sciences 2022, 12, 4986. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional networks for biomedical image segmentation. In Proceedings of the 18th International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI 2015); Springer: Munich, Germany, 2015; pp. 234–241. [Google Scholar]
- Laakso, J.; Himanen, L.; Homm, H.; Morooka, E. V.; Jäger, M. O. J.; Todorović, M.; Rinke, P. Updates to the DScribe library: New descriptors and derivatives. The Journal of Chemical Physics 2023, 158. [Google Scholar] [CrossRef]
- Shapiro, A. Monte Carlo sampling methods. Handbooks in Operations Research and Management Science 2003, 10, 353–425. [Google Scholar]
- Kingma, D. P. Adam: A method for stochastic optimization. arXiv preprint 2014, arXiv:1412.6980. [Google Scholar]
- Kumar, V. Pruning Distorted Images in MNIST Handwritten Digits. arXiv preprint 2023, arXiv:2307.14343. [Google Scholar]
- Hodson, T. O.; Over, T. M.; Foks, S. S. Mean squared error, deconstructed. Journal of Advances in Modeling Earth Systems 2021, 13, e2021MS002681. [Google Scholar]
- Zenodo Dataset. Available: https://zenodo.org/records/14916887.
- Klawohn, S.; Darby, J. P.; Kermode, J. R.; Csányi, G.; Caro, M. A.; Bartók, A. P. Gaussian approximation potentials: Theory, software implementation and application examples. The Journal of Chemical Physics 2023, 159. [Google Scholar]
| 1 | A set of functions is orthonormal if it satisfies for (orthogonality) and (normalization). |
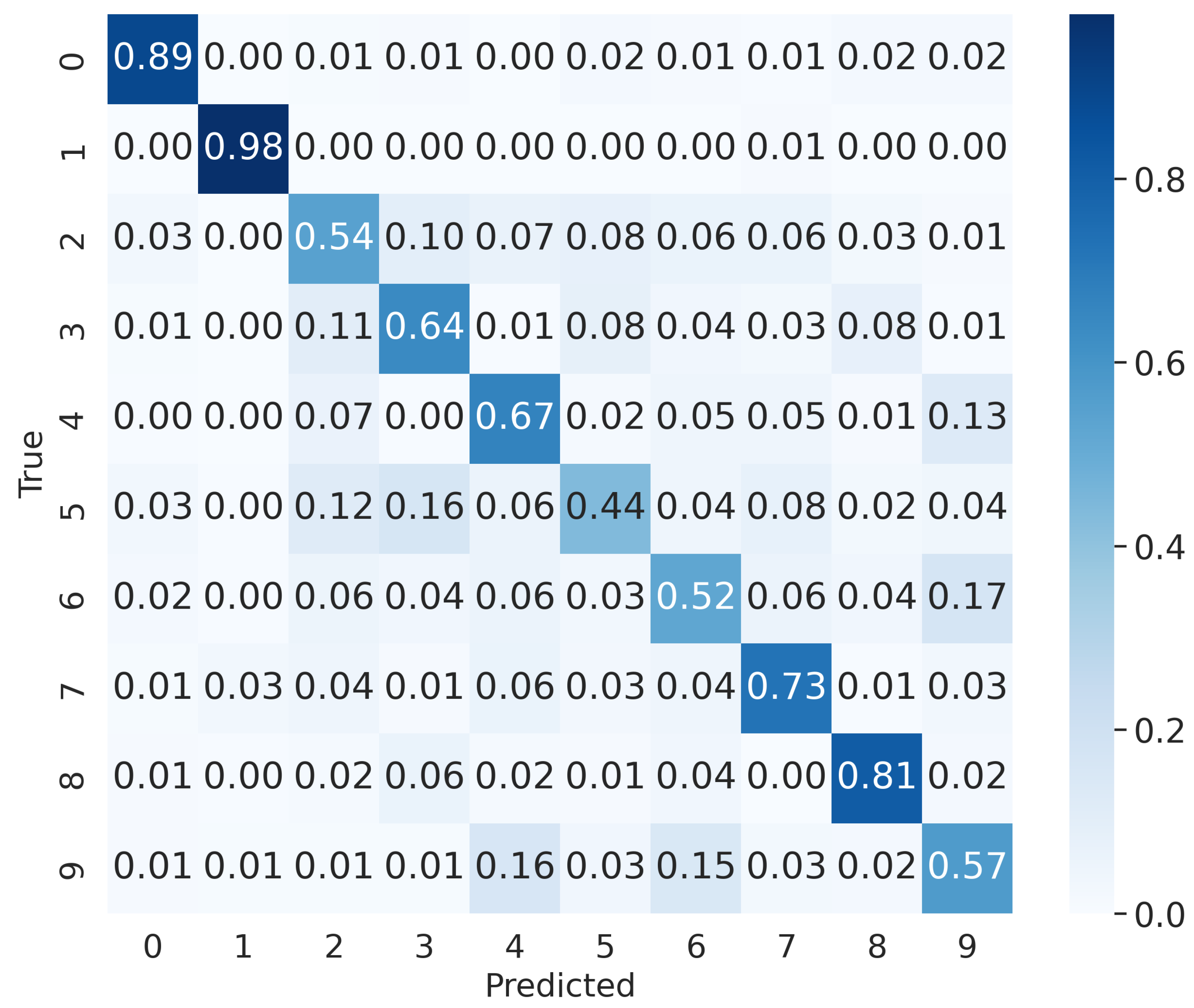


| Accuracy | Recall | Precision | F1 Score |
|---|---|---|---|
| 0.6863 | 0.6863 | 0.6821 | 0.6832 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




