Submitted:
22 January 2025
Posted:
23 January 2025
You are already at the latest version
Abstract
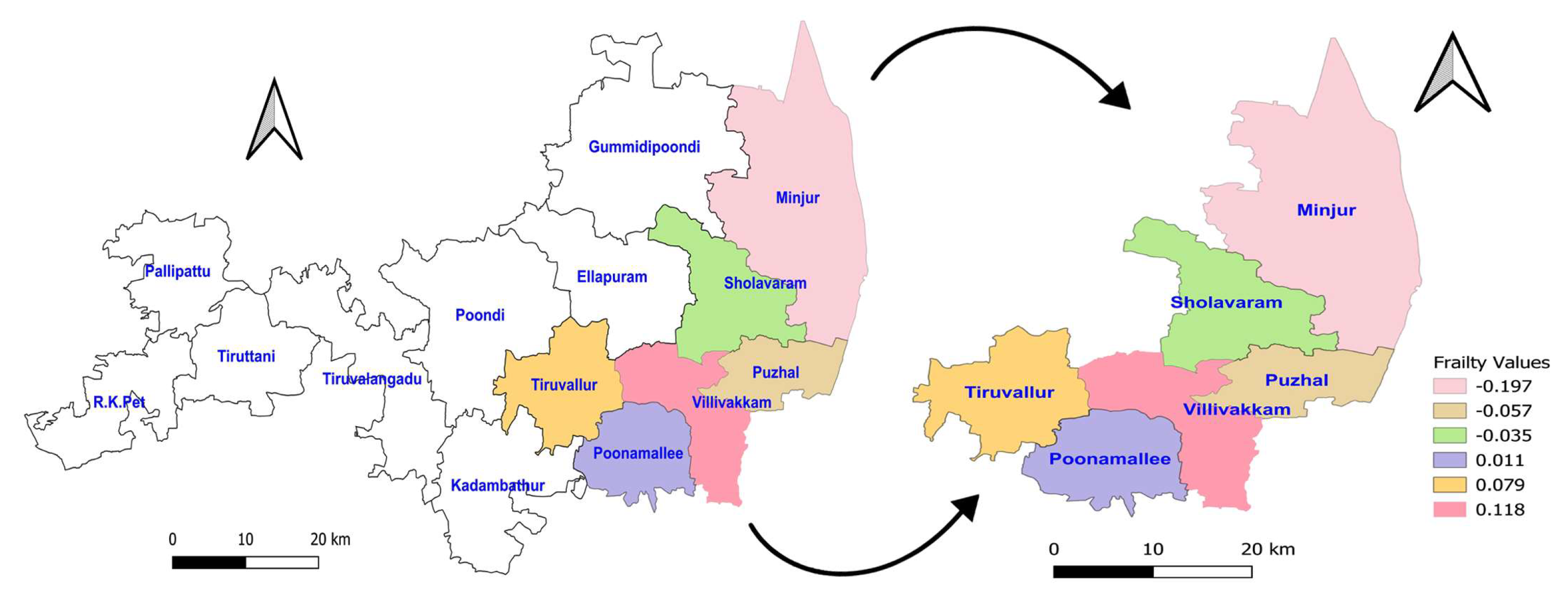
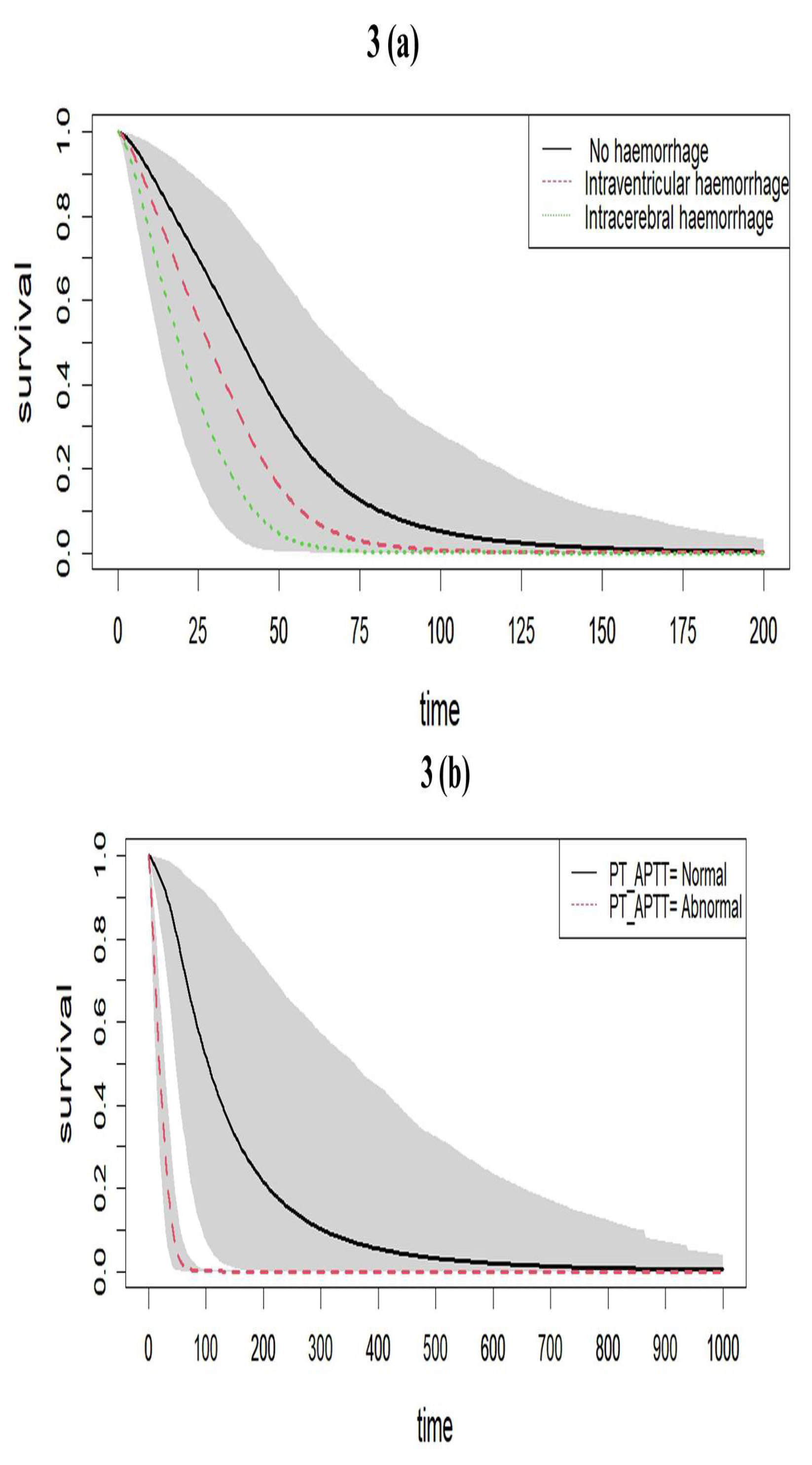
Background: Globally, Neonatal fungal sepsis (NFS) is a leading cause of neonatal mortality, particularly among vulnerable populations in neonatal intensive care units (NICU). The use of spatial frailty models with a Bayesian approach to identify hotspots and risk factors for neonatal deaths due to fungal sepsis has not been explored before. Methods: A cohort of 80 neonates admitted to the NICU and diagnosed with fungal sepsis through blood culture at a Government Hospital in Tamil Nadu, India, during 2018-2020 was considered for this study. The Bayesian spatial frailty models using parametric distributions, including Log-logistic, Log-normal, and Weibull proportional hazard (PH) models were employed to identify associated risk factors for deaths of NFS and hotspot areas using R software. Results: The spatial parametric frailty models were found to be good models for ana-lysing NFS data. Abnormal levels of activated thromboplastin had a significantly higher risk of death across all PH models (Log-logistic, Hazard Ratio (HR), 95% Credible In-terval (CI): 22.12, (5.40,208.08); Log-normal: 20.87, (5.29,123.23); Weibull: 18.49, (5.60,93.41). Haemorrhage had a higher risk of death for the Log-normal (1.65, (1.05,2.75)) and Weibull models (1.75, (1.07,3.12)). Villivakkam, Tiruvallur, and Poo-namallee blocks were identified as high-risk areas. Conclusions: The spatial parametric frailty models proved effective in identifying these risk factors and quantifying their association with mortality. The findings from this study underline the importance of early detection and management of risk factors to improve survival outcomes in neonates with fungal sepsis.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data Source
2.2. Setting and Study design
2.3. Ethics Approval
2.3. Statistical Analysis
3. Results
3.1. Demographic and Clinical Details for NFS Data
3.2. Posterior Estimates of Spatial Frailty of Three PH Models
4. Discussion
5. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| NFS | Neonatal Fungal Sepsis |
| NS | Neonatal Sepsis |
| PH | Proportional Hazard |
| NFHS-5 | National Family Health Survey - 5 |
| NNMR | Neonatal Mortality Rate |
| NICU | Neonatal Intensive Care Units |
| PT | Prothrombin Time |
| aPTT | activated Partial Thromboplastin Clotting Time |
| PT_APTT | Levels of Activated Thromboplastin Level |
| ICH | Intracranial Haemorrhage |
| TB | Tuberculosis |
| MCMC | Markov chain Monte Carlo |
| GA_weeks | Gestational Age in Weeks |
| B_Weight | Neonates Birth Weight |
| LPML | Log Pseudo Marginal Likelihood |
| DIC | Deviance Information Criterion |
| WAIC | Watanabe Akaike Information Criterion |
| QGIS | Quantum Geographic Information System |
| CI | Credible Intervals |
| HR | Hazard Ratio |
References
- Stoll, B.J.; Hansen, N.; Fanaroff, A.A.; Wright, L.L.; Carlo, W.A.; Ehrenkranz, R.A.; Lemons, J.A.; Donovan, E.F.; Stark, A.R.; Tyson, J.E.; Oh. , W.; Bauer, C.R.; Korones, S.B.; Shankaran, S.; Laptook, A.R.; Stevenson, D.K.; Papile, L.-A.; Poole, W.K. Late-Onset Sepsis in Very Low Birth Weight Neonates: The Experience of the NICHD Neonatal Research Network. Pediatrics 2002, 110, 285–291. [Google Scholar] [CrossRef]
- WHO sepsis technical expert meeting, 16-. https://www.who.int/publications/m/item/who-sepsis-technical-expert-meeting-16-17-january-2018 (accessed 2025-01-09). 17 January.
- Fleischmann, C.; Reichert, F.; Cassini, A.; Horner, R.; Harder, T.; Markwart, R.; Tröndle, M.; Savova, Y.; Kissoon, N.; Schlattmann, P.; Reinhart, K.; Allegranzi, B.; Eckmanns, T. Global Incidence and Mortality of Neonatal Sepsis: A Systematic Review and Meta-Analysis. Arch Dis Child 2021, 106, 745–752. [Google Scholar] [CrossRef] [PubMed]
- Newborn mortality. https://www.who.int/news-room/fact-sheets/detail/newborn-mortality (accessed 2025-01-10.
- NFHS. International Institute for Population Sciences. National Family Health Survey (NFHS-5), 20019-21 https://mohfw.gov.in/sites/default/files/NFHS-5_Phase-II_0.pdf (accessed 2025-01-10).
- Adams-Chapman, I.; Stoll, B.J. Neonatal Infection and Long-Term Neurodevelopmental Outcome in the Preterm Infant. Current Opinion in Infectious Diseases 2006, 19, 290–297. [Google Scholar] [CrossRef] [PubMed]
- Stoll, B.J.; Hansen, N.I.; Adams-Chapman, I.; Fanaroff, A.A.; Hintz, S.R.; Vohr, B.; Higgins, R.D.; National Institute of Child Health and Human Development Neonatal Research Network. Neurodevelopmental and Growth Impairment among Extremely Low-Birth-Weight Infants with Neonatal Infection. JAMA 2004, 292, 2357–2365. [Google Scholar] [CrossRef]
- National Neonatal-Perinatal Database 2002 - NNPD Nodal Center AIIMS Delhi New Delhi; 2005. Available at: https://www.newbornwhocc.org/pdf/nnpd_report_2002-03.PDF Accessed 10 August 2019.
- Adalarasan, N.; Stalin, S.; Venkatasamy, S.; Sridevi, S.; Padmanaban, S.; Chinnaiyan, P. Association and Outcome of Intracranial Haemorrhage in Newborn with Fungal Sepsis- A Prospective Cohort Study. IJNMR 2022. [CrossRef]
- Aswi, A.; Cramb, S.; Duncan, E.; Hu, W.; White, G.; Mengersen, K. Bayesian Spatial Survival Models for Hospitalisation of Dengue: A Case Study of Wahidin Hospital in Makassar, Indonesia. Int J Environ Res Public Health 2020, 17, 878. [Google Scholar] [CrossRef]
- Banerjee, S.; Wall, M.M.; Carlin, B.P. Frailty Modeling for Spatially Correlated Survival Data, with Application to Infant Mortality in Minnesota. Biostatistics 2003, 4, 123–142. [Google Scholar] [CrossRef] [PubMed]
- Balan, T.A.; Putter, H. A Tutorial on Frailty Models. Stat Methods Med Res 2020, 29, 3424–3454. [Google Scholar] [CrossRef] [PubMed]
- Motarjem, K.; Mohammadzadeh, M.; Abyar, A. Geostatistical Survival Model with Gaussian Random Effect. Stat Papers 2020, 61, 85–107. [Google Scholar] [CrossRef]
- Thamrin, S.A. ; Aswi; Ansariadi; Jaya, A.K.; Mengersen, K. Bayesian Spatial Survival Modelling for Dengue Fever in Makassar, Indonesia. Gac Sanit, 1. [CrossRef]
- Pérez-Castro, E.; Guzmán-Martínez, M.; Godínez-Jaimes, F.; Reyes-Carreto, R.; Vargas-de-León, C.; Aguirre-Salado, A.I. Spatial Survival Model for COVID-19 in México. Healthcare 2024, 12, 306. [Google Scholar] [CrossRef] [PubMed]
- Egbon, O.A.; Bogoni, M.A.; Babalola, B.T.; Louzada, F. Under Age Five Children Survival Times in Nigeria: A Bayesian Spatial Modeling Approach. BMC Public Health 2022, 22, 2207. [Google Scholar] [CrossRef]
- Migamba, S.M.; Kisaakye, E.; Komakech, A.; Nakanwagi, M.; Nakamya, P.; Mutumba, R.; Migadde, D.; Kwesiga, B.; Bulage, L.; Kadobera, D.; Ario, A.R. Trends and Spatial Distribution of Neonatal Sepsis, Uganda, 2016-2020. BMC Pregnancy Childbirth 2023, 23, 770. [Google Scholar] [CrossRef] [PubMed]
- Kibret, G.D.; Demant, D.; Hayen, A. Bayesian Spatial Analysis of Factors Influencing Neonatal Mortality and Its Geographic Variation in Ethiopia. PLoS One 2022, 17, e0270879. [Google Scholar] [CrossRef]
- Nunes, C.; Taylor, B.M. Modelling the Time to Detection of Urban Tuberculosis in Two Big Cities in Portugal: A Spatial Survival Analysis. Int J Tuberc Lung Dis 2016, 20, 1219–1225. [Google Scholar] [CrossRef]
- Nazar, E.; Baghishani, H.; Doosti, H.; Ghavami, V.; Aryan, E.; Nasehi, M.; Sharafi, S.; Esmaily, H.; Yazdani Charati, J. Bayesian Spatial Survival Analysis of Duration to Cure among New Smear-Positive Pulmonary Tuberculosis (PTB) Patients in Iran, during 2011–2018. International Journal of Environmental Research and Public Health 2021, 18, 54. [Google Scholar] [CrossRef] [PubMed]
- Su, P.-F.; Sie, F.-C.; Yang, C.-T.; Mau, Y.-L.; Kuo, S.; Ou, H.-T. Association of Ambient Air Pollution with Cardiovascular Disease Risks in People with Type 2 Diabetes: A Bayesian Spatial Survival Analysis. Environ Health 2020, 19, 110. [Google Scholar] [CrossRef]
- Henderson, R.; Shimakura, S.; Gorst, D. Modeling Spatial Variation in Leukemia Survival Data. Journal of the American Statistical Association 2002, 97, 965–972. [Google Scholar] [CrossRef]
- Srinivasan., R.; Ponnuraja., C.; Pradeep., AM.; Moeng., SR.; Sivasamy., R.; Venkatesan, P. Srinivasan. R.; Ponnuraja. C.; Pradeep. AM.; Moeng. SR.; Sivasamy. R.; Venkatesan, P. Bayesian Spatial Cox Proportional Hazard Model For HIV Infected Tuberculosis Cases In Chennai. Global Journal of Pure and Applied Mathematics. ISSN, /: Volume 15, Number 6, pp. 971-980. https, 2019; 15. [Google Scholar]
- Besag, J.; Green, P.J. Spatial Statistics and Bayesian Computation. Journal of the Royal Statistical Society Series B: Statistical Methodology 1993, 55, 25–37. [Google Scholar] [CrossRef]
- Banerjee, S.; Carlin, B.P.; Gelfand, A.E.; Banerjee, S. Hierarchical Modeling and Analysis for Spatial Data; Chapman and Hall/CRC: New York, 2003. [Google Scholar] [CrossRef]
- Kau, J.B.; Keenan, D.C.; Li, X. An Analysis of Mortgage Termination Risks: A Shared Frailty Approach with MSA-Level Random Effects. J Real Estate Finan Econ 2011, 42, 51–67. [Google Scholar] [CrossRef]
- Royston, P.; Parmar, M.K.B. Flexible Parametric Proportional-Hazards and Proportional-Odds Models for Censored Survival Data, with Application to Prognostic Modelling and Estimation of Treatment Effects. Stat Med 2002, 21, 2175–2197. [Google Scholar] [CrossRef] [PubMed]
- Al-Aziz, S.N.; Hassan Muse, A.; Jawa, T.M.; Sayed-Ahmed, N.; Aldallal, R.; Yusuf, M. Bayesian Inference in a Generalized Log-Logistic Proportional Hazards Model for the Analysis of Competing Risk Data: An Application to Stem-Cell Transplanted Patients Data. Alexandria Engineering Journal 2022, 61, 13035–13050. [Google Scholar] [CrossRef]
- Khan, S.A.; Khosa, S.K. Generalized Log-Logistic Proportional Hazard Model with Applications in Survival Analysis. J Stat Distrib App 2016, 3, 16. [Google Scholar] [CrossRef]
- Pourhoseingholi, M.A.; Moghimi-Dehkordi, B.; Safaee, A.; Hajizadeh, E.; Solhpour, A.; Zali, M.R. Prognostic Factors in Gastric Cancer Using Log-Normal Censored Regression Model. Indian J Med Res 2009, 129, 262–267. [Google Scholar]
- Khan, S.A. Exponentiated Weibull Regression for Time-to-Event Data. Lifetime Data Anal 2018, 24, 328–354. [Google Scholar] [CrossRef]
- Zhang, Z. Parametric Regression Model for Survival Data: Weibull Regression Model as an Example. Ann Transl Med 2016, 4, 484. [Google Scholar] [CrossRef] [PubMed]
- Sahu, S.K.; Dey, D.K.; Aslanidou, H.; Sinha, D. A Weibull Regression Model with Gamma Frailties for Multivariate Survival Data. Lifetime Data Anal 1997, 3, 123–137. [Google Scholar] [CrossRef]
- Robert, C.P.; Elvira, V.; Tawn, N.; Wu, C. Accelerating MCMC Algorithms. Wiley Interdiscip Rev Comput Stat 2018, 10, e1435. [Google Scholar] [CrossRef]
- Hastings, W.K. Monte Carlo Sampling Methods Using Markov Chains and Their Applications. Biometrika 1970, 57, 97–109. [Google Scholar] [CrossRef]
- Geman, S.; Geman, D. Stochastic Relaxation, Gibbs Distributions and the Bayesian Restoration of Images*. Journal of Applied Statistics 1993, 20, 25–62. [Google Scholar] [CrossRef]
- Gallagher, K.; Charvin, K.; Nielsen, S.; Sambridge, M.; Stephenson, J. Markov Chain Monte Carlo (MCMC) Sampling Methods to Determine Optimal Models, Model Resolution and Model Choice for Earth Science Problems. Marine and Petroleum Geology 2009, 26, 525–535. [Google Scholar] [CrossRef]
- Musa, G.K.; Chaku, S.E.; Nwaze, N.O.; Adehi, M.U.; Omaku, P.E. Bayesian Accelerated Failure Time Model With Spatial Dependency: Application To Under Five Mortality Rate. Global Journal of Pure and Applied Sciences 2024, 30, 537–549. [Google Scholar] [CrossRef]
- Wang, S.; Zhang, J.; Lawson, A.B. A Bayesian Normal Mixture Accelerated Failure Time Spatial Model and Its Application to Prostate Cancer. Stat Methods Med Res 2016, 25, 793–806. [Google Scholar] [CrossRef]
- Zhou, H.; Hanson, T. A Unified Framework for Fitting Bayesian Semiparametric Models to Arbitrarily Censored Survival Data, Including Spatially Referenced Data. Journal of the American Statistical Association 2018, 113, 571–581. [Google Scholar] [CrossRef]
- Mardhiah, K.; Wan-Arfah, N.; Naing, N.N.; Hassan, M.R.A.; Chan, H.-K. Comparison of Cox Proportional Hazards Model, Cox Proportional Hazards with Time-Varying Coefficients Model, and Lognormal Accelerated Failure Time Model: Application in Time to Event Analysis of Melioidosis Patients. Asian Pacific Journal of Tropical Medicine 2022, 15, 128. [Google Scholar] [CrossRef]
- Daniel, K.; Onyango, N.O.; Sarguta, R.J. A Spatial Survival Model for Risk Factors of Under-Five Child Mortality in Kenya. International Journal of Environmental Research and Public Health 2022, 19, 399. [Google Scholar] [CrossRef]
- Mahanta, K.K.; Hazarika, J.; Barman, M.P.; Rahman, T. An Application of Spatial Frailty Models to Recovery Times of COVID-19 Patients in India under Bayesian Approach. JSR 2021, 65, 150–155. [Google Scholar] [CrossRef]
- Teshnizi, S.H.; Ayatollahi, S.M.T. Comparison of Cox Regression and Parametric Models: Application for Assessment of Survival of Pediatric Cases of Acute Leukemia in Southern Iran. Asian Pac J Cancer Prev 2017, 18, 981–985. [Google Scholar] [CrossRef]
- Gemechu, L.L.; Debusho, L.K. Bayesian Spatial Modelling of Tuberculosis-HIV Co-Infection in Ethiopia. PLoS One 2023, 18, e0283334. [Google Scholar] [CrossRef]
- Saleem, S.; Tikmani, S.S.; Goudar, S.S.; Hwang, K.; Dhaded, S.; Guruprasad, G.; Nadig, N.G.; Kusagur, V.B.; Patil, L.G.C.; Siddartha, E.S.; Yogeshkumar, S.; Somannavar, M.S.; Roujani, S.; Khan, M.; Shaikh, M.; Hanif, M.; Bann, C.M.; McClure, E.M.; Goldenberg, R.L.; Group, the P. S. Neonatal Mortality among Preterm Infants Admitted to Neonatal Intensive Care Units in India and Pakistan: A Prospective Study. BJOG: An International Journal of Obstetrics & Gynaecology 2023, 130, 68–75. [Google Scholar] [CrossRef]
- Mihatov Stefanovic, I. Neonatal Sepsis. Biochemia medica 2011, 21, 276–281. [Google Scholar] [CrossRef]
| Parameters | Min | Q1 | Median | Mean | Q3 | Max | SD | |
| Survived (N=30) | GA_weeks | 28 | 32 | 36.5 | 35.47 | 38 | 42 | 3.69 |
| B_WEIGHT | 1 | 1.49 | 2.25 | 2.11 | 2.46 | 3.3 | 0.62 | |
| Platelet | 2 | 14 | 31 | 56.4 | 62.75 | 252 | 67.31 | |
| Haemorrhage | 0 | 0 | 0 | 0.27 | 0 | 2 | 0.69 | |
| PT_APTT | 0 | 0 | 0 | 0.1 | 0 | 1 | 0.31 | |
| Died (N=50) | GA_weeks | 26 | 31 | 35.5 | 34.5 | 38 | 40 | 4.24 |
| B_WEIGHT | 0.76 | 1.47 | 2.1 | 2.06 | 2.73 | 3.96 | 0.77 | |
| Platelet | 3 | 8.75 | 24.5 | 47 | 64 | 232 | 50.98 | |
| Haemorrhage | 0 | 0 | 1 | 0.86 | 2 | 2 | 0.9 | |
| PT_APTT | 0 | 1 | 1 | 0.96 | 1 | 1 | 0.2 | |
| Total (N=80) | GA_weeks | 26 | 32 | 36 | 34.86 | 38 | 42 | 4.05 |
| B_WEIGHT | 0.76 | 1.49 | 2.2 | 2.08 | 2.72 | 3.96 | 0.72 | |
| Platelet | 2 | 11 | 25.5 | 50.53 | 61.75 | 252 | 57.41 | |
| Haemorrhage | 0 | 0 | 0 | 0.64 | 2 | 2 | 0.88 | |
| PT_APTT | 0 | 0 | 1 | 0.64 | 1 | 1 | 0.48 |
| Model | Parameters | Mean | Median | SD | 95% CI (Mean) | |
| Lower | Upper | |||||
| Log-logistic | GA_Weeks | 0.02 | 0.02 | 0.07 | -0.12 | 0.16 |
| B_WEIGHT | 0.12 | 0.13 | 0.36 | -0.59 | 0.81 | |
| Platelet | 0 | 0 | 0 | -0.01 | 0.01 | |
| Haemorrhage | 0.56 | 0.58 | 0.3 | -0.02 | 1.16 | |
| PT_APTT** | 3.1 | 3.03 | 0.84 | 1.69 | 5.34 | |
| Log-normal | GA_Weeks | 0.03 | 0.03 | 0.07 | -0.1 | 0.16 |
| B_WEIGHT | 0.06 | 0.07 | 0.34 | -0.64 | 0.67 | |
| Platelet | 0 | 0 | 0 | -0.01 | 0.01 | |
| Haemorrhage** | 0.5 | 0.5 | 0.26 | 0.05 | 1.01 | |
| PT_APTT** | 3.04 | 2.98 | 0.79 | 1.67 | 4.81 | |
| Weibull | GA_Weeks | 0.01 | 0.01 | 0.07 | -0.14 | 0.15 |
| B_WEIGHT | 0.18 | 0.2 | 0.33 | -0.48 | 0.78 | |
| Platelet | 0 | 0 | 0 | -0.01 | 0.01 | |
| Haemorrhage** | 0.56 | 0.56 | 0.27 | 0.07 | 1.14 | |
| PT_APTT** | 2.92 | 2.87 | 0.74 | 1.72 | 4.54 | |
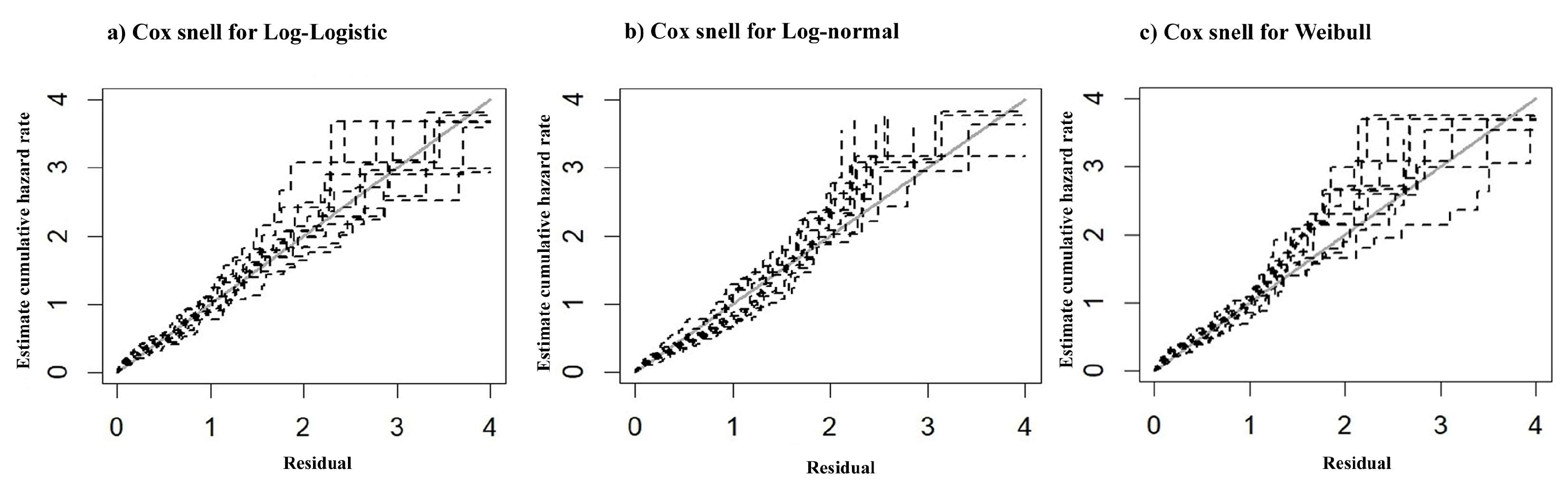
| Model | LPML | DIC | WAIC |
| Log-logistic PH Model | 221.65 | 441.33 | 442.83 |
| Log-normal PH Model | 221.89 | 442.05 | 443.51 |
| Weibull PH Model | 220.68 | 439.64 | 440.76 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




