Submitted:
13 December 2024
Posted:
16 December 2024
You are already at the latest version
Abstract
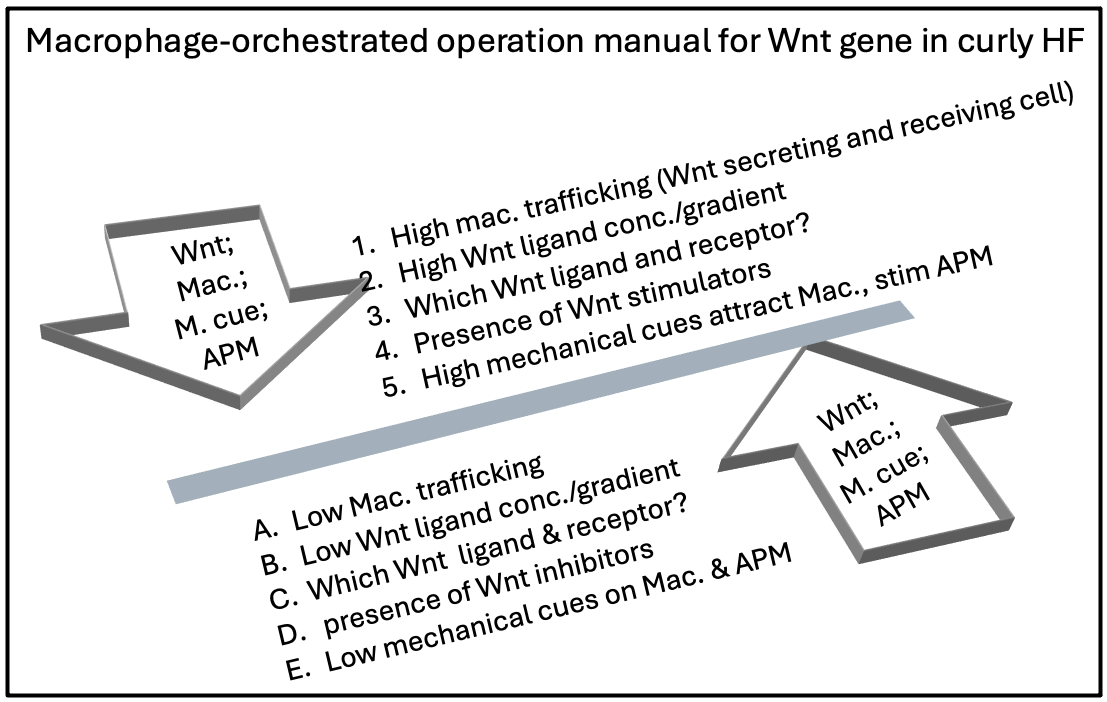
Curly hair is a genetic trait that is controlled by major developmental programs. A curly hair genomewide association studies (GWAS) have uncovered evidence suggesting that hair curl variation is very complex, where many genes are involved, each having a modest effect on hair curl. The adult hair cycle retains an element of developmental dynamics reflected in the crosstalk between the mesenchymal and epithelial elements, so each time a hair follicle goes through rebirth in the hair cycle, all the factors that govern the curl must be re-established. The developmental genes involved in shaping hair curl include trichohyalin, EDAR and WNT10a. Wnt10a is a preserved development gene associated with hair morphology and cycling, with a family of 19 Wnt ligands that bind 10 receptors. Different combinational types between Wnt ligands and receptors determines the distinct downstream pathways directing cell function. Now, it is time to move from GWAS to functional genomics to identify in which physiologic contexts this regulation occurs and investigate what tissues and cell types are involved. In this hypothesis I am proposing that macrophages are the one relevant cell type that regulate perifollicular Wnt gene activity! Macrophages are resident in the perifollicular dermal sheath of the hair follicle; it secretes and receive Wnt; recruited in early embryonic development by Wnt ligands, that first crosstalk! also recruited by mechanical cues that we propose to be generated by a convoluted dermo-epidermal junction in an African skin that has triple the length of a Caucasian skin. Macrophage-secreted Wnt is orchestrating the aesthetic function of the follicle and shape the curly hair shaft. This hypothesis is answering questions about when, what and how Wnt gene activation can shape the curly hair types. Further studies of the diversity of macrophage ligand(s)-receptor(s) interactions with the HFSC to determine hair shape diversity needs collaboration with AI specialists using available software like Interactome and Volcano plot.
Keywords:
Highlights
- GWAS studies of low and high curl patterns found that the Wnt gene is linked to curly hair.
- Going from GWAS to the function requires the identification of a causal cell that can be probed and modulated to have a clinical significance.
- This hypothesis pinpoint macrophages as the causative cell that secrete- and receive-Wnt ligands and is resident in the perifollicular dermal niche;
- This hypothesis considers microscopical architectural findings in African skin where the convoluted dermal-epidermal junction length is threefold that in Caucasian skin, possibly creating mechanical cues that attract macrophages to the dermis.
- Multiple variables are generated by perifollicular macrophages and can create a wide range of curly hair types;
- One novel variable is the number of perifollicular macrophages present in range, close to the HFSCs, as Wnt traveling distance is limited and hence the HF size and its diameter is an additional variable.
- These variables are the answers to which, how and when Wnt gene is activated.
- Artificial intelligence and immune-fluorescence research are required to further elucidate the curly hair gene.
- The field is highly specialized and this research needs collaboration of specialists in AI in drug development, protein-protein interactions and in silico computer-assisted ligand-receptor interactions and macrophage typing studies.
- Immunofluorescence studies of different types of curly hair samples extracted as a hair follicular unit using Wnt ligand-specific antibodies.
Background
A. The Genetics of Curly Hair: GWAS Studies
B. Functional Anatomy of Human Scalp Hair Follicle
B-1. The Pilosebaceous Unit is the HF's Structural Unit
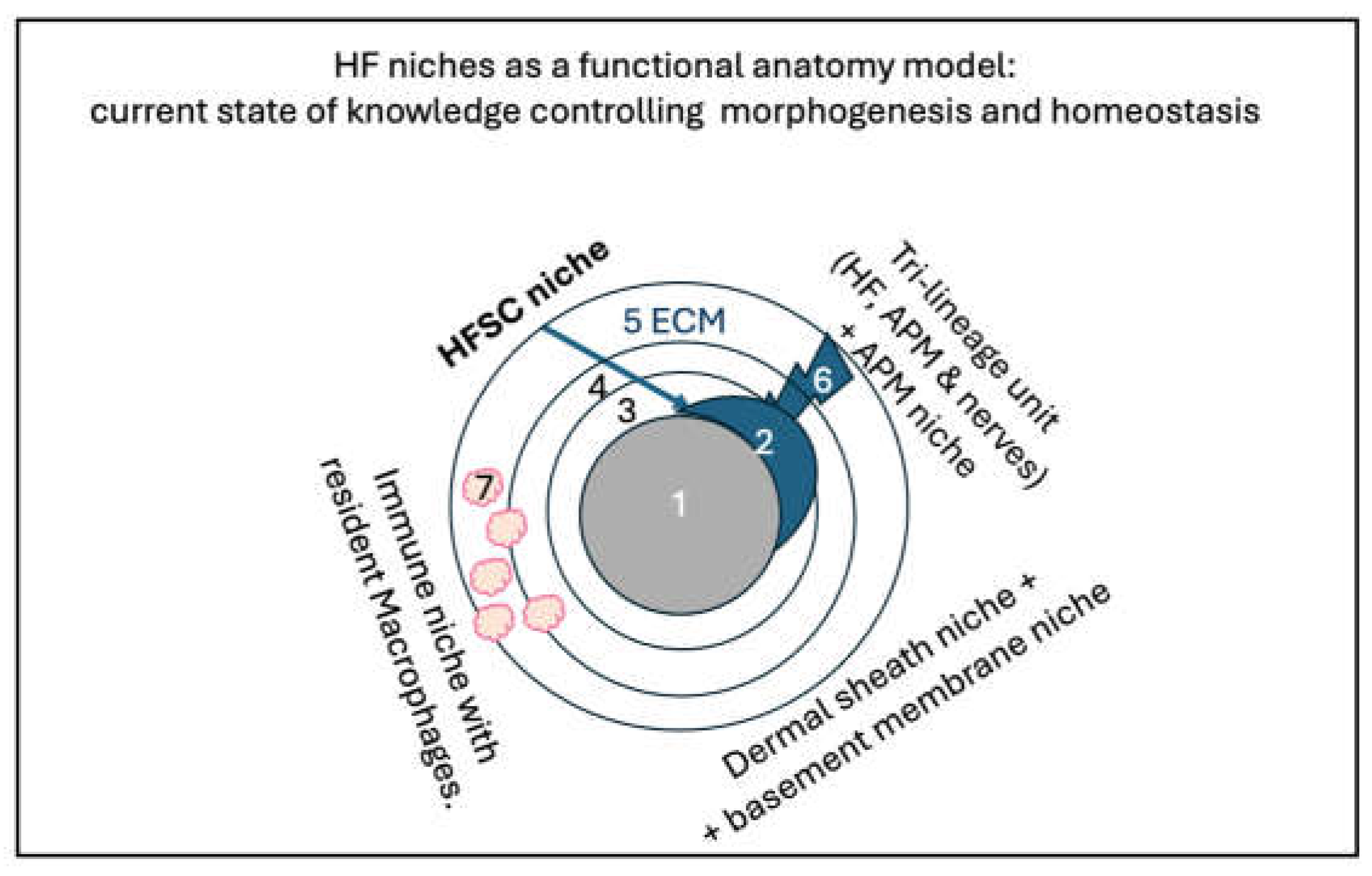
B-2. The Hair Follicle Multiple Niche Architecture
B-3.Perifollicular Macrophages Secrete and Receive Wnt
B-4. Dermal-Epidermal Crosstalk During Embryogenesis and Adult Homeostasis
B-5. The Hair Follicle Microenvironment
The hypothesis: Curly Hair from GWAS to Functional Genomics: Wnt-Secreting and -Receiving Macrophages Orchestrate curly and coily hair types.
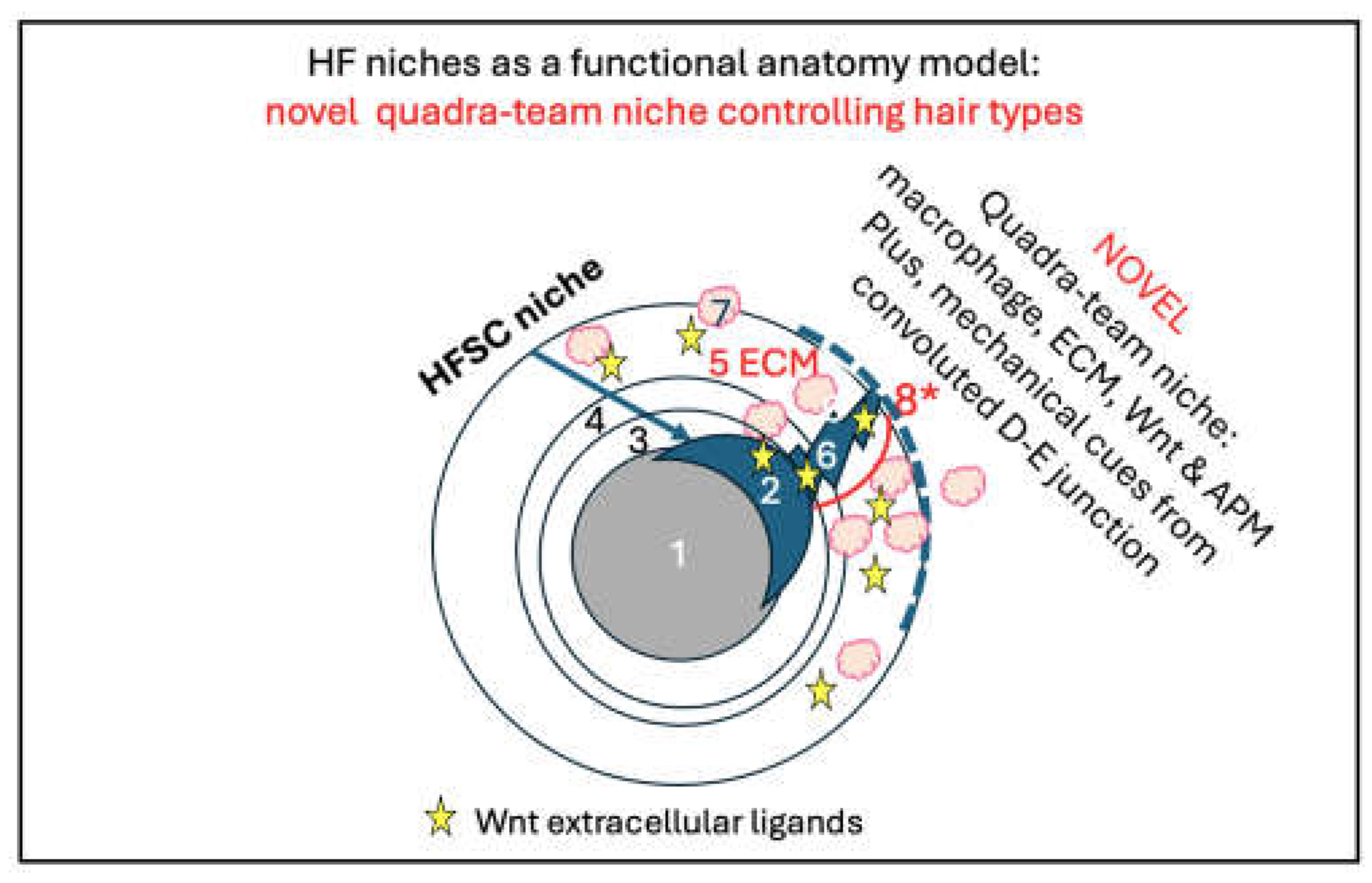
C. How Was the “Quadra-Team Niche” Idea Born?
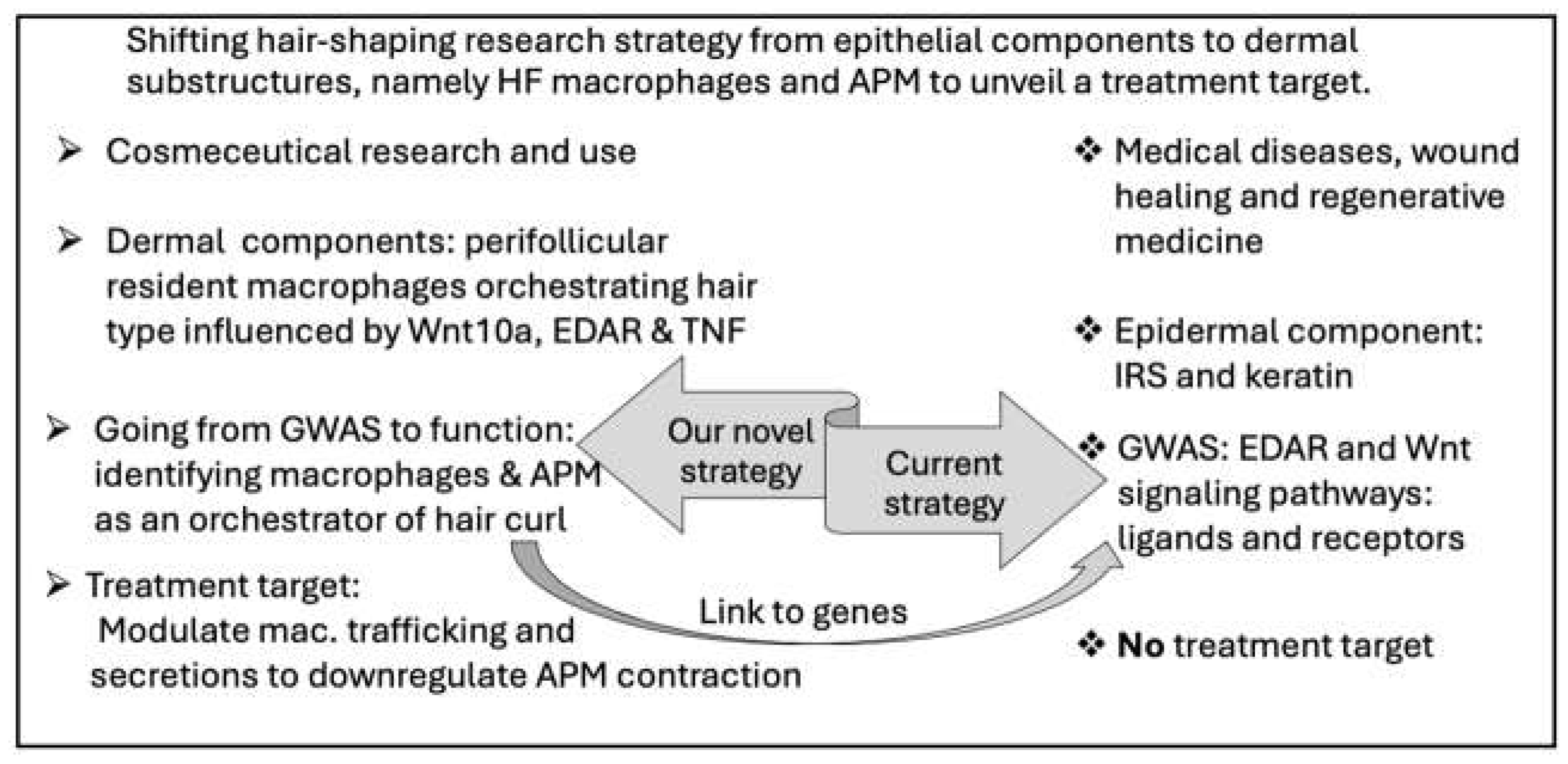
D. Shifting Curly Hair Research Strategy and Starting Cosmetic Immunology: Macrophages Keep Us Healthy and Beautiful!
D-1. The Dermal-Epidermal Junction and ECM Are the Generator of Mechanical Cues
D-2. Mechanical Cues Increase Macrophage Migration: A Novel Variable That Shape the Hair!
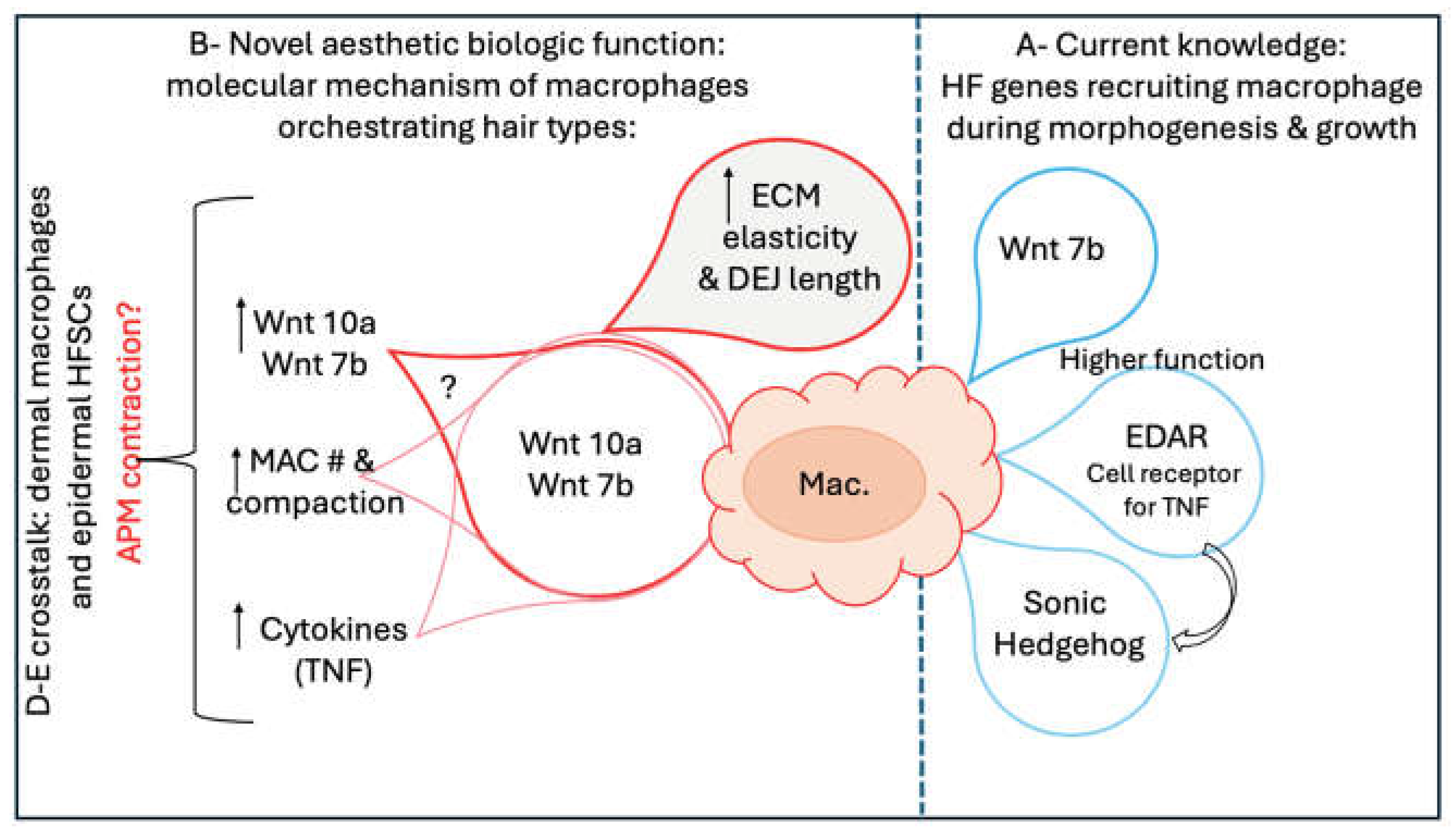
E. Novel Wnt Gene Language: The Crosstalk Between Macrophages and Curly Hair Genes!
E-2. Arrector Pili Muscle Is the Novel Effector Substructure of the Hair Follicle
E-3. Are Curly Hair Patterns Orchestrated by Cues from the Convoluted Long D-E Junction?
Testing the Hypothesis
Developing Selective” Arrector Pili Muscle Relaxing Treatment
Conclusion
Author and Affiliation
Intellectual property
Funding
Competing interest
Acknowledgement
References
- Al Jasim N (2024) Curly Hair Follicle is Sculpted by a Contracted Arrector Pili Muscle: A Hypothesis with Treatment Implications. Am J Pharmacol Pharmacother Vol.11 No. S1: 01. [CrossRef]
- Al Jasim, N. Curly Hair Follicle is Sculpted by a Contracted Arrector Pili Muscle. A Hypothesis with Treatment Implications. Preprints 2024, 2024091791. [CrossRef]
- Andl T., Reddy S. T., Gaddapara T., Millar S. E. (2002). WNT signals are required for the initiation of hair follicle development. Dev. Cell, 2002; 2: 643–653. [CrossRef]
- Anthony S Findley, Alan Monziani, Allison L Richards, et al. Functional dynamic genetic effects on gene regulation are specific to a particular cell types and environmental conditions. eLife, 2021; 10:e67077. [CrossRef]
- Cano-Gamez E and Trynka G. From GWAS to Function: Using Functional Genomics to Identify the Mechanisms Underlying Complex Diseases. Front. Genet., (2020); 11:424. [CrossRef]
- Chakrabarti J, Dua-Awereh M, Schumacher M, Engevik A, Hawkins J, Helmrath MA, Zavros Y. Sonic Hedgehog acts as a macrophage chemoattractant during regeneration of the gastric epithelium. NPJ Regen Med. 222 Jan 12;7(1):3. [CrossRef]
- Chen, CL., Huang, WY., Wang, E.H.C. et al. Functional complexity of hair follicle stem cell niche and therapeutic targeting of niche dysfunction for hair regeneration. J Biomed Sci, 2020; 27: 43. [CrossRef]
- Choi BY. Targeting Wnt/β-Catenin Pathway for Developing Therapies for Hair Loss. Int J Mol Sci. 2020 Jul 12;21(14):4915. [CrossRef]
- Christof Niehrs. The complex world of WNT receptor signaling. Nat Rev Mol Cell Biol. 2012;13(12): 767-79. [CrossRef]
- Chu, SY., Chou, CH., Huang, HD. et al. Mechanical stretch induces hair regeneration through the alternative activation of macrophages. Nat Commun 10, 1524 (2019). [CrossRef]
- Cuevas-Diaz Duran, R.; Martinez-Ledesma, E.; Garcia-Garcia, M,; et al. The Biology and Genomics of Human Hair Follicles: A Focus on Androgenetic Alopecia. Int. J.Mol.Sci,.2024 ; 25,2542. [CrossRef]
- Das, S., Chakrabarti, S. Classification and prediction of protein–protein interaction interface using machine learning algorithm. Sci Rep 11, 1761 (2021). [CrossRef]
- Dong Wook Shin. The Molecular Mechanism of Natural Products Activating Wnt/β-Catenin Signaling Pathway for Improving Hair Loss Life, 2022; 12(11), 1856. [CrossRef]
- Du, H., Bartleson, J.M., Butenko, S. et al. Tuning immunity through tissue mechanotransduction. Nat Rev Immunol 23, 174–188 (2023). [CrossRef]
- Elizabeth S. Malsin, Seokjo Kim, Anna P. Lam et al. Macrophages as a source and recipient of Wnt signals. Frontiers in Immunology, 2019: 10: 1813. [CrossRef]
- Ferhan Muneeb, Jonathan A. Hardman and Ralph Paus. Hair growth control by innate immunocytes: Perifollicular macrophages revisit. Experimental Dermatology, 2019; 28: 425-431. [CrossRef]
- Fujiwara H, Ferreira M, Donati G, et al. The basement membrane of hair follicle stem cells is a muscle cell niche. Cell. 2011 Feb 18;144(4):577-89. [CrossRef]
- Gillian E. Westgate, Natalia V. Botchkareva and Desmond J. Tobin. The biology of hair diversity. International Journal of Hair Science, 2013; 35(4): 329-336. [CrossRef]
- Gillian E. Westgate, Rebecca S. Ginger, and Martin R. Green. The biology and genetics of curly hair. Experimental Dermatology, 2017; 26: 483-490. [CrossRef]
- Grubbs H, Nassereddin A, Morrison M. Embryology, Hair. [Updated 2023 May 1]. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2024 Jan-. Available from: https://www.ncbi.nlm.nih.gov/books/NBK534794.
- Human Protein Atlas: WNT10A, Structure and Interaction. https://www.proteinatlas.org/ENSG00000135925-WNT10A/structure+interaction.
- Idowu, O. C.; Markiewicz, E.; Oladele, D. The Genomic Variation in Textured Hair: Implications in Developing a Holistic Hair Care Routine. Preprints 2024, 2024071218. [CrossRef]
- Jahoda, Colin A.B. and Christiano, Angela M. Niche Crosstalk: Intercellular Signals at the Hair Follicle. Cell, 2011; 146(5): 678–681.
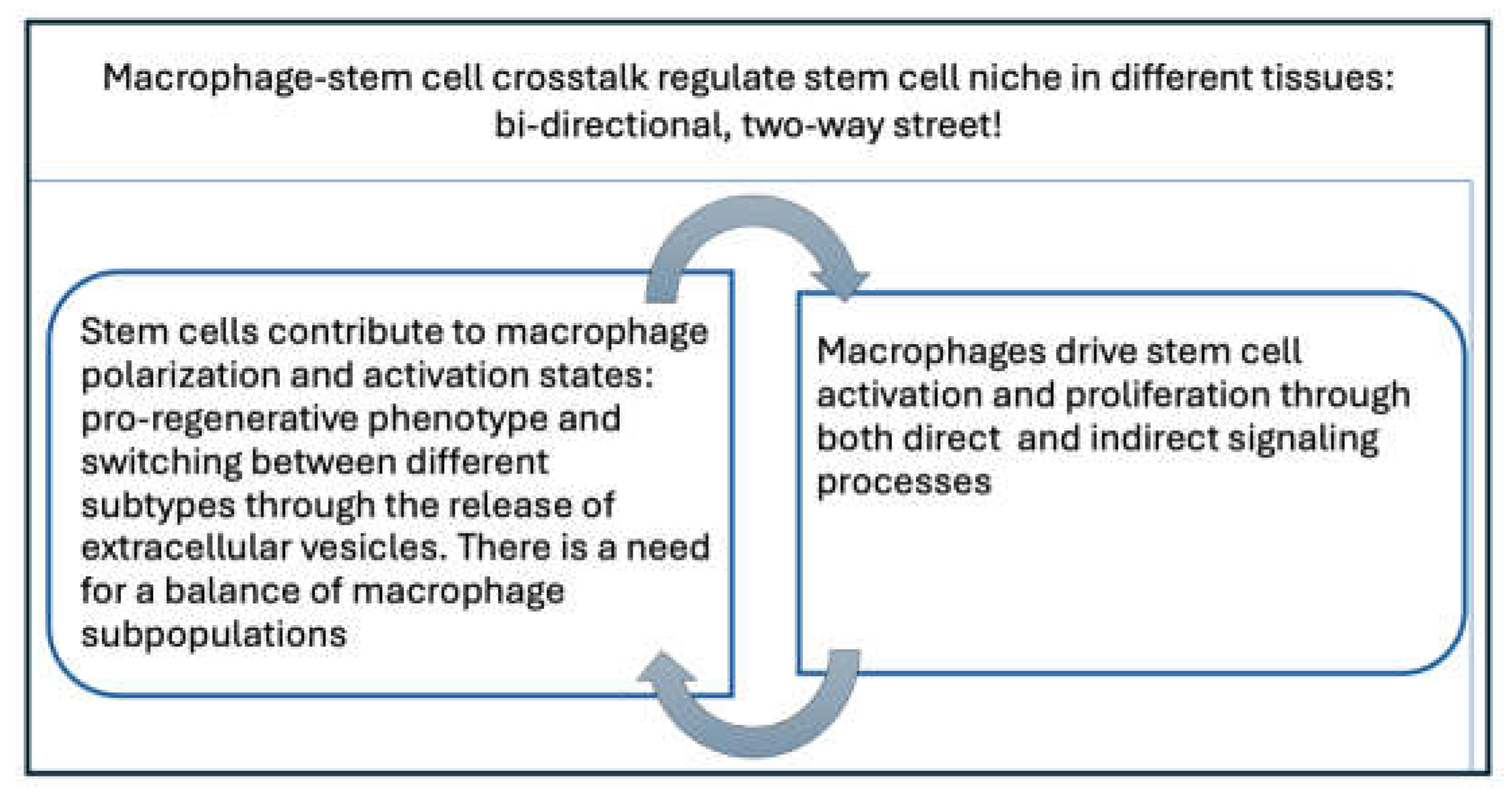
- Jessica D. Manneken, Peter D. Currie. Macrophage-stem cell crosstalk: regulation of stem cell niche. Development: stem cell & regenerative (2023) 150 (8): dev201510. [CrossRef]
- Jesús Cosin-Roger, M Dolores Ortiz-Masia and M Dolores Barrachina. Macrophages as an emerging source of WNT ligands: Relevance in mucosal integrity. Front. Immunol. Sec. Molecular Innate Immunity, 2019; 10. [CrossRef]
- Jia Zhao, Ilya Andreev and Hernandez Moura Silva. Resident tissue macrophages: Key coordinators of tissue homeostasis beyond immunity. Science Immunology, 2024; 9(94). [CrossRef]
- Karunakaran, K. B. Interactome-based framework to translate disease genetic data into biological and clinical insights. Thesis submitted for the PHD degree, University of Reading (2024). https://centaur.reading.ac.uk/119013/1/Karunakaran_thesis.pdf.
- Kefei Nina Li and Tudorita Tumbar. Hair follicle stem cell as a skin-organizing signaling center during adult homeostasis. EMBO J, 2021; 40: e107135. [CrossRef]
- Kiselev A and Park S. Immune niches for hair follicle development and homeostasis. Front Physiol, skin physiology, 2024;15:1397067. [CrossRef]
- Lin X., Liang Zhu and Jing He. Morphogenesis, growth cycle and molecular regulation of hair follicle. Frontiers in cell and development biology, 2022;10: 899095. [CrossRef]
- Liu, J., Xiao, Q., Xiao, J. et al. Wnt/β-catenin signaling function, biological mechanisms, and therapeutic opportunities. Sig Transduct Target Ther 7, 3 (2022). [CrossRef]
- Martel JL, Miao JH, Badri T, et al. Anatomy : Hair Follicle. [Updated 2024 Jun 22]. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2024 Jan-. Available from: https://www.ncbi.nlm.nih.gov/books/NBK470321/.
- Martin Guilliams, Guilhem R. Thierry, Johnny Bonnardel, et al. Establishment and maintenance of the macrophage niche. Immunity, 2020; 52(3): 434-451. [CrossRef]
- Martino PA, Heitman N, Rendl M. The dermal sheath: An emerging component of the hair follicle stem cell niche. Exp Dermatol. 2021; 30(4): 512-521. [CrossRef]
- Mass, E., Nimmerjahn, F., Kierdorf, K. et al. Tissue-specific macrophages: how they develop and choreograph tissue biology. Nat Rev Immunol, 2023; 23, 563–579. [CrossRef]
- Mao MQ, Jing J, Miao YJ, Lv ZF. Epithelial-Mesenchymal Interaction in Hair Regeneration and Skin Wound Healing. Front Med (Lausanne). 2022 Apr 14;9:863786. [CrossRef]
- Michael Wainberg, Nasa Sinnott-Armstrong, Nicholas Mancuso et al. Opportunities and challenges for transcriptome-wide association studies. Nature Genetics, 2019; 51: 592-599. [CrossRef]
- Matamá T, Costa C, Fernandes B, et al. Changing human hair fiber color and shape from the follicle. J Adv Res. 2024 Oct;64:45-65. [CrossRef]
- Ogunbawo, A. R., Henrique A. Mulim, Gabriel S. Campos, et al. Genetic Foundations of Nellore Traits: A Gene Prioritization and Functional Analyses of Genome-Wide Association Study Results. Preprint.org, 14 August 2024. [CrossRef]
- P. C. Agu, C. A. Afiukwa , O. U. Orji. Molecular docking as a tool for the discovery of molecular targets of nutraceuticals in diseases management. Nature Portfolio, Scientific Reports, 2023; 13: 13398. [CrossRef]
- Penelope A., Hirt MD, Ralf Pause MD. Healthy Hair Anatomy, Biology, Morphogenesis, cycling, and Function. Alopecia, 2019; Chapter 1: 1-22. (Author: Mariya Miteva., Elsevier). [CrossRef]
- Piyush Lakhani, Krashn K. Dwivedi, Atul Parashar, et al. Non-Invasive in Vivo Quantification of Directional Dependent Variation in Mechanical Properties for Human Skin. Front. Bioeng. Biotechnol., Sec. Biomechanics, 2021; 9. [CrossRef]
- Rahmani, W., Sinha, S. & Biernaskie, J. Immune modulation of hair follicle regeneration. npj Regen Med 5, 9 (2020). [CrossRef]
- Rompolas P, and Greco V. Stem cell dynamics in the hair follicle niche. Semin Cell Dev Biol., 2014; 25-26: 34-42. [CrossRef]
- Sarah Girardeau-Hubert, Solene Mine, Herve Pageon, et al. The Caucasian and African skin types differ morphologically and functionally in their dermal component. Expérimental Dermatologie; 2009; 18(8): 704-11. [CrossRef]
- Shuaifei Ji, Ziying Zhu, Xiaoyan Sun, et al. Functional hair follicle regeneration: an updated review. Signal Transduct Target Ther. 2021; 6(1):66. http. [CrossRef]
- Shwartz Y, Gonzalez-Celeiro M, Chen CL, et al. Cell Types Promoting Goosebumps Form a Niche to Regulate Hair Follicle Stem Cells. Cell. 2020; 182(3): 578-593.e19. [CrossRef]
- Meli VS, Veerasubramanian PK, Atcha H, Reitz Z, Downing TL, Liu WF. Biophysical regulation of macrophages in health and disease. J Leukoc Biol. 2019 Aug;106(2):283-299. [CrossRef]
- Ya-Chieh Hsu, Lishi Li, and Elaine Fuchs. Emerging interactions between skin stem cells and their niches. Nat Med, 2014; 20(8): 847-856. [CrossRef]
- Zhang, B., Chen, T. Local and systemic mechanisms that control the hair follicle stem cell niche. Nat Rev Mol Cell Biol, 2024; 25: 87–100. [CrossRef]
| Test | Cell type | Ref. |
| Macrophage phenotype classification using machine learning methods (MATLAB) related to expressed markers | Macrophages are heterogenous population including M1 to M2d | Hourani T. et al, 2023 |
| Macrophage autofluorescence | Macrophage phenotypes | Same |
| Molecular docking as a tool for the discovery of molecular targets of nutraceuticals in disease management |
Agu et al., 2019 |
|
|
Prediction of protein-protein interaction interface |
Using machine learning algorithm, molecular docking and post-docking analyses, using docking tools and techniques for the prediction of probable interaction surface. |
Das & Chakrabarti |
|
TWAS |
Integrate GWAS and gene expression datasets to identify gene-trait associations. |
Wainberg et al, 2019 |
| Multi-channel/multi-wavelength flow cytometer | extracted signals of phenotypes | same |
| Single-cell RNA-seq library construction, and bioinformatic analysis of single=cell RNA-seq data |
Mice anagen back skin/ MACs | Rahmani et al., 2020 |
| Putative MAC to HF interactome: Unbiased analysis of top predicted interactions package revealed immune ligands to which DS have transcriptionally expressed receptors | MAC ligands and dermal sheath receptors in cyclic regenerative behavior |
same |
| Unbiased analysis of top predicted interactions package revealed immune ligands to which MAC have transcriptionally expressed receptors | Dermal sheath ligand and MAC receptors |
same |
| Mechanism elucidation: Network biology and molecular interaction mapping discovering etiological genetic factors and functional implication | Breakdown of protein-protein interaction (PPI) network or the “interactome”, due to the genetic variants. And pharmacological effect of drug within the PPI integrating computationally predicted PPIs to circumvent interactome sparsity. There is reference to MACs and Wnt is rheumatoid arthritis | Karunakaran PHD thesis |
| Functional category enrichment analysis of MAC activation in response to mechanical stretch | Volcano plot of differently expressed genes (DEGs) in immune response gene ontology expression value in mice skin | Chu et al., 2019 |
| Potential manipulation of cellular processes through simple mechanical stimulation | A counterbalance between Wnt and BMP-2 and the subsequent two-step mechanism are identified through molecular and genetic analyses. |
Same |
| Define MAC phenotype and subpopulation specification across activated population by transcriptional subtype or effector molecule(s) | Characterize MACs based on different “activation paths” using predictive modelling to integrate multi-tissue data, mitochondrial transfer. | Manneken and Currie 2023; Sanin et al., 2022 |
| direct cell-cell contact visualized by in vivo visualization tools | Understanding the heterogeneity of MAC population, both transcriptionally and behaviorally. |
same |
| Double immunofluorescence with primary antibodies and labelled 2nd antibodies | Whole mount skin preparations | Fujiwara et al., 2011 |
| Quantitative RT-PCR | Total RNA was isolated using RNeasy Mint Kit with an on-column DNase I digestion. RT-PCR was performed using SYBR green super mix. The expression level of target genes was normalized utilizing standard curve method. |
same |
| Bioconductor annotation package “org.Hs.eg.db” for the genome-wide annotation using mapping Entre gene identifier for humans, as the prioritization analysis also used for Homo sapiens interaction network | Functional Analysis Enrichment analysis of the significant prioritized genes was performed using the “Clusterprofiler” |
Ogunbawo et al., 2024 |
| Flow cytometric gating strategy | Used to identify skin myeloid lineages, including macrophages | Liu Z. et al., 2022 |
| Rt.qPCR analysis for the expression of genes encoding ligands for Wnt and BMP pathways | Mice skin | same |
| in order to investigate the transcriptome along the three different axes, we used a study design that combines IPSC technology, high-throughput screening and allele-specific expression analyses. The cell type context has the overall strongest effect on gene expression, splicing, and allelic expression. |
Findley et al., 2021 |
| Type of study | Type of treatment | Ref |
| reviewed treatments targeting Wnt/B-catenin pathway for developing therapies for hair loss | using plant-derived chemicals in various in vitro and in vivo studies, e.g., Ginkgo biloba extract, red ginseng oil and Chinese mallow | Bu Young Choi (2020) |
| studied the molecular mechanism of natural products activating Wnt/β-catenin for improving hair loss | P. Mira nut oil, flavonoids, terpenoids from brown algae and Oleanolic acid (pentacyclic triterpenoid) | Dong Wook Shin (2022) |
|
Same above author in a previous study |
macrophages-extracellular vesicles including Wnt3a and Wnt7b enhanced the proliferation rate of dermal papillary cells. | Dong Wook Shin (2022 |
| new study finds a pool of genes with various levels of transcription associated with different hair shapes that points to epigenetic regulation, which means that gene expression can be affected by environmental factors. | They found that production of curved hair fiber may require proliferation support that can be provided by antioxidants and glutathione metabolites | Matama et al. (2023) |
| Genomic variation in textured hair has implications in developing holistic hair care routine | The review proposes a better understanding of the genetic traits, molecular structure and biomechanics of textured hair to initiate a more effective personalized haircare solution. | (Idowu et al., 2024). |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




