Submitted:
02 November 2024
Posted:
06 November 2024
You are already at the latest version
Abstract
Accurate prediction of pregnancy risk levels is essential for preventing maternal and fetal complications. This study explores the application of ensemble machine learning models combined with oversampling techniques to predict pregnancy risk levels. We utilized a publicly available dataset from the UCI Machine Learning Repository, performing extensive feature engineering, including the introduction of new features like Pulse Pressure and Mean Arterial Pressure. To address class imbalance, we employed the Adaptive Synthetic Sampling (ADASYN) method. We conducted comprehensive hyperparameter tuning to enhance model performance and achieve optimal predictive results. Additional evaluation metrics, including sensitivity, specificity, and precision-recall curves, were used to assess model per- formance comprehensively. Our findings demonstrate that the Voting Classifier, particularly when combined with ADASYN oversampling and optimized hyperpa- rameters, achieves an accuracy of 87.19% and a macro F1 score of 87.66%, effec- tively distinguishing between ’low risk’ and ’mid risk’ pregnancy cases. This work contributes to the field by enhancing prediction accuracy, providing insights into important features influencing pregnancy risk, and addressing ethical considerations in deploying machine learning models in healthcare.
Keywords:
1. Introduction
1.1. Recent Advances in Pregnancy Risk Prediction
2. Materials and Methods
2.1. Dataset Description
Features Included
Class Distribution Before Resampling
Data Ethics and Permissions
2.2. Data Preprocessing
2.2.1. Missing Values
2.2.2. Outlier Detection and Removal
2.2.3. Label Encoding
2.2.4. Feature Scaling
2.3. Feature Engineering and Exploratory Data Analysis (EDA)
2.3.1. Incorporation of Cardiovascular Features
2.3.2. Additional Engineered Features
2.3.3. Correlation Analysis
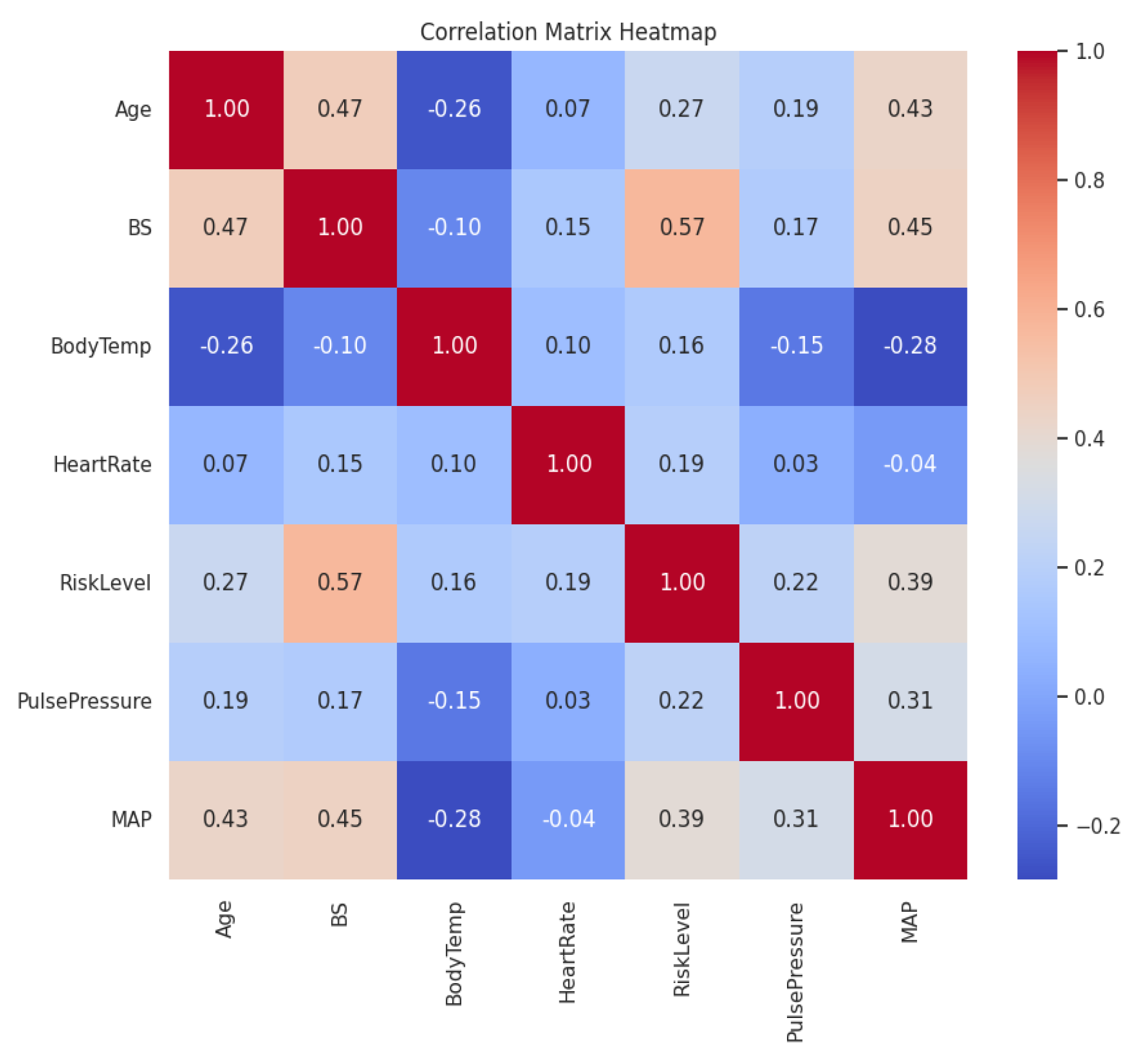
2.3.4. Visualization of Feature Distributions
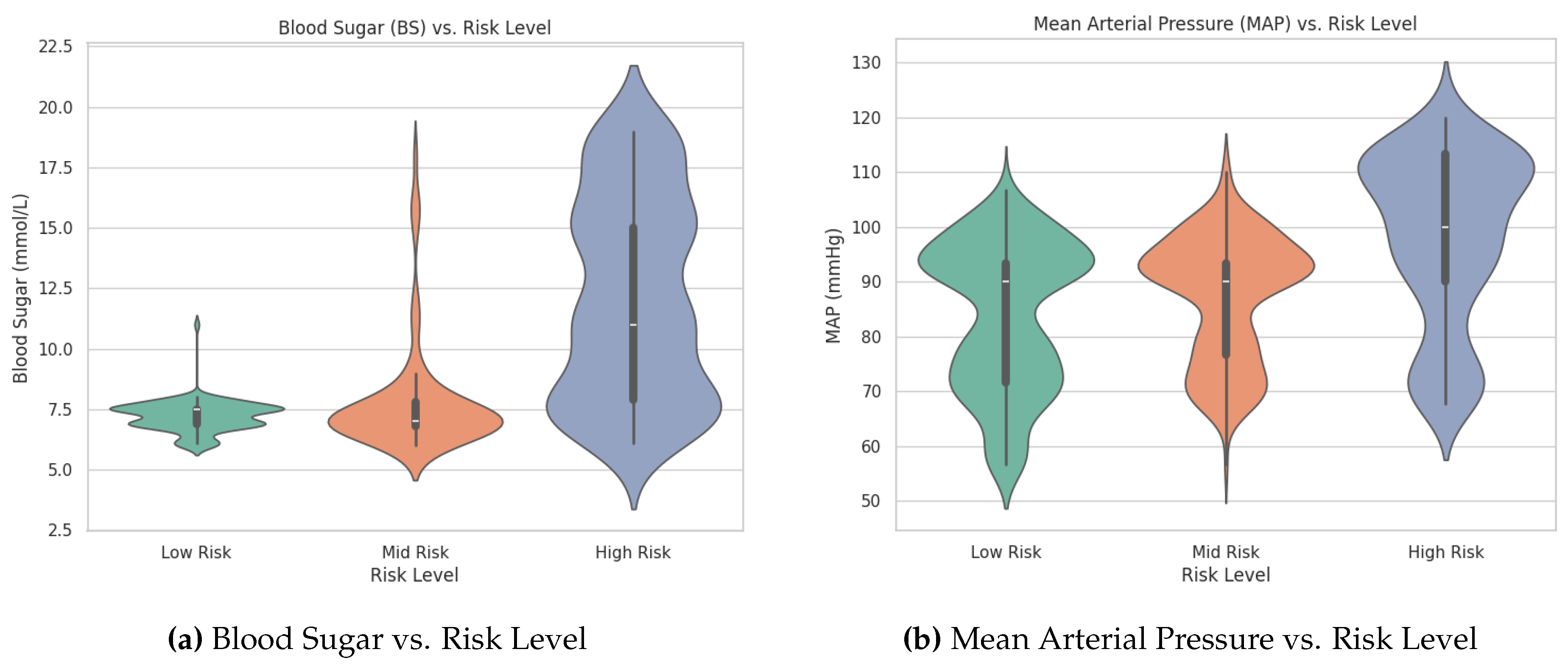
2.3.5. One-Way ANOVA Tests
2.4. Handling Class Imbalance
2.4.1. Oversampling Method Used
2.4.2. Class Distribution After ADASYN
2.4.3. Model Development
2.4.4. Machine Learning Algorithms
2.4.5. Hyperparameter Tuning
2.4.6. Evaluation Metrics
2.5. Statistical Analysis
2.6. Cross-Validation
3. Results
3.1. Random Forest
3.1.1. Evaluation Metrics Explained
Precision
Recall
F1-score
Support
Macro Average vs. Weighted Average
3.1.2. Interpretation and Considerations
3.1.3. Summary of Results
3.2. XGBoost
3.2.1. Interpretation of Results
3.3. Voting Classifier
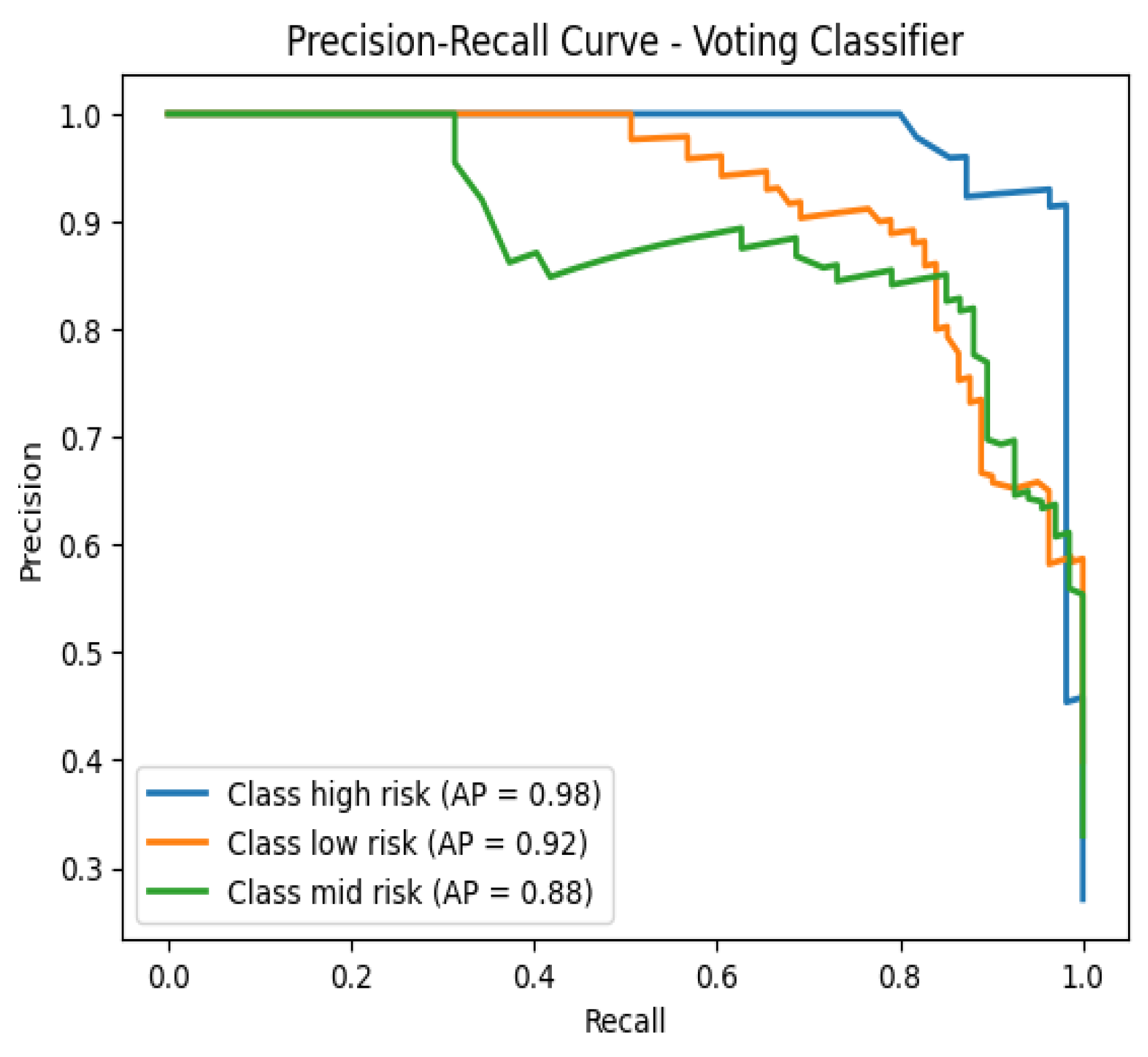
3.4. Cross-Validation Results
3.5. AUC-ROC Analysis
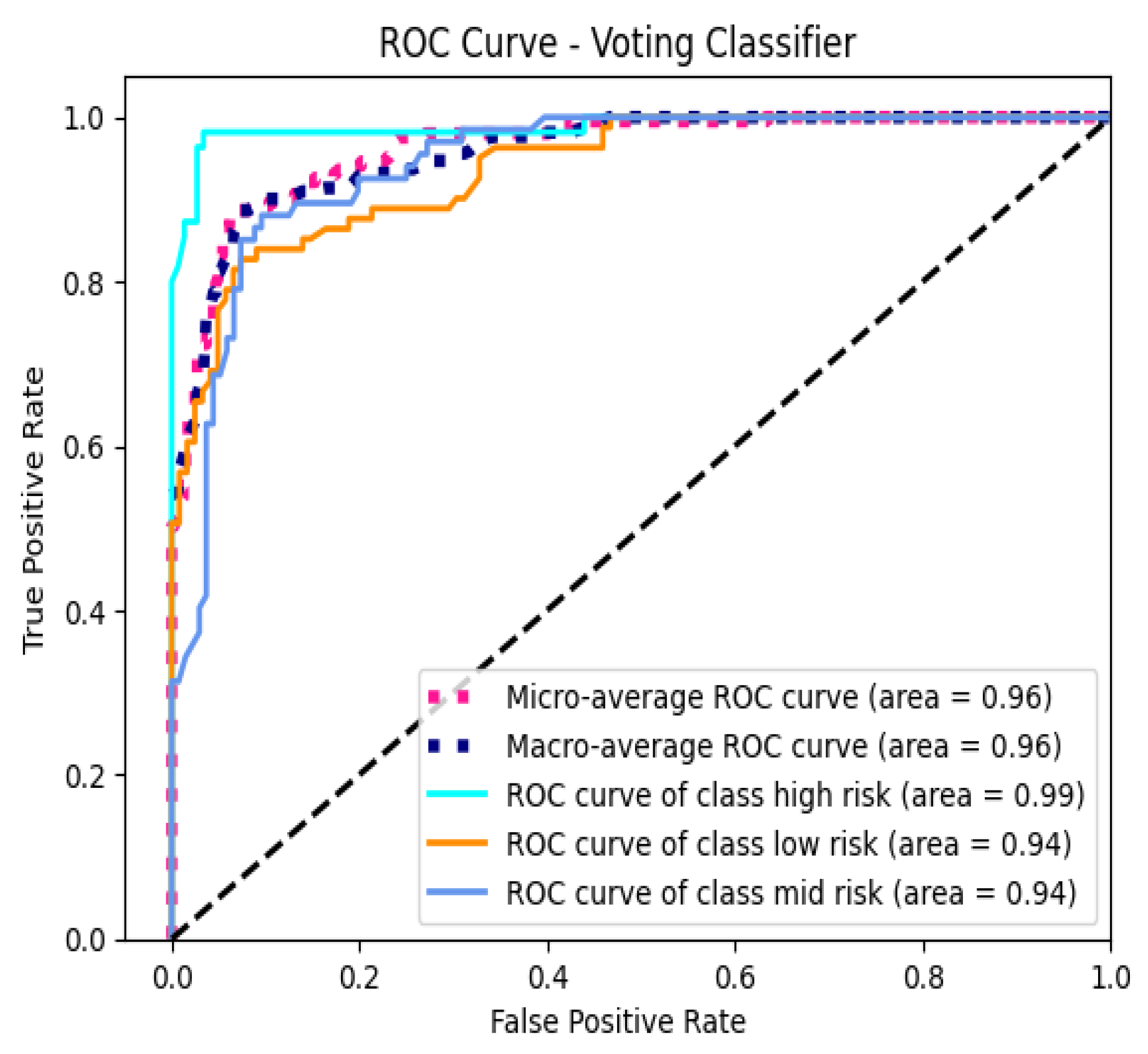
3.6. Feature Importance Analysis
3.7. Misclassification Analysis
| No. | Age | BS | BodyTemp | HeartRate | PP | MAP | True → Predicted |
|---|---|---|---|---|---|---|---|
| 1 | -1.0297 | -0.5562 | -0.4927 | 0.6814 | -0.5782 | -0.5851 | Low Risk → Mid Risk |
| 2 | 0.9005 | 2.8084 | -0.4927 | -0.5276 | 1.1532 | 0.5478 | High Risk → Mid Risk |
| 3 | 2.2368 | -0.2834 | -0.4927 | 0.0769 | 0.2875 | 0.3212 | High Risk → Low Risk |
| 4 | -1.1040 | -0.8290 | -0.4927 | -0.5276 | -1.4439 | -2.1711 | Mid Risk → Low Risk |
| 5 | -0.5843 | -0.5865 | -0.4927 | 0.3187 | -0.5782 | -1.2648 | Mid Risk → Low Risk |
3.8. Statistical Analysis
3.9. Overall Findings
4. Discussion
4.1. Model Performance and Ensemble Advantage
4.2. Importance of Feature Selection
4.3. Addressing Class Imbalance with ADASYN
4.4. Model Discrimination and AUC-ROC Analysis
4.5. Misclassification Insights
4.6. Statistical Significance and Model Comparison
4.7. Clinical Implications
4.8. Limitations
- Sample Size: The dataset’s relatively small size may limit the generalizability of the findings. Larger datasets encompassing diverse populations are necessary to ensure that the models perform consistently across different demographic and clinical groups.
- Class Imbalance: Although ADASYN effectively addressed class imbalance, synthetic data generation methods may not fully capture the inherent variability of real-world data. This limitation could affect the model’s performance when deployed in varied clinical settings where data distributions may differ.
- Feature Scope: The study was restricted to a specific set of physiological measurements, excluding potentially relevant features such as genetic information, detailed medical histories, lifestyle factors, and socio-economic indicators. Incorporating a broader range of features could enhance the models’ predictive capabilities and provide a more comprehensive risk assessment.
- Risk of Overfitting: Training and testing the models on the same dataset, even with cross-validation, poses a risk of overfitting. Overfitted models may perform well on the training data but fail to generalize to unseen data. External validation using independent datasets is essential to evaluate the models’ robustness and real-world applicability.
- Model Interpretability: While ensemble methods offer superior performance, they are often perceived as "black-box" models, making it challenging to interpret the underlying decision-making processes. Enhancing model interpretability is crucial for clinical acceptance and trust.
- Temporal Dynamics: The dataset did not account for temporal changes in physiological measurements throughout pregnancy. Incorporating time-series data could provide insights into the progression of risk levels and improve predictive accuracy.
- Ethical Considerations: The study utilized anonymized, publicly available data. However, ethical considerations regarding data privacy, informed consent, and potential biases in model predictions need continuous attention, especially when deploying models in sensitive healthcare settings.
5. Future Work
6. Conclusion
Acknowledgments
References
- World Health Organization. Maternal mortality: key facts. https://www.who.int/news-room/fact-sheets/detail/maternal-mortality, 2019. Accessed: [Your Access Date].
- Johnson, J.M.; Khoshgoftaar, T.M. Survey on deep learning with class imbalance. Journal of Big Data 2019, 6, 1–54. [Google Scholar] [CrossRef]
- Kavakiotis, I.; Tsave, O.; Salifoglou, A.; Maglaveras, N.; Vlahavas, I.; Chouvarda, I. Machine learning and data mining methods in diabetes research. Computational and Structural Biotechnology Journal 2017, 15, 104–116. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Chen, Q.; Ma, L. Machine learning in prediction of preterm birth. Journal of Translational Medicine 2019, 17, 18. [Google Scholar]
- Tran, K.A.; et al. Deep learning as a tool for prediction of preeclampsia in pregnant women. npj Digital Medicine 2019, 2, 61. [Google Scholar]
- Say, L.; Chou, D.; Gemmill, A.; Tuncalp, O.; Moller, A.B.; Daniels, J.; Gülmezoglu, A.M.; Temmerman, M.; Alkema, L. Global causes of maternal death: a WHO systematic analysis. The Lancet Global Health 2014, 2, e323–e333. [Google Scholar] [CrossRef] [PubMed]
- He, H.; Garcia, E.A. Learning from imbalanced data. IEEE Transactions on Knowledge and Data Engineering 2009, 21, 1263–1284. [Google Scholar] [CrossRef]
- Sakr, S.; et al. Predicting preeclampsia: A comparison of machine learning techniques. Healthcare 2021, 9, 1703. [Google Scholar] [CrossRef]
- Sufriyana, H.; Wu, Y.W.; Su, E.C.Y.; et al. Comparison of machine learning approaches to classify maternal anemia levels. BMC Medical Informatics and Decision Making 2020, 20, 172. [Google Scholar]
- Zhang, J.; et al. Hybrid deep learning model for predicting gestational diabetes mellitus. IEEE Access 2022, 10, 13303–13312. [Google Scholar]
- Calwin, C. Predicting Pregnancy Risk Levels with Machine Learning. https://www.kaggle.com/code/calwin9/predicting-pregnancy-risk-levels-with-machine-lear, 2021. Accessed: [Your Access Date].
- Dua, D.; Graff, C. UCI Machine Learning Repository. https://archive.ics.uci.edu/ml/datasets/Maternal+Health+Risk+Data+Set, 2020. Accessed: [Your Access Date].
- Loerup, L.; Pullon, R.M.C.; Birks, J.; Fleming, S.; Mackillop, L.; Tarassenko, L. Trends of blood pressure and heart rate in normal pregnancies: a systematic review and meta-analysis. BMC Medicine 2019, 17, 167. [Google Scholar] [CrossRef] [PubMed]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; Vanderplas, J.; Passos, A.; Cournapeau, D.; Brucher, M.; Perrot, M.; Duchesnay, E. Scikit-learn: Machine Learning in Python. Journal of Machine Learning Research 2011, 12, 2825–2830. [Google Scholar]
- James, G.; Witten, D.; Hastie, T.; Tibshirani, R. An Introduction to Statistical Learning: With Applications in R; Springer Texts in Statistics, Springer, 2013. [CrossRef]
- Hall, J.E. Guyton and Hall Textbook of Medical Physiology, 13th ed.; Elsevier Health Sciences, 2016.
- Mayet, J.; Hughes, A. Cardiac and vascular pathophysiology in hypertension. Heart 2003, 89, 1104–1109. [Google Scholar] [CrossRef] [PubMed]
- Montgomery, D.C. Design and Analysis of Experiments, 9th ed.; Wiley, 2017.
- Moore, D.S.; McCabe, G.P.; Craig, B.A. Introduction to the Practice of Statistics, 9th ed.; W. H. Freeman, 2017.
- He, H.; Bai, Y.; Garcia, E.A.; Li, S. ADASYN: Adaptive Synthetic Sampling Approach for Imbalanced Learning. IEEE International Joint Conference on Neural Networks, 2008, pp. 1322–1328. [CrossRef]
- Gaso, M.S.; Mekuria, R.R.; Khan, A.; Gulbarga, M.I.; Tologonov, I.; Sadriddin, Z. Utilizing Machine and Deep Learning Techniques for Predicting Re-admission Cases in Diabetes Patients. Proceedings of the International Conference on Computer Systems and Technologies 2024; Association for Computing Machinery: New York, NY, USA, 2024; CompSysTech ’24, p. 76–81. [CrossRef]
- Tologonov, I.; Mekuria, R.R.; Istamov, K.; Gaso, M.S. Detection of Tuberculosis Using Convolutional Neural Network. EasyChair Preprint 1 3500, EasyChair, 2024. [Google Scholar]
- Breiman, L. Random Forests. Machine Learning 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, 2016, pp. 785–794. [CrossRef]
- Kuncheva, L.I. Combining Pattern Classifiers: Methods and Algorithms; Wiley-Interscience, 2004.
| Feature | F-statistic | p-value |
|---|---|---|
| Blood Sugar (BS) | 331.16 | < 0.0001 |
| Mean Arterial Pressure (MAP) | 98.79 | < 0.0001 |
| Pulse Pressure (PP) | 32.26 | < 0.0001 |
| Class | Precision | Recall | F1-score | Support |
|---|---|---|---|---|
| High Risk | 0.91 | 0.96 | 0.94 | 55 |
| Low Risk | 0.86 | 0.80 | 0.83 | 81 |
| Mid Risk | 0.80 | 0.82 | 0.81 | 67 |
| Macro Average | 0.86 | 0.86 | 0.86 | 203 |
| Weighted Average | 0.85 | 0.85 | 0.85 | 203 |
| Class | Precision | Recall | F1-score | Support |
|---|---|---|---|---|
| High Risk | 0.91 | 0.93 | 0.92 | 55 |
| Low Risk | 0.90 | 0.79 | 0.84 | 81 |
| Mid Risk | 0.78 | 0.88 | 0.83 | 67 |
| Macro Avg | 0.86 | 0.87 | 0.86 | 203 |
| Weighted Avg | 0.86 | 0.86 | 0.86 | 203 |
| Class | Precision | Recall | F1-score | Support |
|---|---|---|---|---|
| High Risk | 0.91 | 0.96 | 0.94 | 55 |
| Low Risk | 0.88 | 0.83 | 0.85 | 81 |
| Mid Risk | 0.83 | 0.85 | 0.84 | 67 |
| Macro Avg | 0.87 | 0.88 | 0.88 | 203 |
| Weighted Avg | 0.87 | 0.87 | 0.87 | 203 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




