Submitted:
20 October 2024
Posted:
21 October 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
Patient’s Characteristics
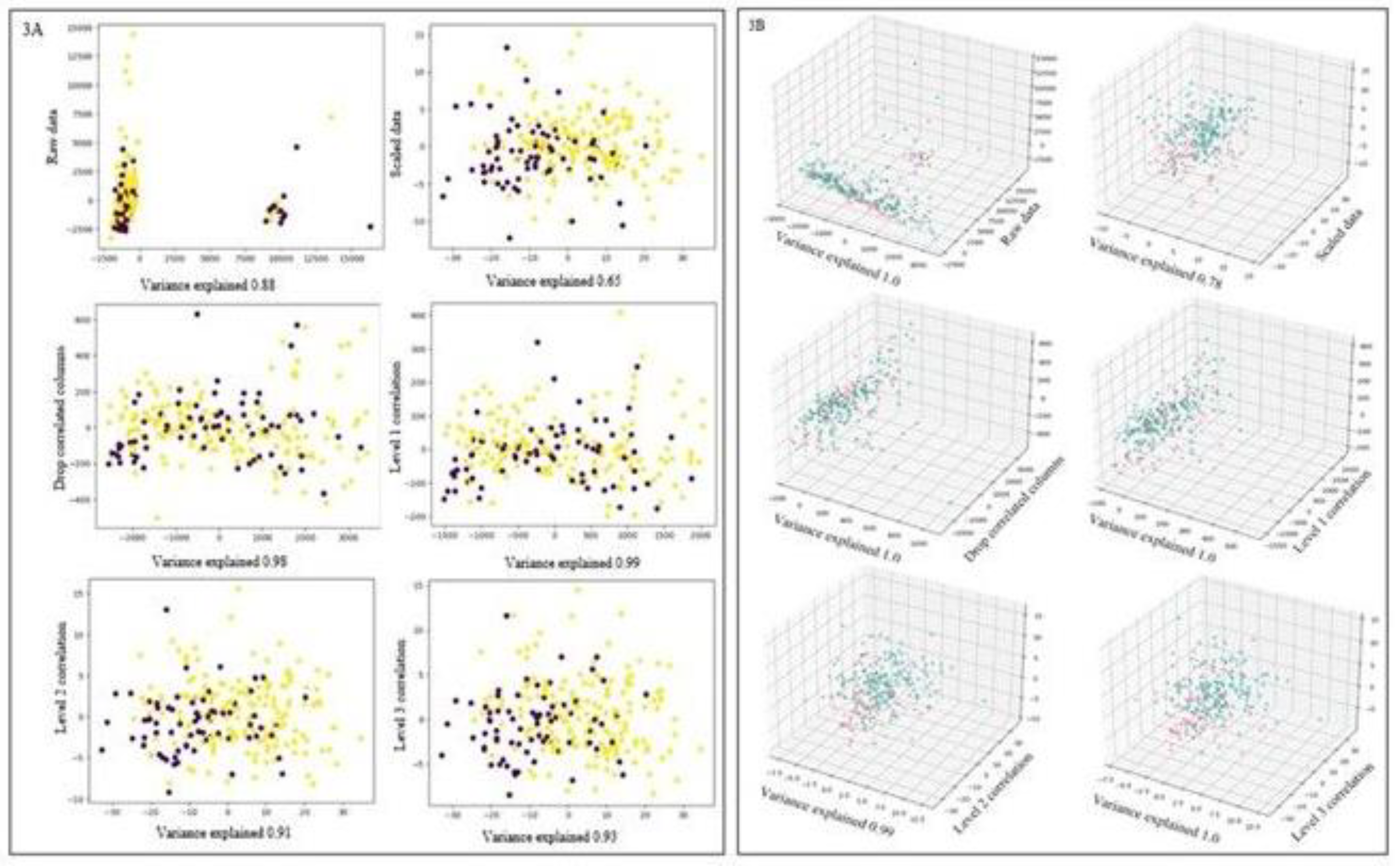
Metabolomics Profiling and Model Creation
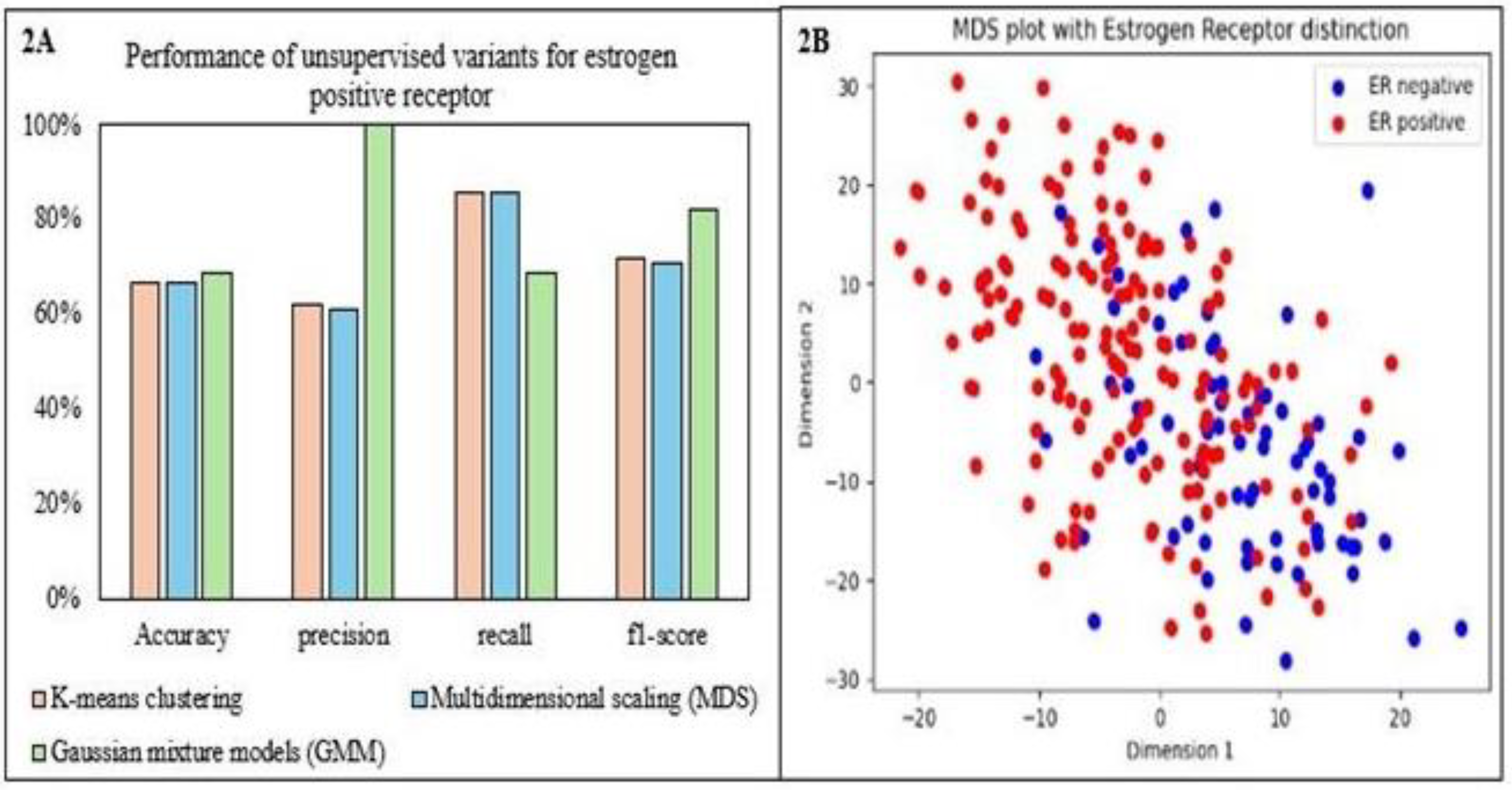
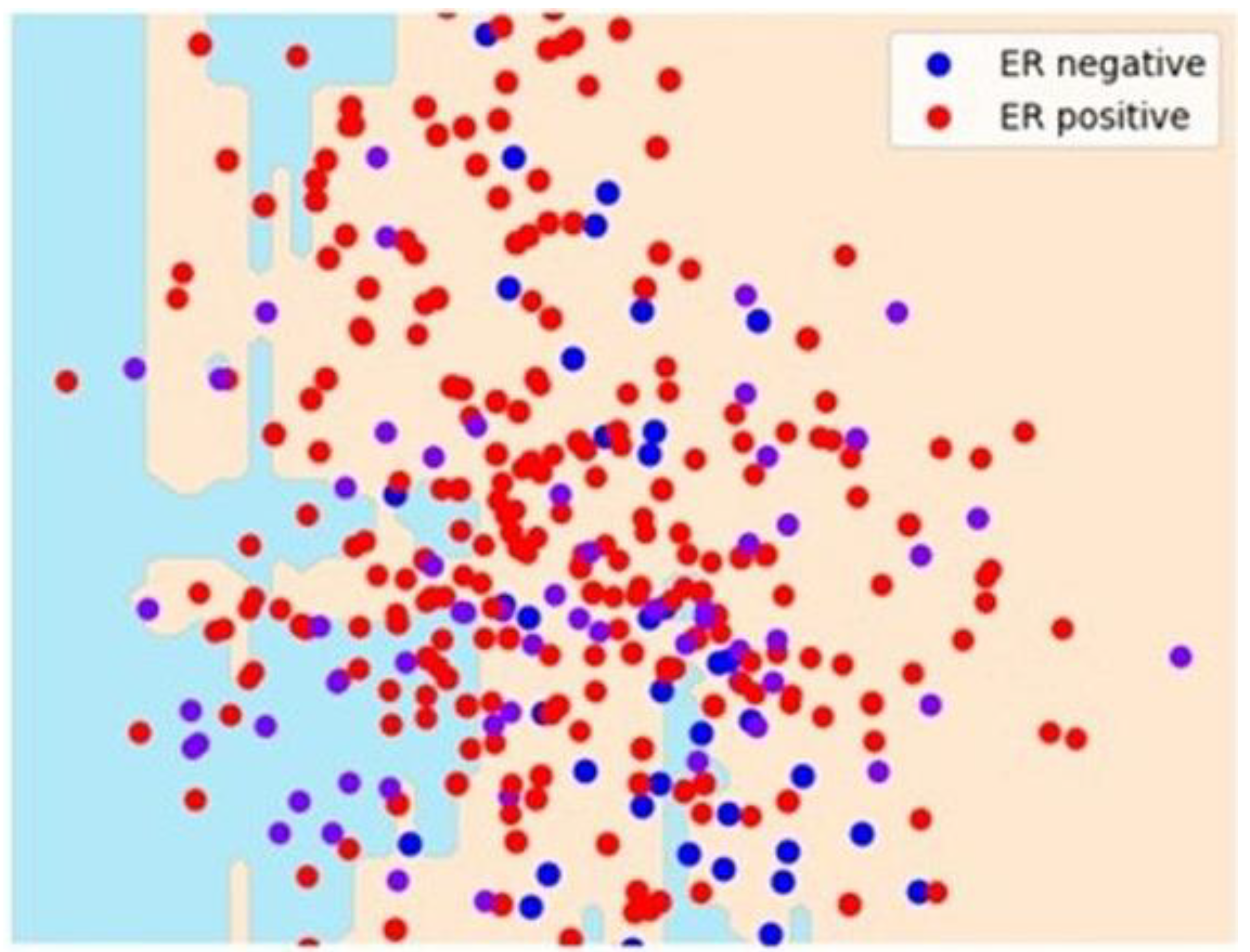
Unsupervised Models
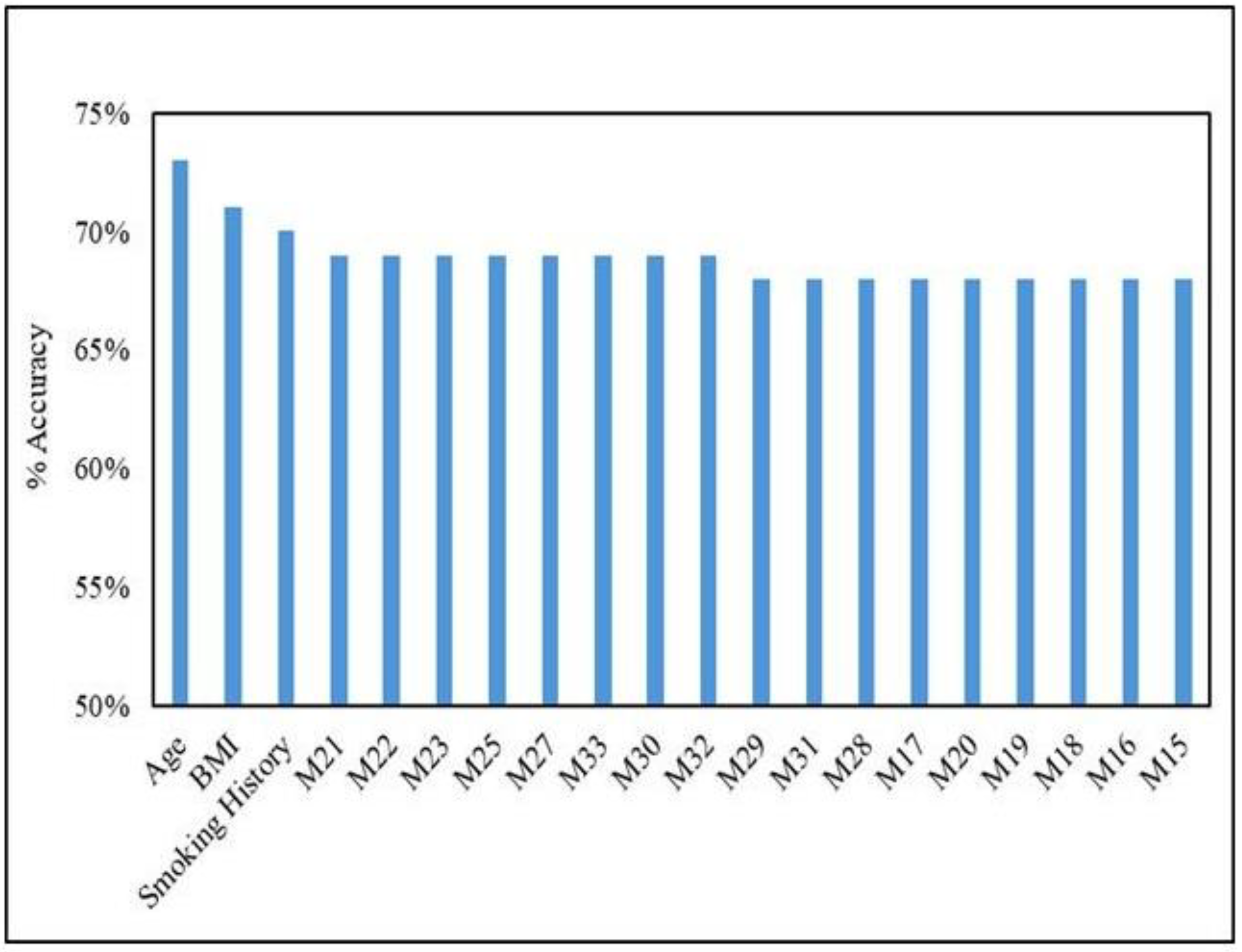
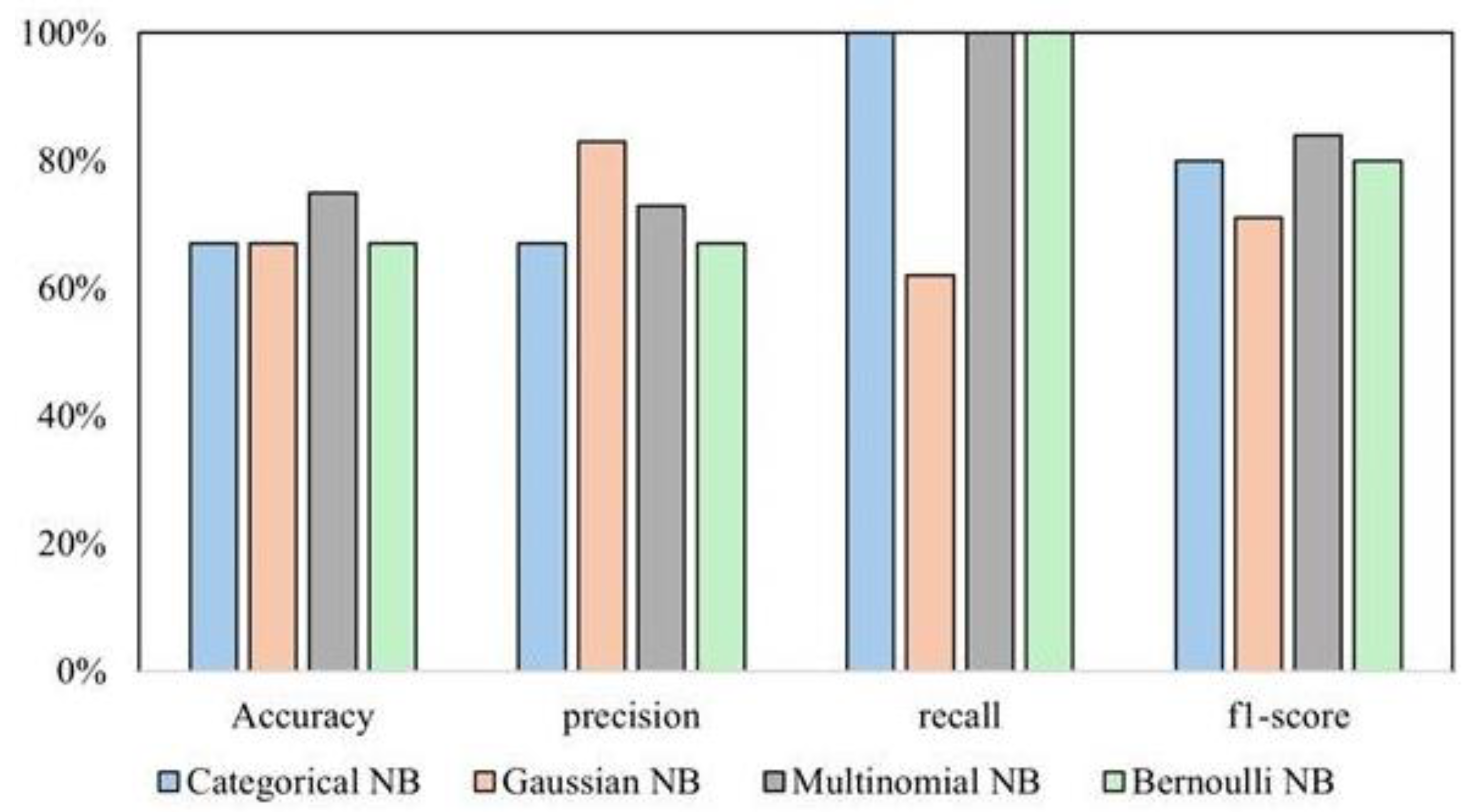
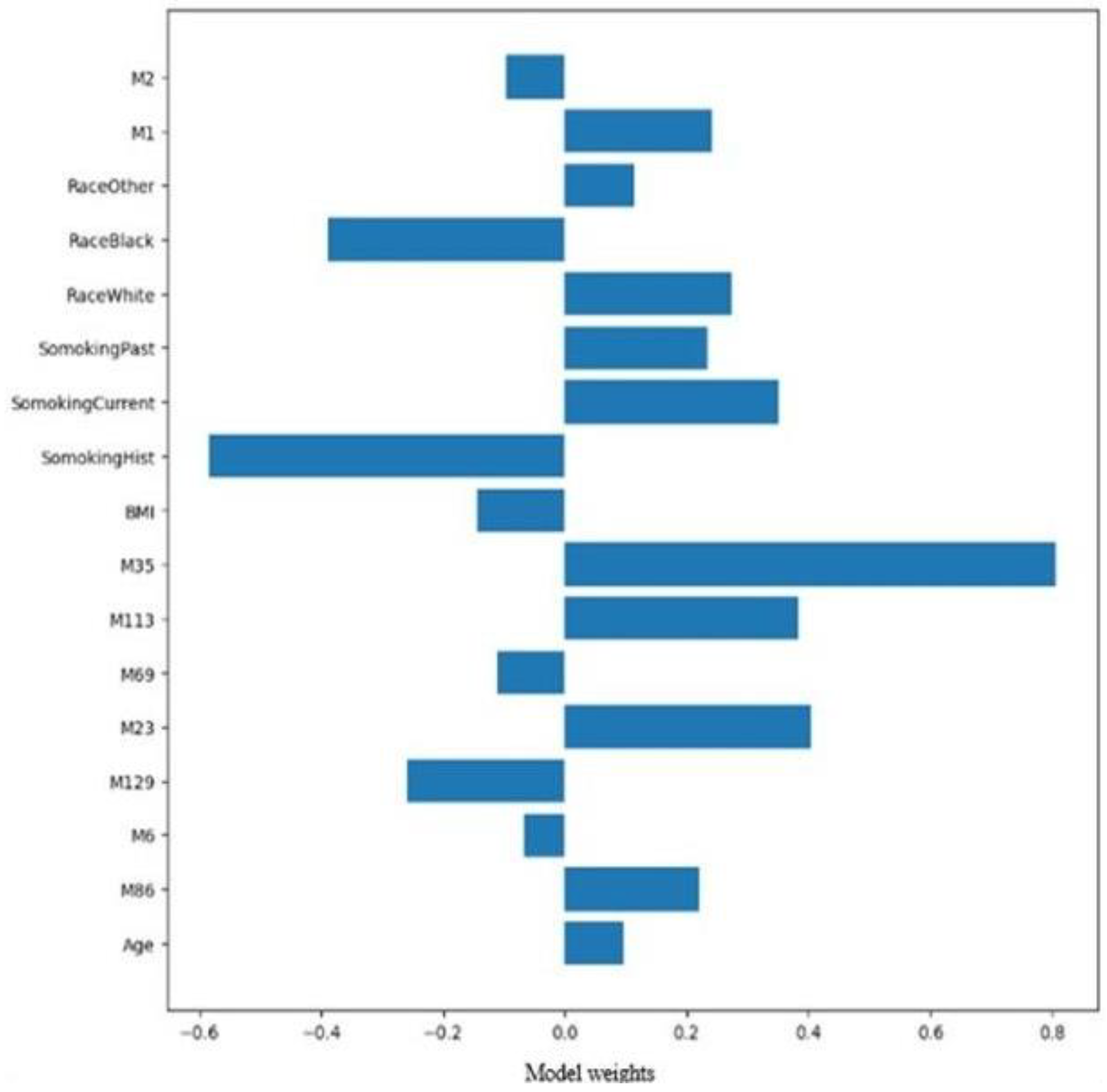
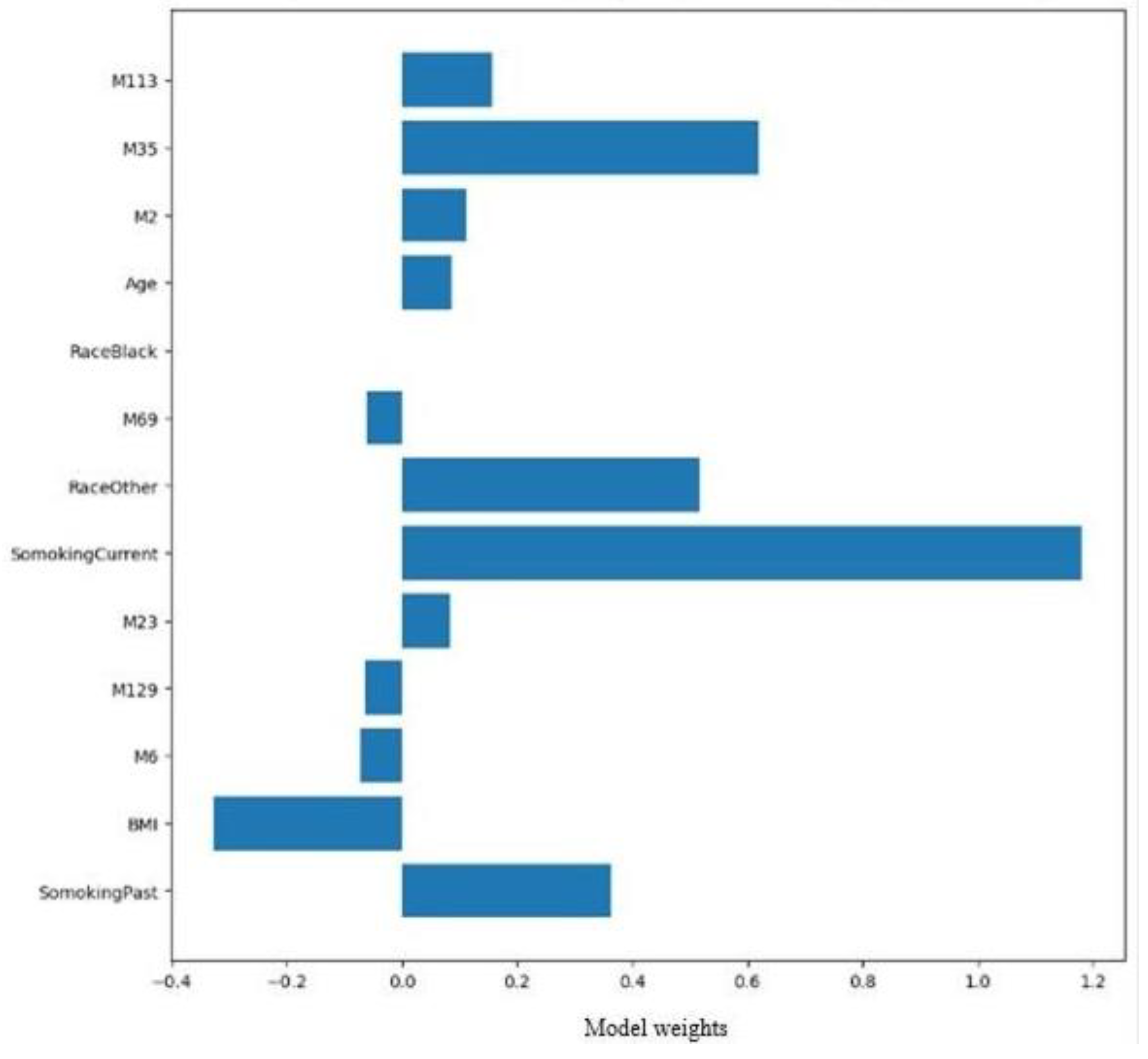
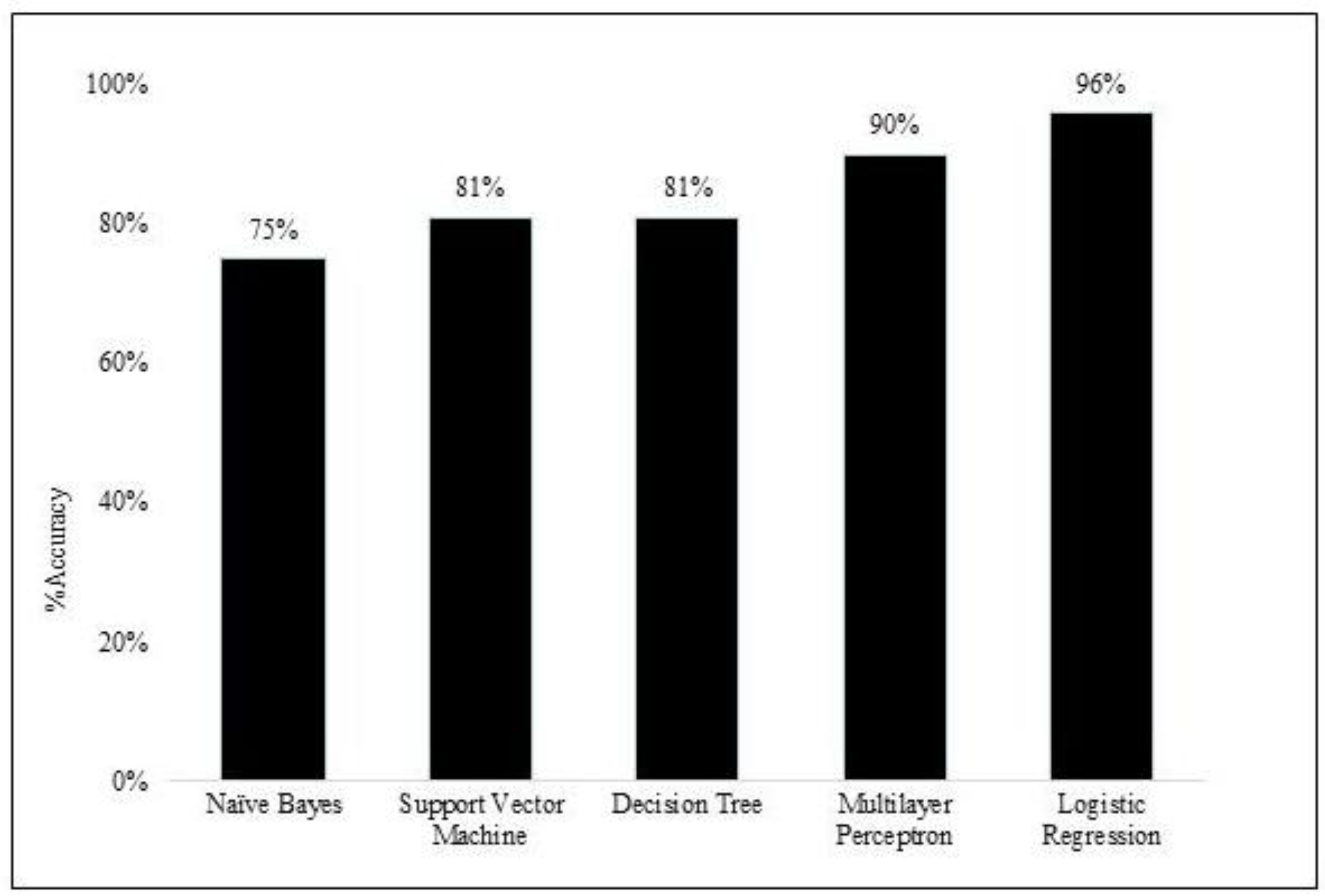
Supervised Models
3. Discussion
4. Materials and Methods
Participants and LC-HRMS Analysis
Data Processing
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Fuhrman, B.J.; Schairer, C.; Gail, M.H.; Boyd-Morin, J.; Xu, X.; Sue, L.Y.; Buys, S.S.; Isaacs, C.; Keefer, L.K.; Veenstra, T.D.; Berg, C.D.; Hoover, R.N.; Ziegler, R.G. Estrogen metabolism and risk of breast cancer in postmenopausal women. J Natl Cancer Inst 2012, 104, 326–339. [Google Scholar] [CrossRef] [PubMed]
- Parise, C.A.; Caggiano, V. Breast cancer survival is defined by the er/pr/her2 subtypes and a surrogate classification according to tumor grade and immunohistochemical biomarkers. J Cancer Epidemiol 2014, 2014, 469251. [Google Scholar] [CrossRef] [PubMed]
- Cicatiello, L.; Mutarelli, M.; Grober, O.M.; Paris, O.; Ferraro, L.; Ravo, M.; Tarallo, R.; Luo, S.; Schroth, G.P.; Seifert, M.; Zinser, C.; Chiusano, M.L.; Traini, A.; De Bortoli, M.; Weisz, A. Estrogen receptor alpha controls a gene network in luminal-like breast cancer cells comprising multiple transcription factors and microRNAs. Am J Pathol 2010, 176, 2113–2130. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.; Lu, Y.; Malone, K.E.; Marchbanks, P.A.; Deapen, D.M.; Spirtas, R.; Burkman, R.T.; Strom, B.L.; McDonald, J.A.; Folger, S.G.; Simon, M.S.; Sullivan-Halley, J.; Press, M.F.; Bernstein, L. Mortality risk of black women and white women with invasive breast cancer by hormone receptors, HER2, and p53 status. BMC Cancer 2013, 13, 225. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.; Lu, Y.; Marchbanks, P.A.; Folger, S.G.; Strom, B.L.; McDonald, J.A.; Simon, M.S.; Weiss, L.K.; Malone, K.E.; Burkman, R.T.; Sullivan-Halley, J.; Deapen, D.M.; Press, M.F.; Bernstein, L. Quantitative measures of estrogen receptor expression in relation to breast cancer-specific mortality risk among white women and black women. Breast Cancer Res 2013, 15, R90. [Google Scholar] [CrossRef]
- Abubakar, M.; Figueroa, J.; Ali, H.R.; Blows, F.; Lissowska, J.; Caldas, C.; Easton, D.F.; Sherman, M.E.; Garcia-Closas, M.; Dowsett, M.; Pharoahz, P.D. Combined quantitative measures of ER, PR, HER2, and KI67 provide more prognostic information than categorical combinations in luminal breast cancer. Mod Pathol 2019, 32, 1244–1256. [Google Scholar] [CrossRef]
- Khande, T.A.; Joshi, A.R.; Khandeparkar, S.G.S.; Kulkarni, M.M.; Gogate, B.P.; Kakade, A.R.; Sahu, P.D.; Khillare, C.D. Study of ER, PR, HER2/neu, p53, and Ki67 expression in primary breast carcinomas and synchronous metastatic axillary lymph nodes. Indian J Cancer 2020, 57, 190–197. [Google Scholar]
- Wei, Y.; Jasbi, P.; Shi, X.; Turner, C.; Hrovat, J.; Liu, L.; Rabena, Y.; Porter, P.; Gu, H. Early breast cancer detection using untargeted and targeted metabolomics. J Proteome Res 2021, 20, 3124–3133. [Google Scholar] [CrossRef]
- Jobard, E.; Dossus, L.; Baglietto, L.; Fornili, M.; Lécuyer, L.; Mancini, F.R.; Gunter, M.J.; Trédan, O.; Boutron-Ruault, M.C.; Elena-Herrmann, B.; Severi, G.; Rothwell, J.A. Investigation of circulating metabolites associated with breast cancer risk by untargeted metabolomics: a case-control study nested within the French E3N cohort. Br J Cancer 2021, 124, 1734–1743. [Google Scholar] [CrossRef]
- Syed, A.H.; Khan, T. Evolution of research trends in artificial intelligence for breast cancer diagnosis and prognosis over the past two decades: A bibliometric analysis. Front Oncol 2022, 12, 854927. [Google Scholar] [CrossRef]
- Ahn, J.S.; Shin, S.; Yang, S.A.; Park, E.K.; Kim, K.H.; Cho, S.I.; Ock, C.Y.; Kim, S. Artificial intelligence in breast cancer diagnosis and personalized medicine. J Breast Cancer 2023, 26, 405–435. [Google Scholar] [CrossRef] [PubMed]
- Jasbi, P.; Wang, D.; Cheng, S.L.; Fei, Q.; Cui, J.Y.; Liu, L.; Wei, Y.; Raftery, D.; Gu, H. Breast cancer detection using targeted plasma metabolomics. J Chromatogr B Analyt Technol Biomed Life Sci 2019, 1105, 26–37. [Google Scholar] [CrossRef] [PubMed]
- Alakwaa, F.M.; Chaudhary, K.; Garmire, L.X. Deep learning accurately predicts estrogen receptor status in breast cancer metabolomics data. J Proteome Res 2018, 17, 337–347. [Google Scholar] [CrossRef] [PubMed]
- Gal, J.; Bailleux, C.; Chardin, D.; Pourcher, T.; Gilhodes, J.; Jing, L.; Guigonis, J.M.; Ferrero, J.M.; Milano, G.; Mograbi, B.; Brest, P.; Chateau, Y.; Humbert, O.; Chamorey, E. Comparison of unsupervised machine-learning methods to identify metabolomic signatures in patients with localized breast cancer. Comput Struct Biotechnol J 2020, 18, 1509–1524. [Google Scholar] [CrossRef]
- Henry, N.L.; Hayes, D.F. Cancer biomarkers. Mol Oncol 2012, 6, 140–146. [Google Scholar] [CrossRef]
- Haince, J.F.; Joubert, P.; Bach, H.; Ahmed Bux, R.; Tappia, P.S.; Ramjiawan, B. Metabolomic fingerprinting for the detection of early-stage lung cancer: From the genome to the metabolome. Int J Mol Sci 2022, 23, 1215. [Google Scholar] [CrossRef]
- Kim, Y.; Koo, I.; Jung, B.H.; Chung, B.C.; Lee, D. Multivariate classification of urine metabolome profiles for breast cancer diagnosis. BMC Bioinformatics 2010, 11, S4. [Google Scholar] [CrossRef]
- Hadi, N.I.; Jamal, Q.; Iqbal, A.; Shaikh, F.; Somroo, S.; Musharraf, S.G. Serum metabolomic profiles for breast cancer diagnosis, grading and staging by gas chromatography-mass spectrometry. Sci Rep 2017, 7, 1715. [Google Scholar] [CrossRef]
- Xiao, Y.; Ma, D.; Yang, Y.S.; Yang, F.; Ding, J.H.; Gong, Y.; Jiang, L.; Ge, L.P.; Wu, S.Y.; Yu, Q.; Zhang, Q.; Bertucci, F.; Sun, Q.; Hu, X.; Li, D.Q.; Shao, Z.M.; Jiang, Y.Z. Comprehensive metabolomics expands precision medicine for triple-negative breast cancer. Cell Res 2022, 32, 477–490. [Google Scholar] [CrossRef]
- Rawal, R. Breast cancer prediction using machine learning. JETIR 2020, 7, 13–24. [Google Scholar]
- Hassan, M.M.; Hassan, M.M.; Yasmin, F.; Khan, M.A.R.; Zaman, S. ; Galibuzzaman; Islam, K.K.; Bairagi, A.K.; A comparative assessment of machine learning algorithms with the least absolute shrinkage and selection operator for breast cancer detection and prediction. Decision Analytics J, 2023, 7, 100245. [Google Scholar]
- Kumar, P.; Samiha, S.P.; Gururaj, C.M.; Kumar, R. Prognostic analysis of machine learning techniques for breast cancer. Int J Bioinfor Intell Comput 2022, 1, 72–83. [Google Scholar]
- Rabiei, R.; Ayyoubzadeh, S.M.; Sohrabei, S.; Esmaeili, M.; Atashi, A. Prediction of breast cancer using ML approaches. J Biomed Phys Eng 2022, 12, 297–308. [Google Scholar] [CrossRef] [PubMed]
- Bauer, K.R.; Brown, M.; Cress, R.D.; Parise, C.A.; Caggiano, V. Descriptive analysis of estrogen receptor (ER)-negative, progesterone receptor (PR)-negative, and HER2-negative invasive breast cancer, the so-called triple-negative phenotype: A population-based study from the California Cancer Registry. Cancer 2007, 109, 1721–1728. [Google Scholar] [CrossRef] [PubMed]
- Santaliz-Casiano,A. ; Mehta, D.; Danciu, O.C.; Patel, H.; Banks, L.; Zaidi, A.; Buckley, J.; Rauscher, G.H.; Schulte, L.; Weller, L.R.; Taiym, D.; Liko-Hazizi, E.; Pulliam, N.; Friedewald, S.M.; Khan, S.; Kim, J.J.; Gradishar, W.; Hegerty, S.; Frasor, J.; Hoskins, K.F.; Madak-Erdogan Z. Identification of metabolic pathways contributing to ER+ breast cancer disparities using a machine-learning pipeline. Sci Rep 2023, 13, 12136. [Google Scholar]
- Meti, N.; Saednia, K.; Lagree, A.; Tabbarah, S.; Mohebpour, M.; Kiss, A.; Lu, F.I.; Slodkowska, E.; Gandhi, S.; Jerzak, K.J.; Fleshner, L.; Law, E.; Sadeghi-Naini, A.; Tran, W.T. Machine learning frameworks to predict neoadjuvant chemotherapy response in breast cancer using clinical and pathological features. JCO Clin Cancer Inform 2021, 5, 66–80. [Google Scholar] [CrossRef]
- Subramani, R.; Poudel, S.; Smith, K.D.; Estrada, A.; Lakshmanaswamy, R. Metabolomics of breast cancer: A review. Metabolites 2022, 12, 643. [Google Scholar] [CrossRef]
- Wang, W.; Rong, Z.; Wang, G.; Hou, Y.; Yang, F.; Qiu, M. Cancer metabolites: promising biomarkers for cancer liquid biopsy. Biomark Res 2023, 11, 66. [Google Scholar] [CrossRef]
- Puskulluoglu, M.; Michalak, I.M. The therapeutic potential of natural metabolites in targeting endocrine-independent HER-2-negative breast cancer. Front Pharmacol 2024, 15, 1349242. [Google Scholar] [CrossRef]
| Total cases (N=238) | ER-positive | ER-negative | p-value | |
|---|---|---|---|---|
| Population | Breast Cancer (N=185) | 164 | 21 | |
| Healthy (N=53) | 0 | 0 | ||
| Race | Black | 11 | 7 | > 0.05 |
| White | 147 | 64 | ||
| Other | 6 | 3 | ||
| Smoking | Current | 18 | 5 | < 0.05 |
| Former | 46 | 14 | ||
| Never | 99 | 54 | ||
| Not Stated | 1 | 1 | ||
| Age | Mean (SD) | 56.86 (12.42) | 52.97 (13.46) | < 0.05 |
| BMI | Mean (SD) | 29.77 (7.28) | 30.30 (7.51) | > 0.05 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




