Submitted:
12 September 2024
Posted:
12 September 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Related Works
3. Materials and Methods
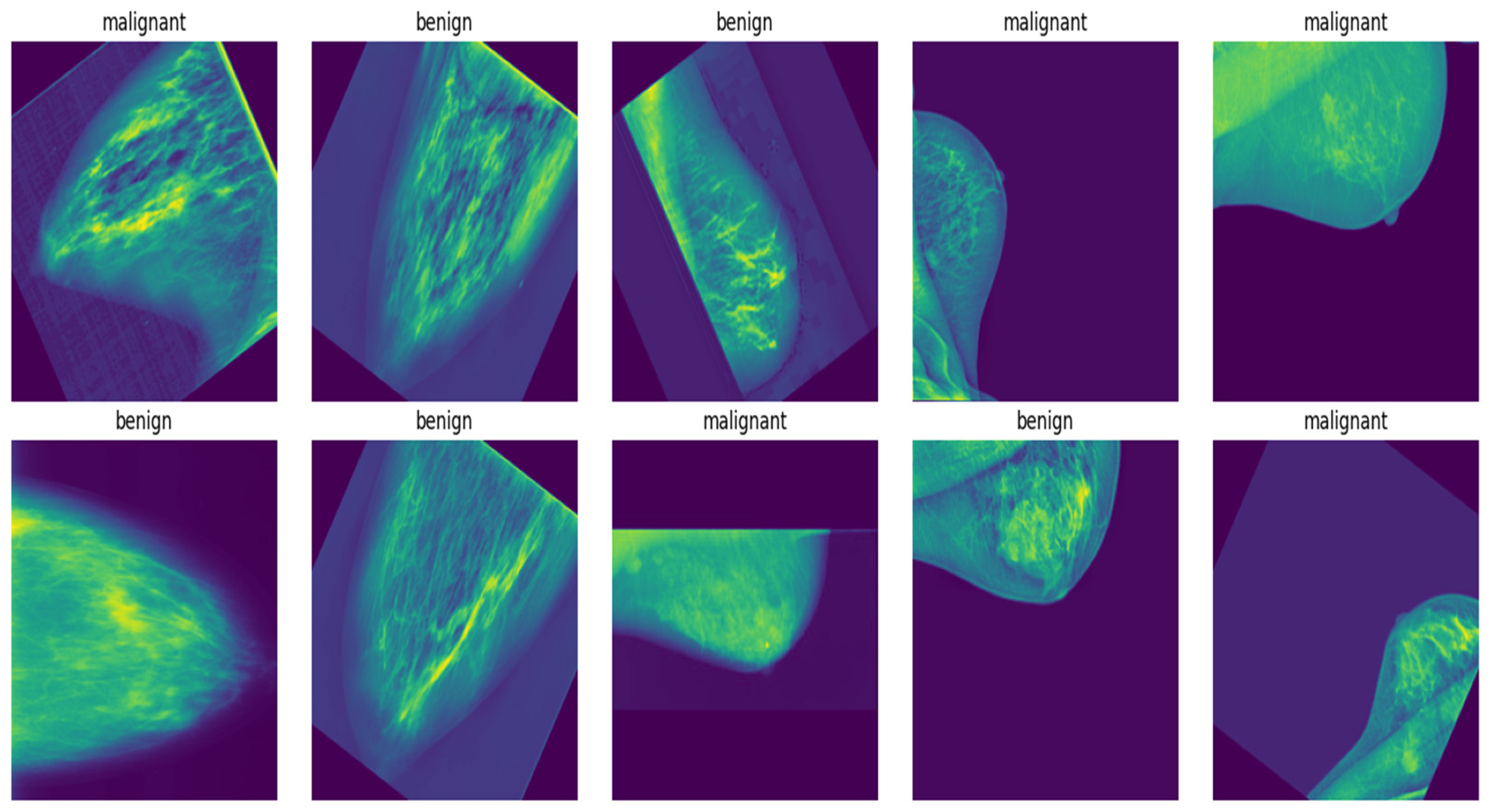
3.1. Data Collection
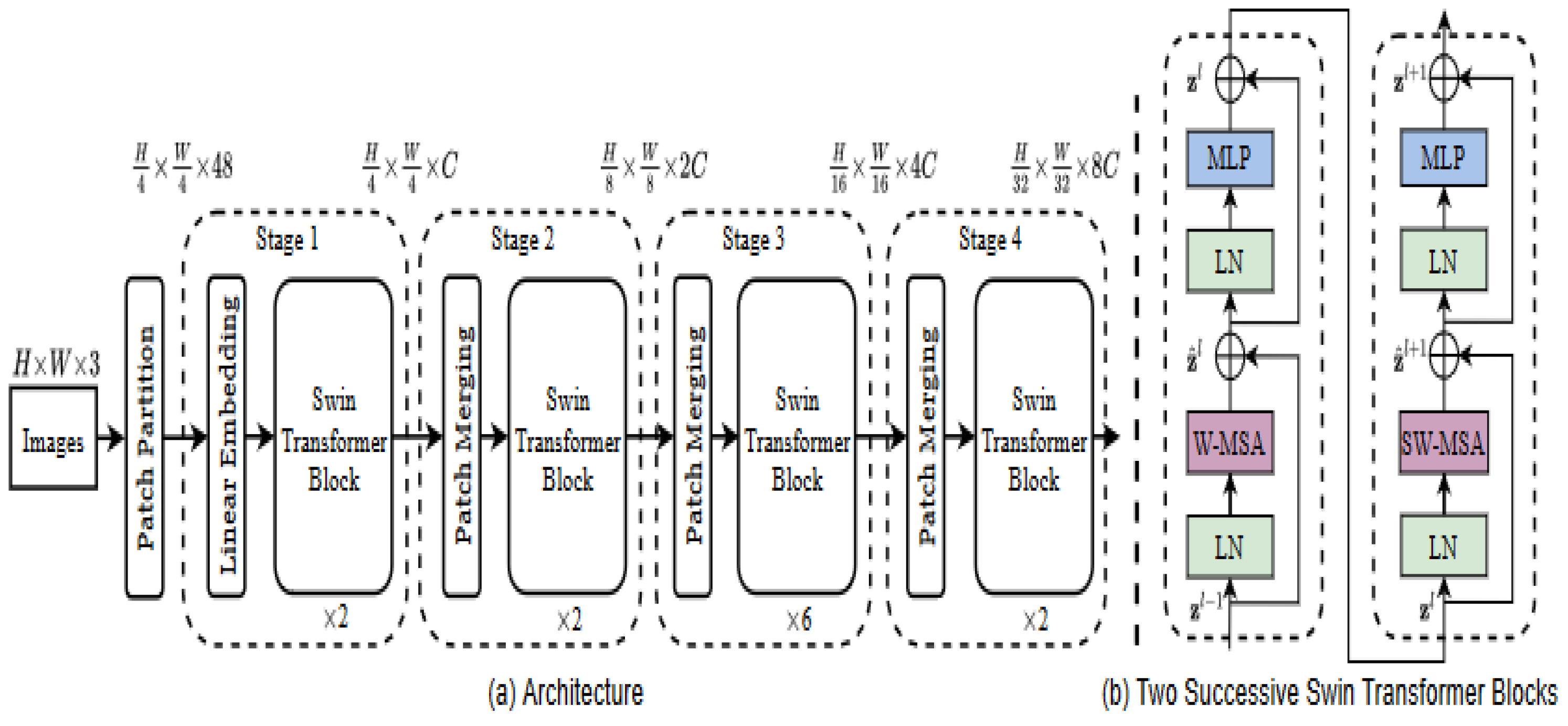
3.2. Shifted Window (SWIN) Transformer
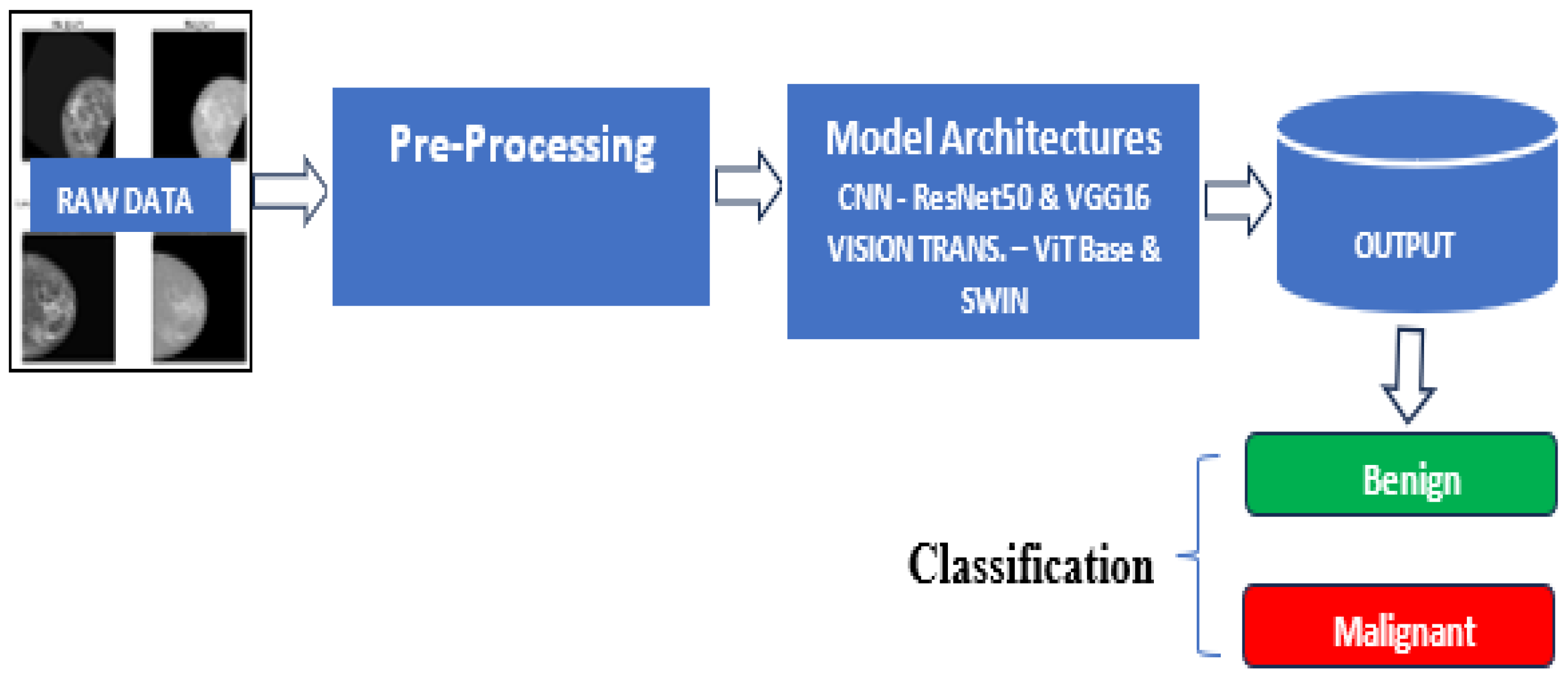
3.3. Model Development
3.3.1. Data Augmentation and Preprocessing Considerations
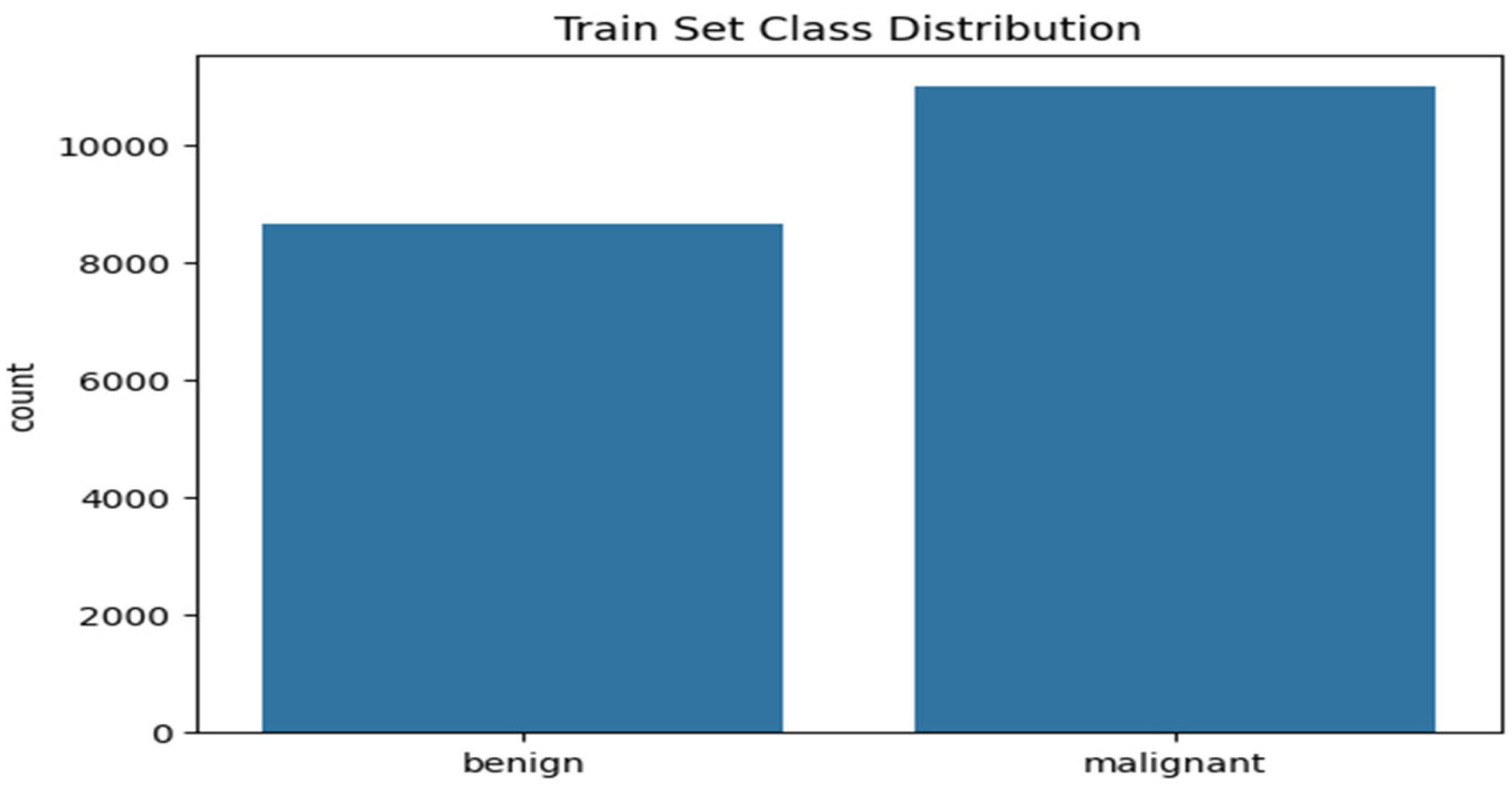
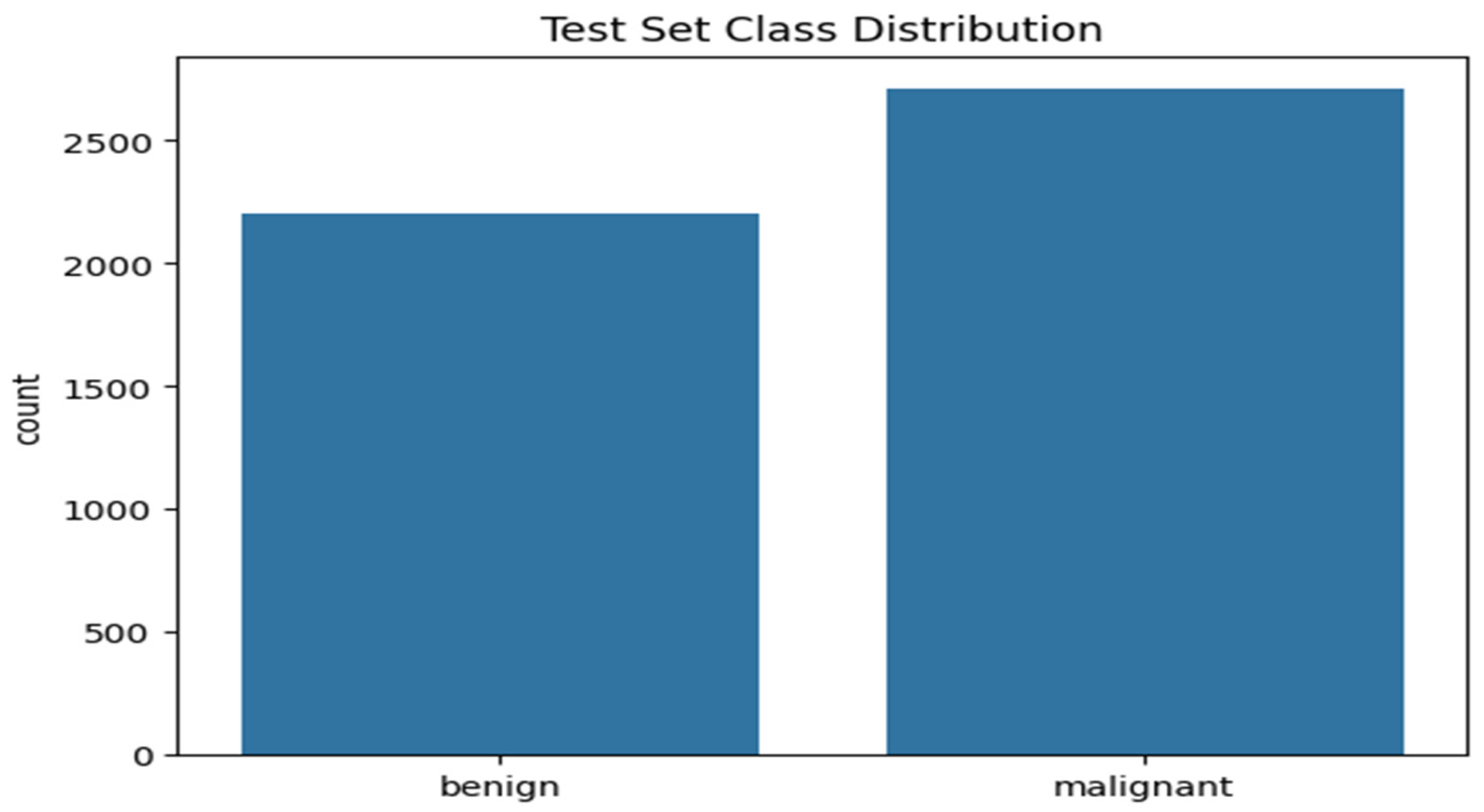
3.3.2. Class Distribution
3.4. Experimental Set-Up
4. Results
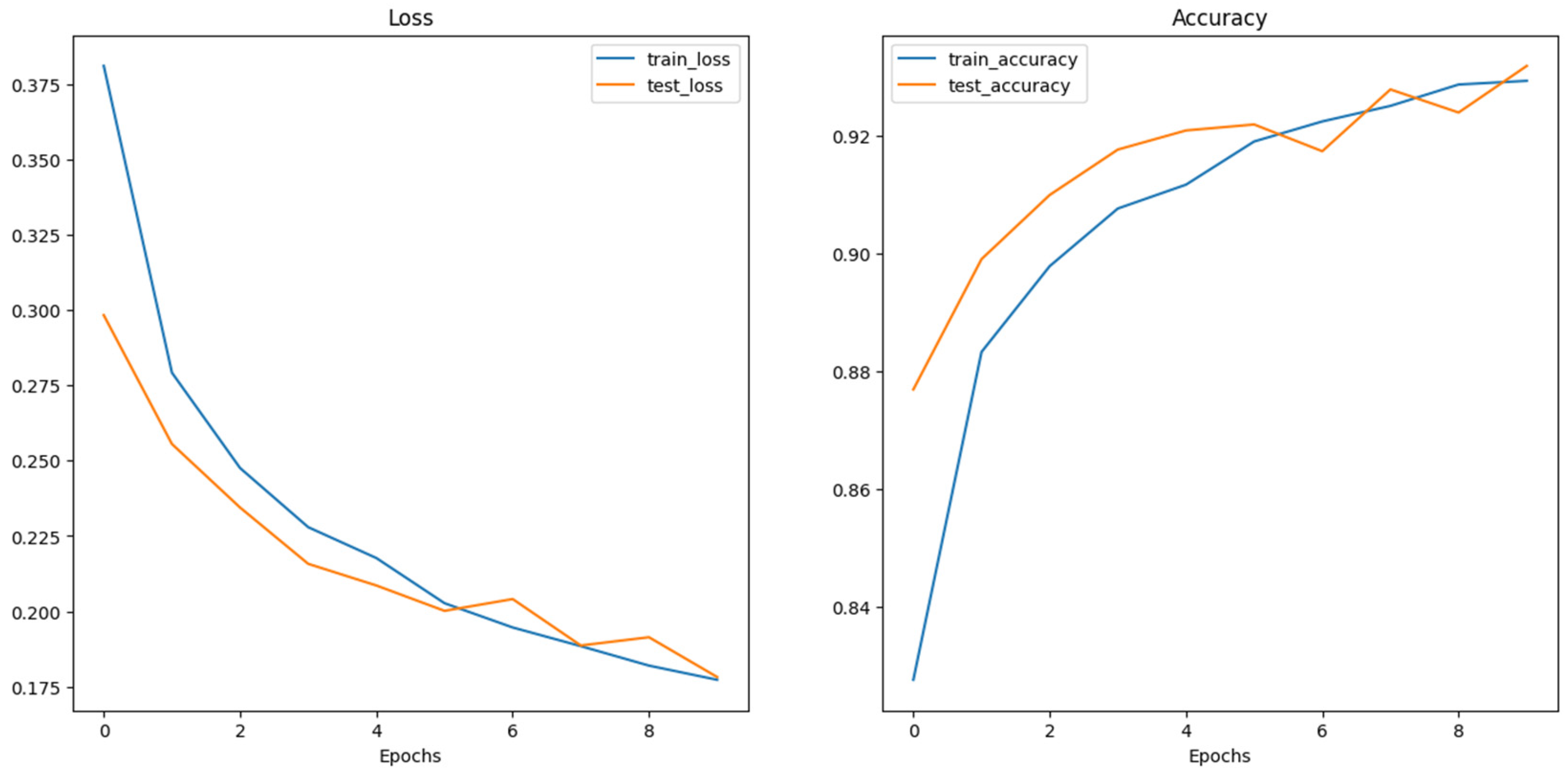
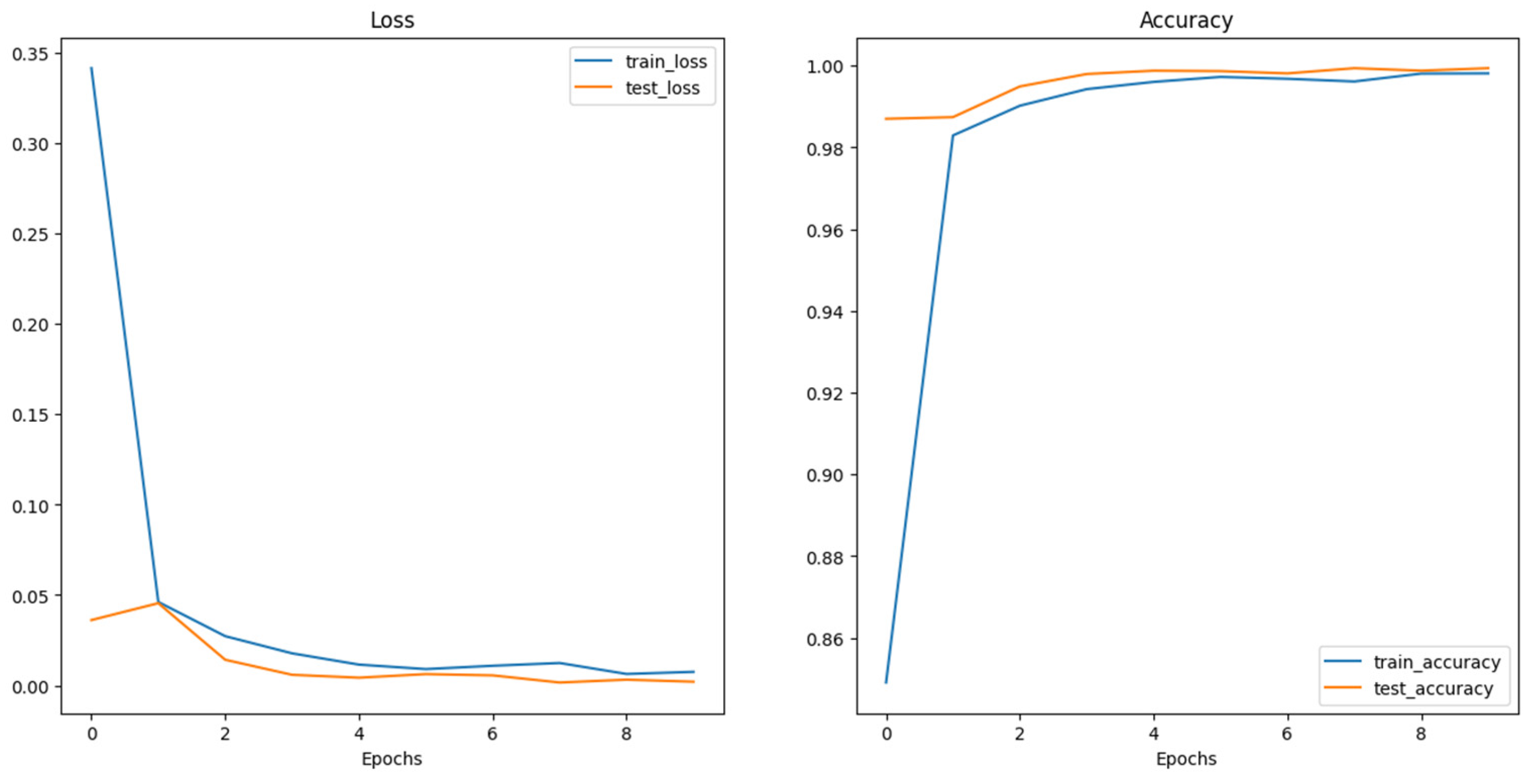
4.1. Model Training
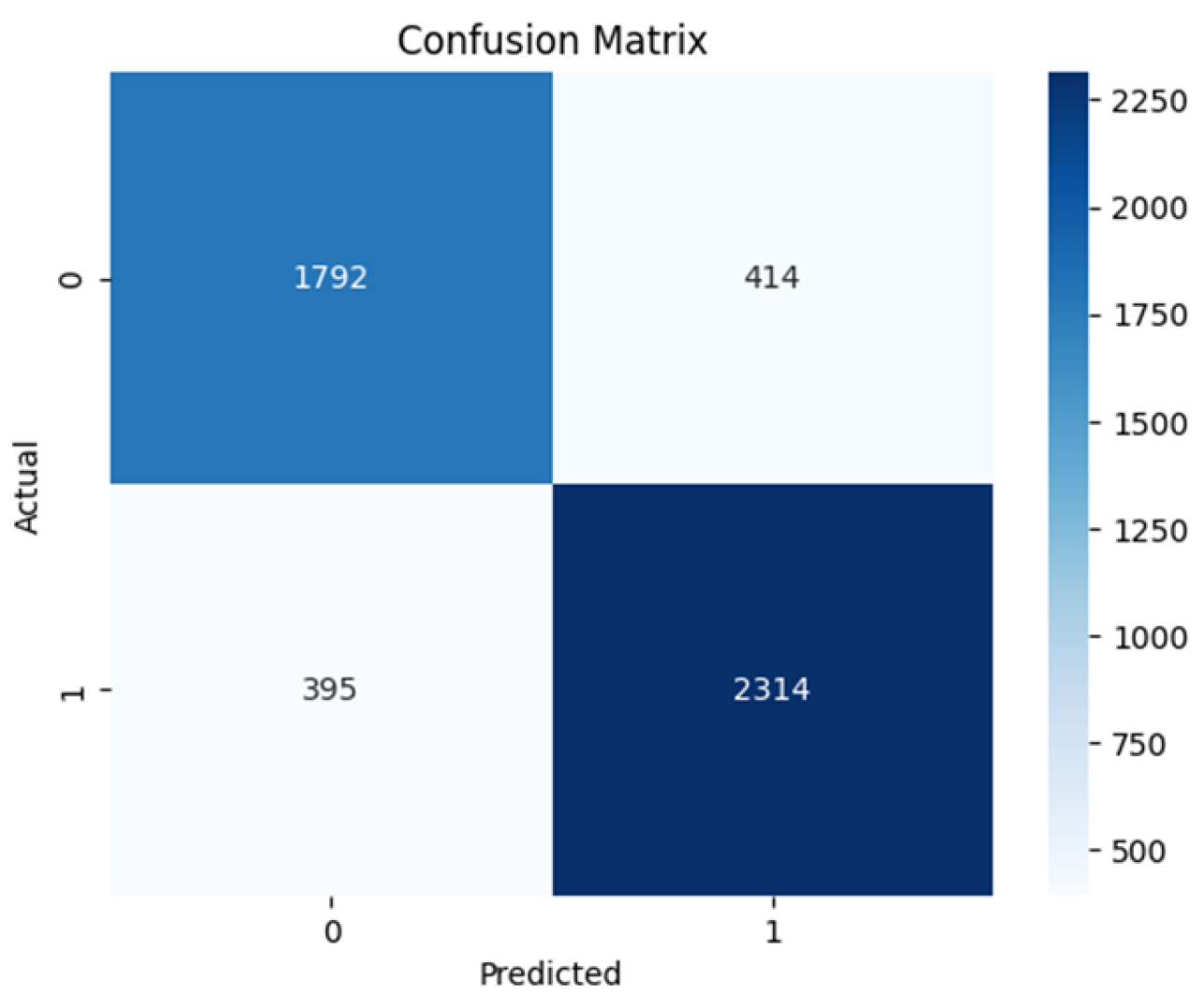
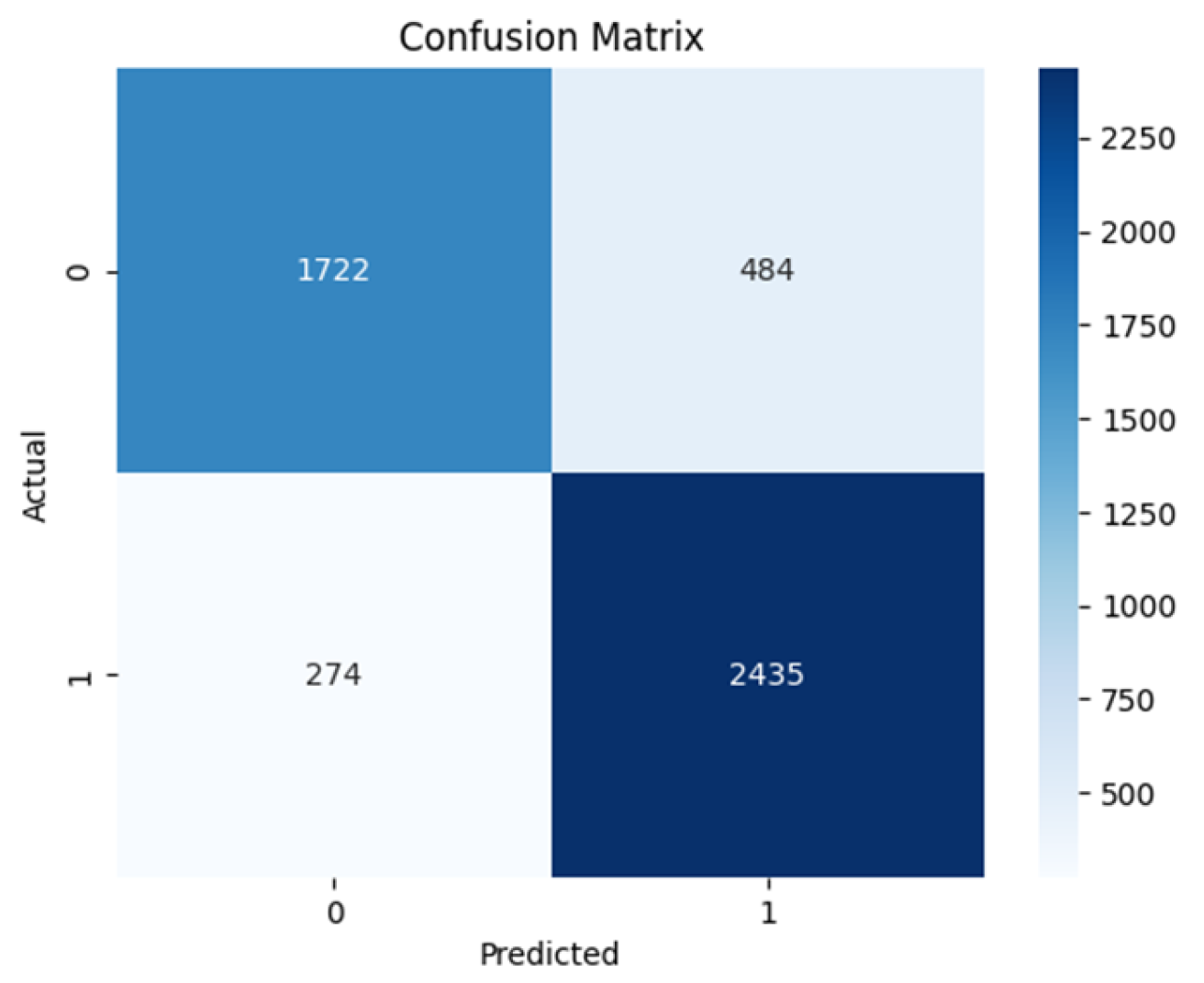
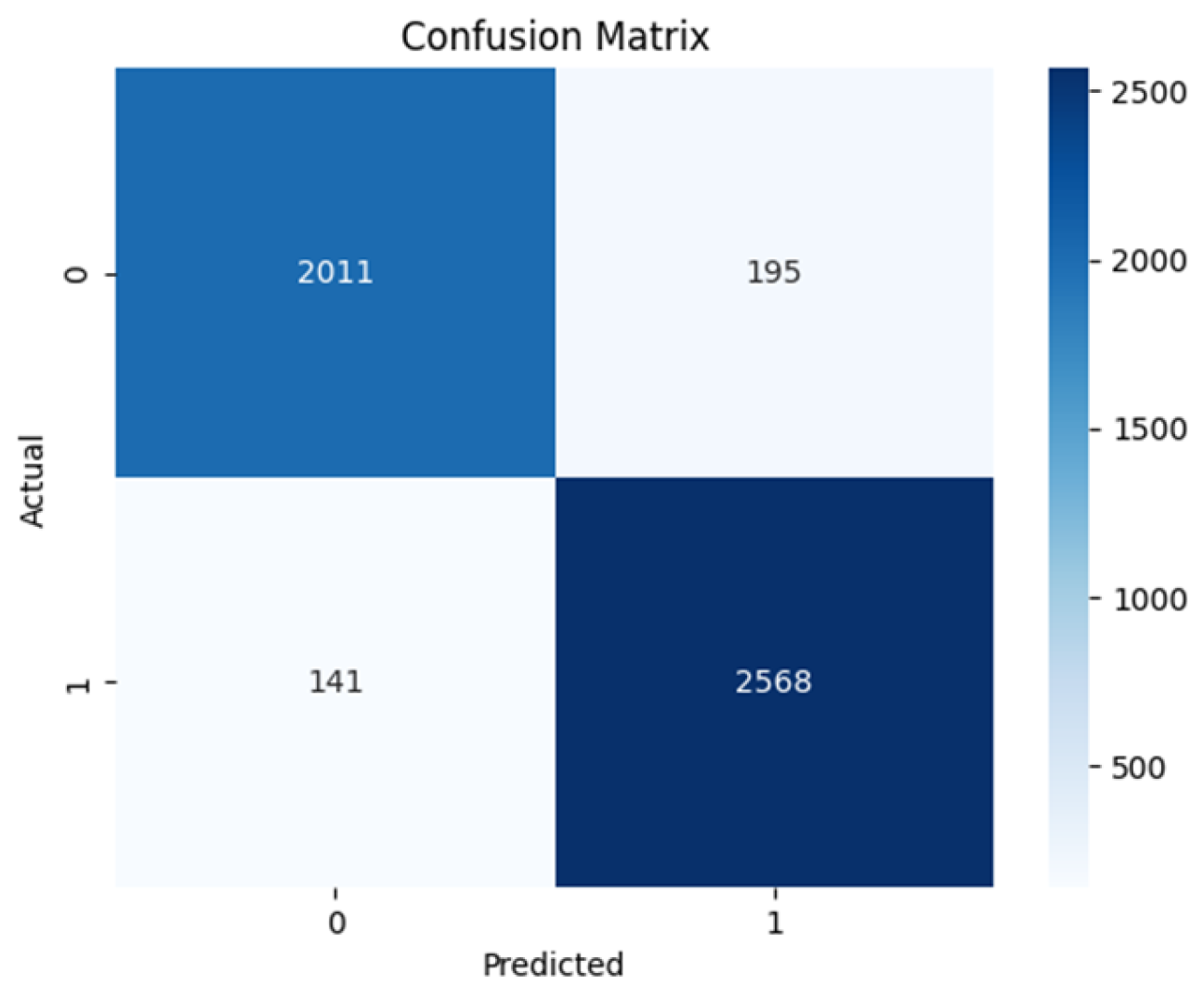
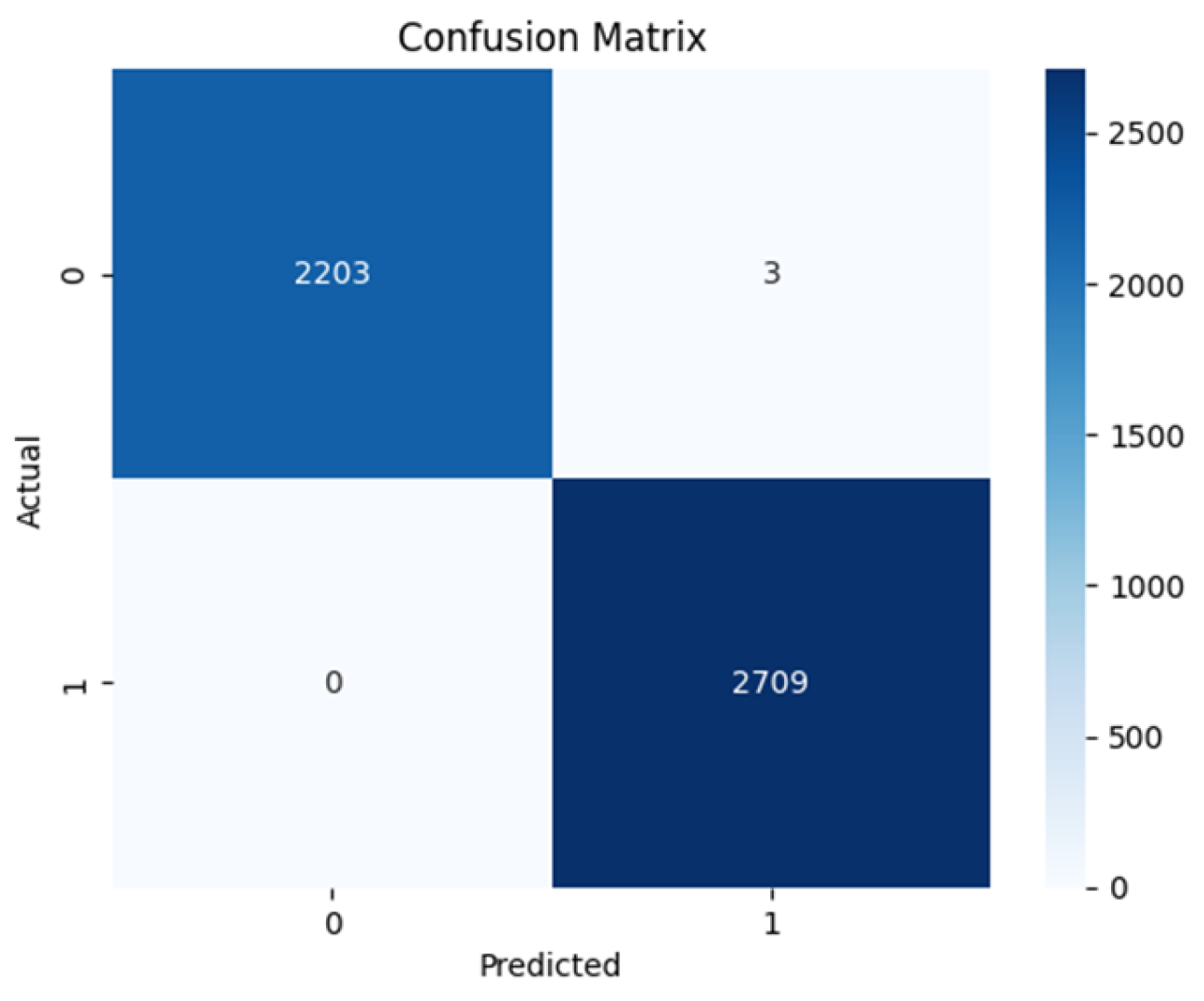
4.2. Confusion Matrix
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Jabeen, K.; Khan, M.A.; Balili, J.; Alhaisoni, M.; Almujally, N.A.; Alrashidi, H.; Tariq, U.; Cha, J.-H. BC2NetRF: Breast Cancer Classification from Mammogram Images Using Enhanced Deep Learning Features and Equilibrium-Jaya Controlled Regula Falsi-Based Features Selection. Diagnostics 2023, 13, 1238. [CrossRef]
- Nasser, M.; Yusof, U.K. Deep Learning Based Methods for Breast Cancer Diagnosis: A Systematic Review and Future Direction. Diagnostics 2023, 13, 161. [CrossRef]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [CrossRef]
- Ayana, G.; Dese, K.; Dereje, Y.; Kebede, Y.; Barki, H.; Amdissa, D.; Husen, N.; Mulugeta, F.; Habtamu, B.; Choe, S.-W. Vision-Transformer-Based Transfer Learning for Mammogram Classification. Diagnostics 2023, 13, 178, Available online: https://discovery.researcher.life/article/vision-transformer-based-transfer-learning-for-mammogram-classification/db5c10858ce13d26a4c879868650ee40. [CrossRef]
- Mohamed, T.I.A.; Ezugwu, A.E.; Fonou-Dombeu, J.V.; Ikotun, A.M.; Mohammed, M. A bio-inspired convolution neural network architecture for automatic breast cancer detection and classification using RNA-Seq gene expression data. Sci. Rep. 2023, 13, 1–19, Available online: https://discovery.researcher.life/article/a-bio-inspired-convolution-neural-network-architecture-for-automatic-breast-cancer-detection-and-classification-using-rna-seq-gene-expression-data/7921b201d096356d952152f6bf074f41. [CrossRef]
- B.M, G.; C.P, S.; T, S. Breast Cancer Diagnosis Using Machine Learning Algorithms - A Survey. Int. J. Distrib. Parallel Syst. 2013, 4, 105–112, Available online: https://discovery.researcher.life/article/breast-cancer-diagnosis-using-machine-learning-algorithms-a-survey/1472f2bc636d352dabc8e3af31f6a66e. [CrossRef]
- Weedon-Fekjaer, H.; Romundstad, P.R.; Vatten, L.J. Modern mammography screening and breast cancer mortality: population study. BMJ 2014, 348, g3701–g3701, Available online: http://www.bmj.com/content/348/bmj.g3701.abstract. [CrossRef]
- Pashayan, N.; Antoniou, A.C.; Ivanus, U.; Esserman, L.J.; Easton, D.F.; French, D.; Sroczynski, G.; Hall, P.; Cuzick, J.; Evans, D.G. Personalized early detection and prevention of breast cancer: ENVISION consensus statement. Nature reviews Clinical oncology 2020, 17, 687–705.
- Chougrad, H.; Zouaki, H.; Alheyane, O. Multi-label transfer learning for the early diagnosis of breast cancer. Neurocomputing 2020, 392, 168–180. [CrossRef]
- Zhou, Z. Breast Cancer Diagnosis with Machine Learning. Highlights Sci. Eng. Technol. 2022, 9, 73–75, Available online: https://discovery.researcher.life/article/breast-cancer-diagnosis-with-machine-learning/72ee85f4e77c3e7fa543d4bd08261845. [CrossRef]
- beyli, E.D. Implementing automated diagnostic systems for breast cancer detection. Expert Syst. Appl. 2007, 33, 1054–1062.
- Heywang-Köbrunner, S.H.; Hacker, A.; Sedlacek, S. Advantages and Disadvantages of Mammography Screening. Breast Care 2011, 6, 2–2. [CrossRef]
- Whang, J.S.; Baker, S.R.; Patel, R.; Luk, L.; Castro III, A. The causes of medical malpractice suits against radiologists in the United States. Radiology 2013, 266, 548–554.
- Zebari, D.A.; Zeebaree, D.Q.; Abdulazeez, A.M.; Haron, H.; Hamed, H.N.A. Improved Threshold Based and Trainable Fully Automated Segmentation for Breast Cancer Boundary and Pectoral Muscle in Mammogram Images. IEEE Access 2020, 8, 203097–203116. [CrossRef]
- Gheflati, B.; Rivaz, H. VISION TRANSFORMERS FOR CLASSIFICATION OF BREAST ULTRASOUND IMAGES. 2022.
- Esteva, A., Kuprel, B., Novoa, R. A., Ko, J., Swetter, S. M., Blau, H. M., Thrun, S., "Dermatologist-level classification of skin cancer with deep neural networks", Nature, Vol. 542, No. 7639, pp. 115-118, 2017. Epub 2017 Jan 25. Erratum in: Nature. 2017 Jun 28;546(7660):686. PMID: 28117445; PMCID: PMC8382232. [CrossRef]
- Liang, T.; Shen, J.; Wang, J.; Liao, W.; Zhang, Z.; Liu, J.; Feng, Z.; Pei, S.; Liu, K. Ultrasound-based prediction of preoperative core biopsy categories in solid breast tumor using machine learning. Quant. Imaging Med. Surg. 2023, 13, 2634–+. [CrossRef]
- Baroni, G.L.; Rasotto, L.; Roitero, K.; Tulisso, A.; Di Loreto, C.; Della Mea, V. Optimizing Vision Transformers for Histopathology: Pretraining and Normalization in Breast Cancer Classification. J. Imaging 2024, 10, 108. [CrossRef]
- Alakwaa, W.; Nassef, M.; Badr, A. Lung Cancer Detection and Classification with 3D Convolutional Neural Network (3D-CNN). Int. J. Adv. Comput. Sci. Appl. 2017, 8. [CrossRef]
- McKinney, S.M.; Sieniek, M.; Godbole, V.; Godwin, J.; Antropova, N.; Ashrafian, H.; Back, T.; Chesus, M.; Corrado, G.S.; Darzi, A.; et al. International evaluation of an AI system for breast cancer screening. Nature 2020, 577, 89–94. [CrossRef]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; van der Laak, J.A.W.M.; van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med. Image Anal. 2017, 42, 60–88. Available online: https://www.sciencedirect.com/science/article/pii/S1361841517301135. [CrossRef]
- Cheng, H.; Shan, J.; Ju, W.; Guo, Y.; Zhang, L. Automated breast cancer detection and classification using ultrasound images: A survey. Pattern Recognit 2010, 43, 299–317.
- Mahoro, E.; Akhloufi, M.A. Applying Deep Learning for Breast Cancer Detection in Radiology. Curr. Oncol. 2022, 29, 8767–8793. [CrossRef]
- Araújo, T.; Aresta, G.; Castro, E.; Rouco, J.; Aguiar, P.; Eloy, C.; Polónia, A.; Campilho, A. Classification of breast cancer histology images using Convolutional Neural Networks. PLOS ONE 2017, 12, e0177544. [CrossRef]
- Albalawi, U.; Manimurugan, S.; Varatharajan, R. Classification of breast cancer mammogram images using convolution neural network. Concurr. Comput. Pr. Exp. 2022, 34. [CrossRef]
- López-Cabrera, J.D.; Rodríguez, L.A.L.; Pérez-Díaz, M. Classification of Breast Cancer from Digital Mammography Using Deep Learning. Intel. Artif. 2020, 23, 56–66. [CrossRef]
- Salama, W.M.; Aly, M.H. Deep learning in mammography images segmentation and classification: Automated CNN approach. Alex. Eng. J. 2021, 60, 4701–4709. [CrossRef]
- Tummala, S.; Kim, J.; Kadry, S. BreaST-Net: Multi-Class Classification of Breast Cancer from Histopathological Images Using Ensemble of Swin Transformers. Mathematics 2022, 10, 4109. [CrossRef]
- Liu, Z.; Lin, Y.; Cao, Y.; Hu, H.; Wei, Y.; Zhang, Z.; Lin, S.; Guo, B. In In Swin transformer: Hierarchical vision transformer using shifted windows; Proceedings of the IEEE/CVF international conference on computer vision; 2021; , pp 10012–10022.
- Mahoro, E.; Akhloufi, M.A. Breast cancer classification on thermograms using deep CNN and transformers. Quant. Infrared Thermogr. J. 2024, 21, 30–49. [CrossRef]
- Abimouloud, M.L.; Bensid, K.; Elleuch, M.; Aiadi, O.; Kherallah, M. In In Mammography breast cancer classification using vision transformers; International Conference on Intelligent Systems Design and Applications; Springer: 2023; , pp 452–461.
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv preprint arXiv:2010.11929 2020.
- Huang, M.-L.; Lin, T.-Y. Dataset of breast mammography images with masses. Data Brief 2020, 31, 105928. [CrossRef]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł; Polosukhin, I. Attention is all you need. Advances in neural information processing systems 2017, 30.
- Bao, H.; Dong, L.; Wei, F.; Wang, W.; Yang, N.; Liu, X.; Wang, Y.; Gao, J.; Piao, S.; Zhou, M. In In Unilmv2: Pseudo-masked language models for unified language model pre-training; International conference on machine learning; PMLR: 2020; , pp 642–652.
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556 2014.
- He, K.; Zhang, X.; Ren, S.; Sun, J. In In Deep residual learning for image recognition; Proceedings of the IEEE conference on computer vision and pattern recognition; 2016; , pp 770–778.
- Xie, F.; Fan, H.; Li, Y.; Jiang, Z.; Meng, R.; Bovik, A. Melanoma Classification on Dermoscopy Images Using a Neural Network Ensemble Model. IEEE Trans. Med Imaging 2016, 36, 849–858. [CrossRef]
- Chollet, F. Deep learning with Python; Simon and Schuster: 2021; .
- Paszke, A.; Gross, S.; Massa, F.; Lerer, A.; Bradbury, J.P.; Chanan, G.; Killeen, T.; Lin, Z.; Gimelshein, N.; Antiga, L. An imperative style, high-performance deep learning library. Adv.Neural Inf.Process.Syst 1912, 32, 8026.
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. CoRR 2015, abs/1512.03385.
- Simonyan, K. Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556 2014.
- Ayana, G.; Dese, K.; Dereje, Y.; Kebede, Y.; Barki, H.; Amdissa, D.; Husen, N.; Mulugeta, F.; Habtamu, B.; Choe, S.-W. Vision-Transformer-Based Transfer Learning for Mammogram Classification. Diagnostics 2023, 13, 178. [CrossRef]
| Dataset Type | No of Benign | No of Malignant | Total images |
| MIAS | 2376 | 1440 | 3816 |
| DDSM | 5970 | 7158 | 13128 |
| IN Breast | 2520 | 5112 | 7632 |
| TOTAL IMAGES | 24,576 | ||
| Model | Sensitivity (TPR) | Specificity (TNR) | NPV | PPV | Precision | Accuracy |
| ResNet50 | 0.858 | 0.794 | 0.863 | 0.834 | 0.837 | 0.829 |
| VGG16 | 0.854 | 0.811 | 0.819 | 0.853 | 0.848 | 0.835 |
| ViT base | 0.948 | 0.913 | 0.934 | 0.929 | 0.929 | 0.932 |
| SWIN | 1.000 | 0.999 | 1.000 | 0.999 | 0.998 | 0.998 |
| ViT scratch | 1.000 | 1.000 | 0 | 0.551 | 0.551 | 0.551 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




