Submitted:
24 August 2024
Posted:
27 August 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
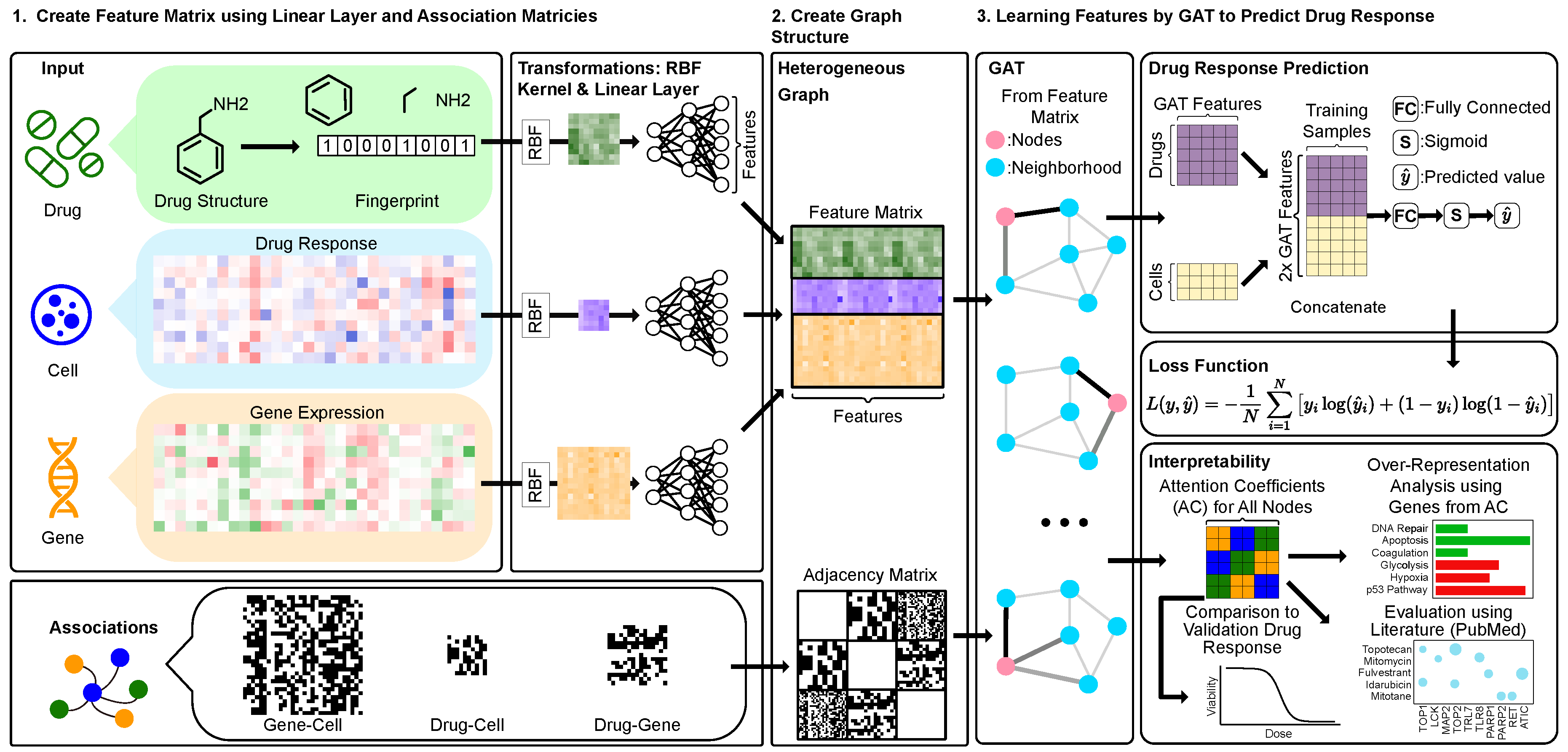
- We create a heterogeneous graph involving DNA-targeting drugs that includes three entity types: genes (i.e., gene expression), cell lines (i.e., drug responses), and drugs (i.e., structures) using the NCI60 [22].
- We evaluate the drug-gene associations suggested by attention coefficients and systematically compare these to existing scientific publication text. We examine the abstracts of journal papers for co-mentions of drug-target relationships from our results.
- Our results propose drug-cell line sensitivity associations based on the attention coefficients derived from the GAT. These associations were then validated through comparison with independent experimental data.
2. Results
2.1. Model Performance
2.1.1. Analysis of GNN Architectures and Their Impact on Performance
2.2. Interpretation
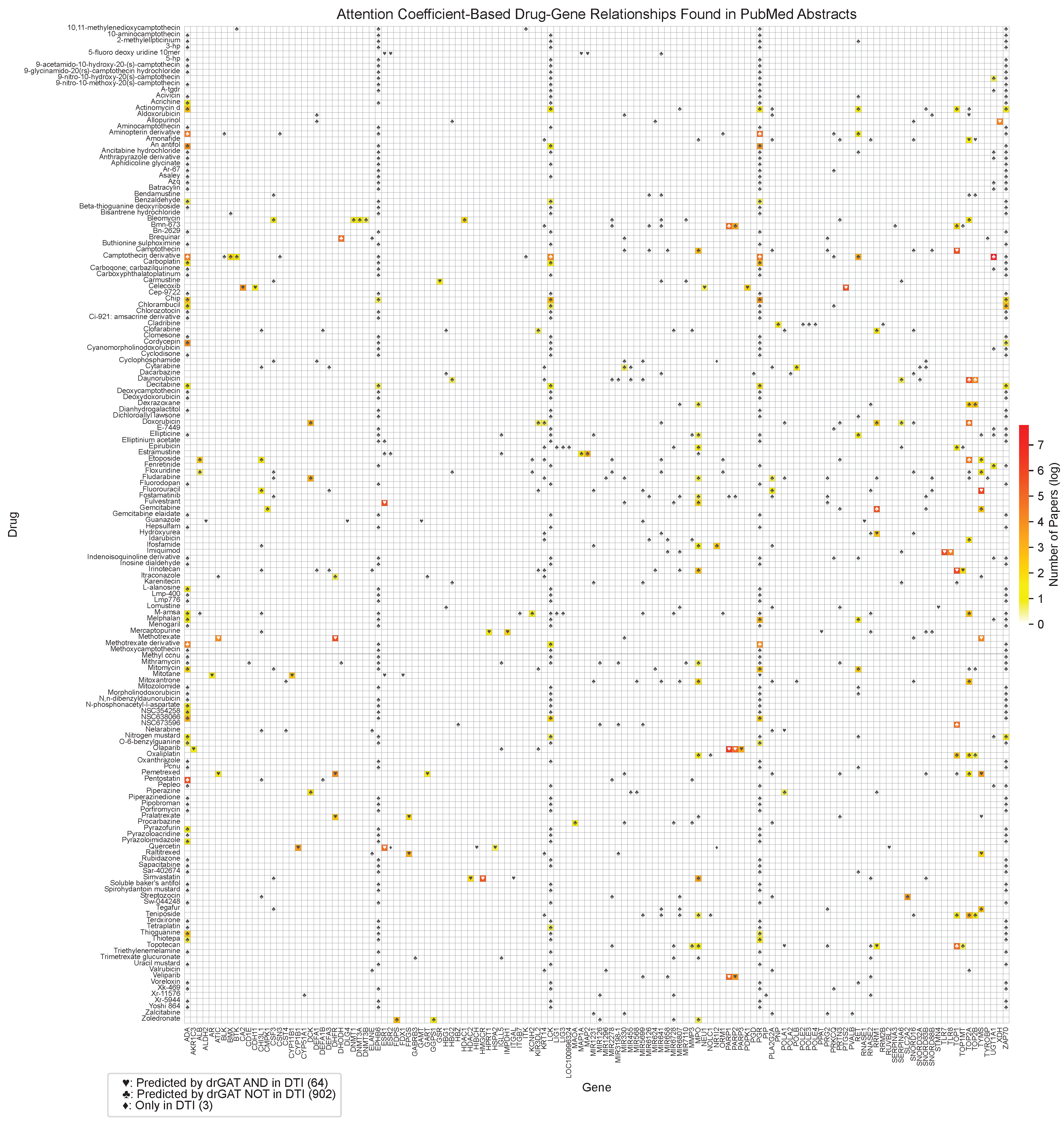
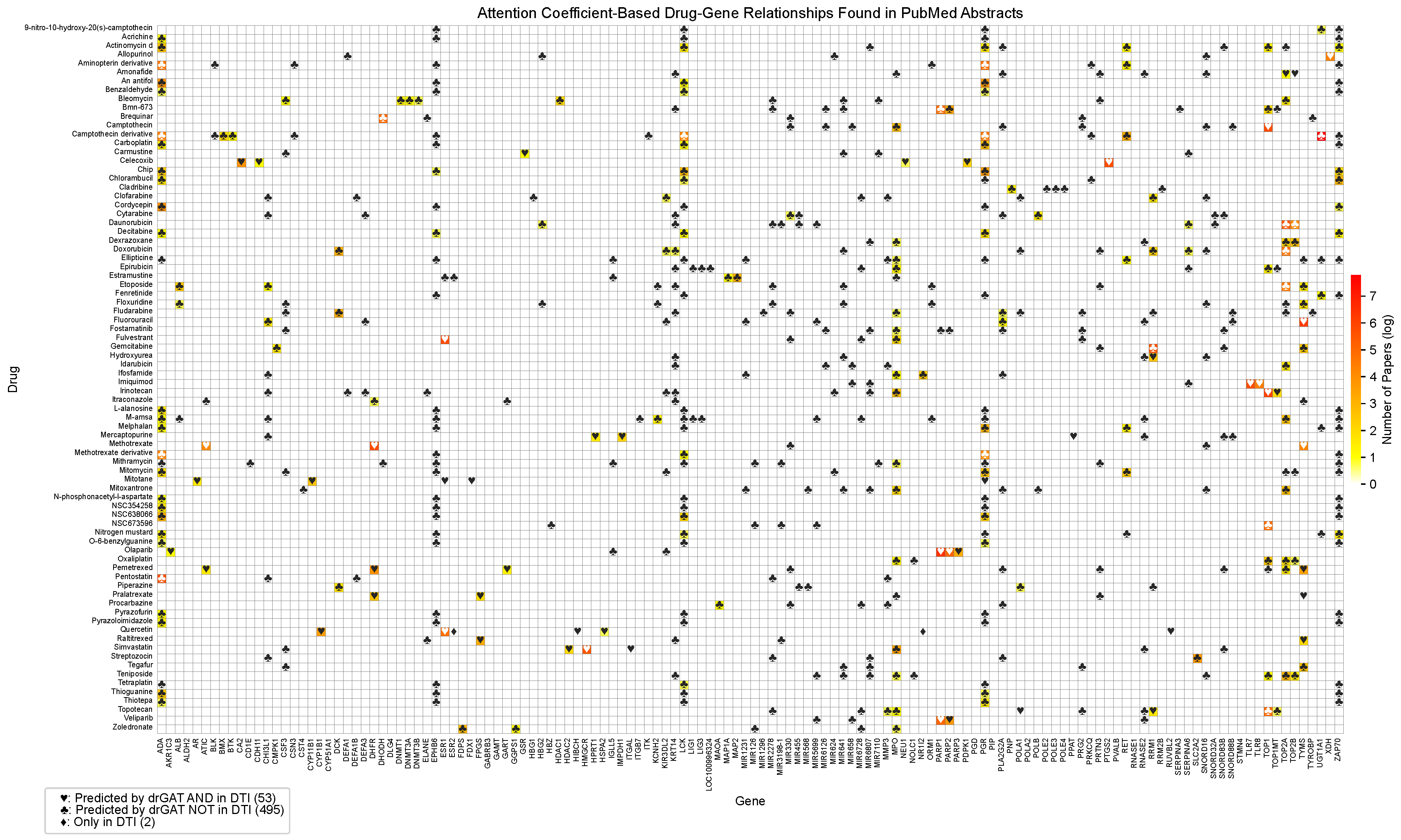
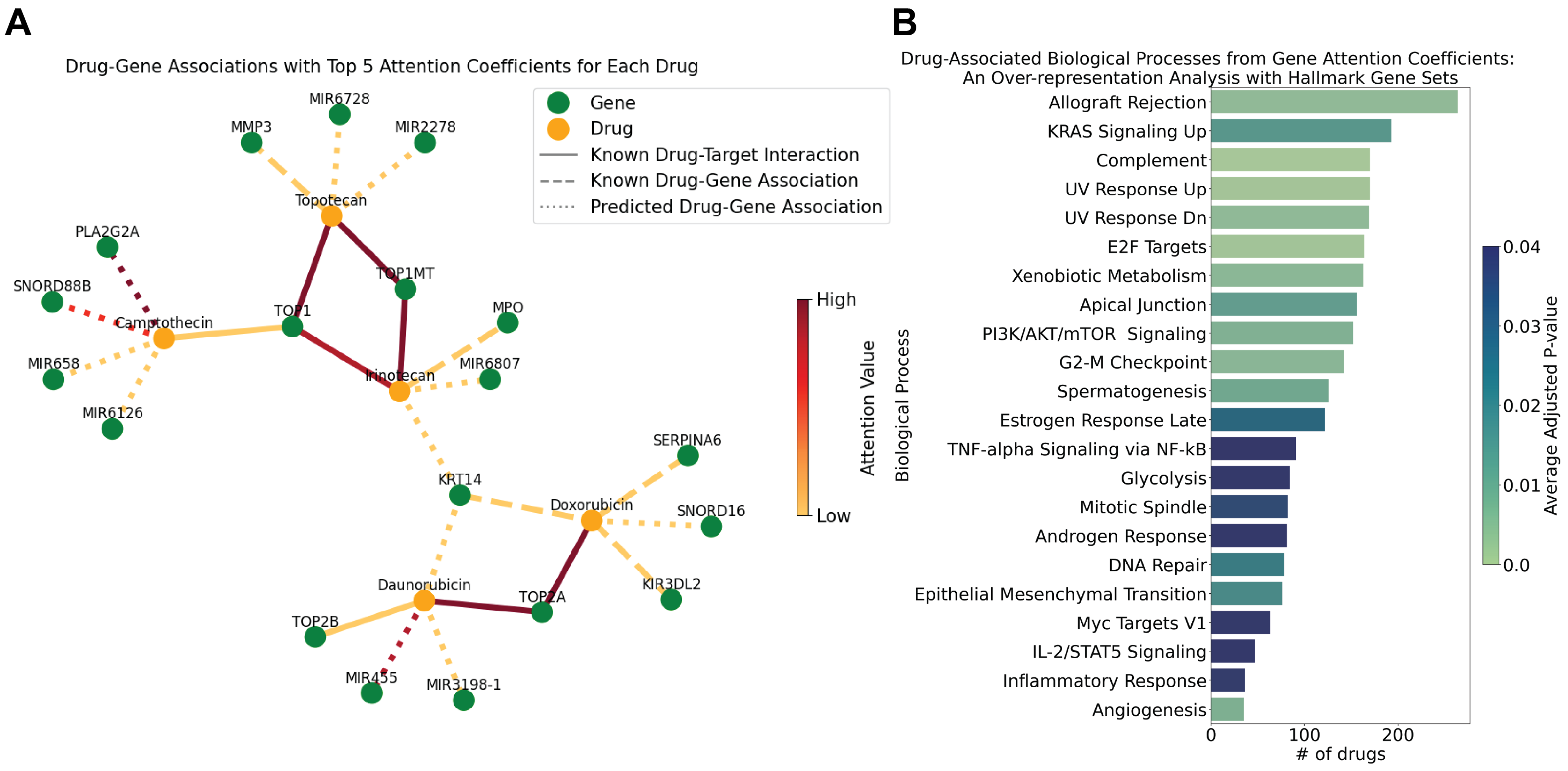
2.2.1. Evaluation of Drug-Gene Associations in Scientific Literature
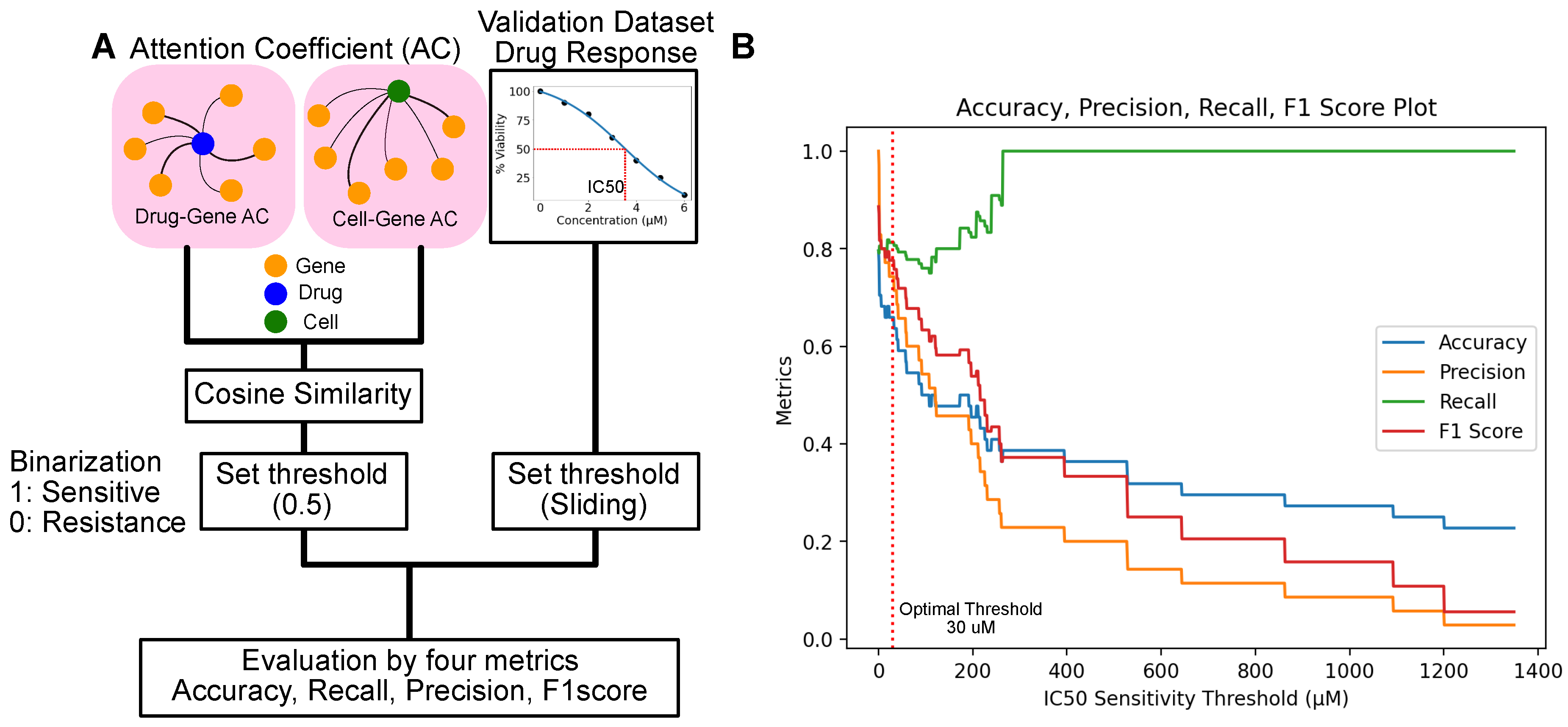
2.2.2. Evaluation of Predicted Drug-Cell Line Response by Comparison with GDSC
2.2.3. Drug-Target Interactions Assessment from Attention Coefficients
2.2.4. Over-Representation Analysis with Attention Coefficients
3. Methods
3.1. Building Input Matrix
3.1.1. Preprocessing
3.1.2. Feature Matrix
3.1.3. Adjacency Matrix
3.1.4. Preventing Data Leakage via Masking
3.2. drGAT Model
3.3. Model Configuration and Training
4. Discussion
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Appendix A. Related Works
Appendix B. Experiment
Appendix B.1. Validation
Appendix B.2. Baseline Methods
Appendix B.3. Comparative GNN Architecture
Appendix B.4. Evaluation Metrics of Prediction Performance
- positive sample and is correctly predicted as positive, also known as True Positive (TP);
- negative samples and is wrongly predicted as positive samples, also known as False Positive (FP);
- negative samples and is correctly predicted as negative samples, also known as True Negative (TN);
- positive samples and is wrongly predicted as negative samples, also known as False Negative (FN).
Appendix B.5. Hyperparameter Tuning
Appendix C. Full Drug-Gene Relationship Table
References
- Azuaje, F. Computational models for predicting drug responses in cancer research. Briefings in bioinformatics 2017, 18, 820–829. [Google Scholar] [CrossRef]
- Kuenzi, B.M.; Park, J.; Fong, S.H.; Sanchez, K.S.; Lee, J.; Kreisberg, J.F.; Ma, J.; Ideker, T. Predicting drug response and synergy using a deep learning model of human cancer cells. Cancer cell 2020, 38, 672–684. [Google Scholar] [CrossRef]
- Chang, Y.T.; Hoffman, E.P.; Yu, G.; Herrington, D.M.; Clarke, R.; Wu, C.T.; Chen, L.; Wang, Y. Integrated identification of disease specific pathways using multi-omics data. bioRxiv 2019, 666065. [Google Scholar]
- Xu, Y.; Liu, X.; Kong, Z.; Wu, Y.; Wang, Y.; Lu, Y.; Gao, H.; Wu, J.; Xu, H. MambaCapsule: Towards Transparent Cardiac Disease Diagnosis with Electrocardiography Using Mamba Capsule Network. arXiv preprint arXiv:2407.20893, arXiv:2407.20893 2024.
- Garnett, M.J.; Edelman, E.J.; Heidorn, S.J.; Greenman, C.D.; Dastur, A.; Lau, K.W.; Greninger, P.; Thompson, I.R.; Luo, X.; Soares, J.; others. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 2012, 483, 570–575. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.T.; Parker, S.J.; Cheng, Z.; Saylor, G.; Van Eyk, J.E.; Yu, G.; Clarke, R.; Herrington, D.M.; Wang, Y. COT: an efficient and accurate method for detecting marker genes among many subtypes. Bioinformatics Advances 2022, 2, vbac037. [Google Scholar]
- Chen, L.; Wu, C.T.; Clarke, R.; Yu, G.; Van Eyk, J.E.; Herrington, D.M.; Wang, Y. Data-driven detection of subtype-specific differentially expressed genes. Scientific reports 2021, 11, 332. [Google Scholar] [CrossRef] [PubMed]
- Leung, M.K.; Delong, A.; Alipanahi, B.; Frey, B.J. Machine learning in genomic medicine: a review of computational problems and data sets. Proceedings of the IEEE 2015, 104, 176–197. [Google Scholar] [CrossRef]
- Cao, C.; Liu, F.; Tan, H.; Song, D.; Shu, W.; Li, W.; Zhou, Y.; Bo, X.; Xie, Z. Deep Learning and Its Applications in Biomedicine. Genomics, Proteomics, Bioinformatics 2018, 16, 17–32. [Google Scholar] [CrossRef]
- Mahmud, M.; Kaiser, M.; Mcginnity, T.; Hussain, A. Deep Learning in Mining Biological Data. Cognitive Computation 2020, 13, 1–33. [Google Scholar] [CrossRef]
- Lipton, Z.C. The mythos of model interpretability: In machine learning, the concept of interpretability is both important and slippery. Queue 2018, 16, 31–57. [Google Scholar] [CrossRef]
- Gilpin, L.H.; Bau, D.; Yuan, B.Z.; Bajwa, A.; Specter, M.; Kagal, L. Explaining explanations: An overview of interpretability of machine learning. 2018 IEEE 5th International Conference on data science and advanced analytics (DSAA). IEEE, 2018, pp. 80–89.
- Kengkanna, A.; Ohue, M. Enhancing Model Learning and Interpretation Using Multiple Molecular Graph Representations for Compound Property and Activity Prediction. 2023 IEEE Conference on Computational Intelligence in Bioinformatics and Computational Biology (CIBCB). IEEE, 2023, pp. 1–8.
- Fu, T.; Huang, K.; Xiao, C.; Glass, L.M.; Sun, J. Hint: Hierarchical interaction network for clinical-trial-outcome predictions. Patterns 2022, 3. [Google Scholar] [CrossRef]
- Fu, T.; Gao, W.; Xiao, C.; Yasonik, J.; Coley, C.W.; Sun, J. Differentiable Scaffolding Tree for Molecular Optimization. International Conference on Learning Representations 2022. [Google Scholar]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention is all you need. Advances in neural information processing systems 2017, 30. [Google Scholar]
- Veličković, P.; Cucurull, G.; Casanova, A.; Romero, A.; Lio, P.; Bengio, Y. Graph attention networks. arXiv preprint arXiv:1710.10903 2017. [Google Scholar]
- Luna, A.; Elloumi, F.; Varma, S.; Wang, Y.; Rajapakse, V.N.; Aladjem, M.I.; Robert, J.; Sander, C.; Pommier, Y.; Reinhold, W.C. CellMiner Cross-Database (CellMinerCDB) version 1.2: Exploration of patient-derived cancer cell line pharmacogenomics. Nucleic acids research 2021, 49, D1083–D1093. [Google Scholar] [CrossRef]
- Luna, A.; Rajapakse, V.N.; Sousa, F.G.; Gao, J.; Schultz, N.; Varma, S.; Reinhold, W.; Sander, C.; Pommier, Y. rcellminer: exploring molecular profiles and drug response of the NCI-60 cell lines in R. Bioinformatics 2016, 32, 1272–1274. [Google Scholar] [CrossRef]
- Yuan, H.; Yu, H.; Wang, J.; Li, K.; Ji, S. On explainability of graph neural networks via subgraph explorations. International Conference on Machine Learning. PMLR, 2021, pp. 12241–12252.
- Yuan, H.; Yu, H.; Gui, S.; Ji, S. Explainability in graph neural networks: A taxonomic survey. IEEE Transactions on Pattern Analysis and Machine Intelligence 2022. [Google Scholar] [CrossRef] [PubMed]
- Shoemaker, R.H. The NCI60 human tumour cell line anticancer drug screen. Nature Reviews Cancer 2006, 6, 813–823. [Google Scholar] [CrossRef]
- Coordinators, N.R. Database resources of the national center for biotechnology information. Nucleic acids research 2012, 41, D8–D20. [Google Scholar] [CrossRef]
- Yang, W.; Soares, J.; Greninger, P.; Edelman, E.J.; Lightfoot, H.; Forbes, S.; Bindal, N.; Beare, D.; Smith, J.A.; Thompson, I.R.; others. Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic acids research 2012, 41, D955–D961. [Google Scholar] [CrossRef]
- Chen, T.; Sun, Y.; Ji, P.; Kopetz, S.; Zhang, W. Topoisomerase IIα in chromosome instability and personalized cancer therapy. Oncogene 2015, 34, 4019–4031. [Google Scholar] [CrossRef]
- Wu, C.T.; Shen, M.; Du, D.; Cheng, Z.; Parker, S.J.; Van Eyk, J.E.; Yu, G.; Clarke, R.; Herrington, D.M.; others. Cosbin: cosine score-based iterative normalization of biologically diverse samples. Bioinformatics Advances 2022, 2, vbac076. [Google Scholar] [CrossRef] [PubMed]
- Liao, C.; Xie, G.; Zhu, L.; Chen, X.; Li, X.; Lu, H.; Xu, B.; Ramot, Y.; Paus, R.; Yue, Z. p53 is a direct transcriptional repressor of keratin 17: lessons from a rat model of radiation dermatitis. Journal of Investigative Dermatology 2016, 136, 680–689. [Google Scholar] [CrossRef] [PubMed]
- Greife, A.; Tukova, J.; Steinhoff, C.; Scott, S.D.; Schulz, W.A.; Hatina, J. Establishment and characterization of a bladder cancer cell line with enhanced doxorubicin resistance by mevalonate pathway activation. Tumor Biology 2015, 36, 3293–3300. [Google Scholar] [CrossRef] [PubMed]
- Saghaeidehkordi, A.; Chen, S.; Yang, S.; Kaur, K. Evaluation of a keratin 1 targeting peptide-doxorubicin conjugate in a mouse model of triple-negative breast cancer. Pharmaceutics 2021, 13, 661. [Google Scholar] [CrossRef]
- De Ronde, J.J.; Lips, E.H.; Mulder, L.; Vincent, A.D.; Wesseling, J.; Nieuwland, M.; Kerkhoven, R.; Vrancken Peeters, M.J.T.; Sonke, G.S.; Rodenhuis, S.; others. SERPINA6, BEX1, AGTR1, SLC26A3, and LAPTM4B are markers of resistance to neoadjuvant chemotherapy in HER2-negative breast cancer. Breast cancer research and treatment 2013, 137, 213–223. [Google Scholar] [CrossRef]
- Han, S.; Lim, K.S.; Blackburn, B.J.; Yun, J.; Putnam, C.W.; Bull, D.A.; Won, Y.W. The Potential of Topoisomerase Inhibitor-Based Antibody–Drug Conjugates. Pharmaceutics 2022, 14, 1707. [Google Scholar] [CrossRef]
- Lu, Y.; Sato, K.; Wang, J. Deep Learning based Multi-Label Image Classification of Protest Activities. arXiv preprint arXiv:2301.04212 2023. [Google Scholar]
- Blaney, S.M.; Tagen, M.; Onar-Thomas, A.; Berg, S.L.; Gururangan, S.; Scorsone, K.; Su, J.; Goldman, S.; Kieran, M.W.; Kun, L.; others. A phase-1 pharmacokinetic optimal dosing study of intraventricular topotecan for children with neoplastic meningitis: A pediatric brain tumor consortium study. Pediatric blood & cancer 2013, 60, 627–632. [Google Scholar]
- Fang, Z.; Liu, X.; Peltz, G. GSEApy: a comprehensive package for performing gene set enrichment analysis in Python. Bioinformatics 2023, 39, btac757. [Google Scholar] [CrossRef]
- Liberzon, A.; Birger, C.; Thorvaldsdóttir, H.; Ghandi, M.; Mesirov, J.P.; Tamayo, P. The molecular signatures database hallmark gene set collection. Cell systems 2015, 1, 417–425. [Google Scholar] [CrossRef]
- Hamidi, M.; Eriz, A.; Mitxelena, J.; Fernandez-Ares, L.; Aurrekoetxea, I.; Aspichueta, P.; Iglesias-Ara, A.; Zubiaga, A.M. Targeting E2F Sensitizes Prostate Cancer Cells to Drug-Induced Replication Stress by Promoting Unscheduled CDK1 Activity. Cancers 2022, 14, 4952. [Google Scholar] [CrossRef]
- Pohl, P.C.; Klafke, G.M.; Carvalho, D.D.; Martins, J.R.; Daffre, S.; da Silva Vaz, I., Jr.; Masuda, A. ABC transporter efflux pumps: a defense mechanism against ivermectin in Rhipicephalus (Boophilus) microplus. International journal for parasitology 2011, 41, 1323–1333. [Google Scholar] [CrossRef] [PubMed]
- Omori, M.; Noro, R.; Seike, M.; Matsuda, K.; Hirao, M.; Fukuizumi, A.; Takano, N.; Miyanaga, A.; Gemma, A. Inhibitors of ABCB1 and ABCG2 overcame resistance to topoisomerase inhibitors in small cell lung cancer. Thoracic Cancer 2022, 13, 2142–2151. [Google Scholar] [CrossRef]
- Chang, Y.T.; Hoffman, E.P.; Yu, G.; Herrington, D.M.; Clarke, R.; Wu, C.T.; Chen, L.; Wang, Y. Integrated identification of disease specific pathways using multi-omics data. bioRxiv 2019, 666065. [Google Scholar]
- Chauhan, M.; Sharma, G.; Joshi, G.; Kumar, R. Epidermal growth factor receptor (EGFR) and its cross-talks with topoisomerases: challenges and opportunities for multi-target anticancer drugs. Current Pharmaceutical Design 2016, 22, 3226–3236. [Google Scholar] [CrossRef] [PubMed]
- Weber, G.F.; Weber, G.F. DNA damaging drugs. Molecular therapies of cancer 2015, 9–112. [Google Scholar]
- Wishart, D.S.; Feunang, Y.D.; Guo, A.C.; Lo, E.J.; Marcu, A.; Grant, J.R.; Sajed, T.; Johnson, D.; Li, C.; Sayeeda, Z.; others. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic acids research 2018, 46, D1074–D1082. [Google Scholar] [CrossRef]
- Morgan, H.L. The generation of a unique machine description for chemical structures-a technique developed at chemical abstracts service. Journal of chemical documentation 1965, 5, 107–113. [Google Scholar] [CrossRef]
- Landrum, G.; others. Rdkit: Open-source cheminformatics software 2016.
- Burbidge, R.; Trotter, M.; Buxton, B.; Holden, S. Drug design by machine learning: support vector machines for pharmaceutical data analysis. Computers & chemistry 2001, 26, 5–14. [Google Scholar]
- Brody, S.; Alon, U.; Yahav, E. How attentive are graph attention networks? arXiv preprint arXiv:2105.14491 2021. [Google Scholar]
- Cai, T.; Luo, S.; Xu, K.; He, D.; Liu, T.y.; Wang, L. Graphnorm: A principled approach to accelerating graph neural network training. International Conference on Machine Learning. PMLR, 2021, pp. 1204–1215.
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv preprint arXiv:1412.6980 2014. [Google Scholar]
- Paszke, A.; Gross, S.; Massa, F.; Lerer, A.; Bradbury, J.; Chanan, G.; Killeen, T.; Lin, Z.; Gimelshein, N.; Antiga, L.; others. Pytorch: An imperative style, high-performance deep learning library. Advances in neural information processing systems 2019, 32. [Google Scholar]
- Fey, M.; Lenssen, J.E. Fast graph representation learning with PyTorch Geometric. arXiv preprint arXiv:1903.02428 2019. [Google Scholar]
- Akiba, T.; Sano, S.; Yanase, T.; Ohta, T.; Koyama, M. Optuna: A next-generation hyperparameter optimization framework. Proceedings of the 25th ACM SIGKDD international conference on knowledge discovery & data mining, 2019, pp. 2623–2631.
- Wong, J.V.; Franz, M.; Siper, M.C.; Fong, D.; Durupinar, F.; Dallago, C.; Luna, A.; Giorgi, J.; Rodchenkov, I.; Babur, Ö.; others. Science Forum: Author-sourced capture of pathway knowledge in computable form using Biofactoid. Elife 2021, 10, e68292. [Google Scholar] [CrossRef]
- Bachman, J.A.; Gyori, B.M.; Sorger, P.K. Automated assembly of molecular mechanisms at scale from text mining and curated databases. Molecular Systems Biology 2023, 19, e11325. [Google Scholar] [CrossRef]
- Rodchenkov, I.; Babur, O.; Luna, A.; Aksoy, B.A.; Wong, J.V.; Fong, D.; Franz, M.; Siper, M.C.; Cheung, M.; Wrana, M.; others. Pathway Commons 2019 Update: integration, analysis and exploration of pathway data. Nucleic acids research 2020, 48, D489–D497. [Google Scholar] [CrossRef]
- Lu, Y.; Wu, C.T.; Parker, S.J.; Chen, L.; Saylor, G.; Van Eyk, J.E.; Herrington, D.M.; Wang, Y. COT: an efficient Python tool for detecting marker genes among many subtypes. bioRxiv 2021, 2021-01. [Google Scholar]
- Peng, W.; Chen, T.; Dai, W. Predicting drug response based on multi-omics fusion and graph convolution. IEEE Journal of Biomedical and Health Informatics 2021, 26, 1384–1393. [Google Scholar] [CrossRef]
- Lu, Y.; Wang, H.; Wei, W. Machine Learning for Synthetic Data Generation: a Review. arXiv preprint arXiv:2302.04062 2023. [Google Scholar]
- Du, D.; Bhardwaj, S.; Parker, S.J.; Cheng, Z.; Zhang, Z.; Van Eyk, J.E.; Yu, G.; Clarke, R.; Herrington, D.M.; others. ABDS: tool suite for analyzing biologically diverse samples. bioRxiv 2023. [Google Scholar]
- Lu, Y. Multi-omics Data Integration for Identifying Disease Specific Biological Pathways. PhD thesis, Virginia Tech, 2018.
- Zhang, B.; Fu, Y.; Zhang, Z.; Clarke, R.; Van Eyk, J.E.; Herrington, D.M.; Wang, Y. DDN2. 0: R and Python packages for differential dependency network analysis of biological systems. bioRxiv 2021, 2021-04. [Google Scholar]
- Fu, Y.; Lu, Y.; Wang, Y.; Zhang, B.; Zhang, Z.; Yu, G.; Liu, C.; Clarke, R.; Herrington, D.M.; Wang, Y. DDN3. 0: Determining significant rewiring of biological network structure with differential dependency networks. Bioinformatics 2024, btae376. [Google Scholar] [CrossRef]
- Lu, Y.; Chen, T.; Hao, N.; Rechem, C.V.; Chen, J.; Fu, T. Uncertainty quantification and interpretability for clinical trial approval prediction. Health Data Science 2024. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Hao, N.; Lu, Y.; Van Rechem, C. Uncertainty Quantification on Clinical Trial Outcome Prediction. arXiv preprint arXiv:2401.03482 2024. [Google Scholar]
- Chen, J.; Hu, Y.; Wang, Y.; Lu, Y.; Cao, X.; Lin, M.; Xu, H.; Wu, J.; Xiao, C.; Sun, J.; others. Trialbench: Multi-modal artificial intelligence-ready clinical trial datasets. arXiv preprint arXiv:2407.00631 2024. [Google Scholar]
- An, X.; Chen, X.; Yi, D.; Li, H.; Guan, Y. Representation of molecules for drug response prediction. Briefings in Bioinformatics 2022, 23, bbab393. [Google Scholar] [CrossRef]
- Lu, Y.; Hu, Y.; Li, C. DrugCLIP: Contrastive Drug-Disease Interaction For Drug Repurposing. arXiv preprint arXiv:2407.02265 2024. [Google Scholar]
- Xu, B.; Lu, Y.; Li, C.; Yue, L.; Wang, X.; Hao, N.; Fu, T.; Chen, J. SMILES-Mamba: Chemical Mamba Foundation Models for Drug ADMET Prediction. arXiv preprint arXiv:2408.05696 2024. [Google Scholar]
- Wang, L.; Li, X.; Zhang, L.; Gao, Q. Improved anticancer drug response prediction in cell lines using matrix factorization with similarity regularization. BMC cancer 2017, 17, 1–12. [Google Scholar] [CrossRef]
- Zhang, F.; Wang, M.; Xi, J.; Yang, J.; Li, A. A novel heterogeneous network-based method for drug response prediction in cancer cell lines. Scientific reports 2018, 8, 1–9. [Google Scholar] [CrossRef]
- Koren, Y.; Bell, R.; Volinsky, C. Matrix factorization techniques for recommender systems. Computer 2009, 42, 30–37. [Google Scholar] [CrossRef]
- Kipf, T.N.; Welling, M. Semi-supervised classification with graph convolutional networks. The International Conference on Learning Representations (ICLR) 2016. [Google Scholar]
- Bolton, E.E.; Wang, Y.; Thiessen, P.A.; Bryant, S.H. PubChem: integrated platform of small molecules and biological activities. In Annual reports in computational chemistry; Elsevier, 2008; Vol. 4, pp. 217–241.
- Li, M.; Wang, Y.; Zheng, R.; Shi, X.; Li, Y.; Wu, F.X.; Wang, J. DeepDSC: a deep learning method to predict drug sensitivity of cancer cell lines. IEEE/ACM transactions on computational biology and bioinformatics 2019, 18, 575–582. [Google Scholar] [CrossRef]
- Jin, H.; Song, Q.; Hu, X. Auto-keras: An efficient neural architecture search system. Proceedings of the 25th ACM SIGKDD international conference on knowledge discovery & data mining, 2019, pp. 1946–1956.
- Wang, C.; Wu, Q.; Weimer, M.; Zhu, E. Flaml: A fast and lightweight automl library. Proceedings of Machine Learning and Systems 2021, 3, 434–447. [Google Scholar]
- Gilmer, J.; Schoenholz, S.S.; Riley, P.F.; Vinyals, O.; Dahl, G.E. Neural message passing for quantum chemistry. International conference on machine learning. PMLR, 2017, pp. 1263–1272.
- Shi, Y.; Huang, Z.; Feng, S.; Zhong, H.; Wang, W.; Sun, Y. Masked label prediction: Unified message passing model for semi-supervised classification. arXiv preprint arXiv:2009.03509 2020. [Google Scholar]
| Method | Description | Data Structure | Interpretability | Accuracy | Precision | Recall | F1 score | |
| Baseline | Deep DSC | AE | DF, GE | - | 0.548 ± 0.000 | 0.516 ± 0.000 | 0.481 ± 0.000 | 0.498 ± 0.000 |
| MOFGCN | GNNs | DF, GE, MT, CNV | - | 0.499 ± 0.001 | 0.487 ± 0.001 | 0.478 ± 0.002 | 0.482 ± 0.001 | |
| Random Forest | Tree | DF, GE | Feature Importance | 0.743 ± 0.003 | 0.720 ± 0.003 | 0.734 ± 0.006 | 0.727 ± 0.004 | |
| LightGBM | Tree | DF, GE | Feature Importance | 0.766 ± 0.000 | 0.791± 0.000 | 0.676 ± 0.000 | 0.729 ± 0.000 | |
| AutoKeras | DNN | PCA+DF, PCA+GE |
- | 0.733 ± 0.007 | 0.709 ± 0.011 | 0.724 ± 0.015 | 0.716 ± 0.007 | |
| drGAT | MPNN | GNNs | DCG | - | 0.620 ± 0.126 | 0.623 ± 0.129 | 0.791± 0.182 | 0.664 ± 0.036 |
| GCN | GNNs | DCG | - | 0.678 ± 0.036 | 0.720 ± 0.022 | 0.506 ± 0.119 | 0.587 ± 0.082 | |
| GAT | GNNs | DCG | Attention | 0.763 ± 0.018 | 0.730 ± 0.040 | 0.790 ± 0.045 | 0.756 ± 0.009 | |
| GATv2 | GNNs | DCG | Attention | 0.779± 0.002 | 0.775 ± 0.013 | 0.741 ± 0.023 | 0.757± 0.006 | |
| Graph Transformer | GNNs | DCG | Attention | 0.764 ± 0.01 | 0.739 ± 0.024 | 0.766 ± 0.025 | 0.752 ± 0.008 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




