Submitted:
16 August 2024
Posted:
26 August 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Structure-Activity Relationships
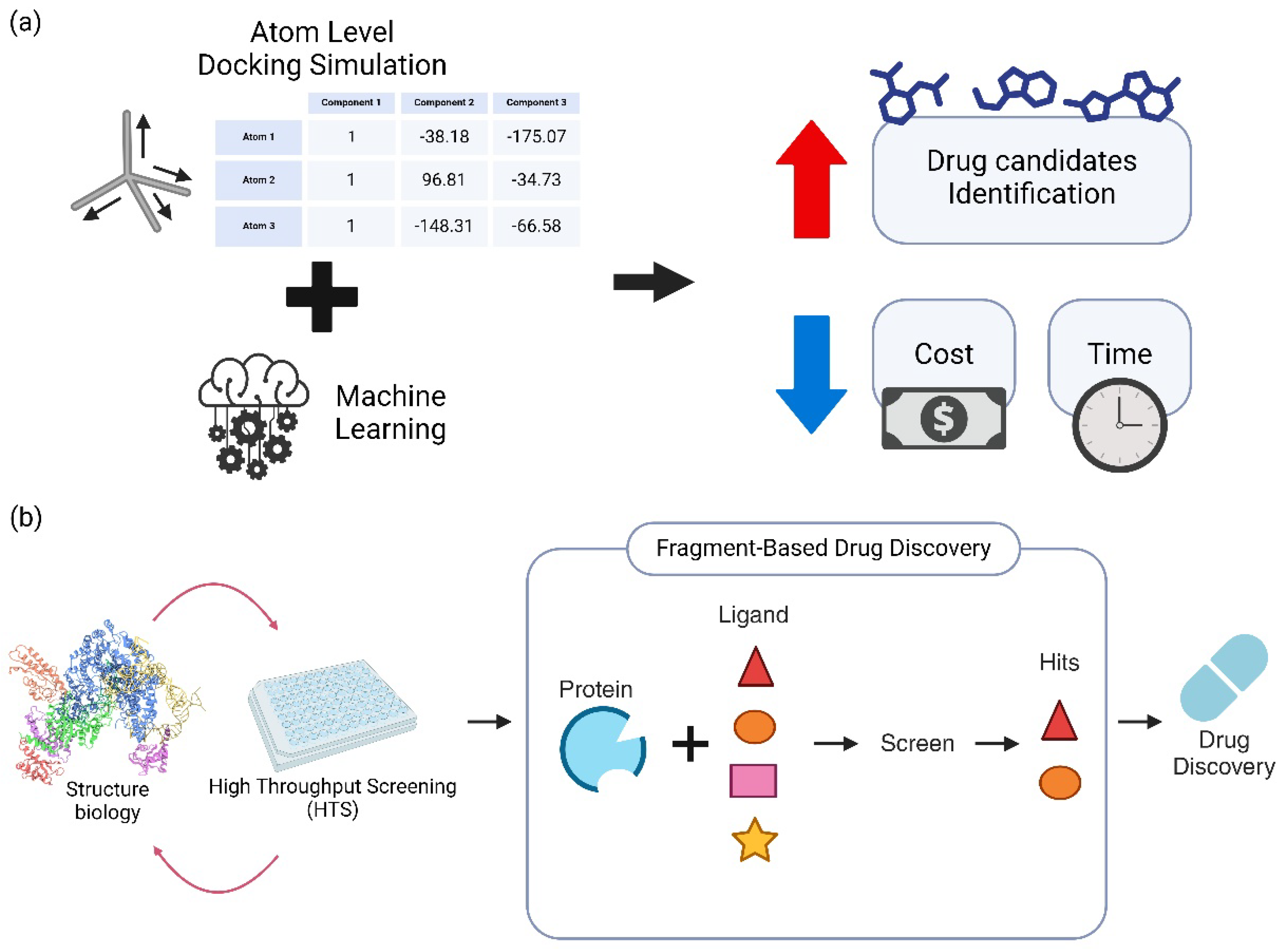
2.1. Computational SAR Models
2.2. Fragment-Based Drug Design
3. Biochemical and Pharmacological Targets
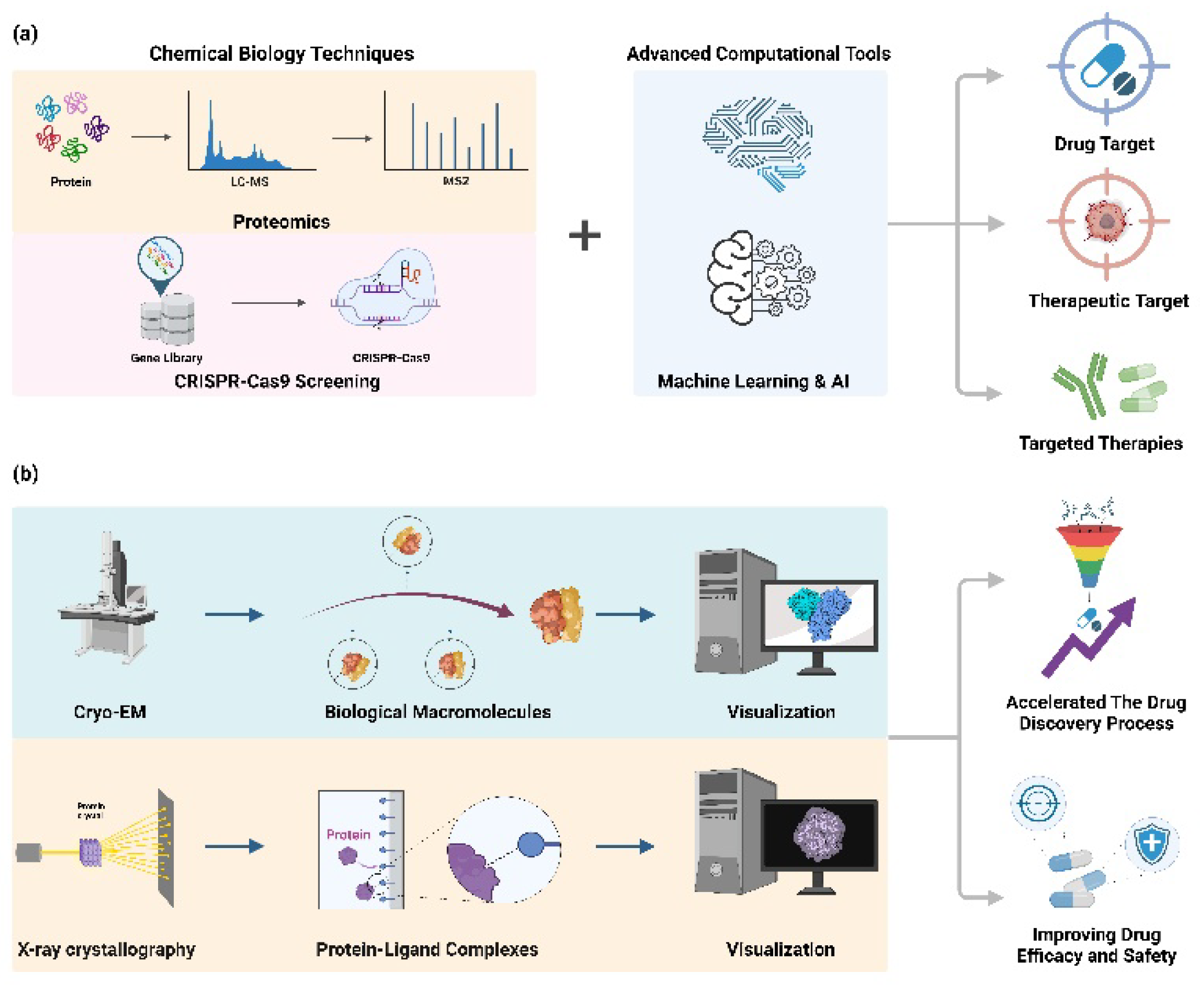
3.1. Target Identification
3.2. Mechanism of Action
4. Drug Design and Synthetic Chemistry
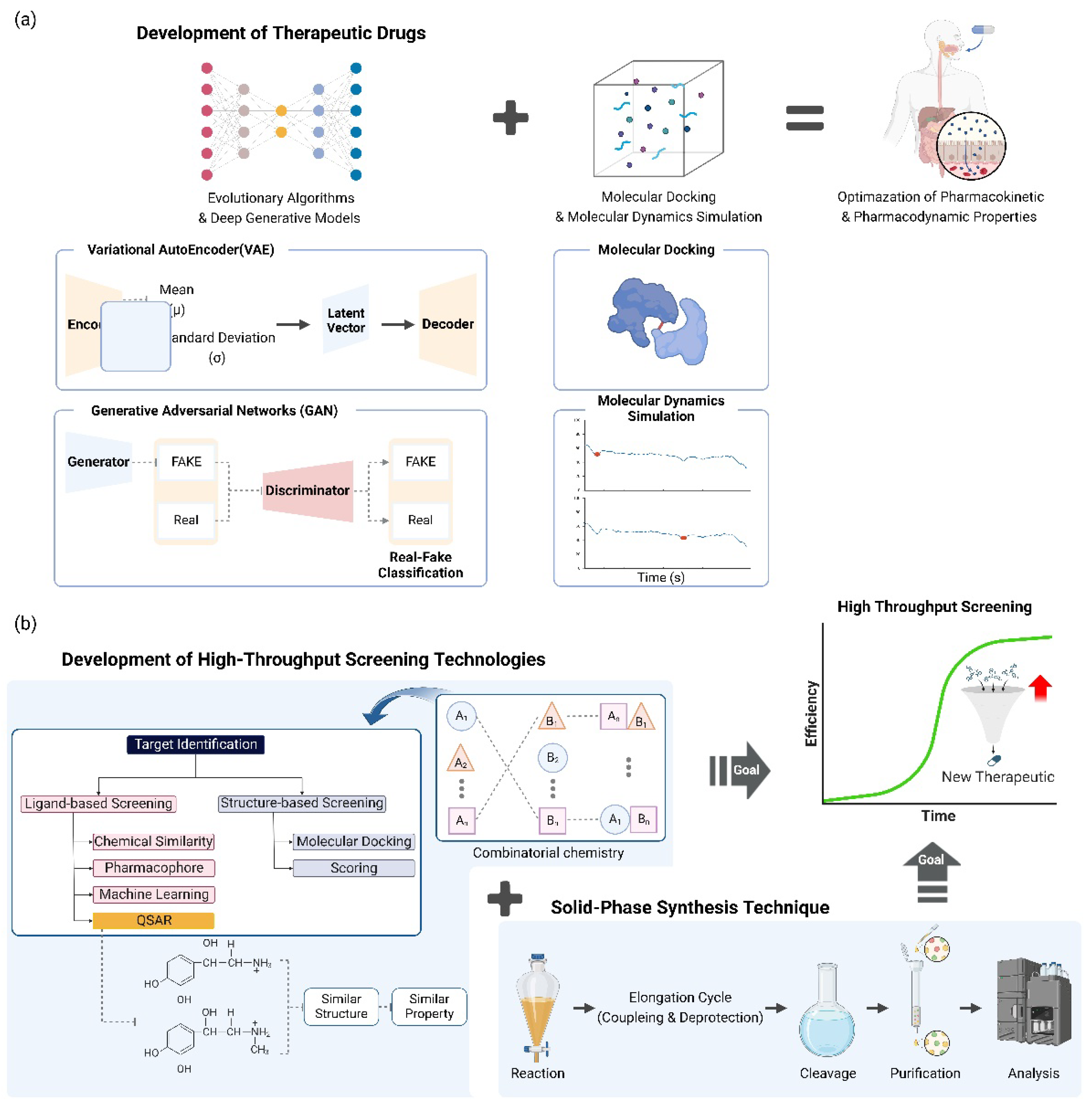
4.1. Automated De Novo Drug Design
4.2. Combinatorial Synthetic Chemistry
5. Virtual Screening
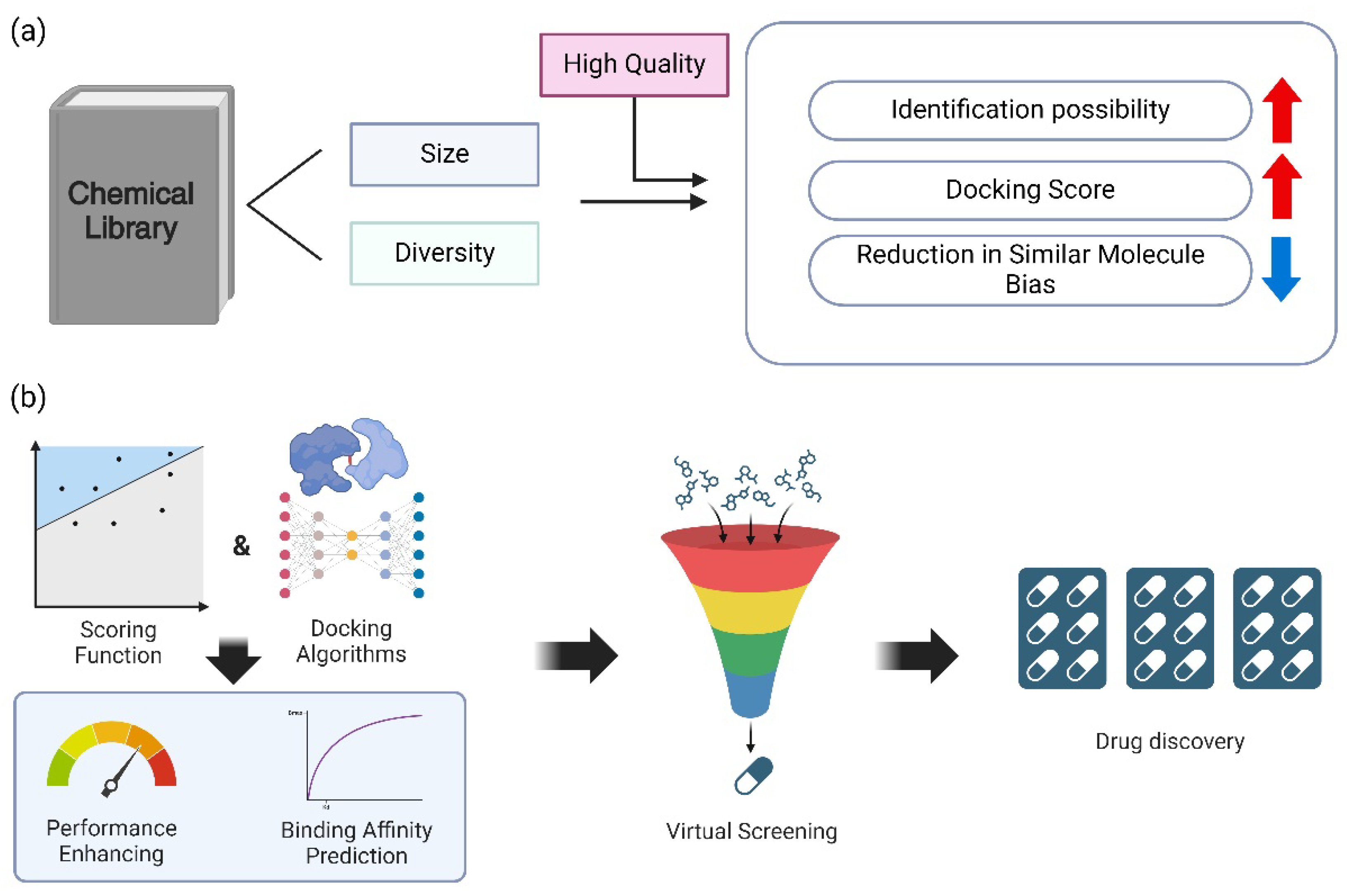
5.1. Library Size and Diversity
5.2. Scoring Functions and Docking Algorithms
6. Drug Safety and Toxicology
6.1. Predictive Toxicology
6.2. Drug Toxicity Mechanisms
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Guengerich, F.P. Mechanisms of drug toxicity and relevance to pharmaceutical development. Drug Metab Pharmacokinet 2011, 26, 3–14. [Google Scholar] [CrossRef] [PubMed]
- Pognan, F.; Beilmann, M.; Boonen, H.C.M.; Czich, A.; Dear, G.; Hewitt, P.; Mow, T.; Oinonen, T.; Roth, A.; Steger-Hartmann, T.; et al. The evolving role of investigative toxicology in the pharmaceutical industry. Nat Rev Drug Discov 2023, 22, 317–335. [Google Scholar] [CrossRef]
- Nguyen, N.; Jennen, D.; Kleinjans, J. Omics technologies to understand drug toxicity mechanisms. Drug Discov Today 2022, 27, 103348. [Google Scholar] [CrossRef] [PubMed]
- Cavasotto, C.N.; Scardino, V. Machine Learning Toxicity Prediction: Latest Advances by Toxicity End Point. ACS Omega 2022, 7, 47536–47546. [Google Scholar] [CrossRef]
- Tran, T.T.V.; Surya Wibowo, A.; Tayara, H.; Chong, K.T. Artificial Intelligence in Drug Toxicity Prediction: Recent Advances, Challenges, and Future Perspectives. J Chem Inf Model 2023, 63, 2628–2643. [Google Scholar] [CrossRef]
- Ingber, D.E. Human organs-on-chips for disease modelling, drug development and personalized medicine. Nat Rev Genet 2022, 23, 467–491. [Google Scholar] [CrossRef] [PubMed]
- Baker, T.K.; Van Vleet, T.R.; Mahalingaiah, P.K.; Grandhi, T.S.P.; Evers, R.; Ekert, J.; Gosset, J.R.; Chacko, S.A.; Kopec, A.K. The Current Status and Use of Microphysiological Systems by the Pharmaceutical Industry: The International Consortium for Innovation and Quality Microphysiological Systems Affiliate Survey and Commentary. Drug Metab Dispos 2024, 52, 198–209. [Google Scholar] [CrossRef]
- Rowland Yeo, K.; Gil Bergland, E.; Chen, Y. Dose Optimization Informed by PBPK Modeling: State-of-the Art and Future. Clin Pharmacol Ther 2024. [Google Scholar] [CrossRef]
- Chen, C.; Wang, J.; Pan, D.; Wang, X.; Xu, Y.; Yan, J.; Wang, L.; Yang, X.; Yang, M.; Liu, G.P. Applications of multi-omics analysis in human diseases. MedComm (2020) 2023, 4, e315. [Google Scholar] [CrossRef]
- Marques, L.; Costa, B.; Pereira, M.; Silva, A.; Santos, J.; Saldanha, L.; Silva, I.; Magalhaes, P.; Schmidt, S.; Vale, N. Advancing Precision Medicine: A Review of Innovative In Silico Approaches for Drug Development, Clinical Pharmacology and Personalized Healthcare. Pharmaceutics 2024, 16. [Google Scholar] [CrossRef]
- Ivanisevic, T.; Sewduth, R.N. Multi-Omics Integration for the Design of Novel Therapies and the Identification of Novel Biomarkers. Proteomes 2023, 11. [Google Scholar] [CrossRef]
- Chicco, D.; Cumbo, F.; Angione, C. Ten quick tips for avoiding pitfalls in multi-omics data integration analyses. PLoS Comput Biol 2023, 19, e1011224. [Google Scholar] [CrossRef]
- Guha, R. On exploring structure-activity relationships. Methods Mol Biol 2013, 993, 81–94. [Google Scholar] [CrossRef]
- Béquignon, O.J.M.; Gómez-Tamayo, J.C.; Lenselink, E.B.; Wink, S.; Hiemstra, S.; Lam, C.C.; Gadaleta, D.; Roncaglioni, A.; Norinder, U.; Water, B.v.d.; et al. Collaborative SAR Modeling and Prospective In Vitro Validation of Oxidative Stress Activation in Human HepG2 Cells. Journal of Chemical Information and Modeling 2023, 63, 5433–5445. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.H.; Thai, Q.M.; Pham, M.Q.; Minh, P.T.H.; Phung, H.T.T. Machine learning combines atomistic simulations to predict SARS-CoV-2 Mpro inhibitors from natural compounds. Molecular Diversity 2023, 28, 553–561. [Google Scholar] [CrossRef] [PubMed]
- Batool, M.; Ahmad, B.; Choi, S. A Structure-Based Drug Discovery Paradigm. International Journal of Molecular Sciences 2019, 20. [Google Scholar] [CrossRef] [PubMed]
- Sliwoski, G.; Kothiwale, S.; Meiler, J.; Lowe, E.W.; Barker, E.L. Computational Methods in Drug Discovery. Pharmacological Reviews 2013, 66, 334–395. [Google Scholar] [CrossRef]
- Naithani, U.; Guleria, V. Integrative computational approaches for discovery and evaluation of lead compound for drug design. Frontiers in Drug Discovery 2024, 4. [Google Scholar] [CrossRef]
- Kirsch, P.; Hartman, A.M.; Hirsch, A.K.H.; Empting, M. Concepts and Core Principles of Fragment-Based Drug Design. Molecules 2019, 24. [Google Scholar] [CrossRef]
- Li, Q. Application of Fragment-Based Drug Discovery to Versatile Targets. Frontiers in Molecular Biosciences 2020, 7. [Google Scholar] [CrossRef]
- Chan, B.W.G.L.; Lynch, N.B.; Tran, W.; Joyce, J.M.; Savage, G.P.; Meutermans, W.; Montgomery, A.P.; Kassiou, M. Fragment-based drug discovery for disorders of the central nervous system: designing better drugs piece by piece. Frontiers in Chemistry 2024, 12. [Google Scholar] [CrossRef]
- Bon, M.; Bilsland, A.; Bower, J.; McAulay, K. Fragment-based drug discovery—the importance of high-quality molecule libraries. Molecular Oncology 2022, 16, 3761–3777. [Google Scholar] [CrossRef] [PubMed]
- Mallakuntla, M.K.; Togre, N.S.; Santos, D.B.; Tiwari, S. Implications of Fragment-Based Drug Discovery in Tuberculosis and HIV. Pharmaceuticals 2022, 15. [Google Scholar] [CrossRef]
- Bon, M.; Bilsland, A.; Bower, J.; McAulay, K. Fragment-based drug discovery-the importance of high-quality molecule libraries. Mol Oncol 2022, 16, 3761–3777. [Google Scholar] [CrossRef]
- Erlanson, D.A.; Fesik, S.W.; Hubbard, R.E.; Jahnke, W.; Jhoti, H. Twenty years on: the impact of fragments on drug discovery. Nat Rev Drug Discov 2016, 15, 605–619. [Google Scholar] [CrossRef] [PubMed]
- Chan, B.; Lynch, N.B.; Tran, W.; Joyce, J.M.; Savage, G.P.; Meutermans, W.; Montgomery, A.P.; Kassiou, M. Fragment-based drug discovery for disorders of the central nervous system: designing better drugs piece by piece. Front Chem 2024, 12, 1379518. [Google Scholar] [CrossRef]
- Villemagne, B.; Faion, L.; Tangara, S.; Willand, N. Recent advances in Fragment-based strategies against tuberculosis. Eur J Med Chem 2023, 258, 115569. [Google Scholar] [CrossRef]
- Jost, M.; Weissman, J.S. CRISPR Approaches to Small Molecule Target Identification. ACS Chem Biol 2018, 13, 366–375. [Google Scholar] [CrossRef] [PubMed]
- Schenone, M.; Dancik, V.; Wagner, B.K.; Clemons, P.A. Target identification and mechanism of action in chemical biology and drug discovery. Nat Chem Biol 2013, 9, 232–240. [Google Scholar] [CrossRef]
- Chen, X.; Wang, Y.; Ma, N.; Tian, J.; Shao, Y.; Zhu, B.; Wong, Y.K.; Liang, Z.; Zou, C.; Wang, J. Target identification of natural medicine with chemical proteomics approach: probe synthesis, target fishing and protein identification. Signal Transduct Target Ther 2020, 5, 72. [Google Scholar] [CrossRef]
- Hedl, T.J.; San Gil, R.; Cheng, F.; Rayner, S.L.; Davidson, J.M.; De Luca, A.; Villalva, M.D.; Ecroyd, H.; Walker, A.K.; Lee, A. Proteomics Approaches for Biomarker and Drug Target Discovery in ALS and FTD. Front Neurosci 2019, 13, 548. [Google Scholar] [CrossRef]
- Tabana, Y.; Babu, D.; Fahlman, R.; Siraki, A.G.; Barakat, K. Target identification of small molecules: an overview of the current applications in drug discovery. BMC Biotechnol 2023, 23, 44. [Google Scholar] [CrossRef] [PubMed]
- Woo, J.H.; Shimoni, Y.; Yang, W.S.; Subramaniam, P.; Iyer, A.; Nicoletti, P.; Rodriguez Martinez, M.; Lopez, G.; Mattioli, M.; Realubit, R.; et al. Elucidating Compound Mechanism of Action by Network Perturbation Analysis. Cell 2015, 162, 441–451. [Google Scholar] [CrossRef]
- Maveyraud, L.; Mourey, L. Protein X-ray Crystallography and Drug Discovery. Molecules 2020, 25. [Google Scholar] [CrossRef]
- Kausar, S.; Said Khan, F.; Ishaq Mujeeb Ur Rehman, M.; Akram, M.; Riaz, M.; Rasool, G.; Hamid Khan, A.; Saleem, I.; Shamim, S.; Malik, A. A review: Mechanism of action of antiviral drugs. Int J Immunopathol Pharmacol 2021, 35, 20587384211002621. [Google Scholar] [CrossRef]
- Lees, J.A.; Dias, J.M.; Han, S. Applications of Cryo-EM in small molecule and biologics drug design. Biochem Soc Trans 2021, 49, 2627–2638. [Google Scholar] [CrossRef] [PubMed]
- Van Drie, J.H.; Tong, L. Cryo-EM as a powerful tool for drug discovery. Bioorg Med Chem Lett 2020, 30, 127524. [Google Scholar] [CrossRef]
- Zheng, H.; Handing, K.B.; Zimmerman, M.D.; Shabalin, I.G.; Almo, S.C.; Minor, W. X-ray crystallography over the past decade for novel drug discovery - where are we heading next? Expert Opin Drug Discov 2015, 10, 975–989. [Google Scholar] [CrossRef] [PubMed]
- Cebi, E.; Lee, J.; Subramani, V.K.; Bak, N.; Oh, C.; Kim, K.K. Cryo-electron microscopy-based drug design. Front Mol Biosci 2024, 11, 1342179. [Google Scholar] [CrossRef]
- Banerjee, S.; Banerjee, D.; Singh, A.; Kumar, S.; Pooja, D.; Ram, V.; Kulhari, H.; Saharan, V.A. A Clinical Insight on New Discovered Molecules and Repurposed Drugs for the Treatment of COVID-19. Vaccines (Basel) 2023, 11. [Google Scholar] [CrossRef]
- Zhu, K.F.; Yuan, C.; Du, Y.M.; Sun, K.L.; Zhang, X.K.; Vogel, H.; Jia, X.D.; Gao, Y.Z.; Zhang, Q.F.; Wang, D.P.; et al. Applications and prospects of cryo-EM in drug discovery. Mil Med Res 2023, 10, 10. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Moretti, R.; Meiler, J. Recent Advances in Automated Structure-Based De Novo Drug Design. J Chem Inf Model 2024, 64, 1794–1805. [Google Scholar] [CrossRef] [PubMed]
- Atz, K.; Cotos, L.; Isert, C.; Hakansson, M.; Focht, D.; Hilleke, M.; Nippa, D.F.; Iff, M.; Ledergerber, J.; Schiebroek, C.C.G.; et al. Prospective de novo drug design with deep interactome learning. Nat Commun 2024, 15, 3408. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Dai, H.; Knight, E.; Wu, F.; Li, Y.; Li, T.; Gerstein, M. A survey of generative AI for de novo drug design: new frontiers in molecule and protein generation. Brief Bioinform 2024, 25. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Li, X.; Lam, K.S. Combinatorial chemistry in drug discovery. Curr Opin Chem Biol 2017, 38, 117–126. [Google Scholar] [CrossRef]
- Galloway, W.R.; Isidro-Llobet, A.; Spring, D.R. Diversity-oriented synthesis as a tool for the discovery of novel biologically active small molecules. Nat Commun 2010, 1, 80. [Google Scholar] [CrossRef]
- Sadybekov, A.V.; Katritch, V. Computational approaches streamlining drug discovery. Nature 2023, 616, 673–685. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, K.K.; Ha, H.H.; Kang, N.Y.; Chandran, Y.; Chang, Y.T. Solid phase combinatorial synthesis of a xanthone library using click chemistry and its application to an embryonic stem cell probe. Chem Commun (Camb) 2011, 47, 7488–7490. [Google Scholar] [CrossRef]
- Lyu, J.; Irwin, J.J.; Shoichet, B.K. Modeling the expansion of virtual screening libraries. Nat Chem Biol 2023, 19, 712–718. [Google Scholar] [CrossRef]
- Kuan, J.; Radaeva, M.; Avenido, A.; Cherkasov, A.; Gentile, F. Keeping pace with the explosive growth of chemical libraries with structure-based virtual screening. WIREs Computational Molecular Science 2023, 13. [Google Scholar] [CrossRef]
- Brenk, R.; Schipani, A.; James, D.; Krasowski, A.; Gilbert, I.H.; Frearson, J.; Wyatt, P.G. Lessons learnt from assembling screening libraries for drug discovery for neglected diseases. ChemMedChem 2008, 3, 435–444. [Google Scholar] [CrossRef]
- Grotsch, K.; Sadybekov, A.V.; Hiller, S.; Zaidi, S.; Eremin, D.; Le, A.; Liu, Y.; Smith, E.C.; Illiopoulis-Tsoutsouvas, C.; Thomas, J.; et al. Virtual Screening of a Chemically Diverse “Superscaffold” Library Enables Ligand Discovery for a Key GPCR Target. ACS Chem Biol 2024, 19, 866–874. [Google Scholar] [CrossRef] [PubMed]
- Shamsian, S.; Sokouti, B.; Dastmalchi, S. Benchmarking different docking protocols for predicting the binding poses of ligands complexed with cyclooxygenase enzymes and screening chemical libraries. Bioimpacts 2024, 14, 29955. [Google Scholar] [CrossRef] [PubMed]
- Guedes, I.A.; Pereira, F.S.S.; Dardenne, L.E. Empirical Scoring Functions for Structure-Based Virtual Screening: Applications, Critical Aspects, and Challenges. Front Pharmacol 2018, 9, 1089. [Google Scholar] [CrossRef]
- Vakser, I.A. Protein-protein docking: from interaction to interactome. Biophys J 2014, 107, 1785–1793. [Google Scholar] [CrossRef]
- Leung, C.H.; Ma, D.L. Recent advances in virtual screening for drug discovery. Methods 2015, 71, 1–3. [Google Scholar] [CrossRef]
- Giorgini, M.; Taroncher, M.; Ruiz, M.J.; Rodriguez-Carrasco, Y.; Tolosa, J. In Vitro and Predictive Computational Toxicology Methods for the Neurotoxic Pesticide Amitraz and Its Metabolites. Brain Sci 2023, 13. [Google Scholar] [CrossRef] [PubMed]
- Sharma, B.; Chenthamarakshan, V.; Dhurandhar, A.; Pereira, S.; Hendler, J.A.; Dordick, J.S.; Das, P. Accurate clinical toxicity prediction using multi-task deep neural nets and contrastive molecular explanations. Sci Rep 2023, 13, 4908. [Google Scholar] [CrossRef] [PubMed]
- Najjar, A.; Kramer, N.; Gardner, I.; Hartung, T.; Steger-Hartmann, T. Editorial: Advances in and applications of predictive toxicology: 2022. Front Pharmacol 2023, 14, 1257423. [Google Scholar] [CrossRef]
- Barnes, D.A.; Firman, J.W.; Belfield, S.J.; Cronin, M.T.D.; Vinken, M.; Janssen, M.J.; Masereeuw, R. Development of an adverse outcome pathway network for nephrotoxicity. Arch Toxicol 2024, 98, 929–942. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



