5. Classification Using Decision Trees
In [
24], we dealt with the question of whether it is possible to classify RBC stiffness using deep learning based on analyzing the video recording of the flow of RBCs, and what accuracy the classification can be achieve in this way. Video recording is relatively easy to obtain. However, if we had available more detailed data on cell movement, which can be obtained, e.g., by recording the flow from several sides or using different sensors, it can be expected that the classification accuracy would be improved.
If we use video images directly for classification, it is natural to choose deep neural networks as the model. However, in the case where we have specific properties of the captured cell available as data, describing its shape, speed, and changes in these properties over time, it makes sense to apply models that are suitable for working with tabular data. The simulation from which we generated the videos can be as well used to generate such data. Before investing into experiments with expensive sensors, we can thus verify the classification accuracy that can theoretically be expected with such a procedure. At the same time, we will be able to compare the classification ability of individual models and also evaluate which predictors have a significant impact on classification accuracy. This can be important when designing real experiments.
5.1. Used Predictors
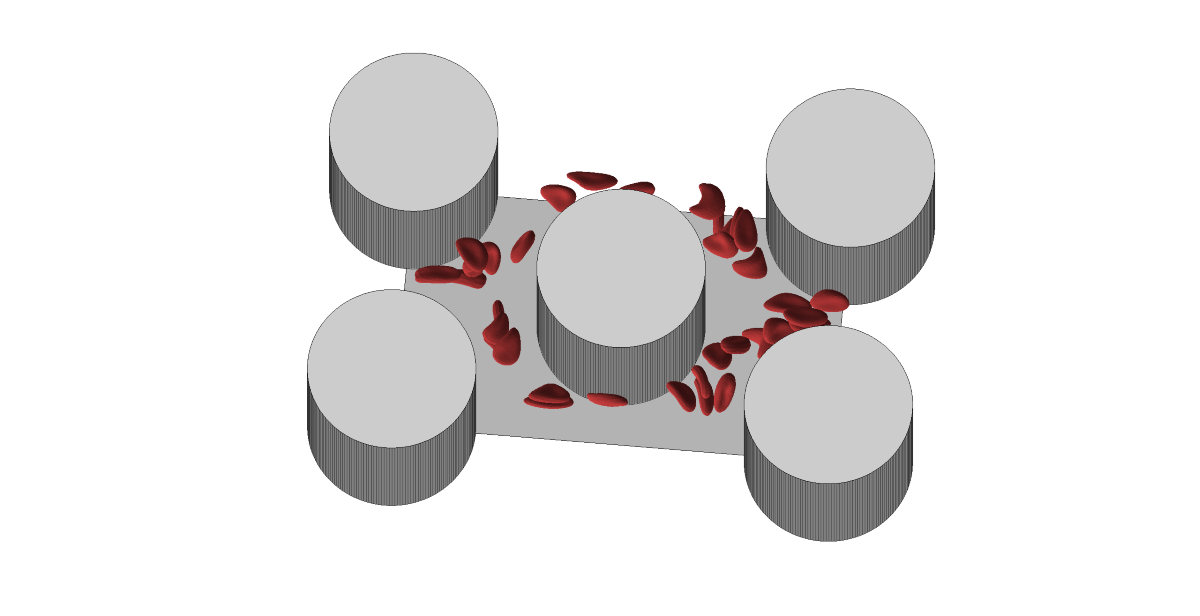
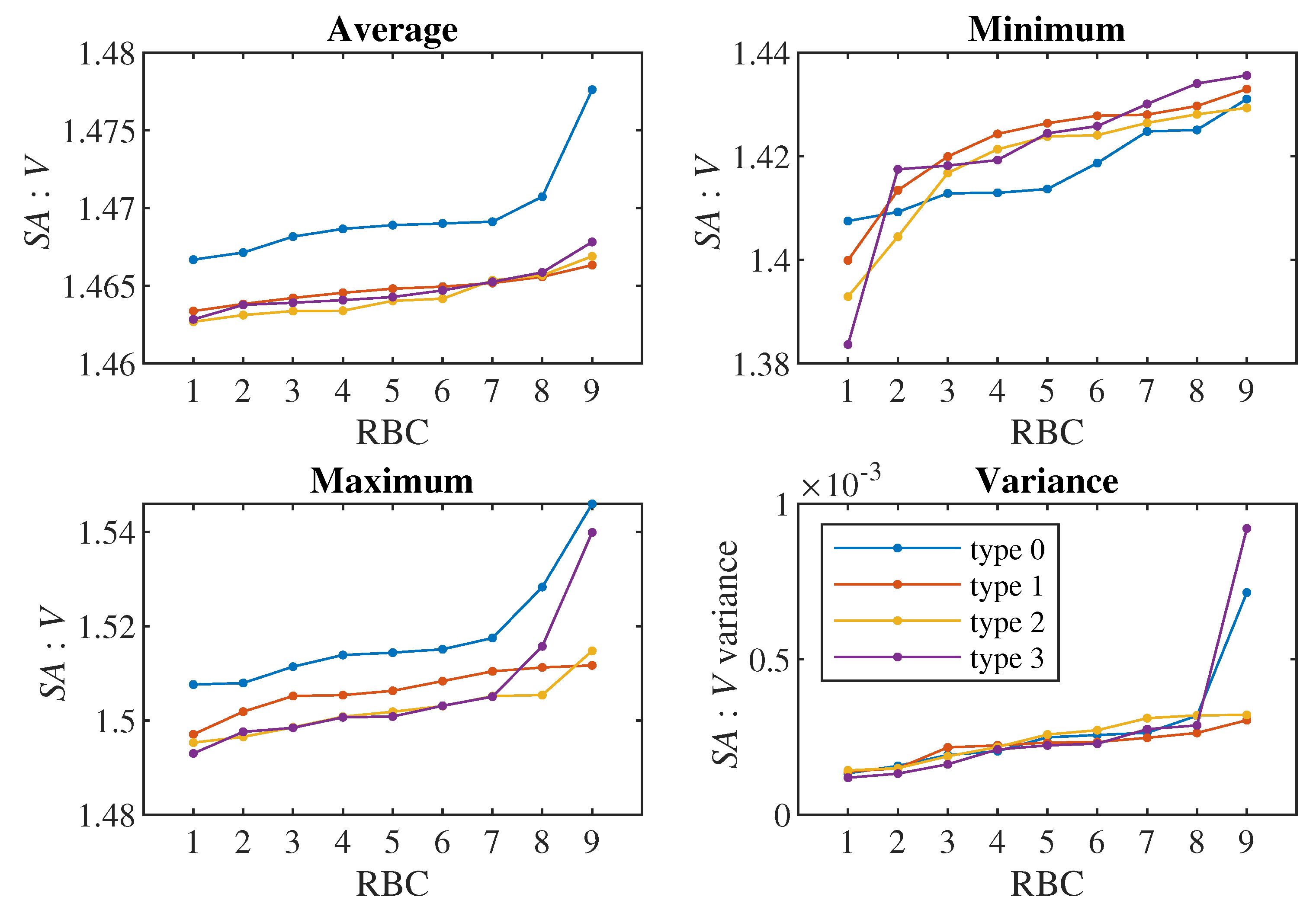
We use cell triangulation to calculate predictors from the simulation. The output of the simulation for each cell is the position of each of the 374 triangulation nodes, determined by three coordinates in three-dimensional space. Various characteristics can be calculated from this data, which we subsequently use as predictors in classification. In total, we created 41 predictors. We divided them into several sets according to the estimated complexity of obtaining them from a real experiment. Based on this, we created 6 tests ordered according to the number of predictors we use. In the first test, we use only the most easily obtainable predictors, while in the final test, we use all of them:
The 1st set consists of only two predictors: the dimensions of the rectangle in which the monitored cell is located, i.e., dimension of the cell in the direction of the x-axis and in the direction of the y-axis. These values can be easily obtained from a (static) snapshot of the cell.
In the 2nd set, the additional predictors are velocities and changes in the velocity of the cell in the x- and y-directions. These data can be calculated from several consecutive images.
The 3rd set also includes dimensions and velocities in the z-axis direction. These values can be obtained from images taken from a different angle.
To the 4th set, we added the cell axis length and the maximum and minimum diameter of the cell equator, which can potentially be determined from multi-angle images.
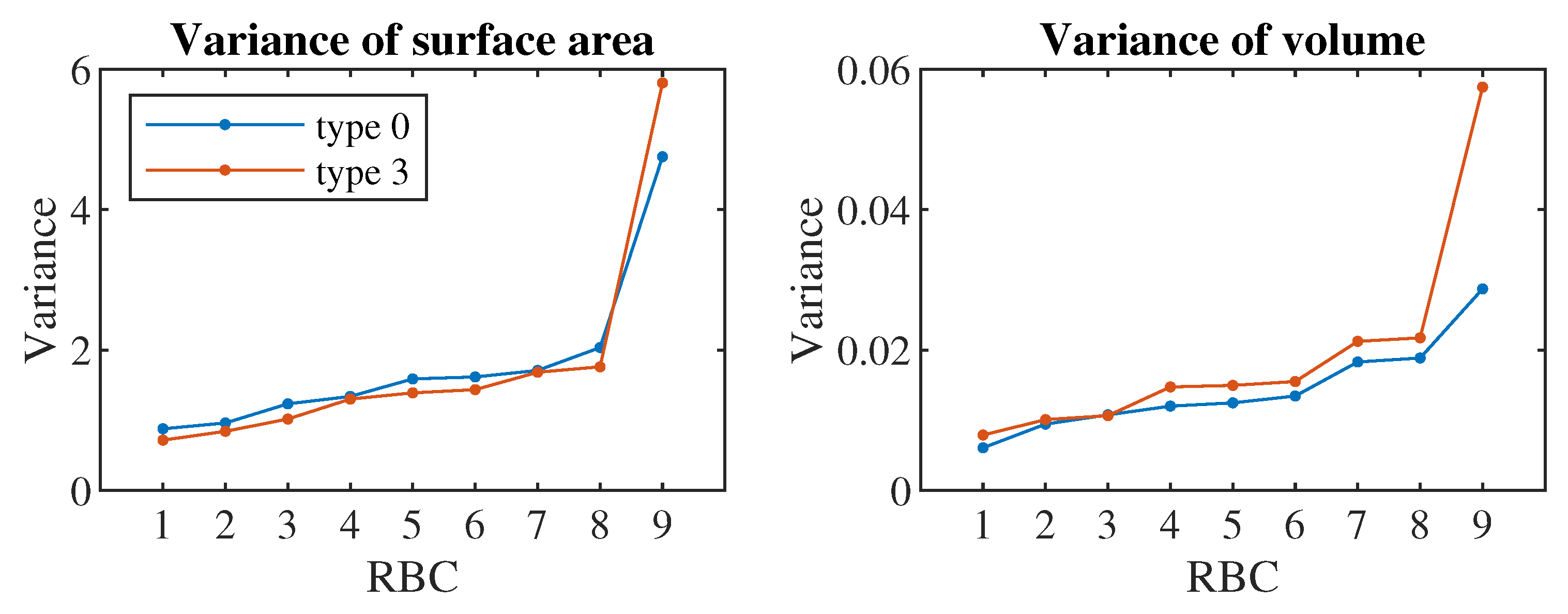
The 5th set additionally contains predictors that can be calculated from the complete cell triangulation: cell surface, cell volume and means, standard deviations and skewness coefficients calculated from the lengths of all triangulation edges, angles formed by every two triangulation triangles and solid angles at all triangulation nodes. Cell triangulation is quite difficult to obtain from a real experiment, it would require the creation of a 3D image of the cell from the scanned flow.
The 6th set also contains means, standard deviations and coefficients of skewness of deviations and absolute deviations of the characteristics added in the 5th set. The deviations are calculated from the cell in a relaxed state, so for the calculation we need to have the basic triangulation of the cell, which in our procedure corresponds to the cell in the first step of the simulation.
The mentioned predictors correspond to the current state of the cell at one monitored moment (in one step of the simulation). During experiments, however, we have the opportunity to observe the cell longer during several steps. It can be expected that the classification accuracy will improve if we track the cell longer. To assess this effect, for each , and 1280, we created a model in which we monitored each cell for simulation steps, with the simulation record saved once per 1000 steps. (Note that one pass of a cell through the channel corresponds to approximately 100000 simulation steps.) Since the values of the individual predictors change during the movement of the cell through the channel, we have S values for each predictor instead of just one value. From these values, we calculated the mean and standard deviation and used them as predictors – in total, we have two predictors for each value described in the bulleted list above (with the exception of the case of , where we only work with static snapshots of the cells).
The technique to extract predictors from series data using statistics like mean, standard deviation and skewness, described both in the previous paragraph and in the 5th and 6th predictor set, is a common approach [
26,
27,
28], which was previously used also for RBC classification [
29].
5.2. Used Machine Learning Tools
Given the nature of the data (number of predictors, high correlation of some predictors), we consider models based on decision trees, which usually achieve the best results for tabular data [
30,
31], to be a suitable tool for classification. We used two of the most popular techniques: random forest and gradient decision trees. For the implementation, we used the Python language and the methods
RandomForestClassifier from the
sklearn library and
XGBClassifier from the
xgboost library.
The goal in this section was not to maximize the accuracy to the highest possible level, therefore we did not optimize the hyperparameters of individual models and were satisfied with the default values.
For each of the six sets of predictors and each of the nine
S values, we trained 2 models using both methods – one for classification into four classes (four types of cell stiffness), the other for classification into two classes (healthy/diseased cell). When evaluating the accuracy of the classification into two classes, in addition to the second model, we also used the first model, in which we combined the three types of stiffness into the result "diseased cell" (similarly, as we did in [
24]).
5.3. Data Preparation
We sampled the simulation for each recorded simulation step, after which followed at least additional steps (so that we could calculate predictors for a given value of S). However, we removed the first 100000 steps of the simulation. The total number of generated samples is thus equal to , where C is the total number of cells in the simulation and N is the total number of recorded simulation steps. (We remind you that every 1000th step is recorded, so the total number of steps is approx. .)
It follows from the above that many generated samples are very similar to each other. The reason is, that the cell changes little between two consecutive steps, moreover, for , two samples recorded close to each other have a large part of the data from which we generate the predictors in common (the last recorded steps of the sample are identical to the first recorded steps of the following sample). This must be kept in mind when dividing the set into a possible training and validation or testing part – it is not appropriate to use data from one simulation in several parts and a new simulation must be used each time.
Due to the tree models used and the omission of hyperparameter optimization, we did not need the validation part, so we we only needed two simulations. We created training samples from the simulation with values of and , the total number of training samples is thus in the range from 81216 (for ) to 35172 (for ). The second simulation, from which we generated test samples, has parameters and , so the number of samples is in a similar range (from 78804 for to 32760 for ).
We recall that the simulations are the same as we used in [
24], that is, there are 9 cells from each class among the 36 cells.
5.4. Classification Results
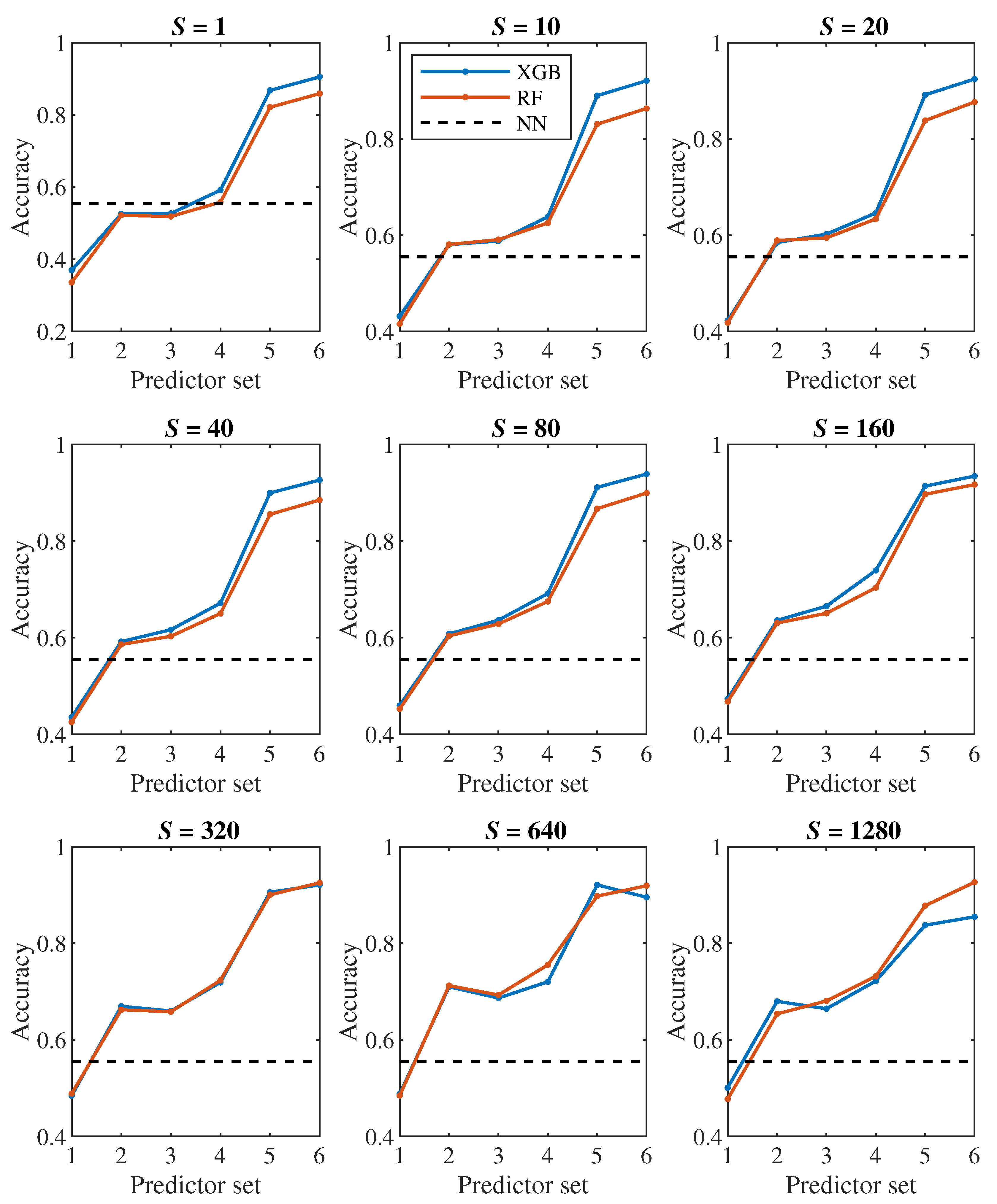
Let us first have a look at the classification accuracy based on the value of
S, that is, let us analyze the effect of the number of steps we monitor each cell. From the plots in
Figure 6,
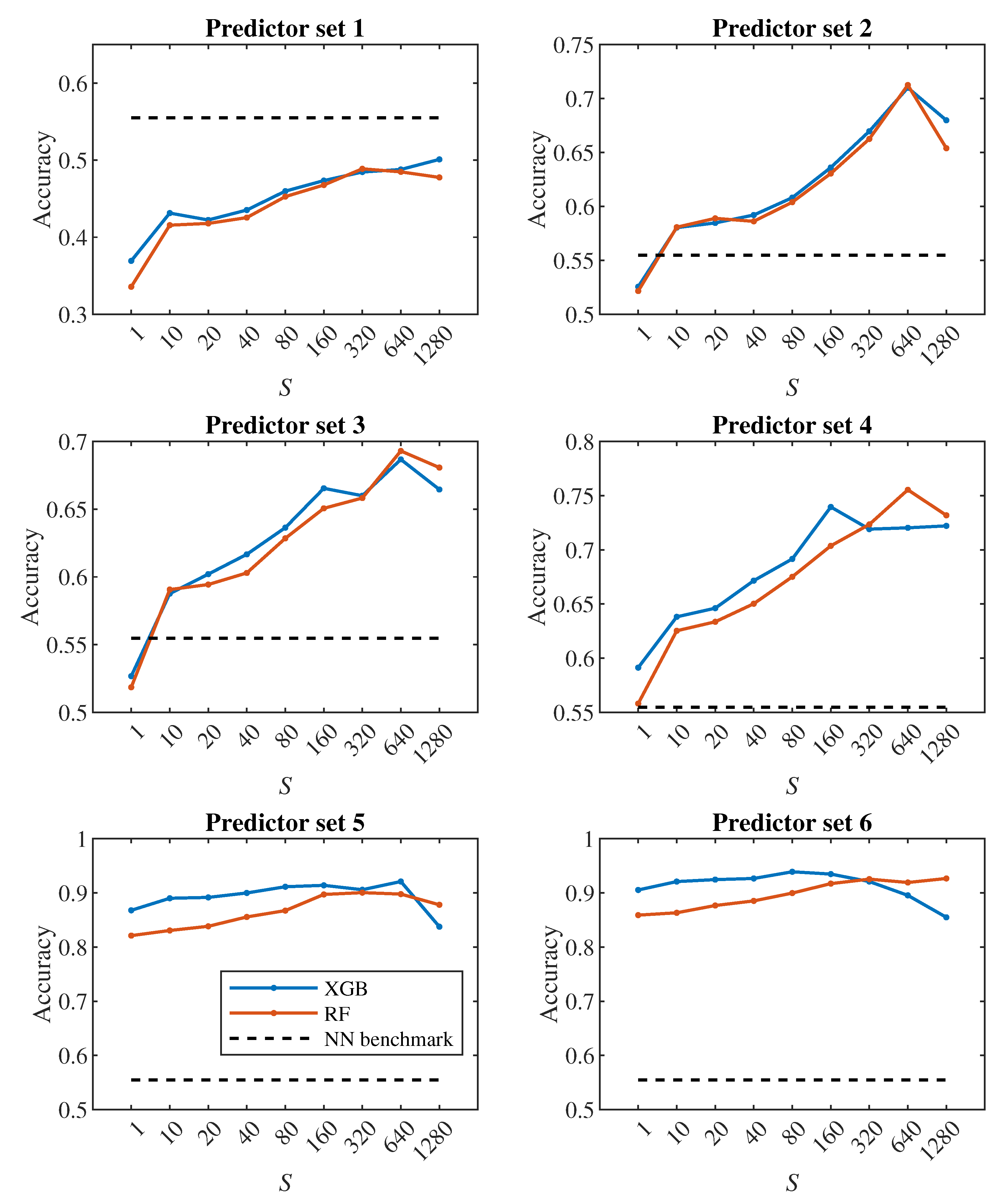
Figure 7, it can be concluded that increasing
S has a positive effect up to the level around the values of
or 320, from which the classification accuracy ceases to improve continuously. Thus, it seems pointless to track the cell for significantly longer than one pass through the channel (approx. 100000 simulation steps).
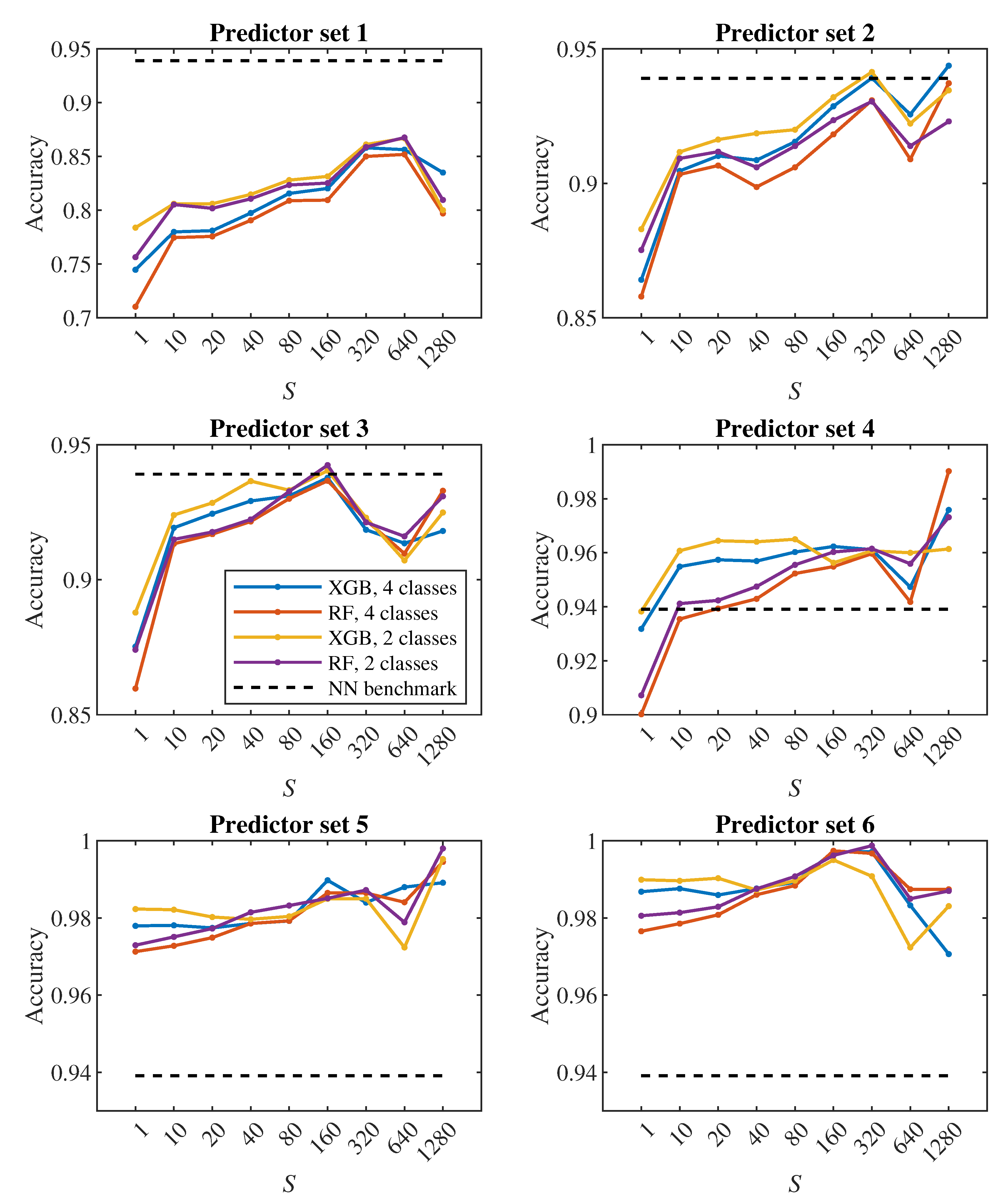
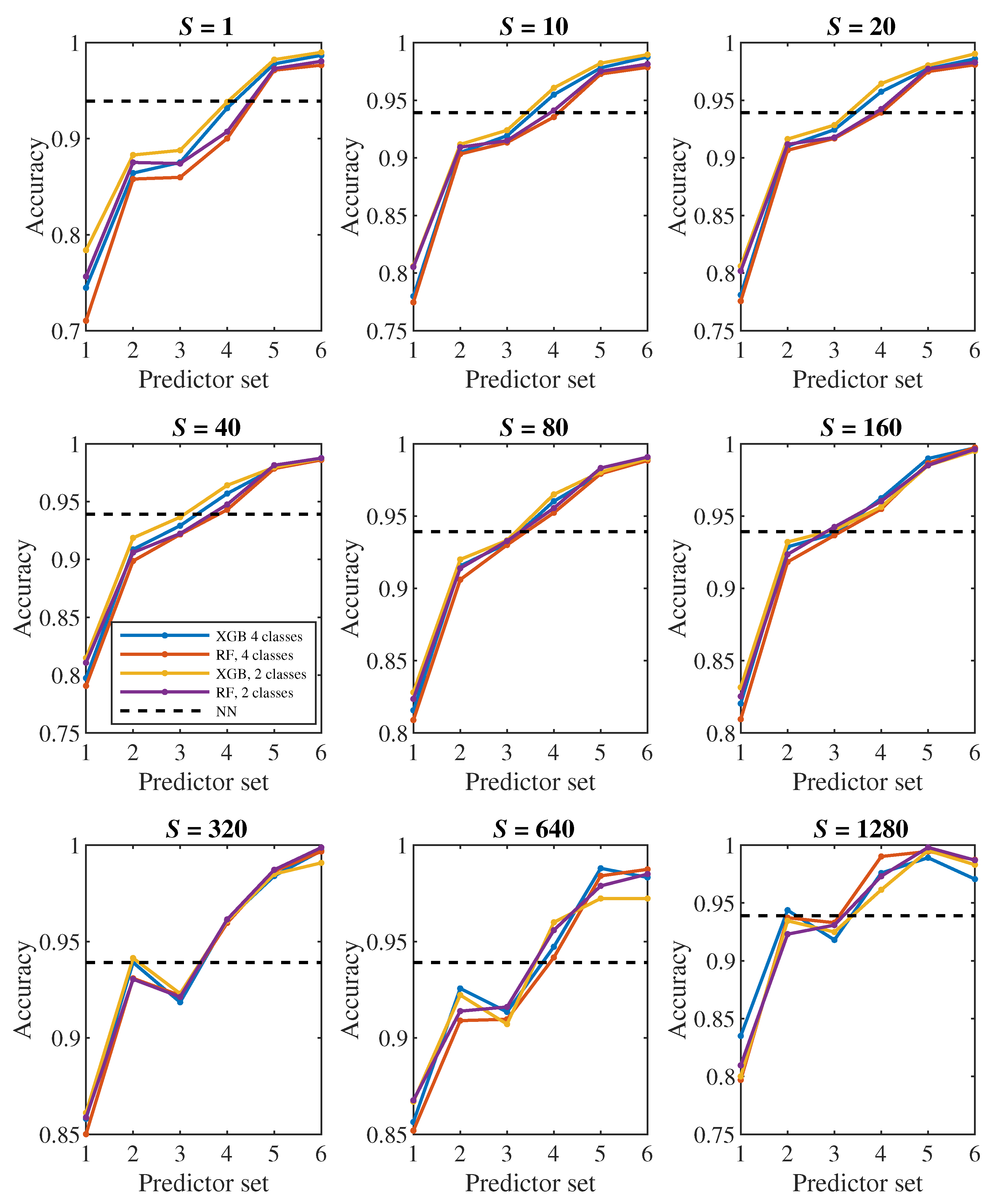
The plots in
Figure 8,
Figure 9 show how the classification accuracy changes depending on which set of predictors we use. Especially when classifying into 4 classes, a significant improvement in accuracy can be observed when moving from the 4th set to the 5th set. On the other side, the effect of the third dimension (the transition from the 2nd set to the 3rd set) does not seem to be very significant.
In all
Figure 6,
Figure 7,
Figure 8 and
Figure 9, the dashed line shows the accuracy we achieved in the classification using deep neural networks in [
24]. Within that approach, we observed each cell during one passage through the channel, which corresponds approximately to a model with
. In
Figure 8 in the middle plot it can be seen that when classifying into 4 classes, we outperformed neural networks result with the 2nd predictor set. Similarly, the middle plot in
Figure 9 shows that in case of 2 classes, we outperformed neural networks with the 4th predictor set.
In [
24], we observed a fairly significant difference in the result between a pair of models created for binary classification (healthy/diseased cell). The model that was trained directly for the classification of 2 classes achieved less accuracy (slightly above 60%) than the model trained for the classification of 4 classes, in which we determined the resulting binary classification only by merging the classes containing diseased cells with three different stiffness levels into one class (accuracy above 90%). This strange phenomenon did not appear when decision trees were used – in
Figure 7,
Figure 9 it can be seen that the accuracy of models trained for the classification of 4 classes (blue and red color) does not differ significantly from the accuracy of models trained directly for binary classification (yellow and purple color).
A comparison of random forest and gradient decision trees does not come out significantly in favor of either method – the results are very similar to each other.
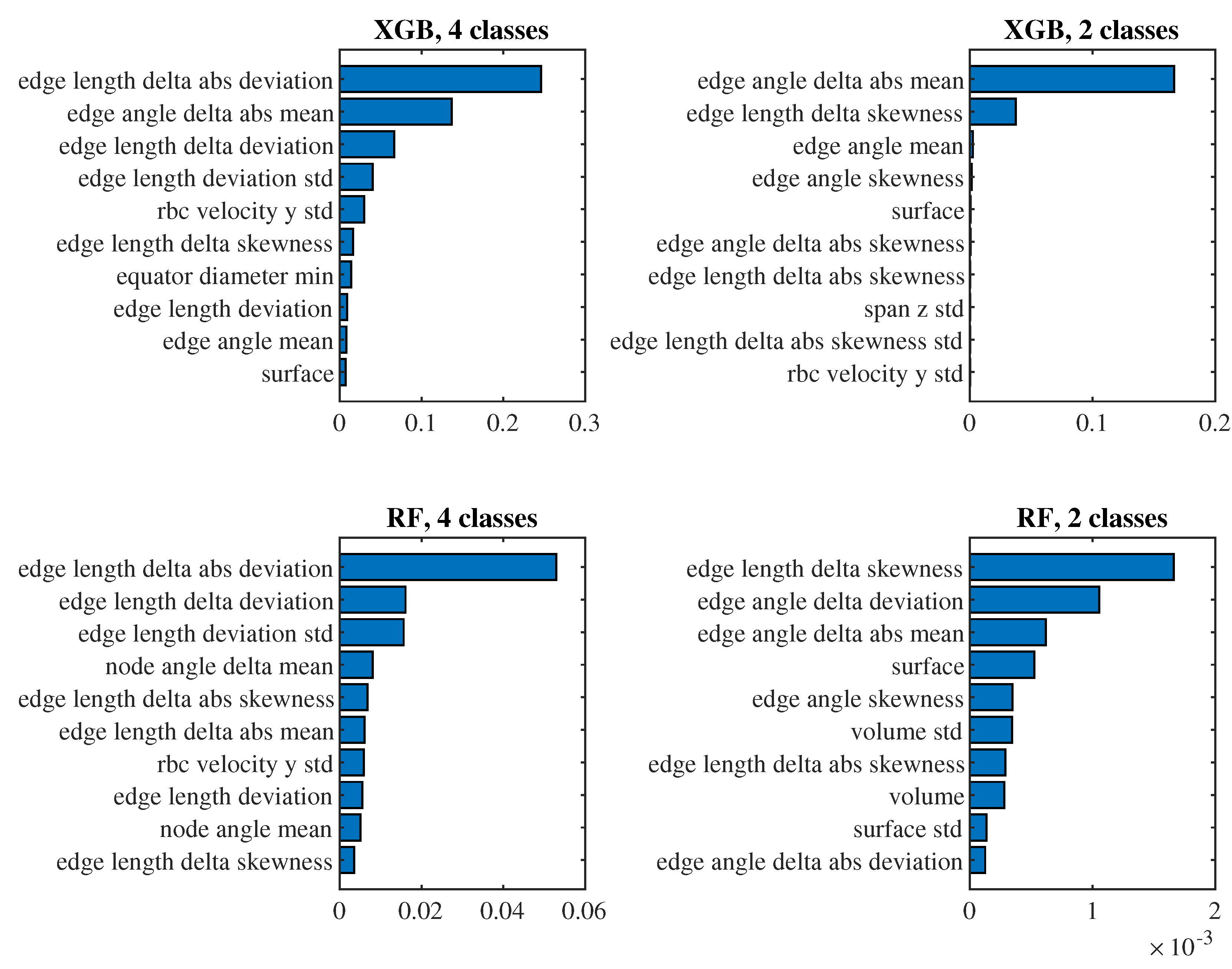
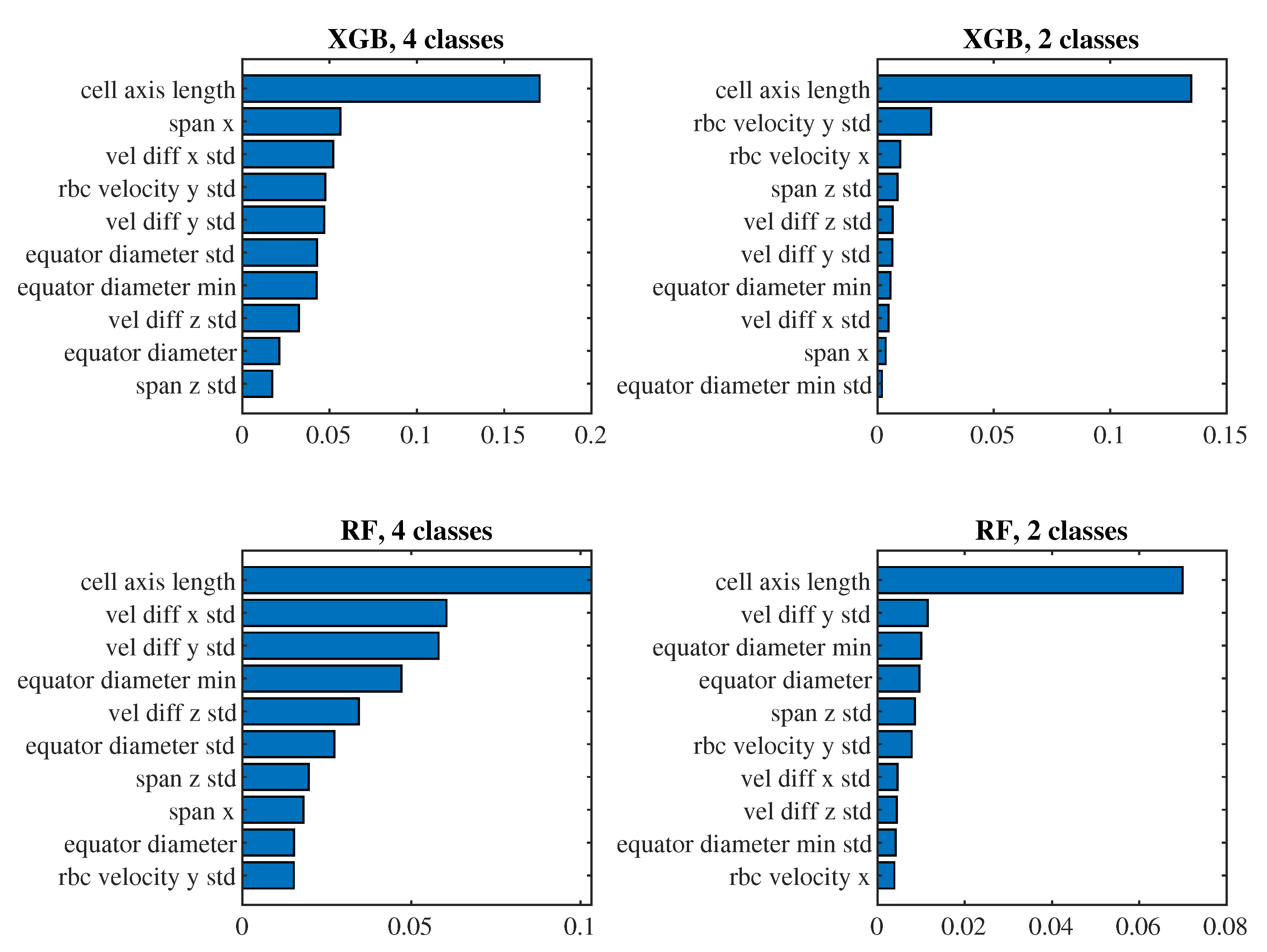
To see which predictors have the biggest impact on the classification, let us take a closer look at the models for
(i.e., those where we observe approximately one passage of a cell through the channel) with the 6th set of predictors and the 4th set of predictors. For these models in
Figure 10 and
Figure 11 we present the significance of the predictors calculated by the
permutation_importance method from the
sklearn library. In the case of the 6th set and classification into 4 classes, the "edge length delta abs deviation" predictor appears to be the most significant in both models, that is, the predictor indicating the standard deviation of the absolute changes in the lengths of the edges of the triangulation against the relaxed state of the cell. When classifying into 2 classes, "edge angle delta abs mean" and "edge length delta skewness" are significant for both models, that is, the average absolute change in the angles at the edges of the triangulation compared to the relaxed state and the coefficient of skewness of the changes in the lengths of the edges of the triangulation compared to the relaxed state. For the simpler 4th set of predictors, the most significant predictor for all models is the mean of the cell axis length.
5.5. Discussion
Our experiments with decision trees reveal insights into the efficiency of various predictor sets and the impact of recorded sequence length on the classification accuracy. Our results suggest that increasing the length of the sequence (S) generally improves classification accuracy up to a certain point, after which the improvements starts to be negligible. This indicates that while longer observation periods can be beneficial, excessively long monitoring may not yield proportional gains in accuracy. Specifically, monitoring cells for approximately one complete passage through the channel appears sufficient for optimal performance.
The comparison between deep neural networks and decision tree-based models reveals that the latter can achieve comparable or better accuracy with simpler predictor sets. This is particularly evident in the classification into four classes, where decision trees outperformed neural networks using the 2nd predictor set. This confirms that decision trees are a proper tool to effectively handle tabular data, providing a more interpretable and efficient alternative to neural networks.
Future research can focus on several directions to further improve the classification of RBC elasticity. One potential way is the integration of various data sources, such as combining video recordings with additional sensors capturing mechanical properties or biochemical markers of RBCs. This could provide a richer dataset and better classification accuracy. Performing experimental validation using actual RBC samples will be essential to confirm the theoretical findings and translate them into practical diagnostic tools.