Submitted:
24 July 2024
Posted:
25 July 2024
You are already at the latest version
Abstract
Keywords:
Introduction
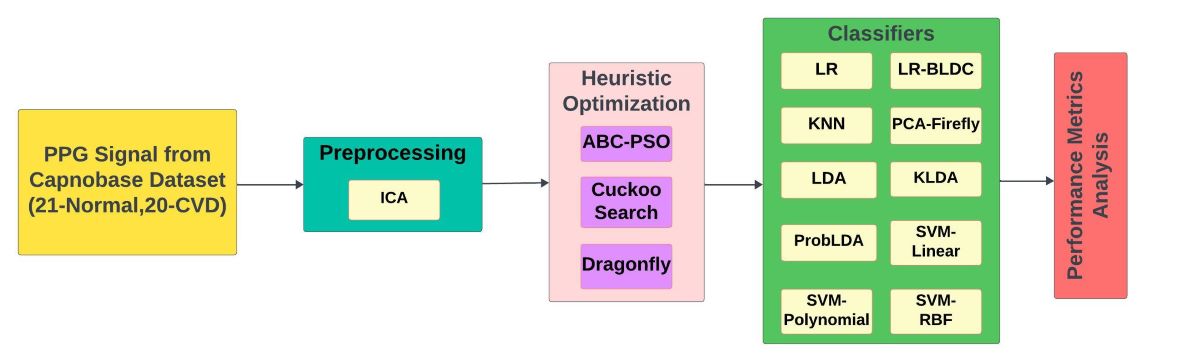
- The study proposes an early detection and intervention method for cardiovascular diseases using PPG signals.
- Three metaheuristic optimization algorithms are used as DR techniques to reduce the dimension of the high-dimensional PPG data.
- The dimensionality-reduced PPG data was then analyzed using ten different classification algorithms to detect the presence of CVD. The classifiers' performance is evaluated using parameters such as accuracy, GDR, MCC, Kappa, error rate, F1 score, and Jaccard index.
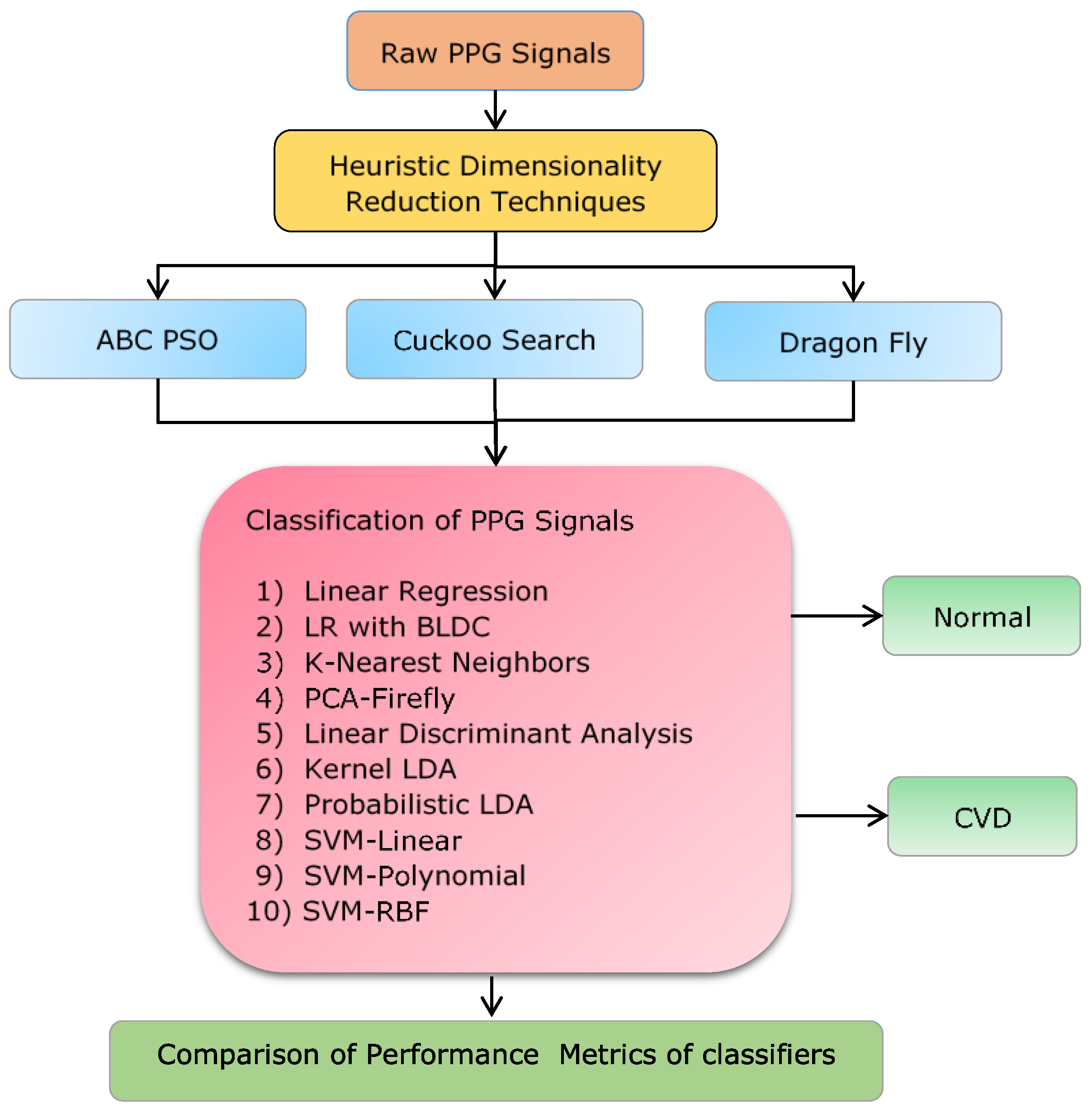
2. Materials and Methods
3. Dimensionality Reduction Techniques:
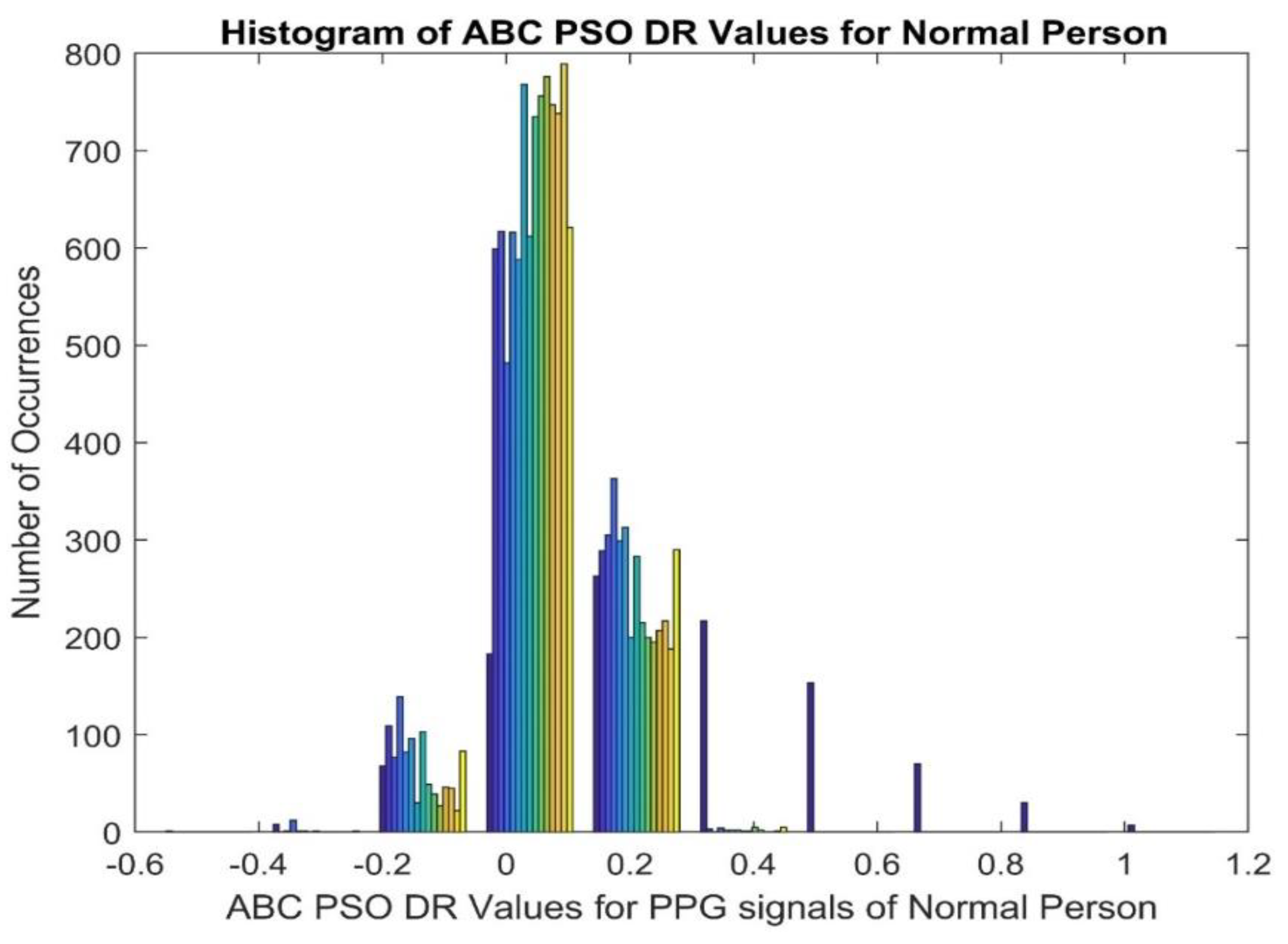
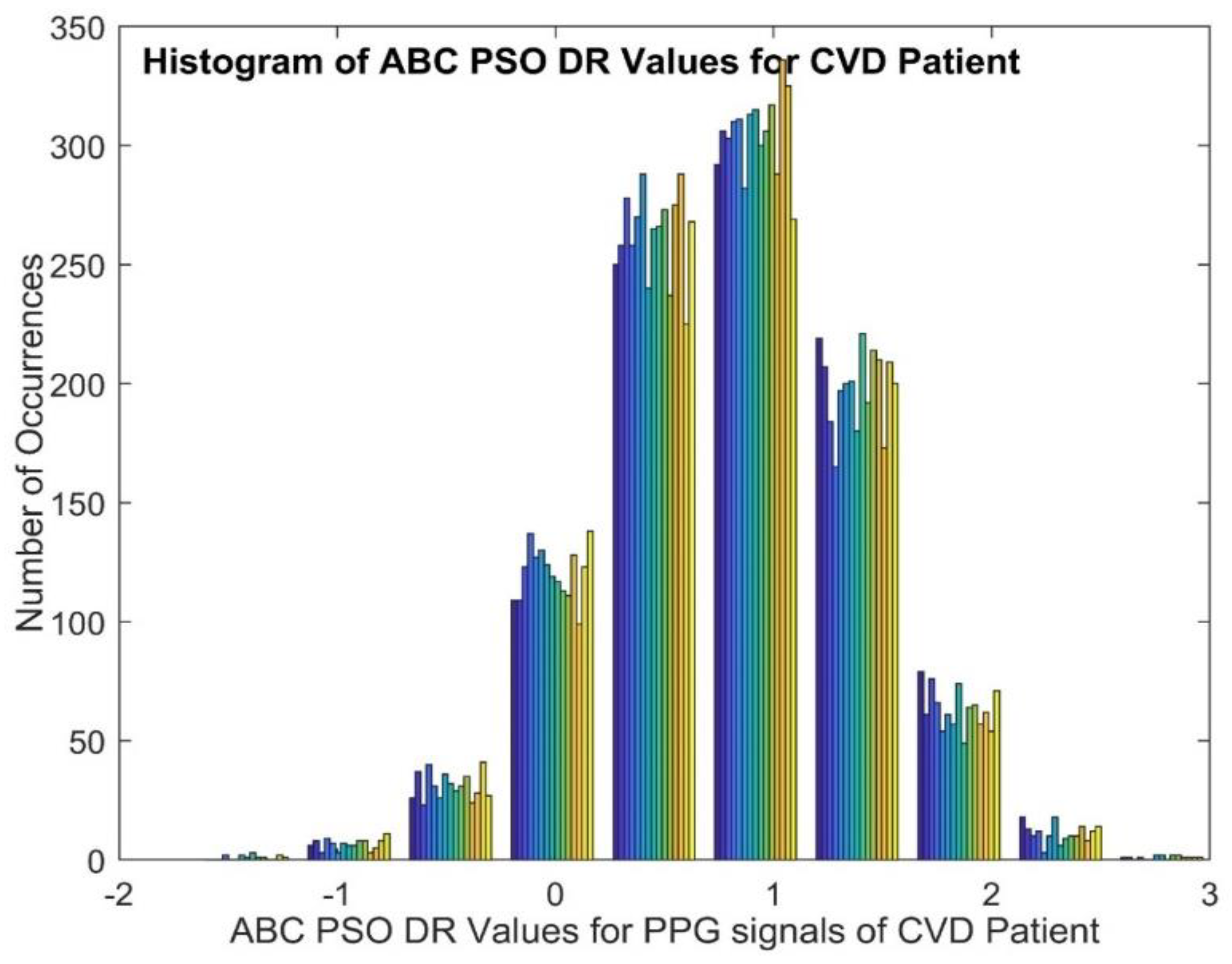
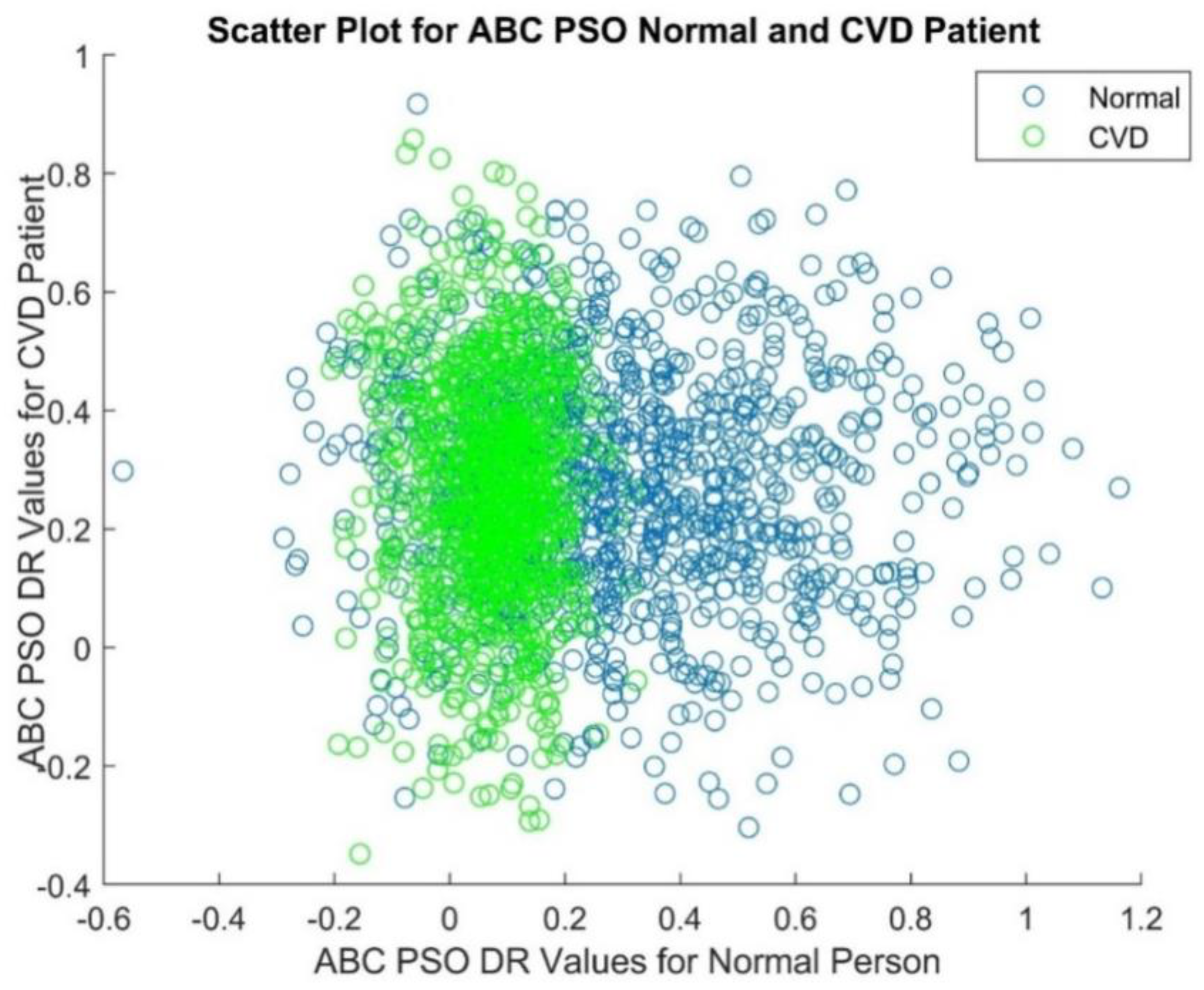
3.1. ABC-PSO (Artificial Bee Colony-Particle Swarm Optimization)
- Initialization: Initialize bee and particle populations with random solutions. Set the number of employed bees, onlooker bees, and scout bees, as well as particle positions and velocities
- Employed Bee Phase: Each employed bee explores new food sources (solutions) using:where is a random number and is a neighboring solution.
- Onlooker Bee Phase: Onlooker bees in the algorithm choose their food sources probabilistically:where is the fitness of solution
- Scout Bee Phase: Abandon poor solutions and have scout bees search for new random solutions.
- PSO Update: Update particle velocities and positions using:where is the personal best position, is the global best position, is the inertia weight, and are random numbers, and are acceleration coefficients.
- Evaluation and Selection: Evaluate new solutions and select the best ones based on fitness.
- Convergence Check: Repeat the steps until stopping criteria, such as maximum iterations or a convergence threshold are met.
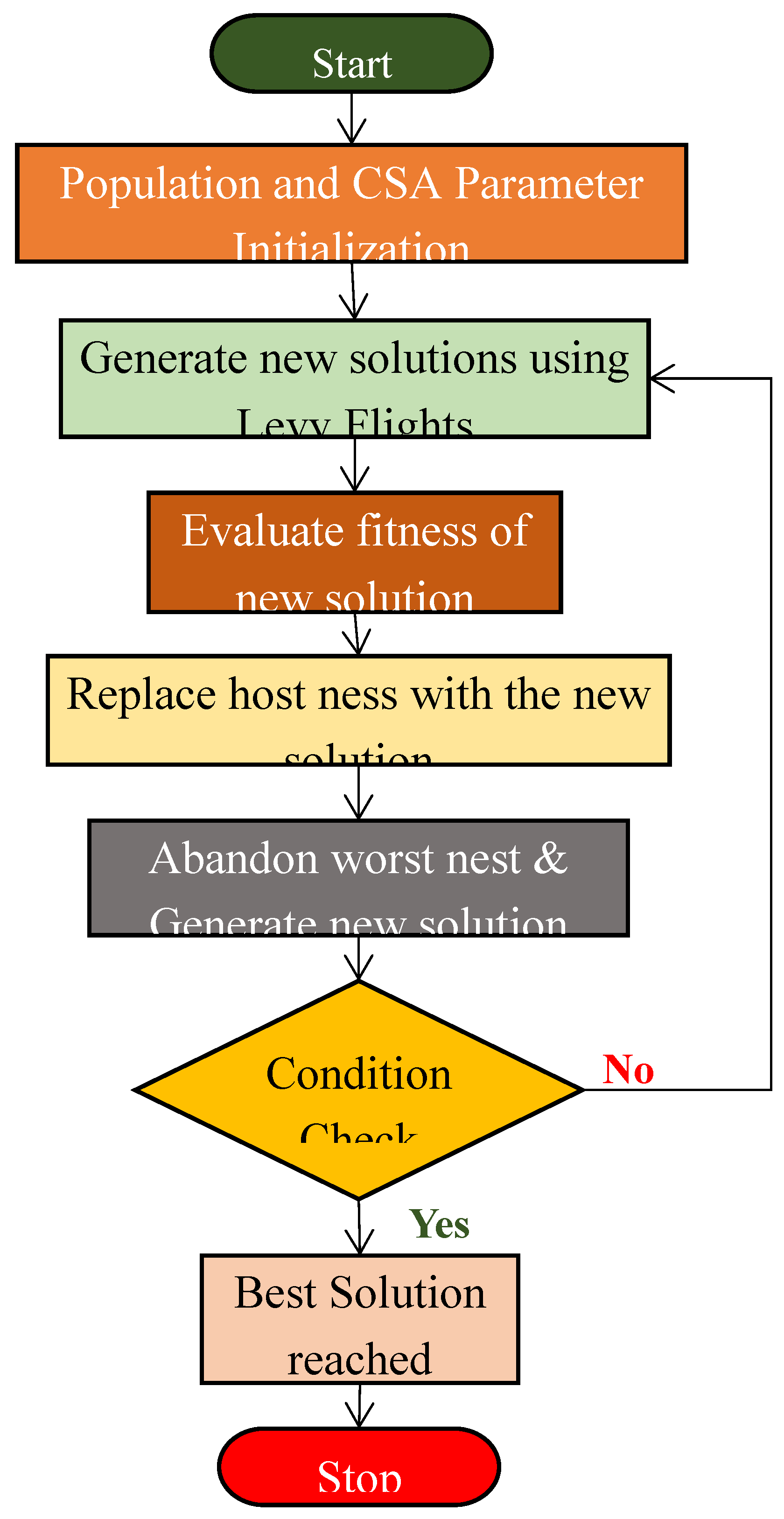
3.2. Cuckoo Search Algorithm (CSA)
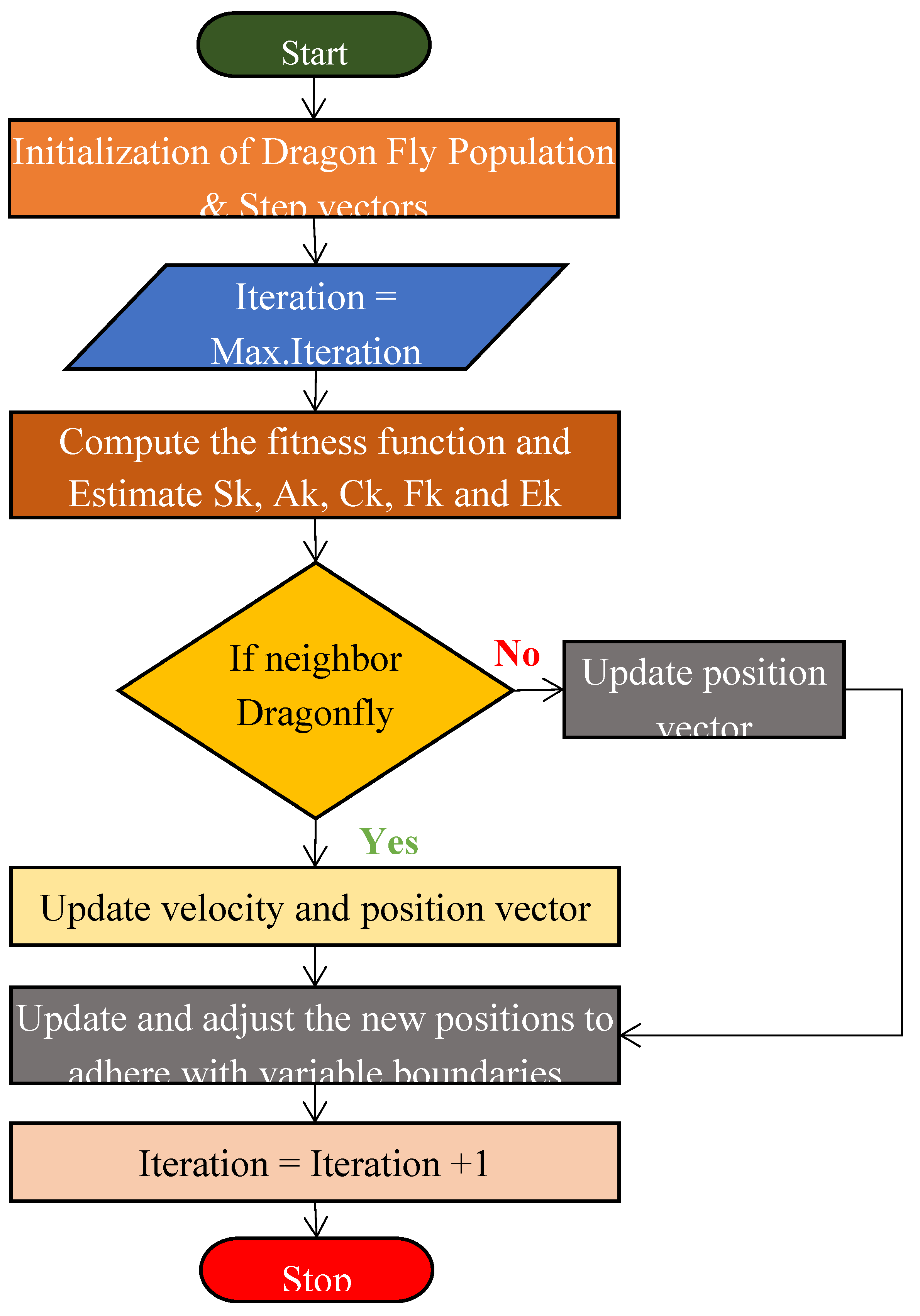
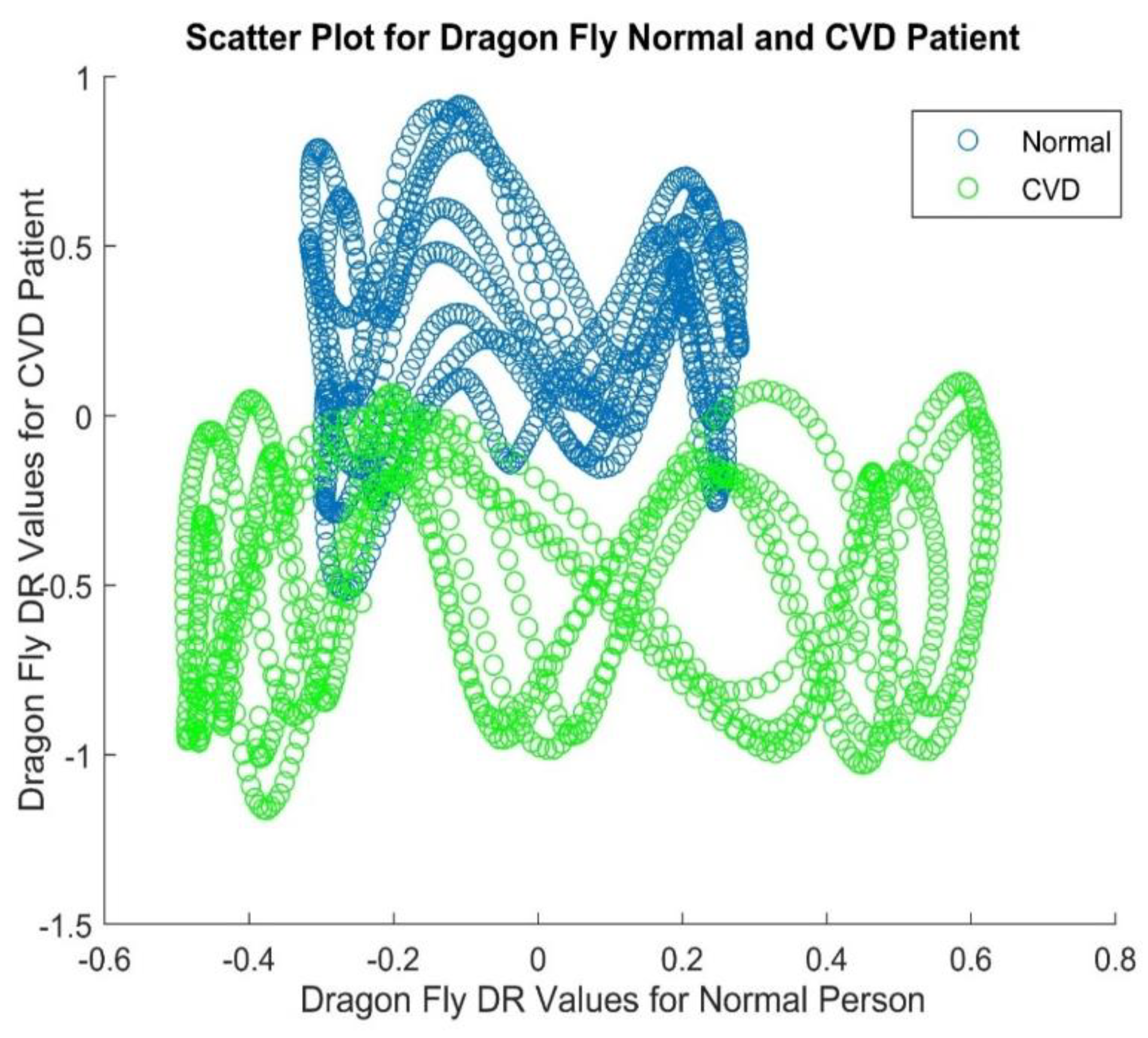
3.3. Dragonfly Algorithm
4. Classifiers for Classification of CVD from Dimensionality Reduced Values
4.1. Linear Regression as a Classifier
4.2. Linear Regression with BLDC
4.3. K-Nearest Neighbor as a Classifier
4.4. PCA-Firefly
4.5. Linear Discriminant Analysis as a Classifier
- Calculate the mean vector for each class
- Calculate the within-class scatter matrix
- Calculate the between-class scatter matrixWhere is the overall mean of the data set.
- Solve the generalized eigenvalue problem
4.6. Kernel LDA as a Classifier
4.7. Probabilistic LDA as a Classifier
4.8. Support Vector Machine as a Classifier
5. Results and Discussion
5.1. Optimal Parameters Selection for Classifiers
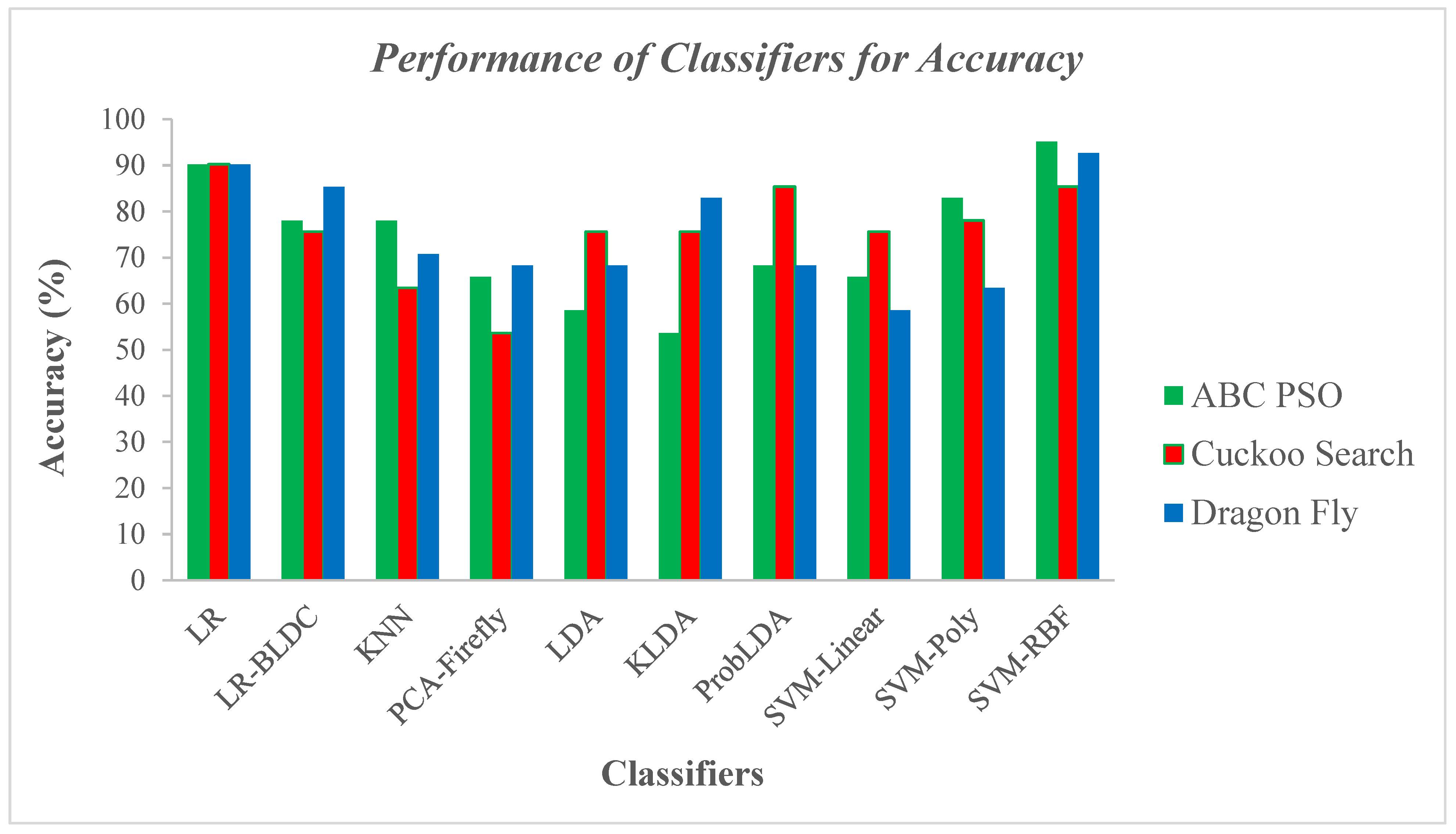
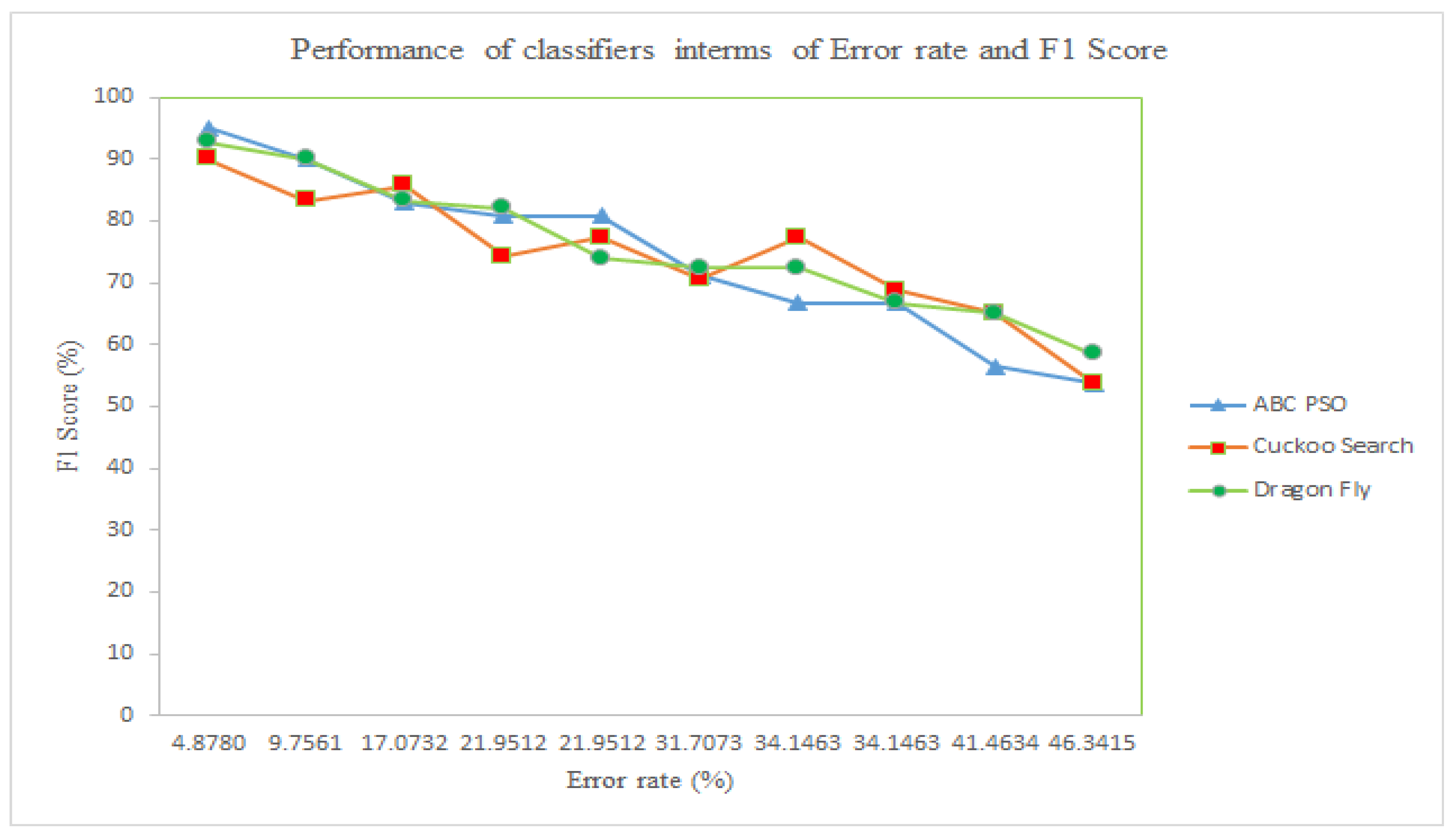
5.2. Performance Analysis of the Classifier
5.3. Analysis of the Computational Complexity of Classifiers
6. Conclusion
References
- Centers for Disease Control and Prevention. The State of Aging and Health in America 2013; Centers for Disease Control and Prevention, US Department of Health and Human Services: Atlanta, GA, USA, 2013. [Google Scholar]
- Kranjec, J, Beguš, S, Geršak, G & Drnovšek, J 2014, ‘Non-contact heart rate and heart rate variability measurements: A review’, Biomedical signal processing and control, vol.13, pp.102-112.
- Miranda, E, Irwansyah, E, Amelga, AY, Maribondang, MM & Salim, M 2016, ‘Detection of cardiovascular disease risk's level for adults using naive Bayes classifier’, Healthcare informatics research, vol.22, no.3, pp.196.
- Majumder S., Mondal T., and Deen M. J., Wearable sensors for remote health monitoring, Sensors. (2017) 17, no. 12, https://doi.org/10.3390/s17010130, 2-s2.0-85009999439. [CrossRef]
- Manogaran G., Shakeel P. M., Fouad H., Nam Y., Baskar S., Chilamkurti N., and Sundarasekar R., Wearable IoT smart-log patch: an edge computing-based bayesian deep learning network system for multi access physical monitoring system, Sensors. (2019) 19, no. 13, https://doi.org/10.3390/s19133030, 2-s2.0-85070464464. [CrossRef]
- Norgeot B., Glicksberg B. S., and Butte A. J., A call for deep-learning healthcare, Nature Medicine. (2019) 25, no. 1, 14–15. [CrossRef]
- Al-Zaben A., Fora M., and Obaidat A., Detection of premature ventricular beats from arterial blood pressure signal, Proceedings of the 2018 IEEE 4th Middle East Conference on Biomedical Engineering (MECBME), 28-30 March 2018, Tunis, Tunisia, IEEE, 17–19.
- Hindia M. N., Rahman T. A., Ojukwu H., Hanafi E. B., and Fattouh A., Enabling remote health-caring utilizing iot concept over LTE-femtocell networks, PLoS One. (2016) 11, no. 5, e0155077, https://doi.org/10.1371/journal.pone.0155077, 2-s2.0-84968677659. [CrossRef]
- Savkar A., Khatate P., and Patil C., Study on techniques involved in tourniqueteless blood pressure measurement using PPG, Proceedings of the 2018 Second International Conference on Intelligent Computing and Control Systems (ICICCS), 14-15 June 2018, Madurai, India, IEEE, 170–172.
- Elgendi M., Fletcher R., Liang Y., Howard N., Lovell N. H., Abbott D., Lim K., and Ward R., The use of photoplethysmography for assessing hypertension, NPJ digital medicine. (2019) 2, no. 1, https://doi.org/10.1038/s41746-019-0136-7. [CrossRef]
- Allen J., Photoplethysmography and its application in clinical physiological measurement, Physiological Measurement. (2007) 28, no. 3, R1–R39, https://doi.org/10.1088/0967-3334/28/3/r01, 2-s2.0-34247467602. [CrossRef]
- Gu-Young J., Yu K.-H., and Nam-Gyun K., Continuous blood pressure monitoring using pulse wave transit time, Proceedings of the ICCAS 2005 International conference on Control, June 2-5, Gyeonggi-Do, Korea, Automation and systems, 834–837.
- Shin, H.; Min, S. Feasibility study for the non-invasive blood pressure estimation based on ppg morphology: normotensive subject study. Biomed. Eng. Online 2017, 16, 10. [Google Scholar] [CrossRef] [PubMed]
- J.L. Moraes, M.X. Rocha, G.G.Vasconcelos, J.E. Vasconcelos Filho, de Albuquerque V.H.C. de Albuquerque, A.R. Alexandria, Advances in photopletysmography signal analysis for biomedical applications, Sensors 18(6) (2018) 1–26. [CrossRef]
- Shintomi, A.; Izumi, S.; Yoshimoto, M.; Kawaguchi, H. Effectiveness of the heartbeat interval error and compensation method on heart rate variability analysis. Healthc. Technol. Lett. 2022, 9, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Rubins, U. Finger and ear photoplethysmogram waveform analysis by fitting with Gaussians. Med Biol. Eng. 2008, 46, 1271–1276. [Google Scholar] [CrossRef] [PubMed]
- Moshawrab, M.; Adda, M.; Bouzouane, A.; Ibrahim, H.; Raad, A. Smart Wearables for the Detection of Cardiovascular Diseases: A Systematic Literature Review. Sensors 2023, 23, 828. [Google Scholar] [CrossRef] [PubMed]
- Reisner, A., Shaltis, P.A., McCombie, D., Asada, H.H. (2008). Utility of the photoplethysmogram in circulatory monitoring. Anesthesiology, 108 (5), 950- 958.
- Neha; Kanawade, R.; Tewary, S.; Sardana, H.K. Photoplethysmography based arrhythmia detection and classification. In Proceedings of the 6th International Conference on Signal Processing and Integrated Networks (SPIN), Noida, India, 7–8 March 2019; pp. 944–948. [CrossRef]
- Paradkar, N.; Chowdhury, S.R. Coronary artery disease detection using photoplethysmography. In Proceedings of the 39th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Jeju, Republic of Korea, 11–15 July 2017; pp. 100–103. [Google Scholar] [CrossRef]
- Prabhakar, S.K.; Rajaguru, H.; Lee, S.W. Metaheuristic-based dimensionality reduction and classification analysis of PPG signals for interpreting cardiovascular disease. IEEE Access 2019, 7, 165181–165206. [Google Scholar] [CrossRef]
- Palanisamy, Sivamani, and Harikumar Rajaguru. "Machine learning techniques for the performance enhancement of multiple classifiers in the detection of cardiovascular disease from PPG signals." Bioengineering 10.6 (2023): 678.
- Rajaguru, H.; Shankar, M.G.; Nanthakumar, S.P.; Murugan, I.A. Performance analysis of classifiers in detection of CVD using PPG signals. In AIP Conference Proceedings; AIP Publishing LLC: Melville, NY, USA, 2023; Volume 2725, p. 020002. [Google Scholar] [CrossRef]
- Al Fahoum, A.S.; Abu Al-Haija, A.O.; Alshraideh, H.A. Identification of Coronary Artery Diseases Using Photoplethysmography Signals and Practical Feature Selection Process. Bioengineering 2023, 10, 249. [Google Scholar] [CrossRef] [PubMed]
- Prabhakar, S.K.; Rajaguru, H.; Kim, S.H. Fuzzy-inspired photoplethysmography signal classification with bioinspired optimization for analyzing cardiovascular disorders. Diagnostics 2020, 10, 763. [Google Scholar] [CrossRef] [PubMed]
- Ihsan, M.F.; Mandala, S.; Pramudyo, M. Study of Feature Extraction Algorithms on Photoplethysmography (PPG) Signals to Detect Coronary Heart Disease. In Proceedings of the International Conference on Data Science and Its Applications (ICoDSA), Bandung, Indonesia, 6–7 July 2022; pp. 300–304. [Google Scholar] [CrossRef]
- Liu, Z.; Zhou, B.; Jiang, Z.; Chen, X.; Li, Y.; Tang, M.; Miao, F. Multiclass Arrhythmia Detection and Classification from Photoplethysmography Signals Using a Deep Convolutional Neural Network. J. Am. Heart Assoc. 2022, 11, e023555. [Google Scholar] [CrossRef] [PubMed]
- Limon, M. F. A., Shiblee, M. F. H., Rouf, A., & Iqbal, M. S. (2023, December). Hybrid ABC-PSO Algorithm based Static Synchronous Compensator for Enhancing Power System Stability. In 2023 10th IEEE International Conference on Power Systems (ICPS) (pp. 1-6). IEEE.
- Yang, Xin-She, and Suash Deb. "Cuckoo search via Lévy flights." In 2009 World congress on nature & biologically inspired computing (NaBIC), pp. 210-214. Ieee, 2009.
- Mirjalili, S. Dragonfly algorithm: A new meta-heuristic optimization technique for solving single-objective, discrete, and multi-objective problems. Neural Comput. Appl. 2016, 27, 1053–1073. [Google Scholar] [CrossRef]
- Rahman, C. M., & Rashid, T. A. (2019). Dragonfly algorithm and its applications in applied science survey. Computational Intelligence and Neuroscience, 2019(1), 9293617.
- Hasan, M. A., Hasan, M. K., & Mottalib, M. A. (2015). Linear regression–based feature selection for microarray data classification. International journal of data mining and bioinformatics, 11(2), 167-179.
- Zhou W, Liu Y, Yuan Q, Li X,2013, ‘Epileptic seizure detection using lacunarity and Bayesian linear discriminant analysis in intracranial EEG. IEEE Transactions on Biomedical Engineering, vol. 60, no. 12, pp. 3375-3381.
- Pandey, A., & Jain, A. (2017). Comparative analysis of KNN algorithm using various normalization techniques. International Journal of Computer Network and Information Security, 10(11), 36. Pandey, A., & Jain, A. (2017). Comparative analysis of KNN algorithm using various normalization techniques. International Journal of Computer Network and Information Security, 10(11), 36.
- Granato, D, Santos, JS,Escher, GB, Ferreira, BL & Maggio, RM 2018,‘Use of principal component analysis (PCA) and hierarchical cluster analysis (HCA) for multivariate association between bioactive compounds and functional properties in foods: A critical perspective’, Trends in Food Science & Technology,vol. 72, pp. 83-90.
- Moazenzadeh, R, Mohammadi, B, Shamshirband, S & Chau, KW 2018, ‘Coupling a firefly algorithm with support vector regression to predict evaporation in northern Iran’, Engineering Applications of Computational Fluid Mechanics, vol. 12, no. 1, pp. 584-597.
- Zhao, H., Lai, Z., Leung, H., Zhang, X. (2020). Linear Discriminant Analysis. In: Feature Learning and Understanding. Information Fusion and Data Science. Springer, Cham. [CrossRef]
- Li, S., Zhang, H., Ma, R., Zhou, J., Wen, J., & Zhang, B. (2023). Linear discriminant analysis with generalized kernel constraint for robust image classification. Pattern Recognition, 136, 109196.
- Ioffe, S. (2006). Probabilistic Linear Discriminant Analysis. In: Leonardis, A., Bischof, H., Pinz, A. (eds) Computer Vision – ECCV 2006. ECCV 2006. Lecture Notes in Computer Science, vol 3954. Springer, Berlin, Heidelberg. [CrossRef]
- Srunitha, K & Padmavathi, S 2016, ‘Performance of SVM classifier for image based soil classification’, In 2016 International Conference on Signal Processing, Communication, Power and Embedded System (SCOPES), pp. 411-415, IEEE.
- Ramamoorthy, K.; Rajaguru, H. Exploitation of Bio-Inspired Classifiers for Performance Enhancement in Liver Cirrhosis Detection from Ultrasonic Images. Biomimetics 2024, 9, 356. [Google Scholar] [CrossRef] [PubMed]
- Hosseini, Z.S.; Zahedi, E.; Attar, H.M.; Fakhrzadeh, H.; Parsafar, M.H. Discrimination between different degrees of coronary artery disease using time-domain features of the finger photoplethysmogram in response to reactive hyperemia. Biomed. Signal Process. Control 2015, 18, 282–292. [Google Scholar] [CrossRef]
- Miao, K.H.; Miao, J.H. Coronary heart disease diagnosis using deep neural networks. Int. J. Adv. Comput. Sci. Appl. 2018, 9, 1–8. [Google Scholar] [CrossRef]
- Shobitha, S.; Sandhya, R.; Ali, M.A. Recognizing cardiovascular risk from photoplethysmogram signals using ELM. In Pro-ceedings of the Second International Conference on Cognitive Computing and Information Processing (CCIP), Mysore, In-dia, 12–13 August 2016; pp. 1–5. [CrossRef]
- Soltane, M.; Ismail, M.; Rashid, Z.A. Artificial Neural Networks (ANN) approach to PPG signal classification. Int. J. Comput. Inf. Sci. 2004, 2, 58–65. [Google Scholar]
| Parameters | Heuristic Algorithms | ||
| ABC-PSO | CSA | DFA | |
| Population Size | 200 | 200 | 200 |
| Control parameters | Inertia weight :0.45 Acceleration coefficients and |
Probability =0.4 Step Size α=1.5 |
Separation alignment cohesion Attraction Distraction |
| Algorithm | Swarm intelligence With Hybrid | Levy flight | Swarm intelligence |
| Stopping Criteria | Training MSE of 10-5 | Training MSE of 10-5 | Training MSE of 10-5 |
| Number of iteration | 200 | 200 | 200 |
| Local Minima Problem | Available in ABC. With proper selection of and in the PSO algorithm through trial and error method. The local minima problem will be solved. | No local minima problem | No local minima problem |
| Over fitting | Over fitting is available due to α and β values of ABC. This can be overcome with the proper selection of Weight (w) of PSO Algorithm | Over fitting is not presented | Over fitting is not presented |
|
Dimensionality Reduction Techniques |
Category | Statistical Metrics | ||||||
|---|---|---|---|---|---|---|---|---|
| Mean | Variance | Skewness | Kurtosis | PCC | Sample Entropy | CCA | ||
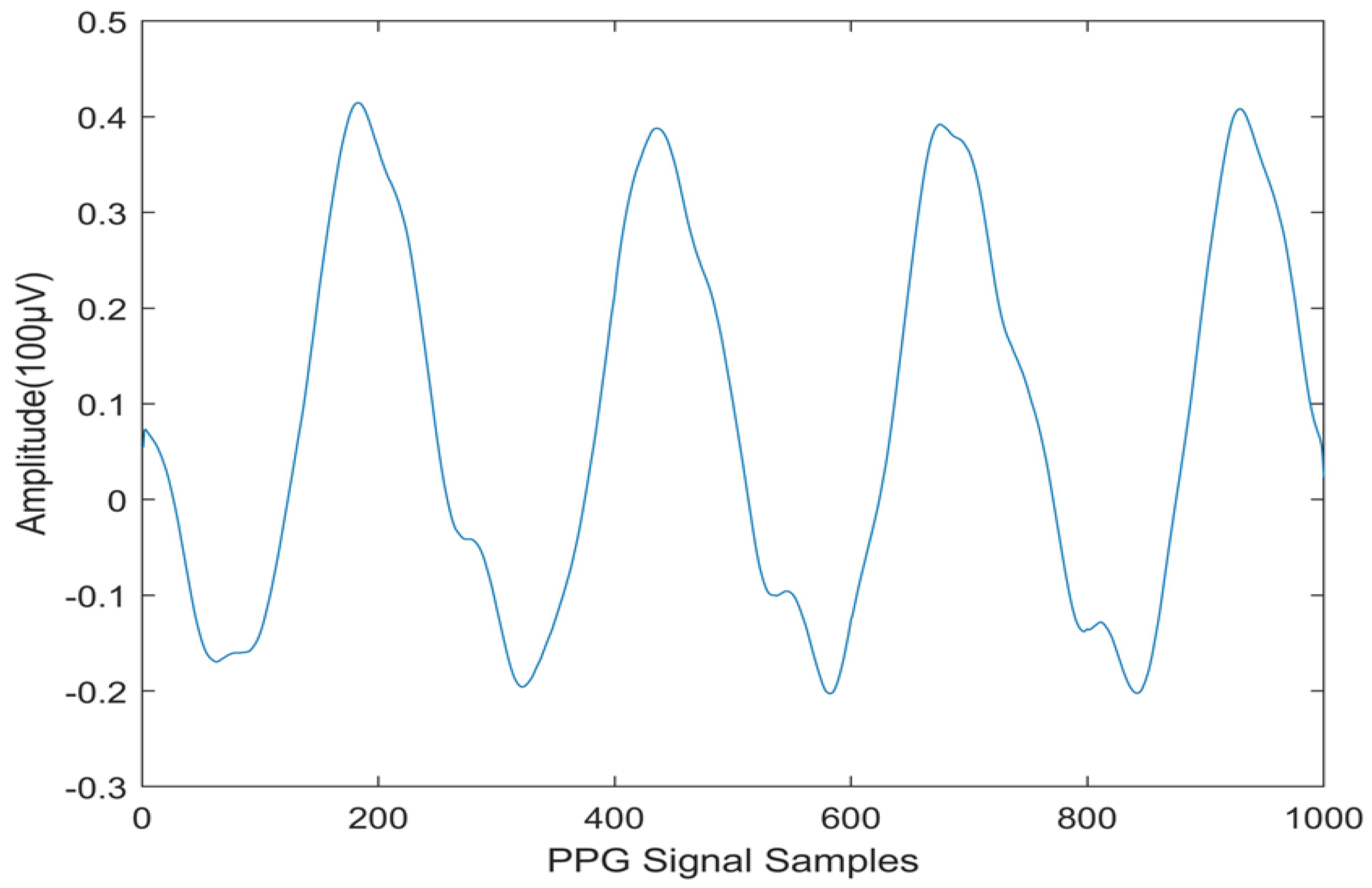
| Normal | 0.0732 | 0.0063 | -0.1165 | 0.2713 | -0.0597 | 9.9494 | 0.1066 | |
| CVD | 0.7872 | 0.3353 | -0.1000 | 0.1435 | 0.0133 | 9.9473 | ||
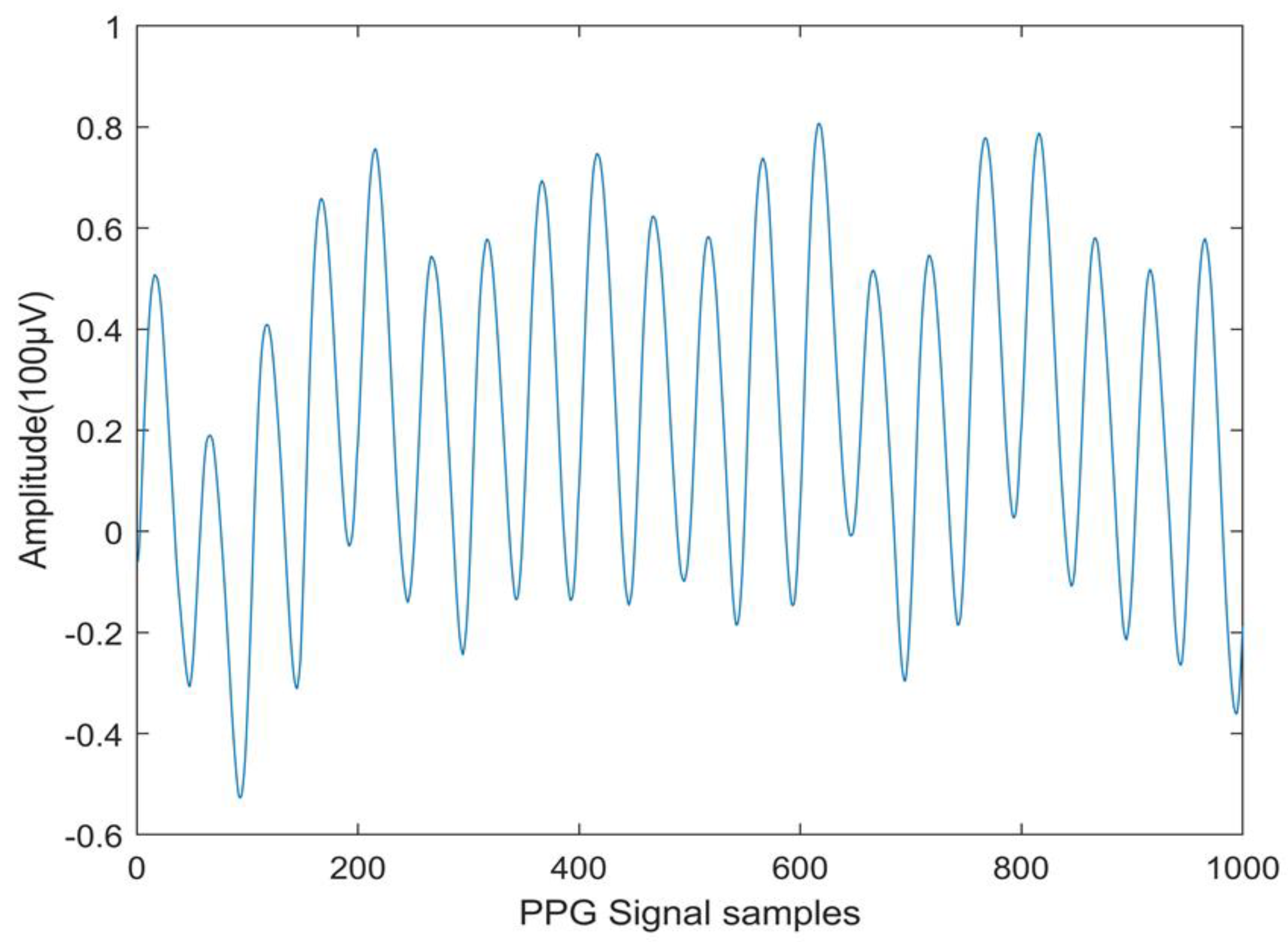
| Cuckoo search | Normal | 0.5236 | 0.0475 | 0.1575 | -0.4556 | 0.3393 | 9.9494 | 0.3674 |
| CVD | 7.8931 | 34.5391 | -0.0901 | -1.7290 | 0.2294 | 4.9919 | ||
| Dragon Fly | Normal | -1.5850 | 378.4756 | -0.0243 | -0.9585 | -0.2145 | 9.9499 | 0.4621 |
| CVD | -3.4728 | 271.9735 | 0.0381 | -0.6919 | 0.1044 | 9.9522 | ||
| Classifiers | ABC PSO | Cuckoo Search | Dragon fly | |||
|---|---|---|---|---|---|---|
| Training MSE | Testing MSE | Training MSE | Testing MSE | Training MSE | Testing MSE | |
| Linear Regression | 3.52E-09 | 2.92E-07 | 5.69E-09 | 1.37E-08 | 5.99E-09 | 1.44E-06 |
| Linear Regression with BDLC | 2.32E-06 | 1.10E-04 | 9.69E-08 | 8.65E-06 | 6.03E-08 | 2.72E-06 |
| KNN (weighted) | 5.72E-08 | 1.44E-06 | 4.60E-06 | 2.81E-03 | 7.02E-08 | 3.24E-06 |
| PCA firefly | 6.65E-07 | 3.80E-05 | 8.45E-06 | 6.08E-03 | 8.69E-06 | 6.25E-05 |
| LDA | 6.69E-05 | 5.48E-03 | 5.05E-06 | 1.44E-05 | 5.54E-06 | 2.70E-05 |
| KLDA | 7.34E-06 | 4.84E-03 | 4.63E-08 | 1.69E-06 | 6.63E-06 | 1.22E-05 |
| ProbLDA | 5.83E-06 | 3.06E-05 | 7.97E-08 | 6.76E-06 | 5.99E-07 | 1.68E-05 |
| SVM (Linear) | 4.05E-06 | 1.69E-04 | 4.85E-08 | 1.82E-06 | 7.89E-06 | 9.03E-03 |
| SVM (Polynomial) | 8.29E-08 | 6.76E-06 | 6.74E-08 | 1.44E-06 | 8.20E-07 | 7.29E-06 |
| SVM(RBF) | 1.92E-10 | 2.45E-09 | 5.38E-07 | 1.85E-05 | 2.45E-10 | 3.62E-09 |
| Classifiers | Optimal Parameters of the Classifiers |
|---|---|
| Linear Regression (LR) | Uniform weight w=0.451, bias:0.003, Criterion: MSE |
| LR with BLDC | The cascading configuration of LR with the following BLDC parameters: Class mean and , Prior probability P(x): 0.5 |
| K-Nearest Neighbors (KNN) | Number of clusters = 2 |
| PCA Firefly |
PCA: A threshold value of 0.72 and decorrelated Eigen vector , using a trial and error training approach Firefly: Initial conditions of = 0.65, = 0.1 For both PCA and firefly, consider MSE of or reaching a maximum of 1000 iterations, whichever comes earliest. Criterion: MSE |
| Linear Discriminant Analysis (LDA) | Weight w=0.56,bias:0.0018 |
| Kernel LDA (KLDA) | Number of clusters:2, w1:0.38, w2:0.642, bias:0.0026±0.0001 |
| Probabilistic LDA(ProbLDA) | Weight w=0.56,bias:0.0018,Assigned Probability>0.5 |
| SVM-Linear | Class weights:0.4 Parameter for Regularization [C]:0.85 Criteria for Convergence : MSE |
| SVM-Polynomial | Parameter for Regularization [C]:0.76 Class weights:0.5 Kernel Function Coefficient [Gamma]:10 Criteria for Convergence : MSE |
| SVM-RBF | Parameter for Regularization [C]:1 Class weights:0.86 Kernel Function Coefficient [Gamma]:100 Criteria for Convergence : MSE |
| DR Techniques |
Classifiers | Accuracy (%) |
GDR (%) | Error rate (%) | Kappa | MCC | F1 Score (%) |
JI (%) |
|---|---|---|---|---|---|---|---|---|
| ABC-PSO | Linear Regression | 90.24 | 89.74 | 9.76 | 0.80 | 0.80 | 90.00 | 81.82 |
| LR-BLDC | 78.05 | 72.73 | 21.95 | 0.56 | 0.60 | 80.85 | 67.86 | |
| K-Nearest Neighbors | 78.05 | 72.73 | 21.95 | 0.56 | 0.60 | 80.85 | 67.86 | |
| PCA Firefly | 65.85 | 57.58 | 34.15 | 0.32 | 0.32 | 66.67 | 50.00 | |
| Linear Discriminant Analysis | 58.54 | 48.48 | 41.46 | 0.17 | 0.17 | 56.41 | 39.29 | |
| Kernel LDA | 53.66 | 38.71 | 46.34 | 0.07 | 0.07 | 53.66 | 36.67 | |
| Probabilistic LDA | 68.29 | 59.38 | 31.71 | 0.37 | 0.38 | 71.11 | 55.17 | |
| SVM-Linear | 65.85 | 57.58 | 34.15 | 0.32 | 0.32 | 66.67 | 50.00 | |
| SVM-Polynomial | 82.93 | 81.08 | 17.07 | 0.66 | 0.66 | 82.93 | 70.83 | |
| SVM-RBF | 95.12 | 95.00 | 4.88 | 0.90 | 0.90 | 95.00 | 90.48 | |
| Cuckoo Search | Linear Regression | 90.24 | 89.74 | 9.76 | 0.80 | 0.80 | 90.00 | 81.82 |
| LR-BLDC | 75.61 | 70.59 | 24.39 | 0.51 | 0.52 | 77.27 | 62.96 | |
| K-Nearest Neighbors | 63.41 | 53.13 | 36.59 | 0.27 | 0.27 | 65.12 | 48.28 | |
| PCA Firefly | 53.66 | 38.71 | 46.34 | 0.07 | 0.07 | 53.66 | 36.67 | |
| Linear Discriminant Analysis | 75.61 | 74.36 | 24.39 | 0.51 | 0.53 | 70.59 | 54.55 | |
| Kernel LDA | 75.61 | 70.59 | 24.39 | 0.51 | 0.52 | 77.27 | 62.96 | |
| Probabilistic LDA | 85.37 | 85.00 | 14.63 | 0.71 | 0.72 | 83.33 | 71.43 | |
| SVM-Linear | 75.61 | 75.00 | 24.39 | 0.51 | 0.55 | 68.75 | 52.38 | |
| SVM-Polynomial | 78.05 | 76.92 | 21.95 | 0.56 | 0.58 | 74.29 | 59.09 | |
| SVM-RBF | 85.37 | 83.78 | 14.63 | 0.71 | 0.71 | 85.71 | 75.00 | |
| Dragon Fly | Linear Regression | 90.24 | 89.74 | 9.76 | 0.80 | 0.80 | 90.00 | 81.82 |
| LR-BLDC | 85.37 | 85.00 | 14.63 | 0.71 | 0.72 | 83.33 | 71.43 | |
| K-Nearest Neighbors | 70.73 | 62.50 | 29.27 | 0.42 | 0.44 | 73.91 | 58.62 | |
| PCA Firefly | 68.29 | 58.06 | 31.71 | 0.37 | 0.39 | 72.34 | 56.67 | |
| Linear Discriminant Analysis | 68.29 | 58.06 | 31.71 | 0.37 | 0.39 | 72.34 | 56.67 | |
| Kernel LDA | 82.93 | 81.58 | 17.07 | 0.66 | 0.66 | 82.05 | 69.57 | |
| Probabilistic LDA | 68.29 | 62.86 | 31.71 | 0.36 | 0.37 | 66.67 | 50.00 | |
| SVM-Linear | 58.54 | 46.88 | 41.46 | 0.17 | 0.17 | 58.54 | 41.38 | |
| SVM-Polynomial | 63.41 | 53.13 | 36.59 | 0.27 | 0.27 | 65.12 | 48.28 | |
| SVM-RBF | 92.68 | 92.50 | 7.32 | 0.85 | 0.85 | 92.68 | 86.36 |
| Classifiers | Heuristic dimensionality reduction techniques | ||
|---|---|---|---|
| ABC-PSO | CSA | DFA | |
| Linear Regression (LR) | |||
| LR with BLDC | |||
| K-Nearest Neighbors (KNN) | |||
| PCA Firefly | |||
| Linear Discriminant Analysis (LDA) | |||
| Kernel LDA (KLDA) | |||
| Probabilistic LDA(ProbLDA) | |||
| SVM-Linear | |||
| SVM-Polynomial | |||
| SVM-RBF | |||
| Sl.no | Authors | Dataset | Number of Subjects | Classifiers | Classes | Accuracy (%) |
|---|---|---|---|---|---|---|
| 1 | Rajaguru et al. [23] 2023 |
Capnobase dataset | Single patient | LR | CVD, Normal | 65.85% |
| 2 | Al Fahoum et al. [24] 2023 |
Internal medicine clinic of Princess Basma Hospital | 200 healthy and 160 with CVD | NB | Normal and abnormal | 89.37% |
| 3 | Prabhakar et al. [25] 2020 |
Capnobase dataset | 28 CVD 14 Normal |
SVM–RBF RBF NN |
CVD, Normal | 95.05% 94.79% |
| 4 | Liu et al. [27] 2022 |
GitHub https://github.com/zdzdliu/PPGArrhythmiaDetection |
45 Subjects | DCNN | CVD, Normal | 85% |
| 5 | Hosseini et al. [42] 2015 |
Tehran Heart Center | 18 Normal 30 CVD |
KNN | Low risk High risk |
81.5% |
| 6 | Miao and Miao [43] 2018 |
Cleveland Clinic Foundation | 303 patients | DNN | CVD, Normal | 83.67% |
| 7 | Shobita et al. [44] 2016 |
Biomedical Research Lab | 30 healthy 30 pathological | ELM | Healthy Risk of CVD |
89.33% |
| 8 | Soltane et al. [45] 2005 |
Seremban Hospital | 114 healthy 56 pathological | ANN | CVD, Normal | 94.70% |
| 9 | This research | Capnobase dataset | 21 Normal 20 CVD |
SVM-RBF | CVD, Normal | 95.12% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




