Submitted:
20 July 2024
Posted:
22 July 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Donkor ES, Muhsen K, Johnson SAM, et al. Multicenter Surveillance of Antimicrobial Resistance among Gram-Negative Bacteria Isolated from Bloodstream Infections in Ghana. Antibiotics (Basel). Jan 27 2023;12(2). [CrossRef]
- Sharma A, Singh A, Dar MA, et al. Menace of antimicrobial resistance in LMICs: Current surveillance practices and control measures to tackle hostility. Journal of Infection and Public Health. 2022/02/01/ 2022;15(2):172-181. [CrossRef]
- Gagliotti C, Balode A, Baquero F, et al. Escherichia coli and Staphylococcus aureus: bad news and good news from the European Antimicrobial Resistance Surveillance Network (EARS-Net, formerly EARSS), 2002 to 2009. Euro Surveill. Mar 17 2011;16(11). [CrossRef]
- Lee H, Yoon EJ, Kim D, et al. Antimicrobial resistance of major clinical pathogens in South Korea, May 2016 to April 2017: first one-year report from Kor-GLASS. Euro Surveill. Oct 2018;23(42). [CrossRef]
- Hattori H, Maeda M, Nagatomo Y, et al. Epidemiology and risk factors for mortality in bloodstream infections: A single-center retrospective study in Japan. Am J Infect Control. Dec 2018;46(12):e75-e79. [CrossRef]
- Musicha P, Cornick JE, Bar-Zeev N, et al. Trends in antimicrobial resistance in bloodstream infection isolates at a large urban hospital in Malawi (1998-2016): a surveillance study. Lancet Infect Dis. Oct 2017;17(10):1042-1052. [CrossRef]
- Tomislav Mestrovic GRA, Lucien R Swetschinski, et al. The burden of bacterial antimicrobial resistance in the WHO European region in 2019: a cross-country systematic analysis. Lancet Public Health. Nov 2022;7(11):e897-e913. [CrossRef]
- Kariuki S, Kering K, Wairimu C, Onsare R, Mbae C. Antimicrobial Resistance Rates and Surveillance in Sub-Saharan Africa: Where Are We Now? Infect Drug Resist. 2022;15:3589-3609. [CrossRef]
- Sutherland T, Mpirimbanyi C, Nziyomaze E, et al. Widespread antimicrobial resistance among bacterial infections in a Rwandan referral hospital. PLOS ONE. 2019;14(8):e0221121. [CrossRef]
- Zhou N, Cheng Z, Zhang X, et al. Global antimicrobial resistance: a system-wide comprehensive investigation using the Global One Health Index. Infectious Diseases of Poverty. 2022/08/23 2022;11(1):92. [CrossRef]
- Deku JG, Dakorah MP, Lokpo SY, et al. The Epidemiology of Bloodstream Infections and Antimicrobial Susceptibility Patterns: A Nine-Year Retrospective Study at St. Dominic Hospital, Akwatia, Ghana. J Trop Med. 2019;2019:6750864. [CrossRef]
- Habyarimana T, Murenzi D, Musoni E, Yadufashije C, F NN. Bacteriological Profile and Antimicrobial Susceptibility Patterns of Bloodstream Infection at Kigali University Teaching Hospital. Infect Drug Resist. 2021;14:699-707. [CrossRef]
- Kallel H, Houcke S, Resiere D, et al. Epidemiology and Prognosis of Intensive Care Unit-Acquired Bloodstream Infection. Am J Trop Med Hyg. Jul 2020;103(1):508-514. [CrossRef]
- Del Bono V, Giacobbe DR. Bloodstream infections in internal medicine. Virulence. 2016/04/02 2016;7(3):353-365. [CrossRef]
- Houssaini ZE, Harrar N, Zerouali K, Belabbes H, Elmdaghri N. [Prevalence of coagulase-negative staphylococci in blood cultures at the Ibn-Rochd University Hospital in Casablanca]. Pan Afr Med J. 2019;33:193. Prévalence des staphylocoques à coagulase négative dans les hémocultures au Centre Hospitalier Universitaire Ibn Rochd de Casablanca. [CrossRef]
- Kern WV, Rieg S. Burden of bacterial bloodstream infection-a brief update on epidemiology and significance of multidrug-resistant pathogens. Clin Microbiol Infect. Feb 2020;26(2):151-157. [CrossRef]
- The burden of bacterial antimicrobial resistance in the WHO European region in 2019: a cross-country systematic analysis. Lancet Public Health. Nov 2022;7(11):e897-e913. [CrossRef]
- Eibach D, Belmar Campos C, Krumkamp R, et al. Extended spectrum beta-lactamase producing Enterobacteriaceae causing bloodstream infections in rural Ghana, 2007-2012. Int J Med Microbiol. Jun 2016;306(4):249-54. [CrossRef]
- Kazmierczak KM, Karlowsky JA, de Jonge BLM, Stone GG, Sahm DF. Epidemiology of Carbapenem Resistance Determinants Identified in Meropenem-Nonsusceptible Enterobacterales Collected as Part of a Global Surveillance Program, 2012 to 2017. Antimicrob Agents Chemother. Jun 17 2021;65(7):e0200020. [CrossRef]
- Ssekatawa K, Byarugaba DK, Wampande E, Ejobi F. A systematic review: the current status of carbapenem resistance in East Africa. BMC Research Notes. 2018/08/31 2018;11(1):629. [CrossRef]
- Jousset AB, Bernabeu S, Bonnin RA, et al. Development and validation of a multiplex polymerase chain reaction assay for detection of the five families of plasmid-encoded colistin resistance. Int J Antimicrob Agents. Mar 2019;53(3):302-309. [CrossRef]
- Leshaba TMS, Mbelle NM, Osei Sekyere J. Current and emerging polymyxin resistance diagnostics: A systematic review of established and novel detection methods. J Appl Microbiol. Jan 2022;132(1):8-30. [CrossRef]
- Abebe W, Tegene B, Feleke T, Sharew B. Bacterial Bloodstream Infections and their Antimicrobial Susceptibility Patterns in Children and Adults in Ethiopia: a 6-Year Retrospective Study. Clin Lab. Nov 1 2021;67(11). [CrossRef]
- Baker S, Thomson N, Weill FX, Holt KE. Genomic insights into the emergence and spread of antimicrobial-resistant bacterial pathogens. Science. May 18 2018;360(6390):733-738. [CrossRef]
- Sartorius, B., Gray, A.P., Weaver, N.D., Aguilar, G.R., Swetschinski, L.R., Ikuta, K.S., Mestrovic, T., Chung, E., Wool, E.E., Han, C. and Hayoon, A.G., 2024. The burden of bacterial antimicrobial resistance in the WHO African region in 2019: a cross-country systematic analysis. The Lancet Global Health, 12(2), pp.e201-e216.
- Remera, E., Rwagasore, E., Muvunyi, C.M. and Ahmed, A., Emergence of the first molecularly-confirmed outbreak of Rift Valley fever among humans in Rwanda, calls for institutionalizing One Health Strategy. IJID One Health.
- Muvunyi, C.M., Ngabonziza, J.C.S., Florence, M., Mukagatare, I., Twagirumukiza, M., Ahmed, A. and Siddig, E.E., 2024. Uncovering the Diversity and Distribution of Fungal Infections in Rwanda: Assessing Risks and Documenting Knowledge and Policy Gaps.
- Fasina, F.O., Fasanmi, O.G., Makonnen, Y.J., Bebay, C., Bett, B. and Roesel, K., 2021. The one health landscape in Sub-Saharan African countries. One Health, 13, p.100325.
- Langfeldt, A., Gold, J.A. and Chiller, T., 2022. Emerging fungal infections: from the fields to the clinic, resistant Aspergillus fumigatus and dermatophyte species: a one health perspective on an urgent public health problem. Current Clinical Microbiology Reports, 9(4), pp.46-51.
- Siddig, E.E., Eltigani, H.F. and Ahmed, A., 2023. The rise of AI: how artificial intelligence is revolutionizing infectious disease control. Annals of Biomedical Engineering, 51(12), pp.2636-2637.
- Allegranzi, B., Kilpatrick, C., Storr, J., Kelley, E., Park, B.J. and Donaldson, L., 2017. Global infection prevention and control priorities 2018–22: a call for action. The Lancet Global Health, 5(12), pp.e1178-e1180.
- World Health Organization, 2020. Guidelines on core components of infection prevention and control programmes at the national and acute health care facility level. World Health Organization. Country Office for Thailand.
- World Health Organization, 2019. Minimum requirements for infection prevention and control programmes.
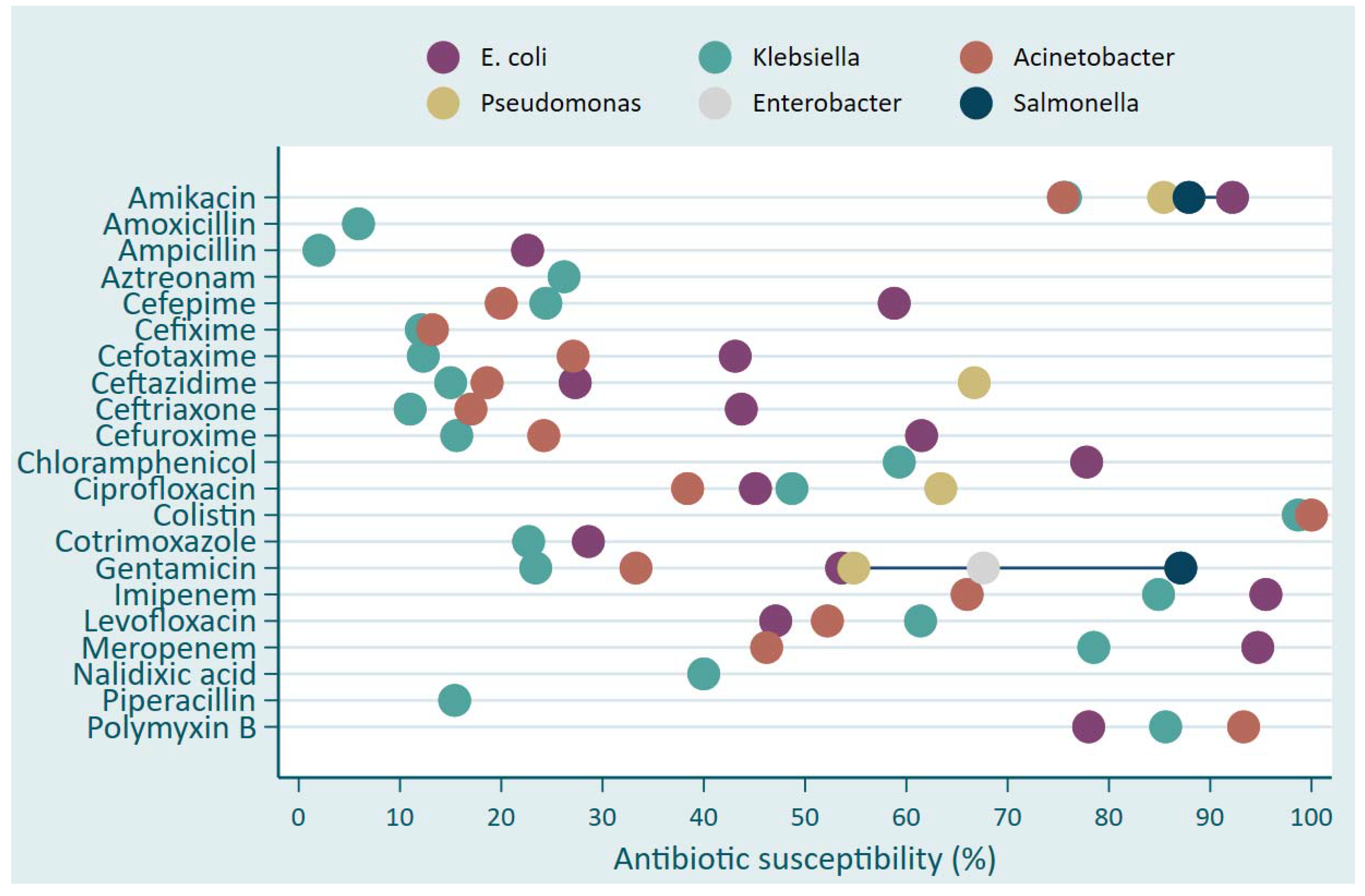
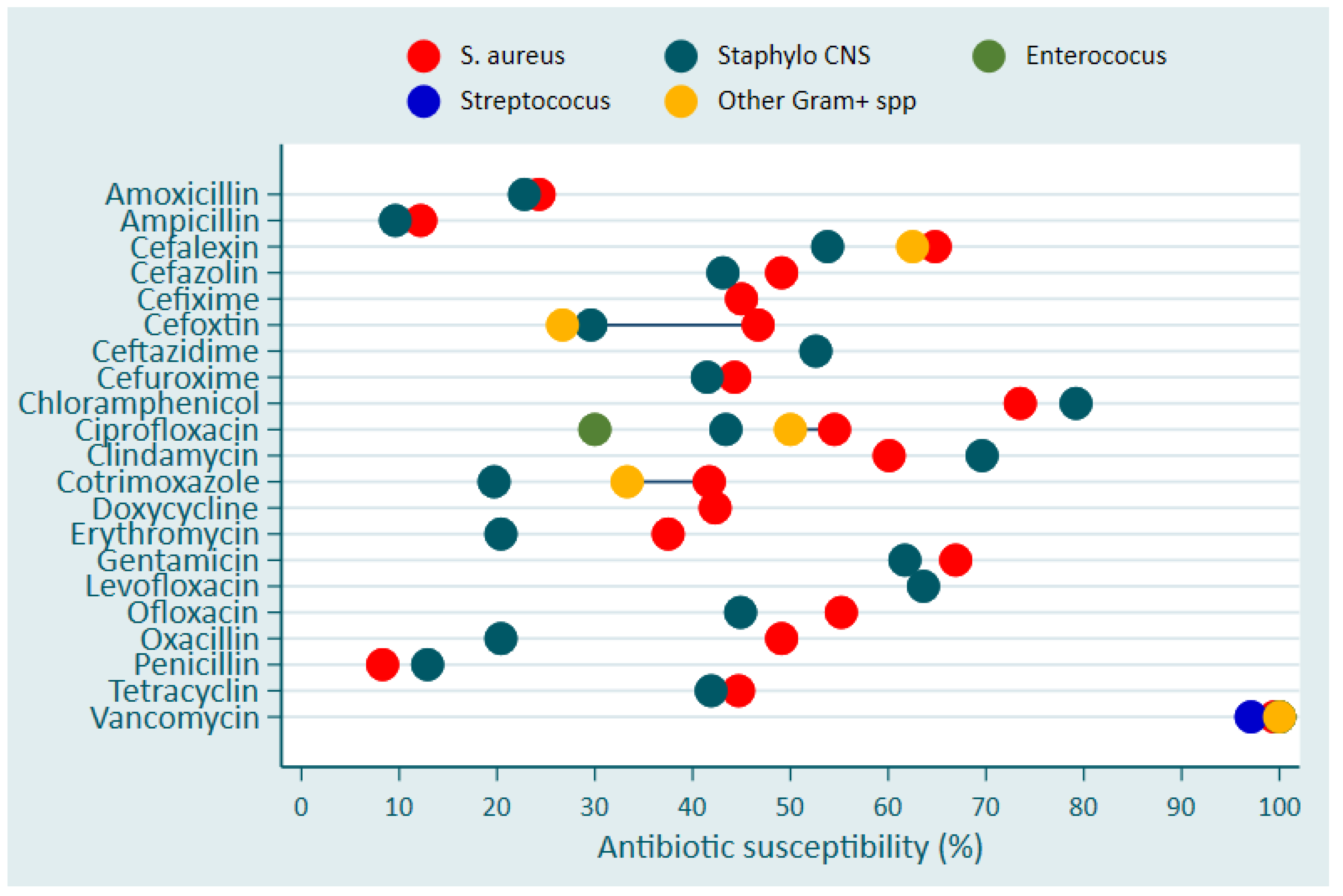
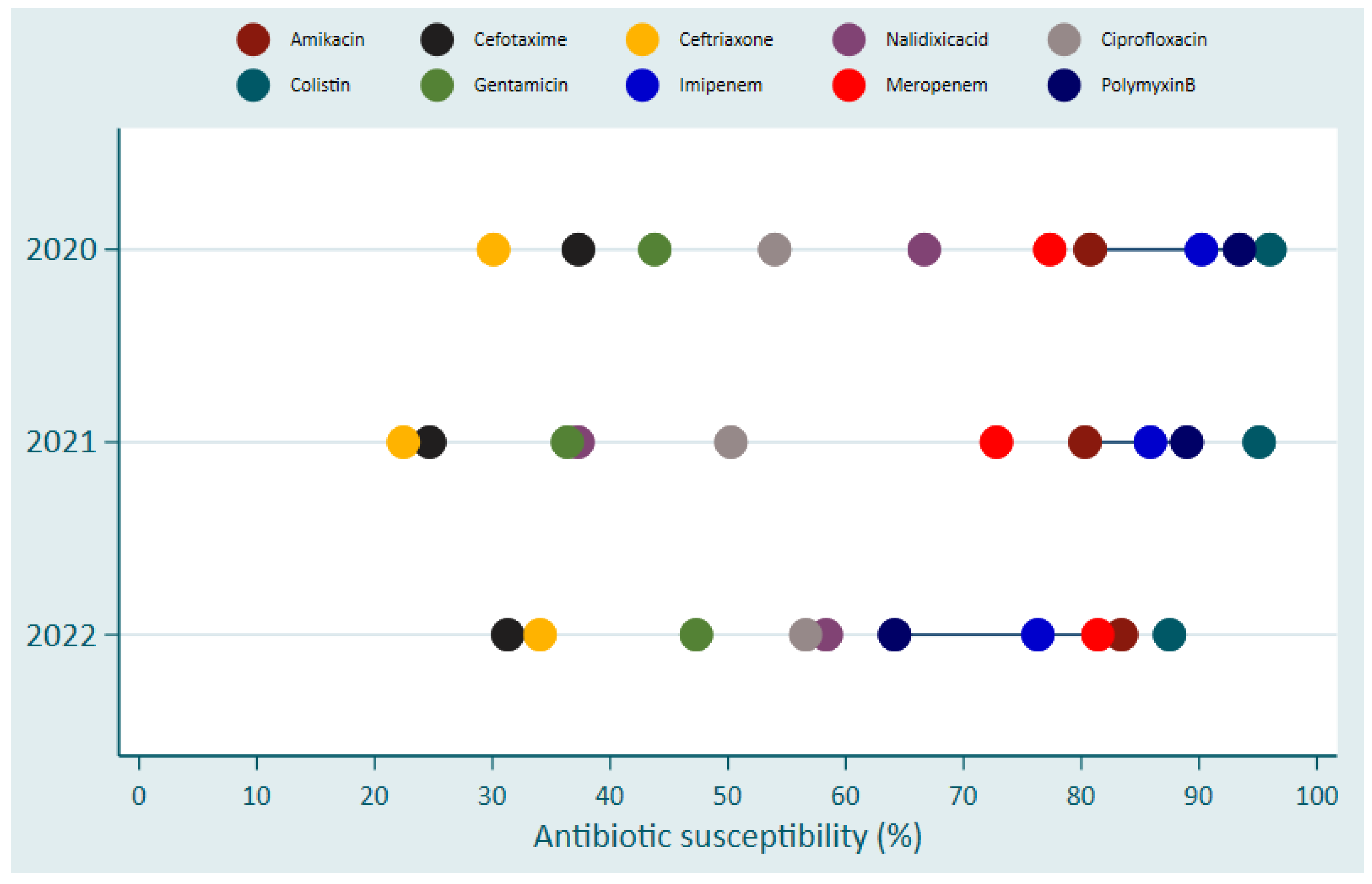
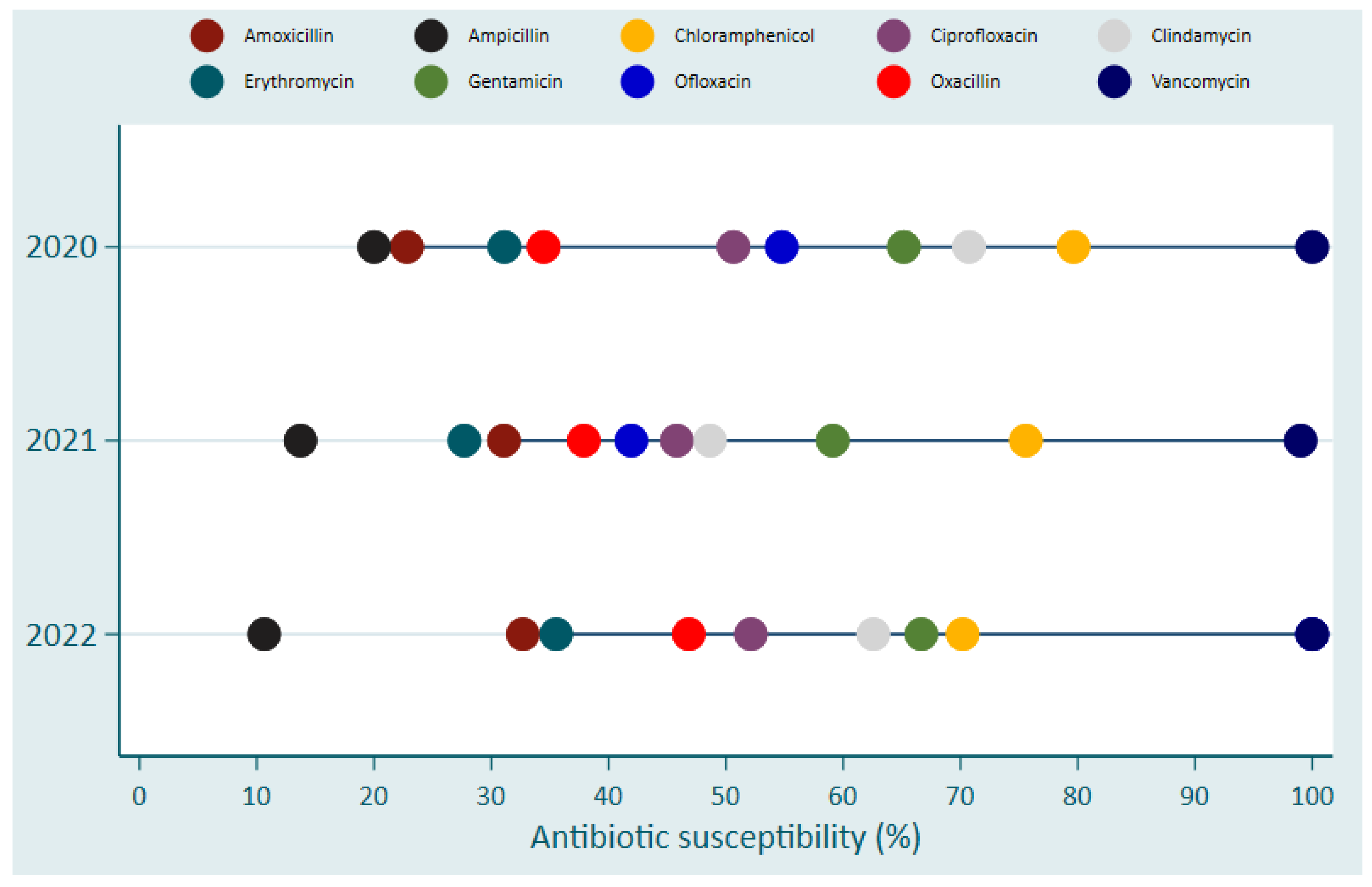
| Pathogens | Hospital | |||
| KFH | CHUK | CHUB | Total | |
| Gram-negative species N (%) | 278 (37.7%) | 328 (44.4%) | 132 (17.9%) | 738 (100.0%) |
| Escherichia coli | 38 (13.7%) | 78 (23.8%) | 15 (11.4%) | 131 (17.8%) |
| Klebsiella spp. | 89 (32.0%) | 152 (46.3%) | 59 (44.7%) | 300 (40.7%) |
| Acinetobacter spp. | 38 (13.7%) | 51 (15.5%) | 22 (16.7%) | 111 (15.0%) |
| Pseudomonas spp. | 28 (10.1%) | 12 (3.7%) | 8 (6.1%) | 48 (6.5%) |
| Enterobacter spp. | 33 (11.9%) | 9 (2.7%) | 5 (3.8%) | 47 (6.4%) |
| Salmonella spp. | 6 (2.2%) | 18 (5.5%) | 18 (13.6%) | 42 (5.7%) |
| Serratia spp. | 17 (6.1%) | 0 (0.0%) | 0 (0.0%) | 17 (2.3%) |
| *Other spp. | 29 (10.4%) | 8 (2.4%) | 5 (3.8%) | 42 (5.7%) |
| Gram-positive species N (%) | 544 (68.5%) | 211 (26.6%) | 39 (4.9%) | 794 (100.0%) |
| Staphylococcus aureus | 177 (32.5%) | 198 (93.8%) | 22 (56.4%) | 397 (50.0%) |
| Staphylococcus (CNS) | 269 (49.4%) | 3 (1.4%) | 11 (28.2%) | 283 (35.6%) |
| Enterococcus spp. | 32 (5.9%) | 4 (1.9%) | 0 (0.0%) | 36 (4.5%) |
| Streptococcus spp. | 24 (4.4%) | 6 (2.8%) | 6 (15.4%) | 36 (4.5%) |
| **Other spp. | 42 (7.7%) | 0 (0.0%) | 0 (0.0%) | 42 (5.3%) |
| Gram-negative isolates N (%) | |||||||||
| Requesting department | Escherichia coli | Klebsiella spp. | Acinetobacter spp. | Pseudomonas spp. | Enterobacter spp. | Salmonella spp. | Serratia spp. | Other spp. | All Gram-negative |
| OPD | 11 (24.4%) | 7 (15.6%) | 10 (22.2%) | 3 (6.7%) | 2 (4.4%) | 9 (20.0%) | 2 (4.4%) | 1 (2.2%) | 45 (6.1%) |
| Int. med. | 36 (30.0%) | 36 (30.0%) | 13 (10.8%) | 8 (6.7%) | 13 (10.8%) | 8 (6.7%) | 0 (0.0%) | 6 (5.0%) | 120 (16.3%) |
| Surgery | 7 (24.1%) | 12 (41.4%) | 5 (17.2%) | 4 (13.8%) | 0 (0.0%) | 0 (0.0%) | 0 (0.0%) | 1 (3.4%) | 29 (3.9%) |
| Emerg. | 19 (22.6%) | 31 (36.9%) | 9 (10.7%) | 4 (4.8%) | 4 (4.8%) | 9 (10.7%) | 3 (3.6%) | 5 (6.0%) | 84 (11.4%) |
| ICU | 15 (10.2%) | 56 (38.1%) | 32 (21.8%) | 10 (6.8%) | 13 (8.8%) | 0 (0.0%) | 5 (3.4%) | 16 (10.9%) | 147 (19.9%) |
| Paed. | 27 (16.0%) | 82 (48.5%) | 23 (13.6%) | 12 (7.1%) | 6 (3.6%) | 15 (8.9%) | 0 (0.0%) | 4 (2.4%) | 169 (22.9%) |
| O&G | 9 (81.8%) | 1 (9.1%) | 0 (0.0%) | 0 (0.0%) | 0 (0.0%) | 1 (9.1%) | 0 (0.0%) | 0 (0.0%) | 11 (1.5%) |
| Neonat. | 6 (7.1%) | 54 (64.3%) | 11 (13.1%) | 4 (4.8%) | 4 (4.8%) | 0 (0.0%) | 0 (0.0%) | 5 (6.0%) | 84 (11.4%) |
| NICU | 1 (3.6%) | 15 (53.6%) | 1 (3.6%) | 1 (3.6%) | 2 (7.1%) | 0 (0.0%) | 5 (17.9%) | 3 (10.7%) | 28 (3.8%) |
| Spec. | 0 (0.0%) | 4 (44.4%) | 4 (44.4%) | 0 (0.0%) | 1 (11.1%) | 0 (0.0%) | 0 (0.0%) | 0 (0.0%) | 9 (1.2%) |
| Others | 0 (0.0%) | 2 (16.7%) | 3 (25.0%) | 2 (16.7%) | 2 (16.7%) | 0 (0.0%) | 2 (16.7%) | 1 (8.3%) | 12 (1.6%) |
| Total | 131 (17.8%) | 300 (40.7%) | 111 (15.0%) | 48 (6.5%) | 47 (6.4%) | 42 (5.7%) | 17 (2.3%) | 42 (5.7%) | 738 (100%) |
| Gram-positive isolates N (%) | ||||||
| Requesting department | Staphylococcus aureus | Staphylococcus (CNS) | Enterococcus spp. | Streptococcus spp. | Other spp. | All Gram-positive |
| OPD | 30 (60.0%) | 19 (38.0%) | 0 (0.0%) | 0 (0.0%) | 1 (2.0%) | 50 (6.3%) |
| Int. med. | 76 (55.5%) | 43 (31.4%) | 5 (3.6%) | 6 (4.4%) | 7 (5.1%) | 137 (17.3%) |
| Surgery | 16 (59.3%) | 4 (14.8%) | 2 (7.4%) | 3 (11.1%) | 2 (7.4%) | 27 (3.4%) |
| Emerg. | 98 (53.0%) | 62 (33.5%) | 10 (5.4%) | 6 (3.2%) | 9 (4.9%) | 185 (23.3%) |
| ICU | 46 (34.3%) | 64 (47.8%) | 15 (11.2%) | 3 (2.2%) | 6 (4.5%) | 134 (16.9%) |
| Paed. | 82 (58.6%) | 38 (27.1%) | 1 (0.7%) | 11 (7.9%) | 8 (5.7%) | 140 (17.6%) |
| O&G | 6 (40.0%) | 8 (53.3%) | 0 (0.0%) | 0 (0.0%) | 1 (6.7%) | 15 (1.9%) |
| Neonat. | 26 (78.8%) | 5 (15.2%) | 0 (0.0%) | 1 (3.0%) | 1 (3.0%) | 33 (4.2%) |
| NICU | 10 (23.3%) | 19 (44.2%) | 3 (7.0%) | 4 (9.3%) | 7 (16.3%) | 43 (5.4%) |
| Spec. | 2 (66.7%) | 1 (33.3%) | 0 (0.0%) | 0 (0.0%) | 0 (0.0%) | 3 (0.4%) |
| Others | 5 (18.5%) | 20 (74.1%) | 0 (0.0%) | 2 (7.4%) | 0 (0.0%) | 27 (3.4%) |
| Total | 397 (50.0%) | 283 (35.6%) | 36 (4.5%) | 36 (4.5%) | 42 (5.3%) | 794 (100%) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




