Submitted:
19 July 2024
Posted:
19 July 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Aim of the Study
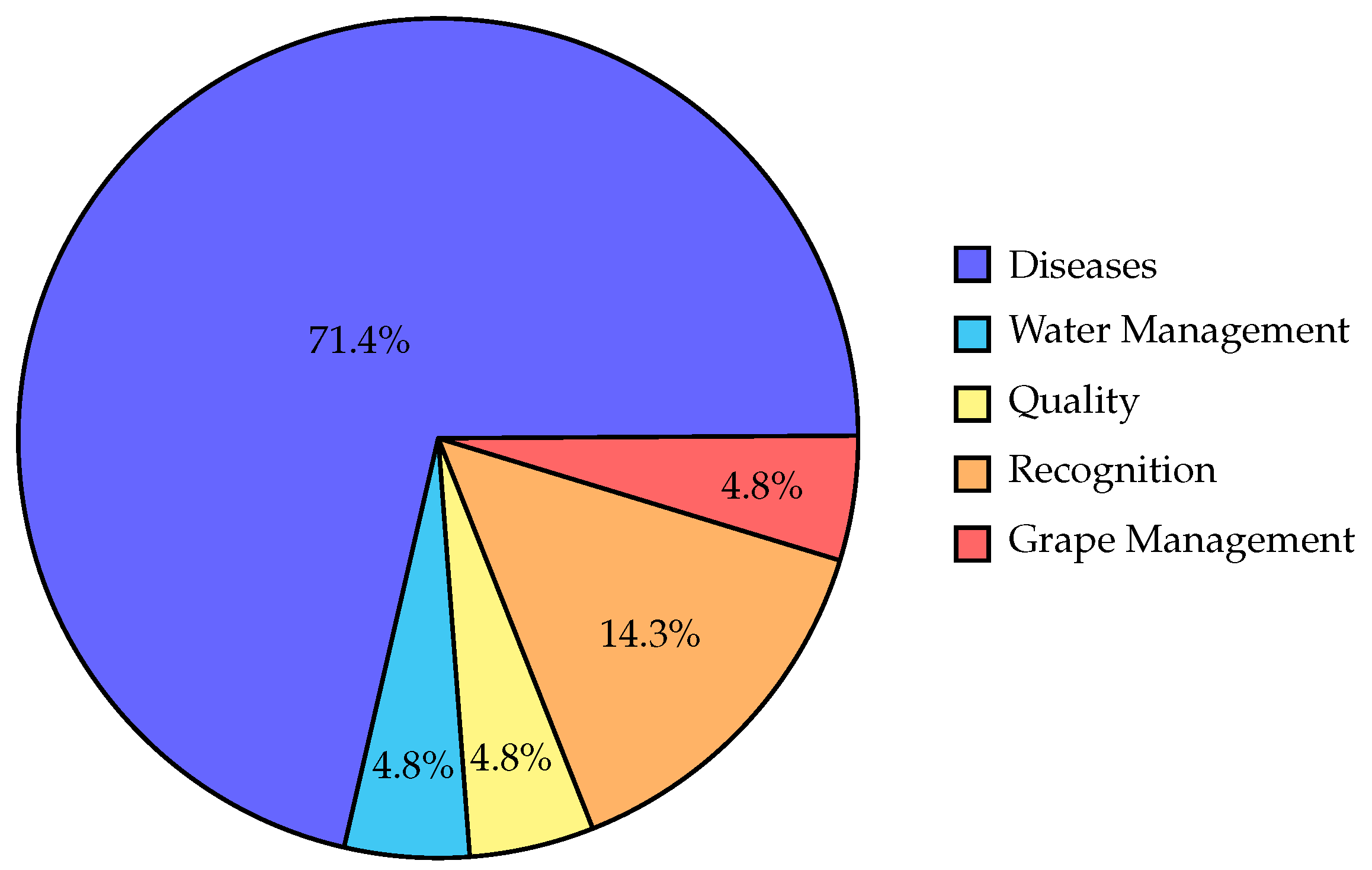
- Knowing that the majority of the research (88%) had been done in the last 5 years, it needed to be organized in one place where they will be compared together.
- Moreover, we organize all the technologies used, the applications of ML in Grapevine research, and correlate them together. As a result, it becomes more likely that researchers will look at the general picture and all perspectives.
- Knowing the current studied ML topics in Grapevine research and comparing them with the ML topics studied in General Agriculture, it’s easier to understand the topics that haven’t been thoroughly studied yet in Grapevine.
- The datasets are organized in the same place so that future researchers can easily utilize them, while hopefully, future works could arise using the same data for comparison reasons.
2. Materials and Methods
2.1. Outline of the Paper
3. Background
3.1. Machine Learning in Agriculture
- Crop Management
- Water Management
- Soil Management
- Livestock Management
- Supervised learning: The input and output are known, and the machine attempts to find the best path to an output given an input; [2];
- Unsupervised learning: No labels are provided, leaving the learning algorithm to generate structure within its input [5];
- Semi-supervised learning: A mixture of labeled and unlabeled data constitute the input data [4];
- Reinforcement learning: Decisions are made towards finding out actions that can lead to the more positive outcome, while it is solely determined by trial and error method and delayed outcome [3].
3.2. Supervised Learning
- Classification: Classification problems arise when the output variable is categorical such as “disease” or “no disease”.
- Regression: Regression problems occur when the output variable is a true continuous value such as stock price prediction.
3.3. Unsupervised Learning
- Clustering: A clustering problem is one in which you want to disclose the underlying groupings in the data, such as grouping animals based on particular characteristics/features such as leg count.
- Association: Here you want to identify association rules, such as those who buy X also buy Y.
3.4. Semi-supervised Learning
3.5. Reinforcement Algorithms
4. Diseases in Grapevine
4.1. Grapevine Yellow (GY)
4.2. Esca
4.3. Downy mildew
4.4. Leafroll
4.5. Pierce‘s
4.6. Root Rot
5. Datasets
- GrapeCS-ML database: https://researchoutput.csu.edu.au/en/datasets/grape-image-database [14] The GrapeCS-ML database consists of images of grape varieties at different stages of development together with the corresponding ground truth data (e.g., pH and Brix) obtained from chemical analysis. The database consists of five datasets for 15 grape varieties taken at several stages of development and includes size and/or Macbeth color references. Altogether, the database contains a total of 2078 images.
- In the paper LDD: A Dataset for Grape Diseases Object Detection and Instance Segmentation [15] The authors created a grapevine disease database with images from Horta’s internal databases, the competition Grapevine Disease Images and from the web, and manual segmentations. It contains 1,092 RGB images of grapes and 17,706 annotations (instances) for the tasks of Object Detection and Instance Segmentation. More specifically, it contains the categories Black Rot with 1180 instances, Esca (Black measles) with 1383 instances, Leaf blight (Isariopsis Leaf Spot) with 1076 instances, and Healthy with 423 instances.
- Grapes-Leaf-Disease-detection repository https://github.com/shreyansh-kothari/Grapes-Leaf-Disease-detection consists of images about grapevine diseases, to be exact, it includes Esca (240 images), Black Rot (210 images), Healthy (220 images), Leaf Blight (210 images).
- In [16] https://github.com/cu-cairlab/iros2022-OnlineDMSeg.git there is a dataset with 282 raw images, classifying them in healthy or with Downey Mildew disease.
-
PlantVillage:A collection of datasets augmented with plantVillage dataset for grapevine diseases. It consists of the categories Black rot (1180 images), Esca (1383 images), Grapevine Yellow (134 images), Healthy (84 images), and Leaf blight (1076 images). They also have edited versions of the images with various operations increasing the total number of them.
- http://dx.doi.org/10.17632/89cnxc58kj.1: A dataset [18] specific about Esca disease in Grapes, consists of 881 healthy RGB images and 887 Esca images.
-
For the paper [19], they also created a dataset using hyperspectral imaging under laboratory lighting, their dataset consisted of 496 images of 248 plants. Also, this dataset distinguishes between symptomatic and asymptomatic leaves.The images in the collection were captured on grapevine leaves from five different varieties: Ak, Ala Idris, Büzgülü, Dimnit, and Nazli each kind of grapevine species has 100 images. This dataset is available to researchers at https://www.kaggle.com/datasets/muratkokludataset/grapevine-leaves-image-dataset https://www.muratkoklu.com/datasets/. The same dataset has been used in [20].
- Some research used also statistical data for grapevine disease (e.g [21]), some statistical data for diseases can be found on the following pages based on grapevines located in Italy http://www.agroambiente.info/ https://agroambiente.info.regione.toscana.it/.
6. Techniques in Grapevine
6.1. Data Format Techniques
6.1.1. RGB Images
- Classification and Mapping: The valuable spectral information gained from the AVIRIS-NG can be used with machine learning algorithms for classification and mapping applications. This may include the mapping of vegetation, surface and open water, and crop types. They can also use the labeled AVIRIS-NG data in other images to automatically classify objects or features.
- Environmental Monitoring: We can use AVIRIS-NG images to estimate environmental parameters such as vegetation health, water quality, pollution, etc. Machine learning algorithms permit the examination of the spectral patterns and correlations in AVIRIS-NG data for the detection and observation of anomalies or changes over time, which is greatly helpful in identifying environmental hazards or changes in ecosystems.
- Disease Detection: Early detection of plant diseases is feasible by training machine learning models on the AVIRIS-NG images. The algorithms detect even minor changes in vegetation health since AVIRIS-NG captures the spectral signatures of vegetation. Thus, farmers and researchers make informed decisions to control the spread of diseases and protect crops.
- Mineral Exploration: AVIRIS-NG data can be used in machine learning applications for mineral exploration and mapping. In the reflectance spectra, there are unique absorption features for rocks and minerals that can be observed from the spectral signatures obtained from the AVIRIS-NG images. By training machine learning algorithms it may be allowed to identify and detect minerals, and hence locate and map deposits of various minerals.
6.1.2. Hyperspectral Images
6.1.3. Multispectral images
6.1.4. Thermal Images
6.1.5. Unmanned Aerial Vehicle (UAV) Images
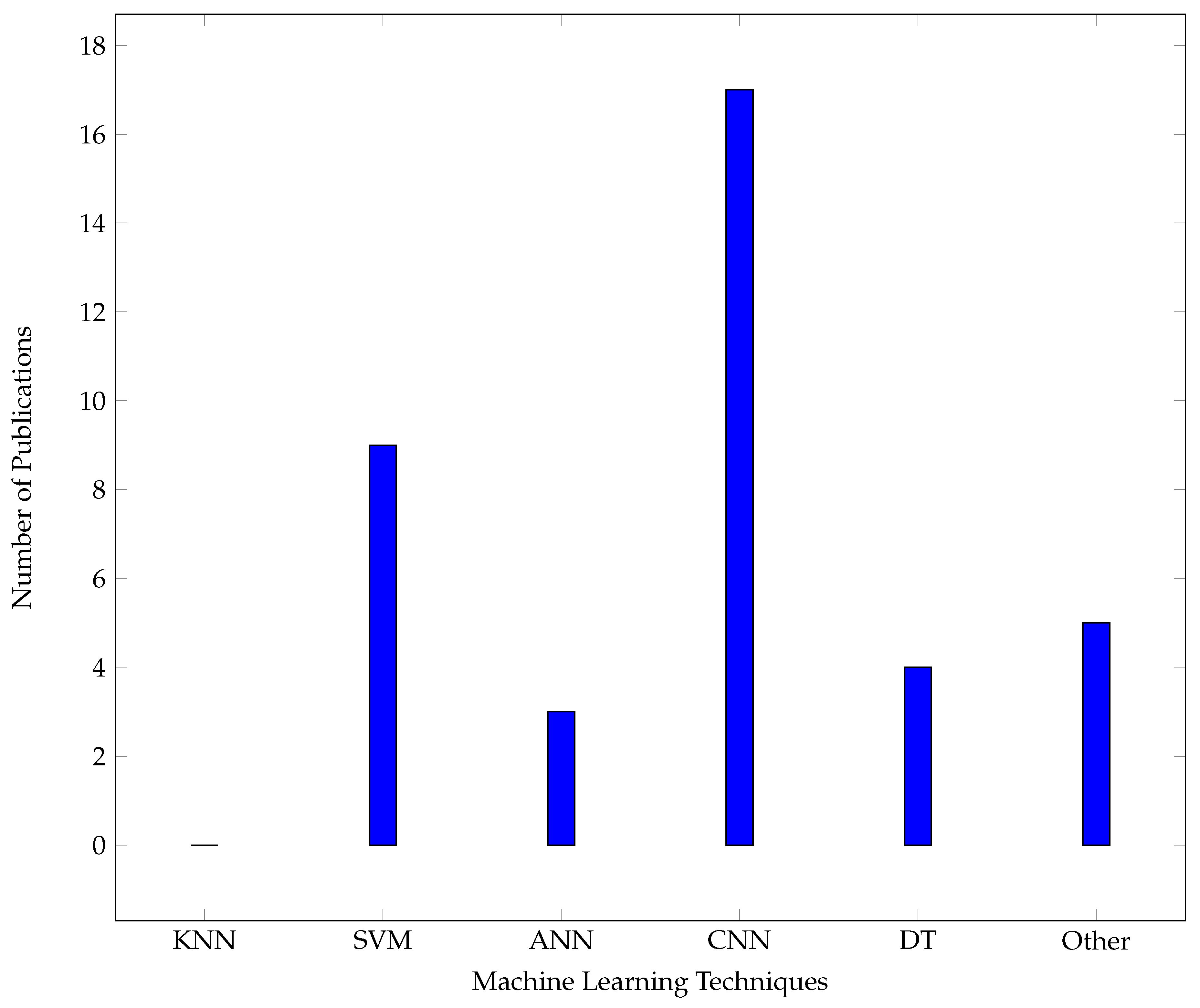
6.2. Machine Learning Techniques
6.2.1. Statistical Measures
6.2.2. K-Nearest Neighbour (KNN)
6.2.3. Naive Bayes (NB)
6.2.4. Support Vector Machine (SVM)
6.2.5. Decision Trees
Random Forest
C5.0
6.2.6. Artificial Neural Networks (ANN)
MultiLayer Perceptron (MLP)
Convolutional Neural Network (CNN)
Single Shot Multibox Detector
- MobileNets are lightweight convolutional neural network models designed to perform deep learning tasks on resource-constrained devices. They are attempting to strike a balance between model size and processing efficiency while maintaining acceptable levels of accuracy. MobileNets do this by the use of depthwise separable convolutions, which divide regular convolutions into two different layers: depthwise and pointwise. Depthwise apply a single filter to each input channel, and thus minimize computational complexity. Pointwise convolutions employ 1x1 convolutions to combine the outputs of depthwise convolutions to capture channel-wise interactions, which results in a reduced number of parameters and calculations required making it suitable for resource-constrained devices. MobileNets offer flexibility through the use of hyperparameters like width multiplier () or resolution multiplier (), which allows for extra trade-offs between model size, processing costs, and accuracy [62].
- Inception-V2 is a deep neural network model designed to deal better with objects that change size across images instead of having to find the optimal kernel. Inception-V2 improves on the original Inception model by reducing computational complexity through the use of factorized convolution techniques. By reducing the number of individual convolutions, Inception-V2 improves runtime performance while maintaining a comparable level of accuracy [63].
6.2.7. Genetic Algorithm (GA) in feature selection
7. Machine Learning Applications to Grapevine
7.1. Diseases
7.1.1. Grapevine Yellow Disease
Detection of grapevine yellows symptoms in Vitis vinifera L. with artificial intelligence
Using Image Texture and Spectral Reflectance Analysis to Detect Yellowness and Esca in Grapevines at Leaf-Level
Detection of Two Different Grapevine Yellows in Vitis vinifera Using Hyperspectral Imaging
7.1.2. Flavescence Dorée Disease
Automatic detection of Flavescense Dorée grapevine disease in hyperspectral images using machine learning
Development of Spectral Disease Indices for ‘Flavescence Dorée’ Grapevine Disease Identification
Assessment of the optimal spectral bands for designing a sensor for vineyard disease detection: the case of ‘Flavescence dorée’
7.1.3. Esca Disease
Evaluating the suitability of hyper- and multispectral imaging to detect foliar symptoms of the grapevine trunk disease Esca in vineyards
A grapevine leaves dataset for early detection and classification of esca disease in vineyards through machine learning
Deep learning approach with colorimetric spaces and vegetation indices for vine diseases detection in UAV images
7.1.4. Mildew Disease
Predicting symptoms of downy mildew, powdery mildew, and gray mold diseases of grapevine through machine learning
Near Real-Time Vineyard Downy Mildew Detection and Severity Estimation
Deep Learning for the Differentiation of Downy Mildew and Spider Mite in Grapevine under Field Conditions
Artificial Intelligence and Novel Sensing Technologies for Assessing Downy Mildew in Grapevine
Early Detection of Grapevine (Vitis Vinifera) Downy Mildew (Peronospora) and Diurnal Variations Using Thermal Imaging
Vine Disease Detection in UAV Multispectral Images Using Optimized Image Registration and Deep Learning Segmentation Approach
7.1.5. Leafroll Disease
Phenotyping grapevine red blotch virus and grapevine leafroll-associated viruses before and after symptom expression through machine-learning analysis of hyperspectral images
Early Detection of Grapevine Leafroll Disease in a Red-Berried Wine Grape Cultivar Using Hyperspectral Imaging
Scalable Early Detection of Grapevine Virus Infection with Airborne Imaging Spectroscopy
7.1.6. Pierce’s Disease
Vision-Based Grapevine Pierce’s Disease Detection System Using Artificial Intelligence
7.1.7. Root Rot Disease
Early Identification of Root Rot Disease by Using Hyperspectral Reflectance: The Case of Pathosystem Grapevine/Armillaria
7.1.8. General Disease Detection System
Bringing Semantics to the Vineyard: An Approach on Deep Learning-Based Vine Trunk Detection
Automatic Grape Leaf Diseases Identification via United Model Based on Multiple Convolutional Neural Networks
Early Detection of Plant Viral Disease Using Hyperspectral Imaging and Deep Learning
Entropy-Controlled Deep Features Selection Framework for Grape Leaf Diseases Recognition
Grape Leaf Disease Detection and Classification Using Machine Learning
A Smart Agricultural Application: Automated Detection of Diseases in Vine Leaves Using Hybrid Deep Learning
A Deep-Learning-Based Real-Time Detector for Grape Leaf Diseases Using Improved Convolutional Neural Networks
VddNet: Vine Disease Detection Network Based on Multispectral Images and Depth Map
UAS-Based Hyperspectral Sensing Methodology for Continuous Monitoring and Early Detection of Vineyard Anomalies
7.2. Water Status Assessment
Vineyard Water Status Assessment Using on-the-go Thermal Imaging and Machine Learning
Vineyard water status estimation using multispectral imagery from a UAV platform and machine learning algorithms for irrigation scheduling management
7.3. Plant Deficiencies
A deep learning algorithm for detection of potassium deficiency in a red grapevine and spraying actuation using a raspberry pi3
7.4. Classification
In-field high throughput grapevine phenotyping with a consumer-grade depth camera
A CNN-SVM study based on selected deep features for grapevine leaves classification
On-the-go hyperspectral imaging under field conditions and machine learning for the classification of grapevine varieties
Image classification for detection of winter grapevine buds in natural conditions using scale-invariant features transform, a bag of features, and support vector machines
Automated grapevine cultivar identification via leaf imaging and deep convolutional neural networks: a proof-of-concept study employing primary Iranian varieties
7.5. Others
Geographical and cultivar features differentiate grape microbiota in Northern Italy and Spain vineyards
Detection of single grapevine berries in images using fully convolutional neural networks
Insect classification and detection in field crops using modern machine learning techniques
Path Planning Algorithms Benchmarking for Grapevines Pruning and Monitoring
A robot system for pruning grape vines
Assessment of Smoke Contamination in Grapevine Berries and Taint in Wines Due to Bushfires Using a Low-Cost E-Nose and an Artificial Intelligence Approach
VineInspector: The Vineyard Assistant
8. Discussion
Funding
References
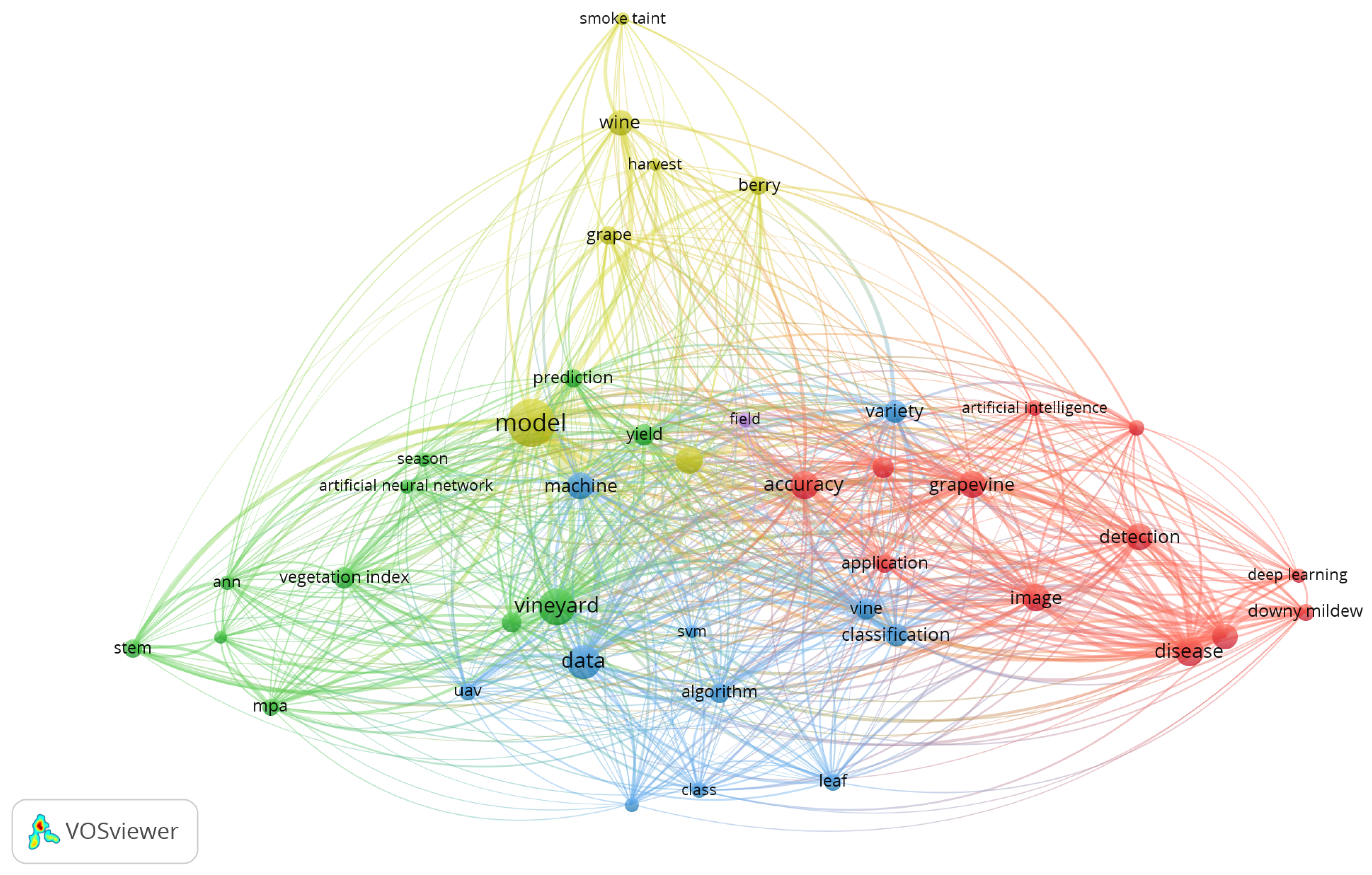
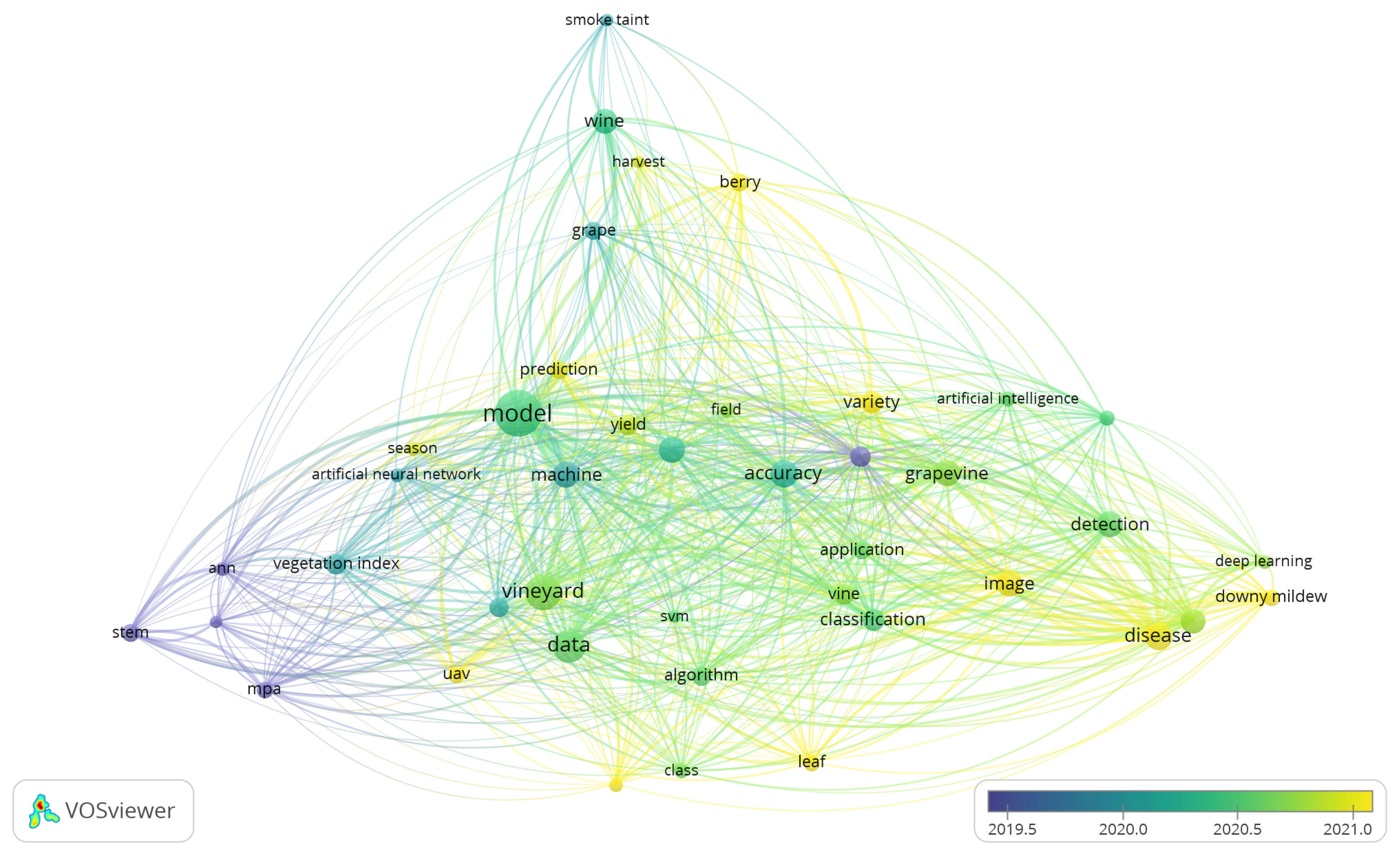
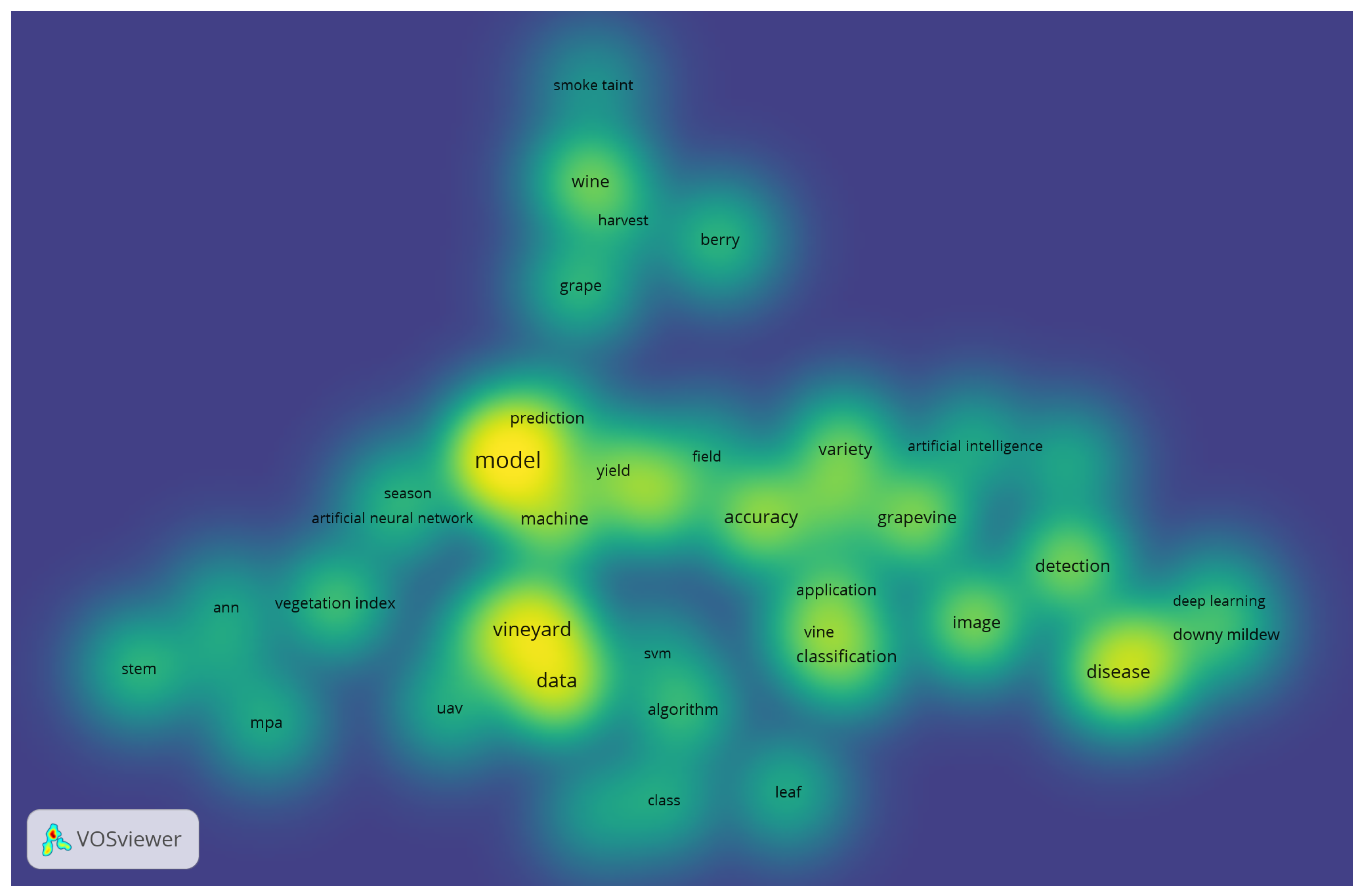
- Van Eck, N.; Waltman, L. Software survey: VOSviewer, a computer program for bibliometric mapping. scientometrics 2010, 84, 523–538. [Google Scholar] [CrossRef] [PubMed]
- Vishnoi, V.K.; Kumar, K.; Kumar, B. Plant disease detection using computational intelligence and image processing. Journal of Plant Diseases and Protection 2021, 128, 19–53. [Google Scholar] [CrossRef]
- Abdulridha, J.; Batuman, O.; Ampatzidis, Y. UAV-based remote sensing technique to detect citrus canker disease utilizing hyperspectral imaging and machine learning. Remote Sensing 2019, 11, 1373. [Google Scholar] [CrossRef]
- Ouhami, M.; Hafiane, A.; Es-Saady, Y.; El Hajji, M.; Canals, R. Computer vision, IoT and data fusion for crop disease detection using machine learning: a survey and ongoing research. Remote Sensing 2021, 13, 2486. [Google Scholar] [CrossRef]
- Sharma, A.; Jain, A.; Gupta, P.; Chowdary, V. Machine learning applications for precision agriculture: A comprehensive review. IEEE Access 2020, 9, 4843–4873. [Google Scholar] [CrossRef]
- Benos, L.; Tagarakis, A.C.; Dolias, G.; Berruto, R.; Kateris, D.; Bochtis, D. Machine learning in agriculture: A comprehensive updated review. Sensors 2021, 21, 3758. [Google Scholar] [CrossRef] [PubMed]
- Fontaine, F.; Gramaje, D.; Armengol, J.; Smart, R.; Nagy, Z.A.; Borgo, M.; Rego, C.; Corio-Costet, M.F. Grapevine trunk diseases. A review, 2016.
- Gessler, C.; Pertot, I.; Perazzolli, M. Plasmopara viticola: a review of knowledge on downy mildew of grapevine and effective disease management. Phytopathologia Mediterranea 2011, 50, 3–44. [Google Scholar]
- Naidu, R.A.; Maree, H.J.; Burger, J.T. Grapevine leafroll disease and associated viruses: a unique pathosystem. Annual Review of Phytopathology 2015, 53, 613–634. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Luscher, J.; Plank, C.M.; Brillante, L.; Cooper, M.L.; Smith, R.J.; Al-Rwahnih, M.; Yu, R.; Oberholster, A.; Girardello, R.; Kurtural, S.K. Grapevine red blotch virus may reduce carbon translocation leading to impaired grape berry ripening. Journal of agricultural and food chemistry 2019, 67, 2437–2448. [Google Scholar] [CrossRef]
- Hopkins, D.; Purcell, A. Xylella fastidiosa: cause of Pierce’s disease of grapevine and other emergent diseases. Plant disease 2002, 86, 1056–1066. [Google Scholar] [CrossRef]
- Cruz, A.C.; El-Kereamy, A.; Ampatzidis, Y. Vision-based grapevine pierce’s disease detection system using artificial intelligence. 2018 ASABE Annual International Meeting. American Society of Agricultural and Biological Engineers, 2018, p. 1.
- Langenhoven, S.D.; Halleen, F.; Spies, C.F.; Stempien, E.; Mostert, L. Detection and quantification of black foot and crown and root rot pathogens in grapevine nursery soils in the Western Cape of South Africa. Phytopathologia Mediterranea 2018, 57, 519–537. [Google Scholar]
- Seng, K.P.; Ang, L.M.; Schmidtke, L.M.; Rogiers, S.Y. Computer vision and machine learning for viticulture technology. IEEE Access 2018, 6, 67494–67510. [Google Scholar] [CrossRef]
- Rossi, L.; Valenti, M.; Legler, S.E.; Prati, A. LDD: A Grape Diseases Dataset Detection and Instance Segmentation. Image Analysis and Processing–ICIAP 2022: 21st International Conference, Lecce, Italy, –27, 2022, Proceedings, Part II. Springer, 2022, pp. 383–393. 23 May. [CrossRef]
- Liu, E.; Gold, K.; Cadle-Davidson, L.; Combs, D.; Jiang, Y. Near Real-Time Vineyard Downy Mildew Detection and Severity Estimation. 2022 IEEE/RSJ International Conference on Intelligent Robots and Systems (IROS), 2022, pp. 9187–9194. [CrossRef]
- Cruz, A.; Ampatzidis, Y.; Pierro, R.; Materazzi, A.; Panattoni, A.; De Bellis, L.; Luvisi, A. Detection of grapevine yellows symptoms in Vitis vinifera L. with artificial intelligence. Computers and electronics in agriculture 2019, 157, 63–76. [Google Scholar] [CrossRef]
- Falaschetti, L. ESCA-dataset, 2021.
- Sawyer, E.; Laroche-Pinel, E.; Flasco, M.; Cooper, M.L.; Corrales, B.; Fuchs, M.; Brillante, L. Phenotyping grapevine red blotch virus and grapevine leafroll-associated viruses before and after symptom expression through machine-learning analysis of hyperspectral images. Frontiers in Plant Science 2023, 14. [Google Scholar] [CrossRef] [PubMed]
- Koklu, M.; Unlersen, M.F.; Ozkan, I.A.; Aslan, M.F.; Sabanci, K. A CNN-SVM study based on selected deep features for grapevine leaves classification. Measurement 2022, 188, 110425. [Google Scholar] [CrossRef]
- Volpi, I.; Guidotti, D.; Mammini, M.; Marchi, S. Predicting symptoms of downy mildew, powdery mildew, and gray mold diseases of grapevine through machine learning. Italian Journal of Agrometeorology.
- Castleman, K.R. Digital image processing; Prentice Hall Press, 1996.
- Rajendran, S.; Aravindan, S.; Rajakumar, T.; Sivakumar, M.; Mohan, K. Hyperspectral Remote Sensing and Spectral Signature Applications; 2009. [CrossRef]
- Kettig, R.L.; Landgrebe, D. Classification of multispectral image data by extraction and classification of homogeneous objects. IEEE Transactions on geoscience Electronics 1976, 14, 19–26. [Google Scholar] [CrossRef]
- Wiecek, B. Review on thermal image processing for passive and active thermography. 2005 IEEE Engineering in Medicine and Biology 27th Annual Conference. IEEE, 2006, pp. 686–689. [CrossRef]
- Al-Saddik, H.; Laybros, A.; Billiot, B.; Cointault, F. Using image texture and spectral reflectance analysis to detect Yellowness and Esca in grapevines at leaf-level. Remote Sensing 2018, 10, 618. [Google Scholar] [CrossRef]
- Bendel, N.; Backhaus, A.; Kicherer, A.; Köckerling, J.; Maixner, M.; Jarausch, B.; Biancu, S.; Klück, H.C.; Seiffert, U.; Voegele, R.T.; others. Detection of two different grapevine yellows in Vitis vinifera using hyperspectral imaging. Remote Sensing 2020, 12, 4151. [Google Scholar] [CrossRef]
- Silva, D.M.; Bernardin, T.; Fanton, K.; Nepaul, R.; Pádua, L.; Sousa, J.J.; Cunha, A. Automatic detection of Flavescense Dorée grapevine disease in hyperspectral images using machine learning. Procedia Computer Science 2022, 196, 125–132. [Google Scholar] [CrossRef]
- Al-Saddik, H.; Simon, J.C.; Cointault, F. Development of spectral disease indices for ‘Flavescence Dorée’grapevine disease identification. Sensors 2017, 17, 2772. [Google Scholar] [CrossRef]
- Al-Saddik, H.; Simon, J.C.; Cointault, F. Assessment of the optimal spectral bands for designing a sensor for vineyard disease detection: the case of ‘Flavescence dorée’. Precision Agriculture 2019, 20, 398–422. [Google Scholar] [CrossRef]
- Bendel, N.; Kicherer, A.; Backhaus, A.; Klück, H.C.; Seiffert, U.; Fischer, M.; Voegele, R.T.; Töpfer, R. Evaluating the suitability of hyper-and multispectral imaging to detect foliar symptoms of the grapevine trunk disease Esca in vineyards. Plant methods 2020, 16, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Alessandrini, M.; Rivera, R.C.F.; Falaschetti, L.; Pau, D.; Tomaselli, V.; Turchetti, C. A grapevine leaves dataset for early detection and classification of esca disease in vineyards through machine learning. Data in Brief 2021, 35, 106809. [Google Scholar] [CrossRef] [PubMed]
- Kerkech, M.; Hafiane, A.; Canals, R. Deep leaning approach with colorimetric spaces and vegetation indices for vine diseases detection in UAV images. Computers and electronics in agriculture 2018, 155, 237–243. [Google Scholar] [CrossRef]
- Gutiérrez, S.; Hernández, I.; Ceballos, S.; Barrio, I.; Díez-Navajas, A.M.; Tardaguila, J. Deep learning for the differentiation of downy mildew and spider mite in grapevine under field conditions. Computers and Electronics in Agriculture 2021, 182, 105991. [Google Scholar] [CrossRef]
- Hernández, I.; Gutiérrez, S.; Ceballos, S.; Iñíguez, R.; Barrio, I.; Tardaguila, J. Artificial intelligence and novel sensing technologies for assessing downy mildew in grapevine. Horticulturae 2021, 7, 103. [Google Scholar] [CrossRef]
- Cohen, B.; Edan, Y.; Levi, A.; Alchanatis, V. Early detection of grapevine (Vitis vinifera) downy mildew (Peronospora) and diurnal variations using thermal imaging. Sensors 2022, 22, 3585. [Google Scholar] [CrossRef] [PubMed]
- Kerkech, M.; Hafiane, A.; Canals, R. Vine disease detection in UAV multispectral images using optimized image registration and deep learning segmentation approach. Computers and Electronics in Agriculture 2020, 174, 105446. [Google Scholar] [CrossRef]
- Gao, Z.; Khot, L.R.; Naidu, R.A.; Zhang, Q. Early detection of grapevine leafroll disease in a red-berried wine grape cultivar using hyperspectral imaging. Computers and Electronics in Agriculture 2020, 179, 105807. [Google Scholar] [CrossRef]
- Romero Galvan, F.E.; Sousa, D.; Pavlick, R.; Aggarwal, S.; Trolley, G.R.; Forrestel, E.J.; Bolton, S.L.; Dokoozlian, N.; Alsina, M.D.M.; Gold, K.M. Scalable early detection of grapevine virus infection with airborne imaging spectroscopy. bioRxiv, 2022. [Google Scholar] [CrossRef]
- Calamita, F.; Imran, H.A.; Vescovo, L.; Mekhalfi, M.L.; La Porta, N. Early identification of root rot disease by using hyperspectral reflectance: The case of pathosystem grapevine/armillaria. Remote Sensing 2021, 13, 2436. [Google Scholar] [CrossRef]
- Aguiar, A.S.; Monteiro, N.N.; Santos, F.N.d.; Solteiro Pires, E.J.; Silva, D.; Sousa, A.J.; Boaventura-Cunha, J. Bringing semantics to the vineyard: An approach on deep learning-based vine trunk detection. Agriculture 2021, 11, 131. [Google Scholar] [CrossRef]
- Ji, M.; Zhang, L.; Wu, Q. Automatic grape leaf diseases identification via UnitedModel based on multiple convolutional neural networks. Information Processing in Agriculture 2020, 7, 418–426. [Google Scholar] [CrossRef]
- Nguyen, C.; Sagan, V.; Maimaitiyiming, M.; Maimaitijiang, M.; Bhadra, S.; Kwasniewski, M.T. Early detection of plant viral disease using hyperspectral imaging and deep learning. Sensors 2021, 21, 742. [Google Scholar] [CrossRef] [PubMed]
- Adeel, A.; Khan, M.A.; Akram, T.; Sharif, A.; Yasmin, M.; Saba, T.; Javed, K. Entropy-controlled deep features selection framework for grape leaf diseases recognition. Expert Systems 2022, 39, e12569. [Google Scholar] [CrossRef]
- Huang, Z.; Qin, A.; Lu, J.; Menon, A.; Gao, J. Grape leaf disease detection and classification using machine learning. 2020 International Conferences on Internet of Things (iThings) and IEEE Green Computing and Communications (GreenCom) and IEEE Cyber, Physical and Social Computing (CPSCom) and IEEE Smart Data (SmartData) and IEEE Congress on Cybermatics (Cybermatics). IEEE, 2020, pp. 870–877. [CrossRef]
- Alkan, A.; Abdullah, M.U.; Abdullah, H.O.; Assaf, M.; Zhou, H. A smart agricultural application: automated detection of diseases in vine leaves usinghybrid deep learning. Turkish Journal of Agriculture and Forestry 2021, 45, 717–729. [Google Scholar] [CrossRef]
- Xie, X.; Ma, Y.; Liu, B.; He, J.; Li, S.; Wang, H. A deep-learning-based real-time detector for grape leaf diseases using improved convolutional neural networks. Frontiers in plant science 2020, 11, 751. [Google Scholar] [CrossRef] [PubMed]
- Kerkech, M.; Hafiane, A.; Canals, R. VddNet: Vine disease detection network based on multispectral images and depth map. Remote Sensing 2020, 12, 3305. [Google Scholar] [CrossRef]
- Adão, T.; Peres, E.; Pádua, L.; Hruška, J.; Sousa, J.J.; Morais, R. UAS-based hyperspectral sensing methodology for continuous monitoring and early detection of vineyard anomalies. Proceedings of the Small Unmanned Aerial Systems for Environmental Research, Vila Real, Portugal.
- Gutiérrez, S.; Diago, M.P.; Fernández-Novales, J.; Tardaguila, J. Vineyard water status assessment using on-the-go thermal imaging and machine learning. PLoS One 2018, 13, e0192037. [Google Scholar] [CrossRef] [PubMed]
- Romero, M.; Luo, Y.; Su, B.; Fuentes, S. Vineyard water status estimation using multispectral imagery from an UAV platform and machine learning algorithms for irrigation scheduling management. Computers and electronics in agriculture 2018, 147, 109–117. [Google Scholar] [CrossRef]
- Ukaegbu, U.; Tartibu, L.; Laseinde, T.; Okwu, M.; Olayode, I. A deep learning algorithm for detection of potassium deficiency in a red grapevine and spraying actuation using a raspberry pi3. 2020 international conference on artificial intelligence, big data, computing and data communication systems (icabcd). IEEE, 2020, pp. 1–6. [CrossRef]
- Milella, A.; Marani, R.; Petitti, A.; Reina, G. In-field high throughput grapevine phenotyping with a consumer-grade depth camera. Computers and electronics in agriculture 2019, 156, 293–306. [Google Scholar] [CrossRef]
- Gutiérrez, S.; Fernández-Novales, J.; Diago, M.P.; Tardaguila, J. On-the-go hyperspectral imaging under field conditions and machine learning for the classification of grapevine varieties. Frontiers in plant science 2018, 9, 1102. [Google Scholar] [CrossRef] [PubMed]
- Pérez, D.S.; Bromberg, F.; Diaz, C.A. Image classification for detection of winter grapevine buds in natural conditions using scale-invariant features transform, bag of features and support vector machines. Computers and electronics in agriculture 2017, 135, 81–95. [Google Scholar] [CrossRef]
- Nasiri, A.; Taheri-Garavand, A.; Fanourakis, D.; Zhang, Y.D.; Nikoloudakis, N. Automated grapevine cultivar identification via leaf imaging and deep convolutional neural networks: a proof-of-concept study employing primary iranian varieties. Plants 2021, 10, 1628. [Google Scholar] [CrossRef] [PubMed]
- Oerke, E.C.; Herzog, K.; Toepfer, R. Hyperspectral phenotyping of the reaction of grapevine genotypes to Plasmopara viticola. Journal of Experimental Botany 2016, 67, 5529–5543. [Google Scholar] [CrossRef] [PubMed]
- Pouzoulet, J.; Scudiero, E.; Schiavon, M.; Rolshausen, P.E. Xylem vessel diameter affects the compartmentalization of the vascular pathogen Phaeomoniella chlamydospore in grapevine. Frontiers in Plant Science 2017, 8, 1442. [Google Scholar] [CrossRef] [PubMed]
- Shruthi, U.; Nagaveni, V.; Raghavendra, B. A review on machine learning classification techniques for plant disease detection. 2019 5th International conference on advanced computing & communication systems (ICACCS). IEEE, 2019, pp. 281–284. [CrossRef]
- Padol, P.B.; Yadav, A.A. SVM classifier-based grape leaf disease detection. 2016 Conference on Advances in signal processing (CASP). IEEE, 2016, pp. 175–179. [CrossRef]
- Gutierrez, S.; Tardaguila, J.; Fernandez-Novales, J.; Diago, M.P. Support vector machine and artificial neural network models for the classification of grapevine varieties using a portable NIR spectrophotometer. PloS one 2015, 10, e0143197. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.; Yang, J.; Li, Z.; Qi, F. Grape disease image classification based on lightweight convolution neural networks and channelwise attention. Computers and Electronics in Agriculture 2020, 178, 105735. [Google Scholar] [CrossRef]
- Ghoury, S.; Sungur, C.; Durdu, A. Real-time diseases detection of grape and grape leaves using faster r-cnn and ssd mobilenet architectures. International conference on advanced technologies, computer engineering and science (ICATCES 2019), 2019, pp. 39–44.
- Chuanlei, Z.; Shanwen, Z.; Jucheng, Y.; Yancui, S.; Jia, C. Apple leaf disease identification using genetic algorithm and correlation-based feature selection method. International Journal of Agricultural and Biological Engineering 2017, 10, 74–83. [Google Scholar]
- Barbedo, J.G.A. Detection of nutrition deficiencies in plants using proximal images and machine learning: A review. Computers and Electronics in Agriculture 2019, 162, 482–492. [Google Scholar] [CrossRef]
- Rangel, B.M.S.; Fernandez, M.A.A.; Murillo, J.C.; Ortega, J.C.P.; Arreguín, J.M.R. KNN-based image segmentation for grapevine potassium deficiency diagnosis. 2016 International Conference on Electronics, Communications and Computers (CONIELECOMP). IEEE, 2016, pp. 48–53. [CrossRef]
- Mezzasalma, V.; Sandionigi, A.; Guzzetti, L.; Galimberti, A.; Grando, M.S.; Tardaguila, J.; Labra, M. Geographical and cultivar features differentiate grape microbiota in Northern Italy and Spain vineyards. Frontiers in microbiology 2018, 9, 946. [Google Scholar] [CrossRef]
- Zabawa, L.; Kicherer, A.; Klingbeil, L.; Milioto, A.; Topfer, R.; Kuhlmann, H.; Roscher, R. Detection of single grapevine berries in images using fully convolutional neural networks. Proceedings of the IEEE/CVF conference on computer vision and pattern recognition workshops, 2019, pp. 0–0.
- Kasinathan, T.; Singaraju, D.; Uyyala, S.R. Insect classification and detection in field crops using modern machine learning techniques. Information Processing in Agriculture 2021, 8, 446–457. [Google Scholar] [CrossRef]
- Magalhães, S.A.; dos Santos, F.N.; Martins, R.C.; Rocha, L.F.; Brito, J. Path planning algorithms benchmarking for grapevines pruning and monitoring. Progress in Artificial Intelligence: 19th EPIA Conference on Artificial Intelligence, EPIA 2019, Vila Real, Portugal, –6, 2019, Proceedings, Part II 19. Springer, 2019, pp. 295–306. 3 September. [CrossRef]
- Botterill, T.; Paulin, S.; Green, R.; Williams, S.; Lin, J.; Saxton, V.; Mills, S.; Chen, X.; Corbett-Davies, S. A robot system for pruning grape vines. Journal of Field Robotics 2017, 34, 1100–1122. [Google Scholar] [CrossRef]
- Fuentes, S.; Summerson, V.; Gonzalez Viejo, C.; Tongson, E.; Lipovetzky, N.; Wilkinson, K.L.; Szeto, C.; Unnithan, R.R. Assessment of smoke contamination in grapevine berries and taint in wines due to bushfires using a low-cost E-nose and an artificial intelligence approach. Sensors 2020, 20, 5108. [Google Scholar] [CrossRef] [PubMed]
- Mendes, J.; Peres, E.; Neves dos Santos, F.; Silva, N.; Silva, R.; Sousa, J.J.; Cortez, I.; Morais, R. VineInspector: The Vineyard Assistant. Agriculture 2022, 12. [Google Scholar] [CrossRef]
| DATASETS | DISEASES | CLASS | INFO | |||||
| HEALTHY | GY | ESCA | MILDEW | LEAFROLL | OTHERS | |||
| 1 | 2078 | Grape Varieties + Chemical Data) |
||||||
| 2 | 423 | 1383 | 1180+1076 | Black Rot + Leaf-blight | ||||
| 3 | 220 | 240 | 210+2010 | Leaf-blight + Black Rot | ||||
| 4 | 282* | *Mentioned the total number of images regardless of healthy or diseased) |
||||||
| 5 | 84 | 134 | 1383 | 1180+1076 | Expanded version of 2 | |||
| 6 | 881 | 887 | ||||||
| 7 | 135 | 156 | 108 + 97 | Red Blotch images and combined Red Blotch and Leafroll images. |
||||
| 8 | Statistical Data | |||||||
| Cite | Data Format Techniques | Notes | ||||||
| RGB | Spectral | Thermal | UAV | |||||
| Hyper | Multi | |||||||
| DISEASE | GY | [17] | ✓ | |||||
| [26] | ✓ | ✓ | PROSPECT model | |||||
| [27] | ✓ | |||||||
| DOREE | [28] | ✓ | Autoencoders - dimensionality reduction |
|||||
| [29] | ✓ | |||||||
| [30] | ✓ | |||||||
| ESCA | [31] | ✓ | ✓ | VNIR and SWIR cameras (from RGB) | ||||
| [32] | ✓ | ImageDataGenerator | ||||||
| [33] | ✓ | ✓ | ||||||
| MILDEW | [21] | Geographical data analysis (statistics) | ||||||
| [16] | ✓ | |||||||
| [34] | ✓ | |||||||
| [35] | ✓ | ✓ | ||||||
| [36] | ✓ | Meteorological data | ||||||
| [37] | ✓ | ✓ | ||||||
| LEAFROLL | [19] | ✓ | Laboratory lighting | |||||
| [38] | ✓ | |||||||
| [39] | ✓ | Airborne imaging (NASA AVIRIS) | ||||||
| PIERCE’S | [12] | ✓ | ||||||
| ROOT ROT | [40] | ✓ | ||||||
| GENERAL | [41] | ✓ | ✓ | |||||
| [42] | ✓ | |||||||
| [43] | ✓ | |||||||
| [44] | ✓ | |||||||
| [45] | ✓ | |||||||
| [46] | ✓ | |||||||
| [47] | ✓ | |||||||
| [48] | ✓ | ✓ | ||||||
| [49] | ✓ | ✓ | Anomaly detection | |||||
| WATER - STATUS | [50] | ✓ | ||||||
| [51] | ✓ | ✓ | ||||||
| PLANT DEFICIENCIES | [52] | ✓ | Raspberry Pi 3 | |||||
| CLASSIFICATION | [53] | ✓ | 3D Depth Camera | |||||
| [20] | ✓ | |||||||
| [54] | ✓ | In motion images | ||||||
| [55] | ✓ | In natural conditions | ||||||
| [56] | ✓ | |||||||
| Cite | Machine Learning Techniques | Best ACC |
Notes | |||||||
| KNN | SVM | ANN | CNN | Decision Trees |
Other | |||||
| DISEASE | GY | [17] | ✓ | 99.33% | ResNet101 | |||||
| [26] | ✓ | 99% | Back Propagation Neural Network |
|||||||
| [27] | rRBF, MLP | 96% | Varies between tests (e.g field, BN-GY) |
|||||||
| DOREE | [28] | ✓ | 83% | |||||||
| [29] | ✓ | 96% | ||||||||
| [30] | ✓ | 96+% | ||||||||
| ESCA | [31] | rRBF | 95% | Varies depending on the type of data and the year |
||||||
| [32] | ✓ | 99.48% | ||||||||
| [33] | ✓ | 95.8% | ||||||||
| MILDEW | [21] | ✗ | C5.0 | 70+% | Better Accuracy when tested diseases separately |
|||||
| [16] | ✓ | 83% | Modified ResNet | |||||||
| [34] | ✓ | 94% | ||||||||
| [35] | ✗ | ✓ | 82% | |||||||
| [36] | ✓ | ✗ | 81.6% | |||||||
| [37] | ✓ | 92+% | SegNet Architecture | |||||||
| LEAFROLL | [19] | ✓ | ✗ | 87% | 3D-CNN, Binary classification on a symptomatic dataset |
|||||
| [38] | ✓ | 89.93% | Least-squares SVM | |||||||
| [39] | ✓ | 87% | Random Forest | |||||||
| PIERCE’S | [12] | ✓ | 99.2% | AlexNet Architecture | ||||||
| ROOT ROT | [40] | NB | 90% | Healthy vs Diseased plants | ||||||
| GENERAL | [41] | ✓ | 84.16% | Average precision, MobileNets, Inseption-V2 |
||||||
| [42] | ✗ | 99.17% | UnitedModels based on 4 models |
|||||||
| [43] | ✓ | ✓ | ✓ | 75% | Preprocessing: SVM Prediction: 3D-CNN, RF |
|||||
| [44] | ✗ | ✓ | 99% | LS-SVM, cosine KNN Several steps from preprocessing to prediction |
||||||
| [45] | ✓ | 98% | Vanilla CNN 100% accuracy with ensemble model |
|||||||
| [46] | ✓ | 92.5% | AlexNet + Transfer Learning |
|||||||
| [47] | ✓ | 81.1% | Focused in speed and multiple Disease Detection |
|||||||
| [48] | ✓ | 93.72% | VddNet | |||||||
| [49] | - | |||||||||
| WATER - STATUS | [50] | ✓ | 65% | Regression models, for prediction |
||||||
| [51] | ✓ | 82.9% | Pattern recognition ANN | |||||||
| PLANT DEFICIENCIES | [52] | ✗ | ✓ | 80% | ||||||
| CLASSIFICATION | [53] | ✓ | 91.52% | VGG19 | ||||||
| [20] | ✓ | ✗ | 97.6% | Cubic SVM | ||||||
| [54] | ✗ | MLP | 99% | |||||||
| [55] | ✓ | 90.8% | ||||||||
| [56] | ✓ | 99.11% | Modified VGG16 | |||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




