Submitted:
15 July 2024
Posted:
16 July 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Theory and Formula
3. Experimental Setup
4. Result Discussions
5. Conclusions
Authors’ Disclosures of Potential Conflicts of Interest
Grant/Funding support
Financial disclosures
Clinical trial registration
Acknowledgements
References
- Fischer, M., Census and evaluation of p53 target genes. Oncogene (2017) 36, 3943–3956.
- Sabapathy, K and Lane, DP (2019) Understanding p53 functions through p53 antibodies. J. Mol. Cell Biol 11, 1-35.
- Campo, E., Cymbalista, F., Ghia, P., Jäger, U., Pospisilova, S., Rosenquist, R., Schuh, A and Stilgenbauer, S.; (2018) TP53 aberrations in chronic lymphocytic leukaemia: an overview of the clinical implications of improved diagnostics. Haematologica103, 1956-1968. [CrossRef]
- Gonzalez, D., Martinez, P., Wade, R., Hockley, S.;,Oscier, D., Matutes, E., Dearden, CE., Richards, SM., Catovsky, D and Morgan, GJ., (2011) Mutational status of the TP53 gene as a predictor of response and survival in patients with chronic lymphocytic leukaemia: results from the LRF CLL4 trial. J Clin Oncol 29, 2223-2229. [CrossRef]
- Cordone, I., Masi, S., Mauro, FR., Soddu, S., Morsilli, O., Valentini, T., Vegna, ML., Guglielmi, C., Mancini, F., Giuliacci, S., Sacchi, A., Mandelli F and Foa, R.; (1998,) p53 expression in B-cell chronic lymphocytic leukaemia: a marker of disease progression and poor prognosis. Blood 91, 4342-4349.
- Hallek, M., Cheson, BD., Catovsky, D., Caligaris-Cappio, F., Dighiero, G., Döhner, H., Hillmen, P., Keating, M., Montserrat, E., Chiorazzi, N., Stilgenbauer, S., Rai, KR., Byrd, JC., Eichhorst, B., O'Brien, S., Robak, T., Seymour, JF and Kipps, TJ.; (2018). I will guidelines for diagnosis, indications for treatment, response assessment, and supportive management of CLL. Blood 21;131(25):2745-2760. [CrossRef]
- Cordone Masi, S., Mauro, FR., Soddu, S and Morsilli O.; (2019) p53 Expression in B-Cell Chronic Lymphocytic Leukemia: A Marker of Disease Progression and Poor Prognosis. Blood 91, 4342-4349.
- Bosch, F and Dalla- Favera., R (2022. Chronic lymphocytic leukaemia: from genetics to treatment. Nature review. Clinical Oncology; www.nature.
- Spiro, S., Galton, D.A.G., Wiltshaw, E and Lohmman., R.C.; (1975) Follicular Lymphoma: A survey of 75 cases with special references to the Syndrome resembling Chronic Lymphocytic Leukemia. Br. J. Cancer, Suppl. II, 60.
- Murnyák, B and Hortobágyi, T.; (2016) Immunohistochemical correlates of TP53 somatic mutations in cancer. Oncotarget 40, :64910-64920.
- Sellmann, L., Carpinteiro, A., Nückel, H., Scholtysik, R and Siemer, D., (2012) p53 protein expression in chronic lymphocytic leukemia. Leuk Lymphoma 53, 1282-1298.
- Allende-Vega, N., Krzywinska, E., Orecchioni,S., Lopez-Royuela, N., Reggiani, F., Talarico, G.; Rossi, JF., Rossignol, R., Hicheri, Y., Cartron, G., Bertolini, F and Villalba, M., (2015) The presence of wild type p53 in haematological cancers improves the efficacy of combinational therapy targeting metabolism. Oncotarget 22, 19228-19245. [CrossRef]
- Demma, M.J., Wong, S., Maxwell, E and Dasmahapatra M., (2004) 24–CP-31398 Restores DNA-binding Activity to Mutant p53 in Vitro but Does Not Affect p53 Homologs p63 and p73. The Journal of Biological Chemistry 279, 4, 45887–45896.
- Carpinteiro A., Nückel H., Scholtysik R and Siemer D., (2012) p53 protein expression in chronic lymphocytic leukemia, Blood Cancer Journal. 53(7):1282-1288.
- S.H. Van der Burg., Cock K and Menon A.G (2001) Long lasting p53-specific T cell memory responses in the absence of anti-p53 antibodies in patients with resected primary colorectal cancer, Eur. J. Immunol. 31, 146–155.
- Kondapuram, S.K., Sarvagalla, S and Coumar, M (2019) Targeting autophagy with small molecules for cancer therapy. Journal of Cancer Metastasis and Treatment 1, 23-25.
- Shahjahani, M.; Mohammadiasl, J.; Noroozi, F.; Seghatoleslami, M.; Shahrabi. S.; Saba, and Saki, N (2015) Molecular basis of chronic lymphocytic leukaemia diagnosis and prognosis. Cell Oncol 38, :93-109. [CrossRef]
| CLL – Age patients |
CLL stage I/II, (n=17 patients) P-53 protein concentration in reactive lymphocytes B |
CLL stage III/IV (n = 3 patients] Percentage of p53 isoform proteins |
p value |
| The patients age ranging from 39 to 85 years. |
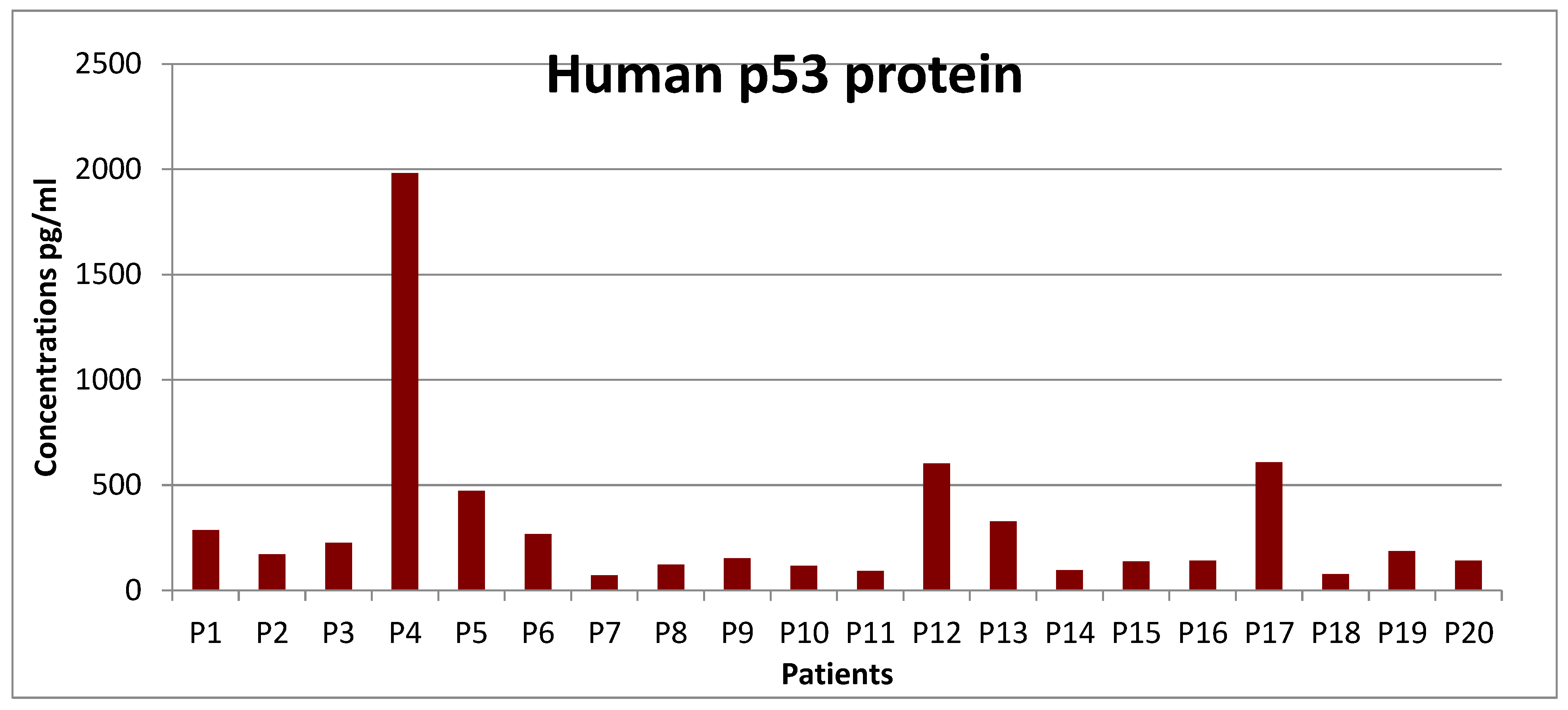
The average p-53 protein concentration in CLL, 16.76 μg / dL |
P-5 izoform proteins with elevated values was present in 15% (3 of 20 cases 2 Men = 50μg / dL and 60μg / dL, respectively 1 Female = 140μg / dL |
p value 0.034 |
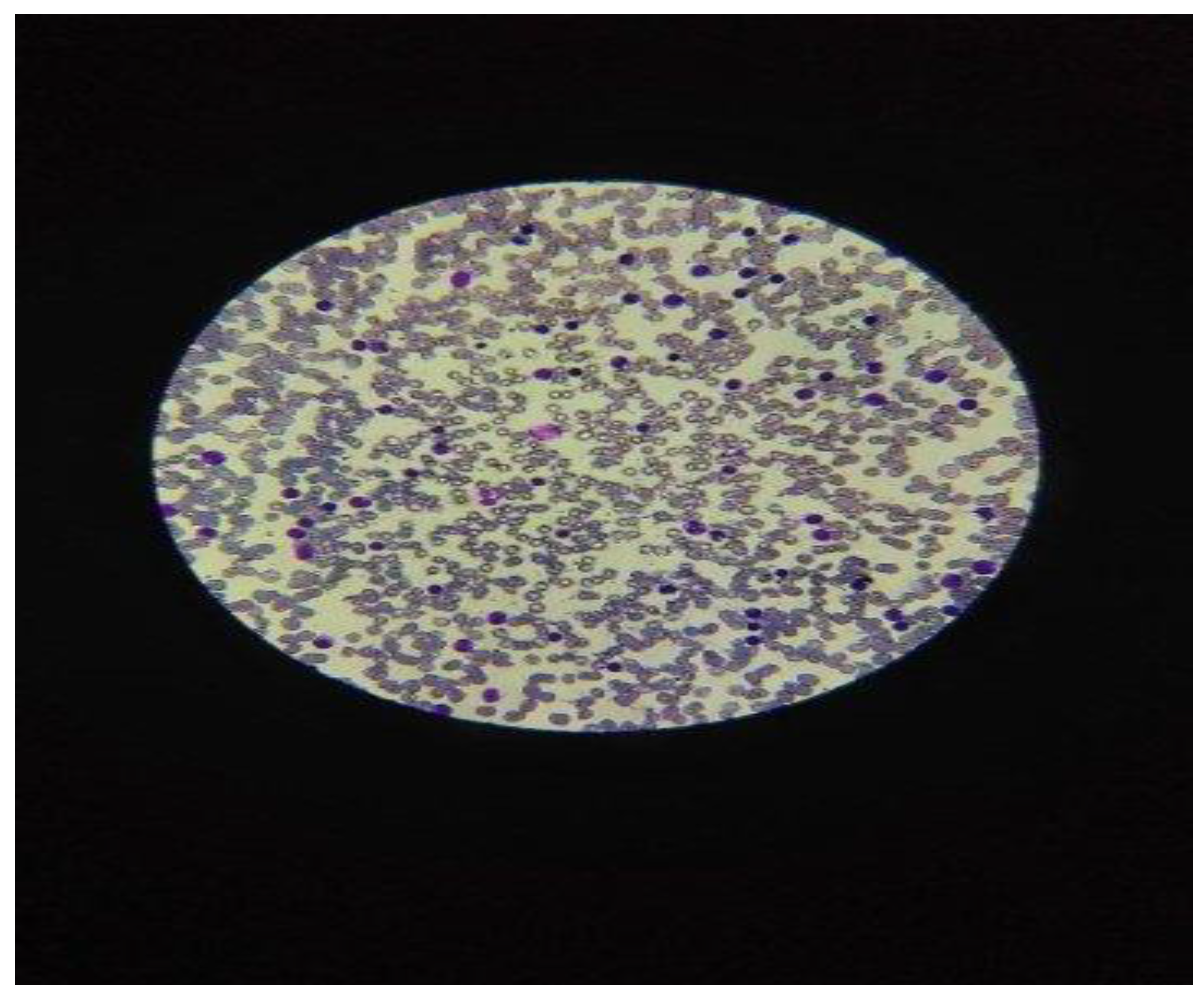
| Hematological parameters in peripheral blood | Mean values of hemogram: No. Leukocytes = 35-50 x 10³/µL Hb = 11.8g / dL; Platelet = 140 x 10³/µL Lymphocytes = 65-80% |
No. Leucocytes = 250-500 x 10³/µL Hb = 8.6g / dL. No. platelets = 45x10³/µL Lymphocytes = 85-90% |
p value 0.05 |
| LLC - Patients | CLL stage I / II, n = 47 patients) P-53 protein concentration in reactive B Lymphocytes |
CLL stage III/IV (n = 140 patients] Percentage of p53 isoform proteins | p value |
|---|---|---|---|
| The age of patients with CLL ranged from 52 to 84 years, with a mean age of 62 years. | In the samples studied at CLL it was 16.7% (7 out of 42 cases). | The percentage of p53 positive cells ranged from 7-32%. | 0.398 |
| Hematological parameters in peripheral blood | No. Leukocyte = 35x10³ / dl Hb = 12.2g / dL Thrombocytes = 140 x 10³/µL Lymphocytes = 75-80% |
Nr. Leukocytes = 350 x10³/µL Hb = 10.8g / dL Platelets= 80 x 10³/µL Lymphocytes = 80-90% |
0.398 |
| P53 proteins form, (positive) n = 7/42 | 15% | 25% | 0.398 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




