Submitted:
14 May 2024
Posted:
14 May 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Trimming Paired-End Reads (Data Preparation, Adapter Trimming, and Quality Filtering)
- 𝑄Q represents the Phred quality score,
- 𝑃P represents the probability of the base being called incorrectly.
2.2. Pairing Trimmed Reads
2.3. Mapping to Host Reference Genome (Reference Genome Selection and Mapping Algorithm)
- Match ScoreMatch Score represents the score assigned to a matching base pair,
- Mismatch PenaltyMismatch Penalty represents the penalty assigned to a mismatched base pair.
2.4. De Novo Assembly of Unmapped Reads
2.5. Sequence Blast Similarity Search from Assembled Contigs
2.5.1. Database Selection
- Nucleotide Database for Viruses: This database contains nucleotide sequences of viruses obtained from various sources, including NCBI GenBank and other curated repositories.
- Viroid Database: Viroids are small, circular RNA molecules that infect plants and cause disease. The viroid database contains sequences of known viroid species and strains.
- Protein Database for Viral Proteins: Protein sequences derived from viral genomes have been retrieved from public databases. These sequences represent the proteome of various viruses and are essential for protein-level analysis.
2.5.2. BLAST Parameters Optimization
- E-value Threshold: The E-value represents the expected number of chance alignments that would occur randomly in a database of a particular size. Lower E-values indicate higher confidence in the match. Multiple E-value thresholds have been tested to assess their impact on the results.
- Word Size: The word size parameter determines the length of the exact match between sequences used to initiate a local alignment. Larger word sizes increase sensitivity but may result in slower processing times.
- Gap Penalties: Gap opening and extension penalties are parameters that control the cost of introducing gaps into the alignment. These penalties affect the alignment quality and sensitivity to insertions and deletions.
2.5.3. BLAST Search using AI Tools
- NcbiblastnCommandline and NcbiblastxCommandline: These functions from the Biopython library are used to interface with the BLAST+ command-line tool.
- Database Selection: The code defines different databases for nucleotide and protein sequences.
- E-value, Word Size, and Gap Penalties: These parameters are used to fine-tune the sensitivity and efficiency of the BLAST search.
- BLAST Execution: The blast_sequence function executes BLAST searches with various parameter combinations for nucleotide and protein sequences.
2.6. Filtering Criteria for BLAST Results and Implementation of Filters
- Alignment Length: This criterion considers the length of the aligned region between the query and subject sequences. Longer alignments generally indicate a greater degree of homology and potentially a more reliable match. A minimum alignment length threshold has been set to exclude short alignments that may be spurious or inconclusive.
- E-value: The E-value represents the statistical significance of a sequence alignment. Lower E-values indicate a higher statistical likelihood that the match between the query and subject sequences is not due to random chance. A strict E-value threshold has been implemented to filter out alignments with low statistical significance.
- Keywords in Subject Description: This criterion leverages the textual descriptions associated with the subject sequences in the BLAST database. Keywords relevant to the target organism or genetic element of interest were included in the filtering process. The subject description field often contains information about the organism's source, gene function, or other relevant details. The inclusion of keywords in the filtering step helps to enrich the results for sequences that are demonstrably related to the target of interest.
- Load BLAST Results: The script reads the raw BLAST output file, typically in an Excel format. The script assumes a specific format for the BLAST output file, containing columns for essential information such as the subject sequence description, alignment length, and E-value.
- Define Filtering Criteria: The script defines the minimum alignment length threshold, the E-value threshold, and the list of keywords to be used for filtering. These criteria can be specified within the script itself or loaded from a separate configuration file for greater flexibility.
- Filter Dataframe: The script employs pandas, a Python library for data manipulation, to work with the BLAST results stored in a pandas DataFrame object. The DataFrame is filtered based on predefined criteria. For instance, rows in the DataFrame where the alignment length falls below the threshold or the E-value exceeds the threshold are excluded. In addition, rows are filtered out where the subject description does not contain any of the specified keywords.
- Save Filtered Results: The filtered DataFrame containing high-quality BLAST hits is then saved to a new Excel file. This file can be used for further analysis or downstream applications.
2.7. Retrieval of Virus Sequences (Accession Number Extraction, Sequence Retrieval using NCBI Entrez Utilities, FASTA Format and Sequence Handling, Saving Fetched Sequences, Error Handling, and Software Dependencies)
2.8. Mapping to Reference Genome (Sequencing Data Preprocessing, Read Mapping, and Mapping Result Assessment) and Consensus Sequence Generation
- -ax map-ont: This aligns reads in splice-aware mode, which is suitable for RNA viruses with potential splicing events.
- -m <reference_genome.fasta>: Specifies the reference genome FASTA file for alignment.
- The mapping results were evaluated using several metrics, including:
- Mapping rate: The percentage of reads that were successfully mapped to the reference genome.
- Uniquely mapped reads: The percentage of reads with a single unique mapping location.
- Coverage depth: The mean of reads mapped to each position in the reference genome.
- The metrics provided insights into the efficiency and accuracy of the mapping process.
- -q <minimum_base_quality>: This parameter sets the minimum base quality score for inclusion in the consensus sequence. For example, -q 30 would include bases with a Phred score ≥ 30).
- -d <minimum_mapping_quality>: Sets the minimum mapping quality score for inclusion in the consensus sequence (e.g., -d 20 for reads with mapping quality ≥ 20).
2.9. SNPs Discovery
- Alignment Loading: The script begins by loading the alignment data generated from the sequencing reads and a reference genome. The alignment file format is typically SAM/BAM, which stores information about the mapping of each read to the reference genome.
- Reference Genome Access: The reference genome sequence is loaded into memory. This reference serves as the basis for the identification of SNPs.
- Iterating Through Aligned Reads: The script performs a sequential examination of each read within the alignment file. Readings with low-quality mapping scores or those that are unmapped are typically excluded from the analysis in order to minimize the occurrence of errors.
- Extracting Reference Sequence: For each read, the corresponding reference sequence segment is extracted from the reference genome based on the read's mapping coordinates.
- Read vs. Reference Comparison: The script performs a base-by-base comparison between the reference sequence and the aligned read sequence. Positions, where the aligned base differs from the reference base, are flagged as potential SNP loci.
- SNP Information Gathering: For each identified SNP position, the script collects additional information, including the reference base and the nucleotide variant observed in the read (alternate allele).
- Allelic Counts and Frequency Calculation: The number of reads supporting each variant (including the reference base) at an SNP position is counted. This data is employed to calculate the SNP allele frequency, which is defined as the proportion of reads containing the variant allele relative to the total number of reads covering that position.
- Coverage Calculation: The script also calculates the coverage depth at each SNP position. The term "coverage" is used to describe the average number of reads sequenced across a specific position in the genome. Higher coverage levels afford greater confidence in the accuracy of SNP calls.
3. Results and Discussion
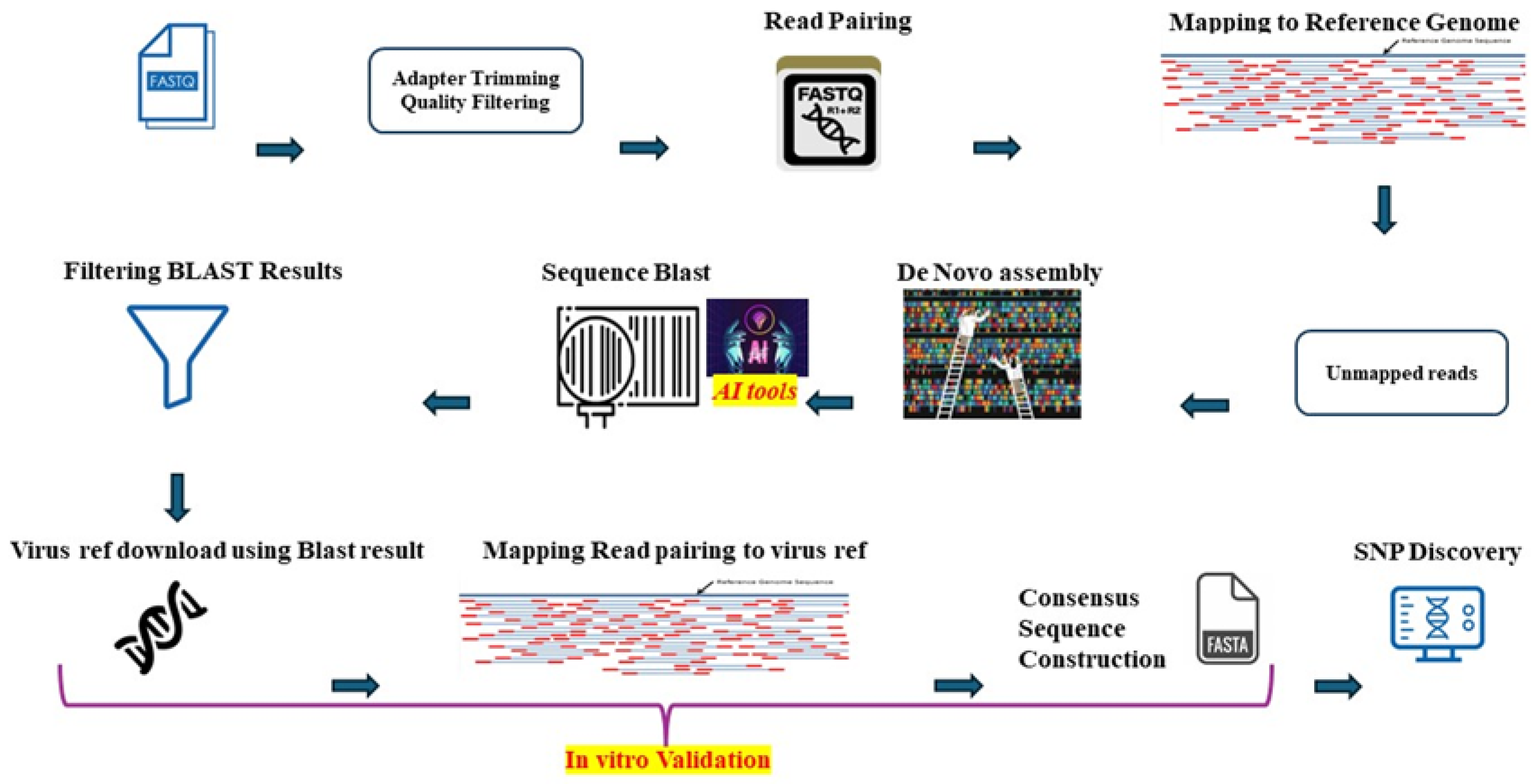
3.1. Pipeline for Detection and Validation
3.2. Quality Control and Adapter Trimming
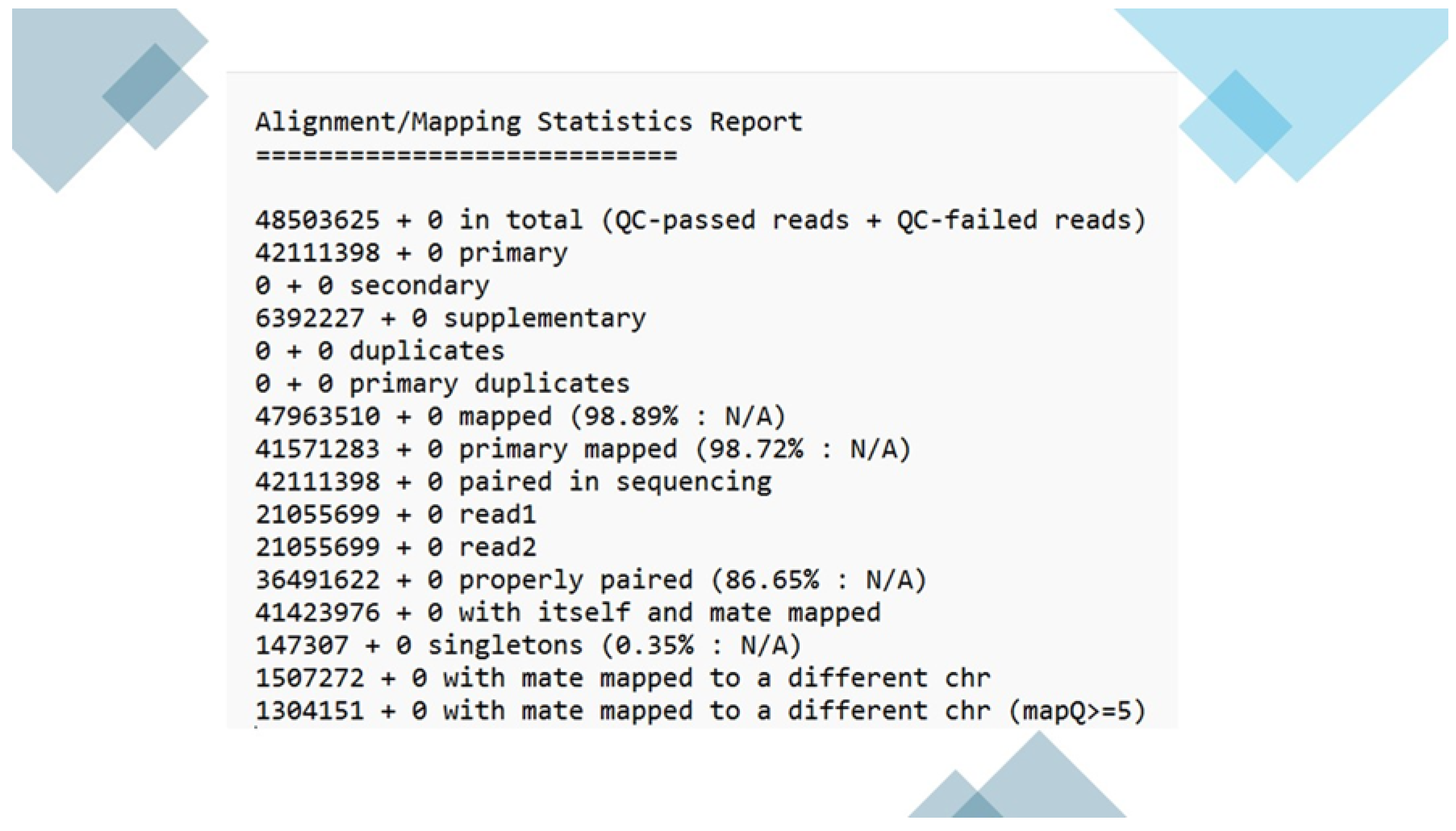
3.3. Paired-End Read Merging and Mapping to host genome and save unmapped reads
3.4. De novo Assembly of Unmapped Reads
3.5. Optimized Viral Sequence Detection: Leveraging AI-Enhanced BLAST in Contig Analysis
3.6. BLAST Parameter Optimization with Algorithmic Efficiency
- E-value Threshold: This value represents the expected number of chance alignments. Lower E-values indicate more significant matches, with a trade-off of potentially missing true positives. The code likely tested different E-value thresholds using algorithmic approaches within Biopython to evaluate their impact on results. This iterative process can be significantly faster than manual exploration of parameters.
- Word Size: The word size defines the minimum length of exact matches used to initiate local alignments. Larger word sizes enhance sensitivity but can lead to slower processing times. Finding the optimal word size depends on the specific dataset and the desired balance between speed and accuracy. Biopython's functionalities allow the code to explore different word size options and select the most efficient value for the data at hand. The study on marine DNA virus communities discusses the impact of k-mer sizes on assembly and taxonomic profiling. This is analogous to the word size parameter in BLAST, where larger word sizes can resolve more repeat regions, akin to larger k-mers providing higher N50 values and average contig lengths [22]. Both parameters are crucial for the accuracy and efficiency of sequence analysis.
- Gap Penalties: Penalties are assigned for introducing gaps (insertions or deletions) in alignments. Adjusting these penalties influences the alignment quality and sensitivity to insertions and deletions within sequences. The code can employ optimization algorithms to find the penalty settings that lead to the most informative alignments for the specific contigs being analyzed.
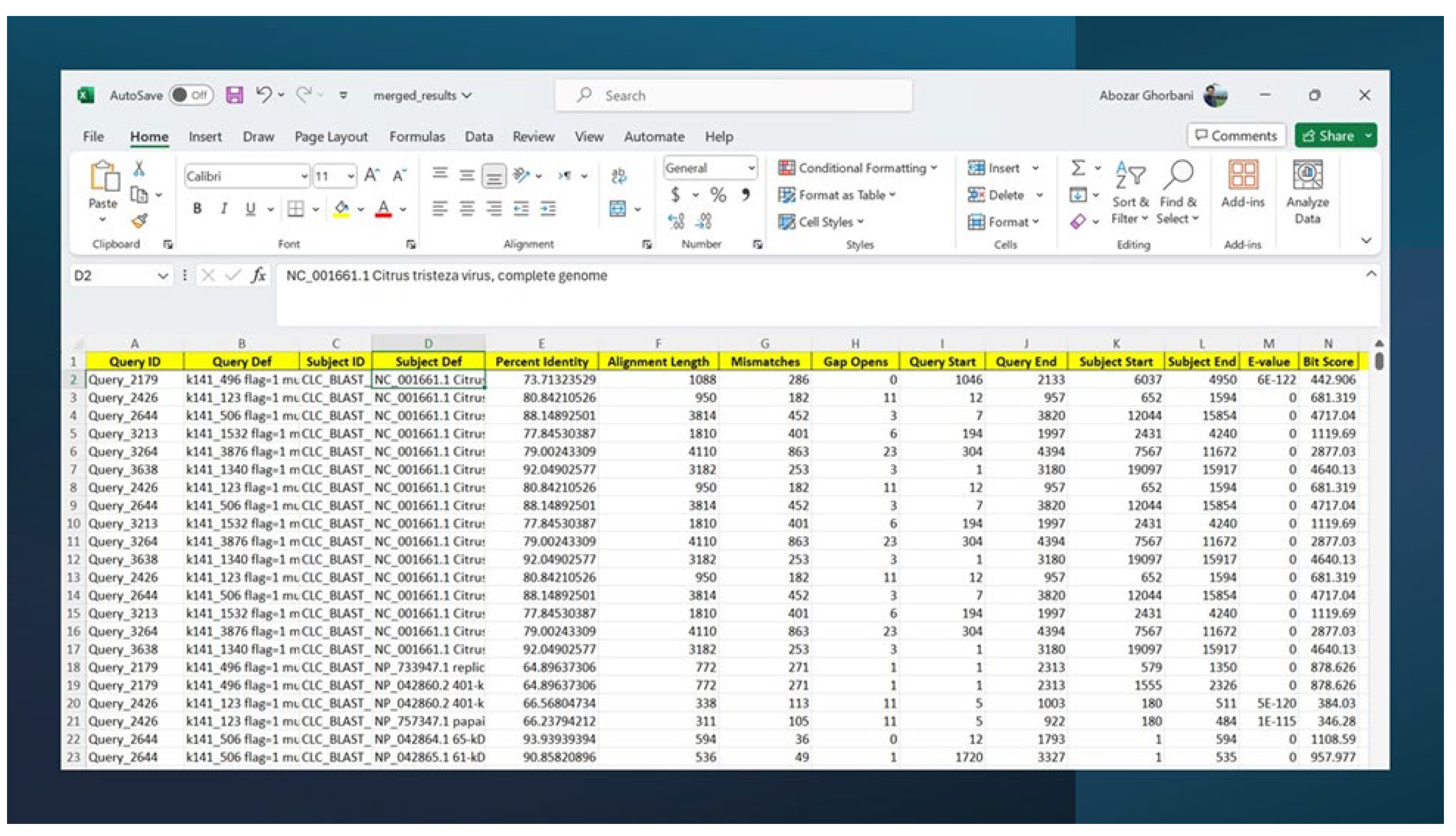
3.7. BLAST Search Results and Evidence for Citrus tristeza virus
3.8. Retrieval of Complete Viral Sequences
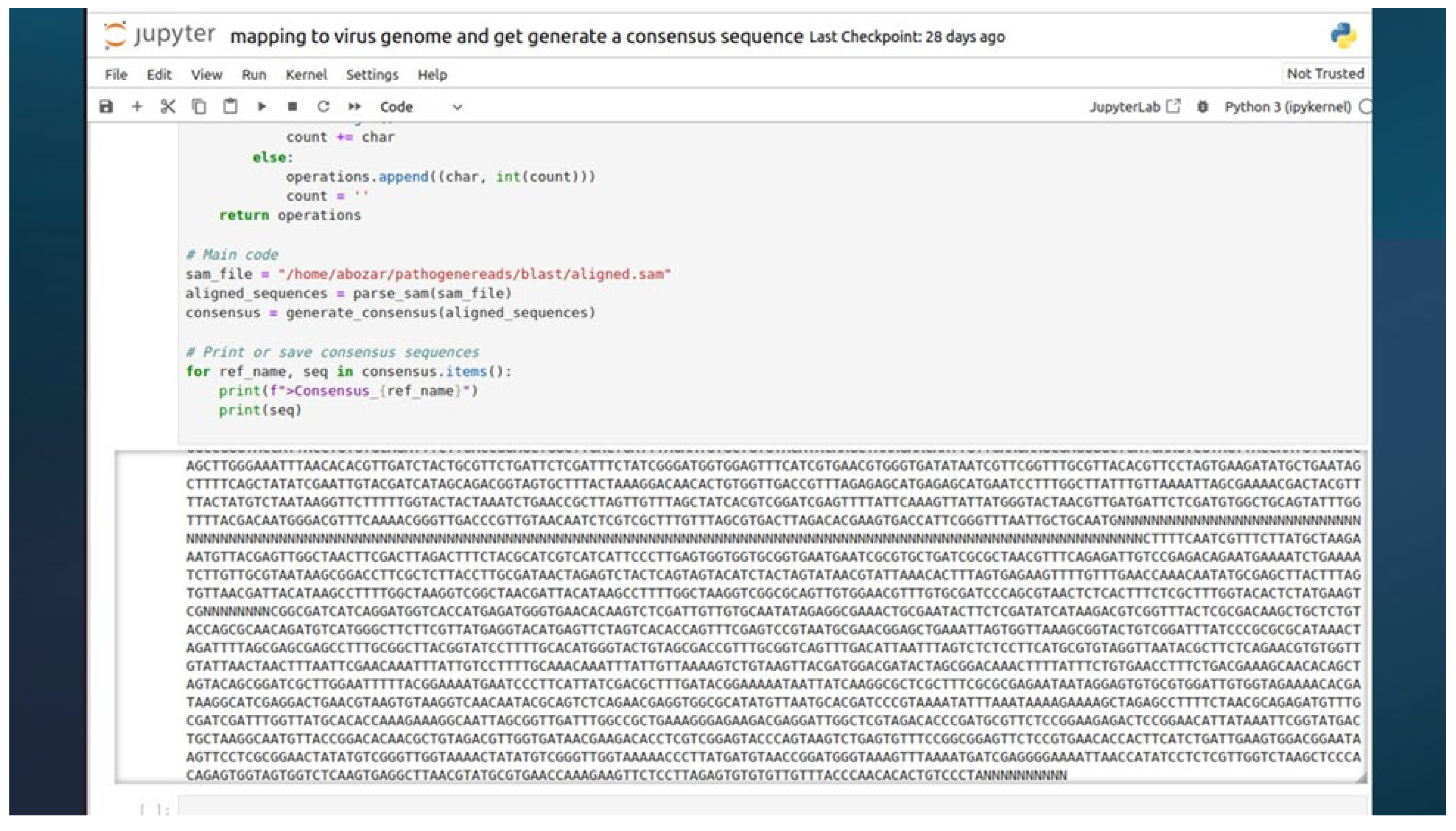
3.9. Mapping and Consensus Sequence Generation
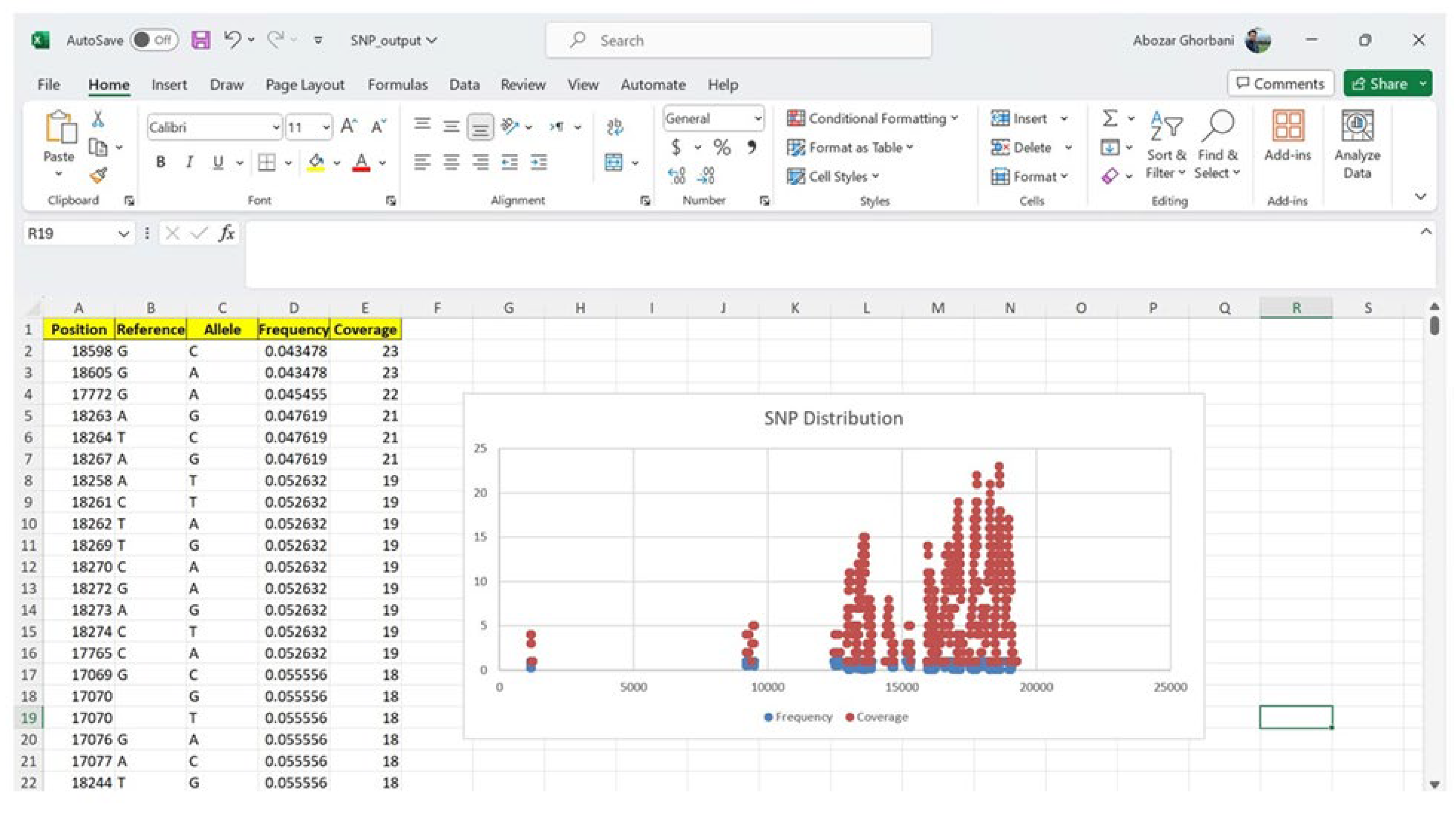
3.10. SNP Discovery Algorithm and Consensus Sequence Analysis
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Cassedy, A., A. Parle-McDermott, and R. O’Kennedy, Virus detection: A review of the current and emerging molecular and immunological methods. Frontiers in Molecular Biosciences, 2021. 8: P. 637559. [CrossRef]
- Zamai, L., Unveiling human non-random genome editing mechanisms activated in response to chronic environmental changes: I. Where might these mechanisms come from and what might they have led to? Cells, 2020. 9(11): P. 2362. [CrossRef]
- Nogales, A. and M. L. DeDiego, Host single nucleotide polymorphisms modulating influenza A virus disease in humans. Pathogens, 2019. 8(4): P. 168. [CrossRef]
- Wang, X.; et al., Detection of respiratory viruses directly from clinical samples using next-generation sequencing: A literature review of recent advances and potential for routine clinical use. Reviews in Medical Virology, 2022. 32(5): P. e2375. [CrossRef]
- Lin, Q., P.K.-H. Tam, and C.S.-M. Tang, Artificial intelligence-based approaches for the detection and prioritization of genomic mutations in congenital surgical diseases. Frontiers in Pediatrics, 2023. 11. [CrossRef]
- Albahri, O.; et al., Systematic review of artificial intelligence techniques in the detection and classification of COVID-19 medical images in terms of evaluation and benchmarking: Taxonomy analysis, challenges, future solutions and methodological aspects. Journal of infection and public health, 2020. 13(10): P. 1381-1396. [CrossRef]
- Cob-Parro, A.C., Y. Lalangui, and R. Lazcano, Fostering Agricultural Transformation through AI: An Open-Source AI Architecture Exploiting the MLOps Paradigm. Agronomy, 2024. 14(2): P. 259. [CrossRef]
- Tuia, D.; et al., Perspectives in machine learning for wildlife conservation. Nature communications, 2022. 13(1): P. 1-15. [CrossRef]
- Ghorbani, A.; et al., Complete genome sequencing and characterization of a potential new genotype of Citrus tristeza virus in Iran. PLoS ONE, 2023. 18(6): P. e0288068. [CrossRef]
- Ghorbani, A., K. Izadpanah, and R.G. Dietzgen, Completed sequence and corrected annotation of the genome of maize Iranian mosaic virus. Archives of virology, 2018. 163: P. 767-770. [CrossRef]
- Gutiérrez, P.; et al., PVDP: A portable open source pipeline for detection of plant viruses in RNAseq data. A case study on potato viruses in Antioquia (Colombia). Physiological and Molecular Plant Pathology, 2021. 113: P. 101604. [CrossRef]
- Guerra, J.V.d.S.; et al., pyKVFinder: An efficient and integrable Python package for biomolecular cavity detection and characterization in data science. BMC bioinformatics, 2021. 22: P. 1-13.
- Ho, T. and I.E. Tzanetakis, Development of a virus detection and discovery pipeline using next generation sequencing. Virology, 2014. 471: P. 54-60. [CrossRef]
- Bush, S.J., Read trimming has minimal effect on bacterial SNP-calling accuracy. Microbial genomics, 2020. 6(12): P. e000434. [CrossRef]
- Alser, M.; et al., Technology dictates algorithms: Recent developments in read alignment. Genome biology, 2021. 22(1): P. 249.
- Rahman, A.; et al., Association mapping from sequencing reads using k-mers. Elife, 2018. 7: P. e32920. [CrossRef]
- Neumann, G.B.; et al., Unmapped short reads from whole-genome sequencing indicate potential infectious pathogens in German Black Pied cattle. Veterinary Research, 2023. 54(1): P. 95. [CrossRef]
- Usman, T.; et al., Unmapped reads from cattle RNAseq data: A source for missing and misassembled sequences in the reference assemblies and for detection of pathogens in the host. Genomics, 2017. 109(1): P. 36-42. [CrossRef]
- Kruppa, J.; et al., Virus detection in high-throughput sequencing data without a reference genome of the host. Infection, Genetics and Evolution, 2018. 66: P. 180-187. [CrossRef]
- Kutnjak, D.; et al., A primer on the analysis of high-throughput sequencing data for detection of plant viruses. Microorganisms, 2021. 9(4): P. 841. [CrossRef]
- Goodacre, N.; et al., A reference viral database (RVDB) to enhance bioinformatics analysis of high-throughput sequencing for novel virus detection. MSphere, 2018. 3(2): P. [CrossRef]
- Kim, K.E.; et al., Optimized metavirome analysis of marine DNA virus communities for taxonomic profiling. Ocean Science Journal, 2022. 57(2): P. 259-268. [CrossRef]
- Roux, S.; et al., iPHoP: An integrated machine learning framework to maximize host prediction for metagenome-derived viruses of archaea and bacteria. PLoS biology, 2023. 21(4): P. e3002083. [CrossRef]
- Torkian, B.; et al., BLAST-QC: Automated analysis of BLAST results. Environmental Microbiome, 2020. 15: P. 1-8. [CrossRef]
- Bhat, N., E.B. Wijaya, and A.A. Parikesit, Use of the “DNAChecker” algorithm for improving bioinformatics research. Makara Journal of Technology, 2019. 23(2): P. 4.
- Zheng, Y.; et al., VirusDetect: An automated pipeline for efficient virus discovery using deep sequencing of small RNAs. Virology, 2017. 500: P. 130-138. [CrossRef]
- de Vries, J.J.; et al., Benchmark of thirteen bioinformatic pipelines for metagenomic virus diagnostics using datasets from clinical samples. Journal of Clinical Virology, 2021. 141: P. 104908.
- Kieft, K., Z. Zhou, and K. Anantharaman, VIBRANT: Automated recovery, annotation and curation of microbial viruses, and evaluation of viral community function from genomic sequences. Microbiome, 2020. 8: P. 1-23. [CrossRef]
- Silva, P.P.; et al., A machine learning-based SNP-set analysis approach for identifying disease-associated susceptibility loci. Scientific Reports, 2022. 12(1): P. 15817. [CrossRef]
- Rollin, J.; et al., Detection of single nucleotide polymorphisms in virus genomes assembled from high-throughput sequencing data: Large-scale performance testing of sequence analysis strategies. PeerJ, 2023. 11: P. e15816. [CrossRef]
| Tools | Need to Install | Description |
| cutadapt | Yes | Trims adapter sequences from Illumina reads. |
| gzip | Likely Installed | Compresses and decompresses files. |
| samtools | Yes | Manipulates alignments in SAM/BAM format. |
| subprocess | Built-in | Python module to run external commands. |
| os | Built-in | Python module for interacting with the operating system. |
| random | Built-in | Python module for generating random numbers. |
| MegaHit | Yes | Performs de novo assembly of sequencing reads. |
| Biopython | Yes | (Biopython: refers to a collection of Python tools for biological analysis) |
| pandas | Yes | Used for data manipulation and analysis. |
|
NCBIBlast n/NCBIBlastx |
Yes | Performs nucleotide or protein similarity searches. |
| Entrez | Likely Installed | Part of Biopython for Entrez database access. |
| SeqIO | Likely Installed | Part of Biopython for sequence input/output. |
| pysam | Yes | For manipulating alignments in SAM/BAM format. |
| Counter | Built-in | Python module for creating collections of key-value pairs. |
| Metric | Read 1 (bp) | Read 2 (bp) | Total (bp) |
|---|---|---|---|
| Total read pairs processed | - | - | 21,066,868 |
| Read with adapter | 1,986,643 (9.4%) | 1,996,373 (9.5%) | - |
| Pairs that were too short | - | - | 11,169 (0.1%) |
| Pairs written (passing filters) | - | - | 21,055,699 (99.9%) |
| Total base pairs processed | 3,160,030,200 | 3,160,030,200 | 6,320,060,400 |
| Quality-trimmed | 3,304,806 | 3,490,032 | 6,794,838 (0.1%) |
| Total written (filtered) | 3,109,982,046 | 3,109,401,003 | 6,219,383,049 (98.4%) |
| Parameter | Value |
|---|---|
| Final contigs number | 3919 |
| Total contigs | 1773988 bp |
| Minimum contigs lengths | 269 bp |
| Maximum contigs lengths | 4394 bp |
| Avrage contigs lenghts | 452 bp |
| N50 | 429 bp |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




