Submitted:
09 May 2024
Posted:
13 May 2024
You are already at the latest version
Abstract
Keywords:
Introduction
Materials and Methods
Statistical Analyses
Results
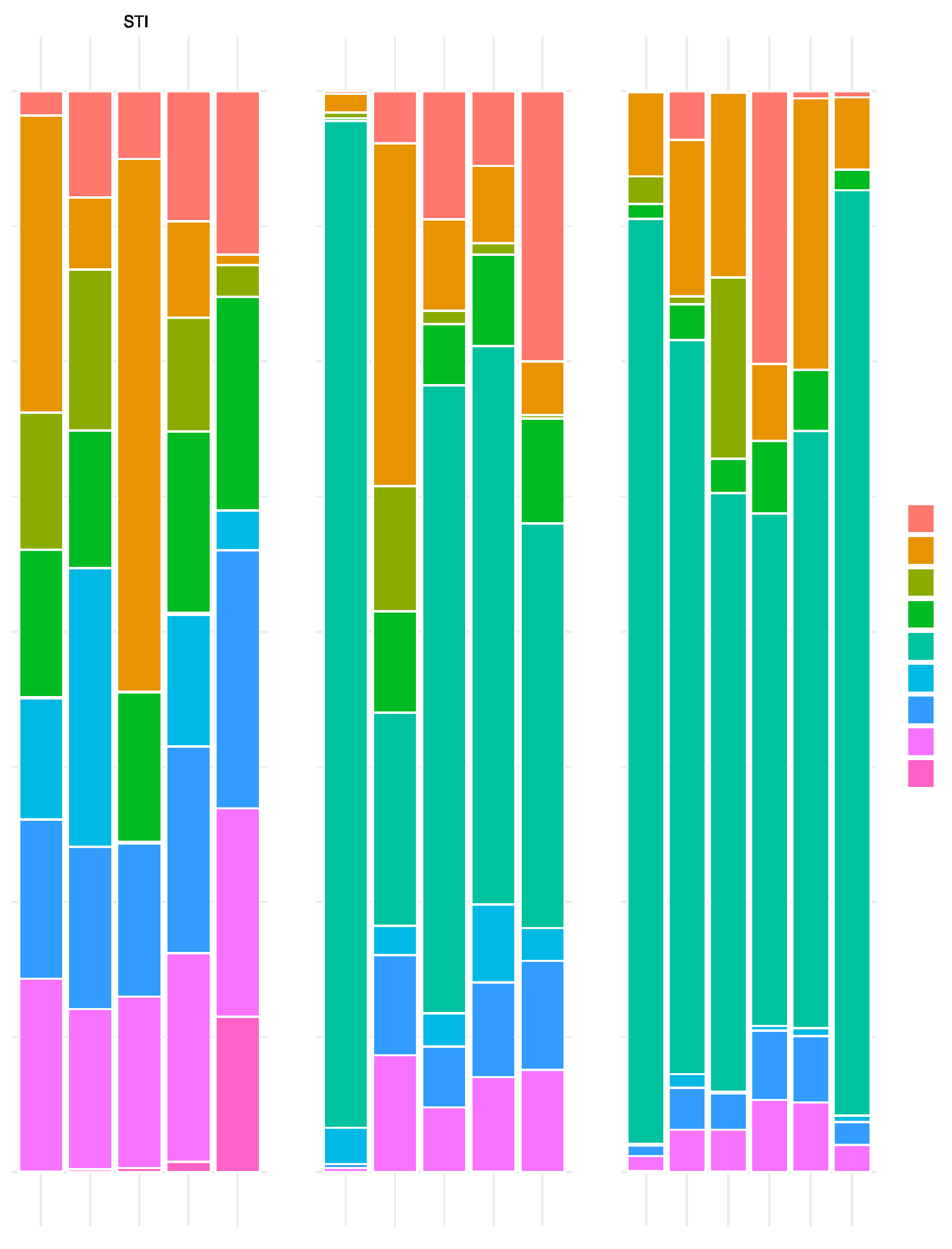
L. acidophilus Dominated the Microbial Community Structure of GAD85 and NCK56 Treated Mice
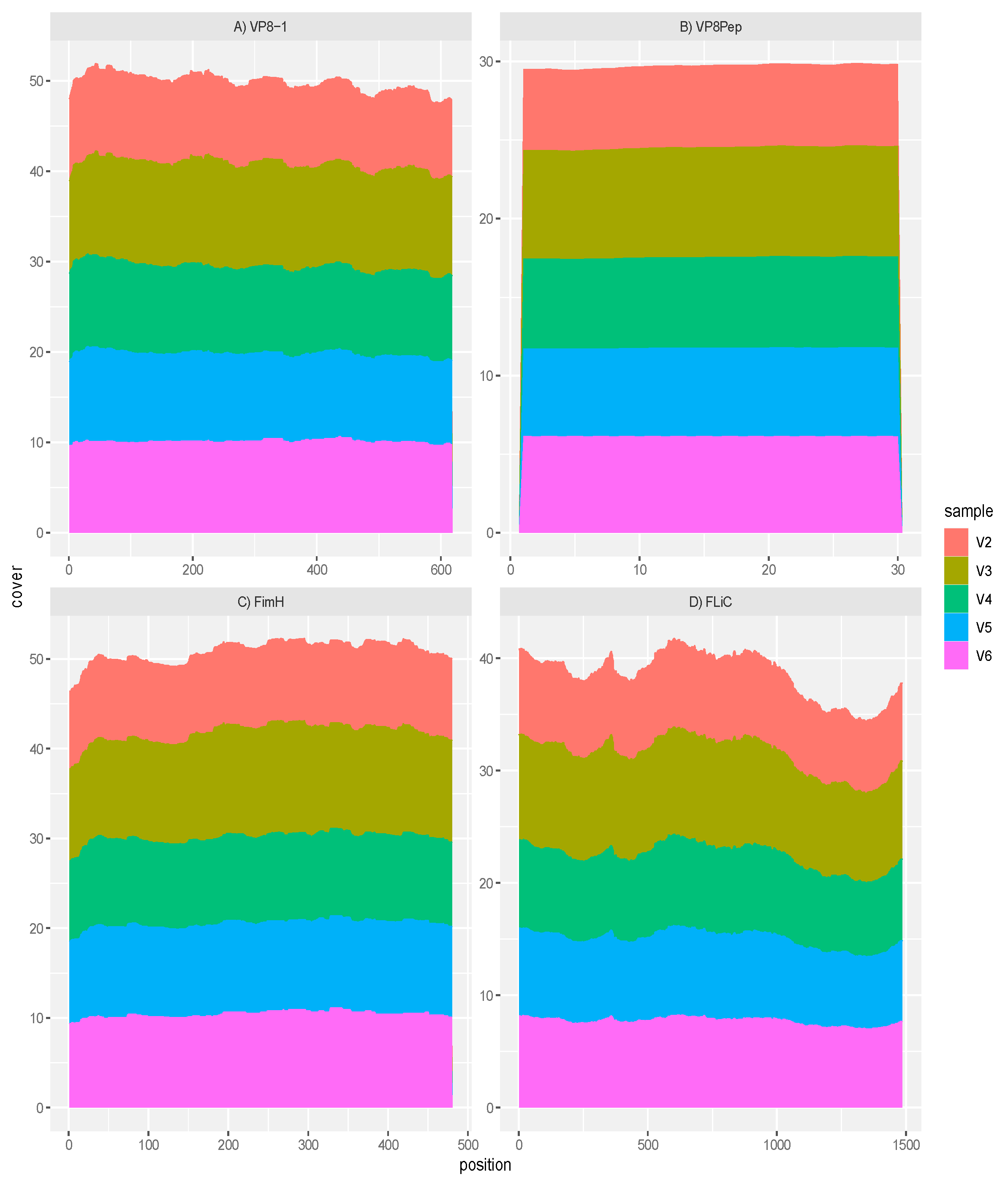
GAD85 Expresses the VP8 Antigens and the FimH and FliC Adjuvants
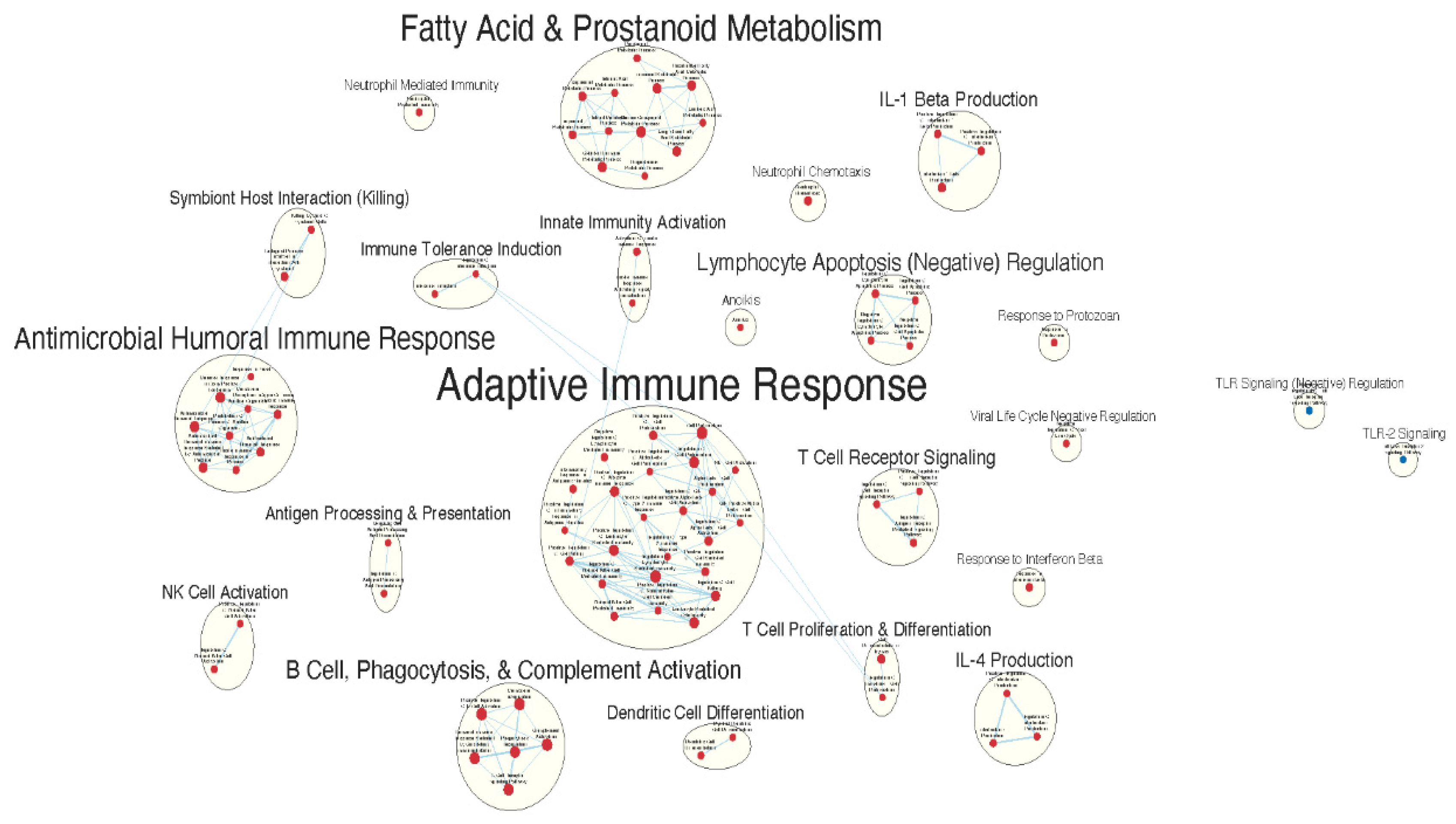
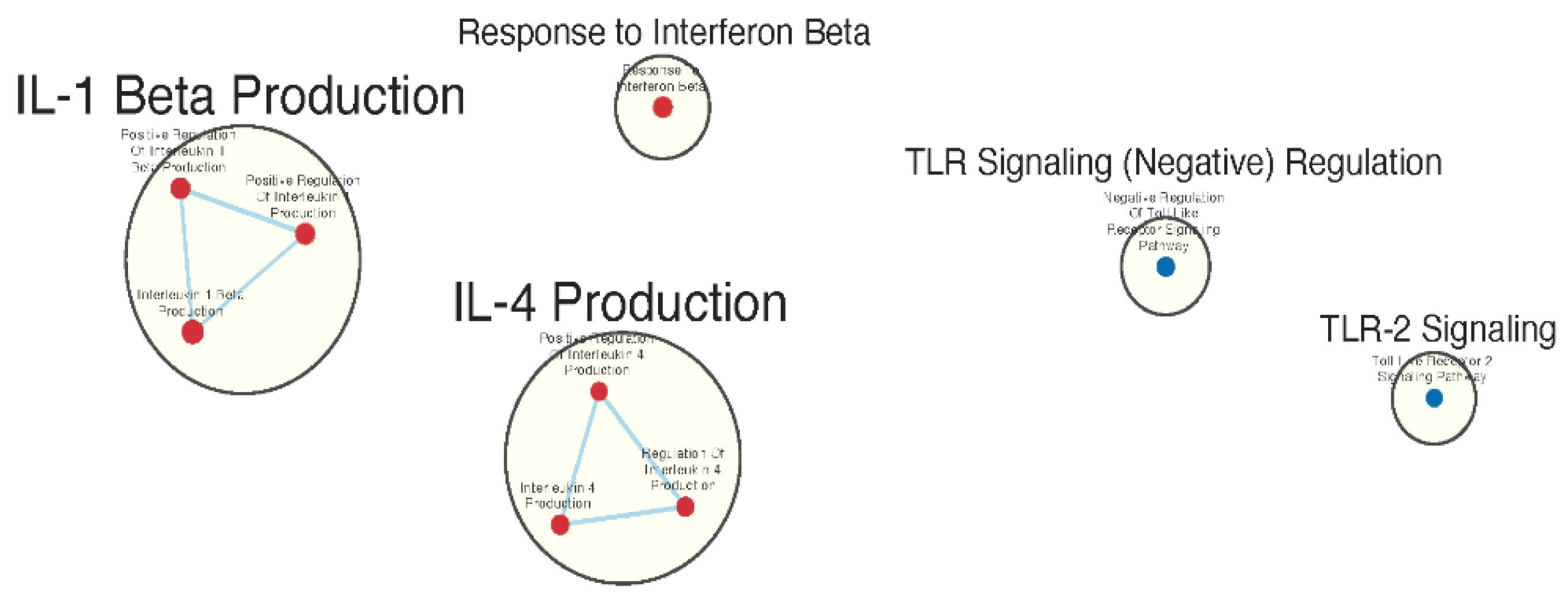
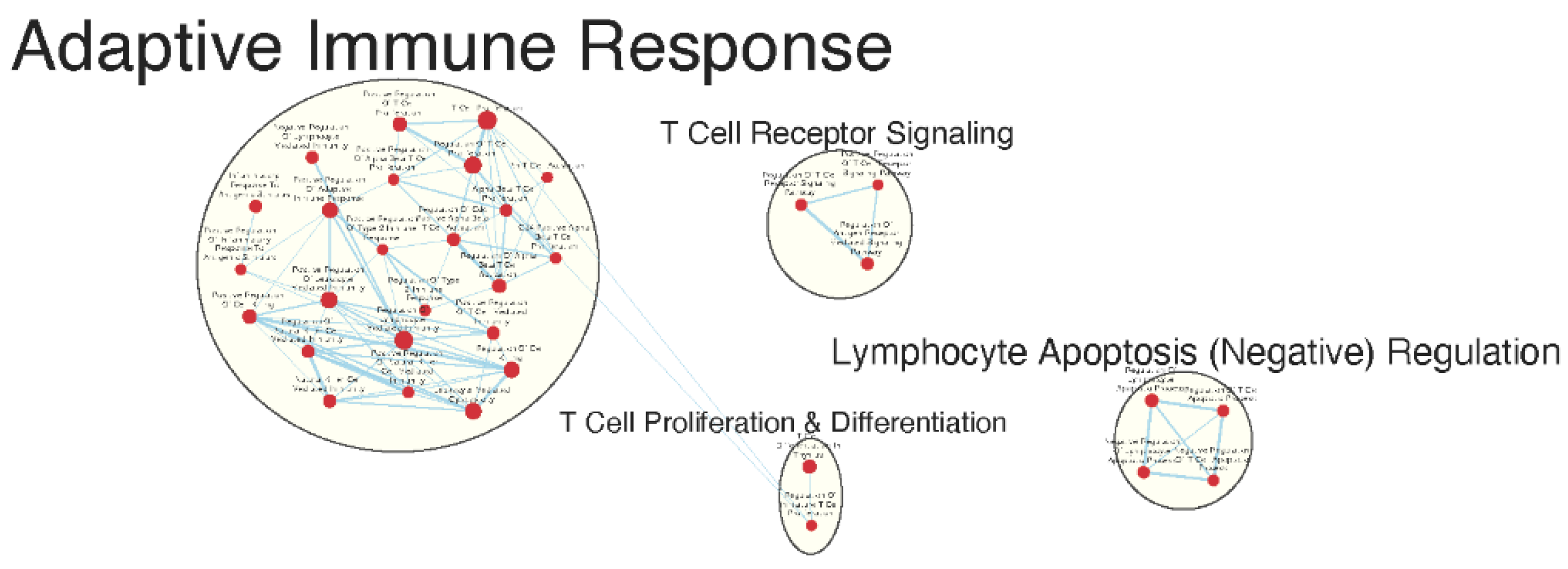
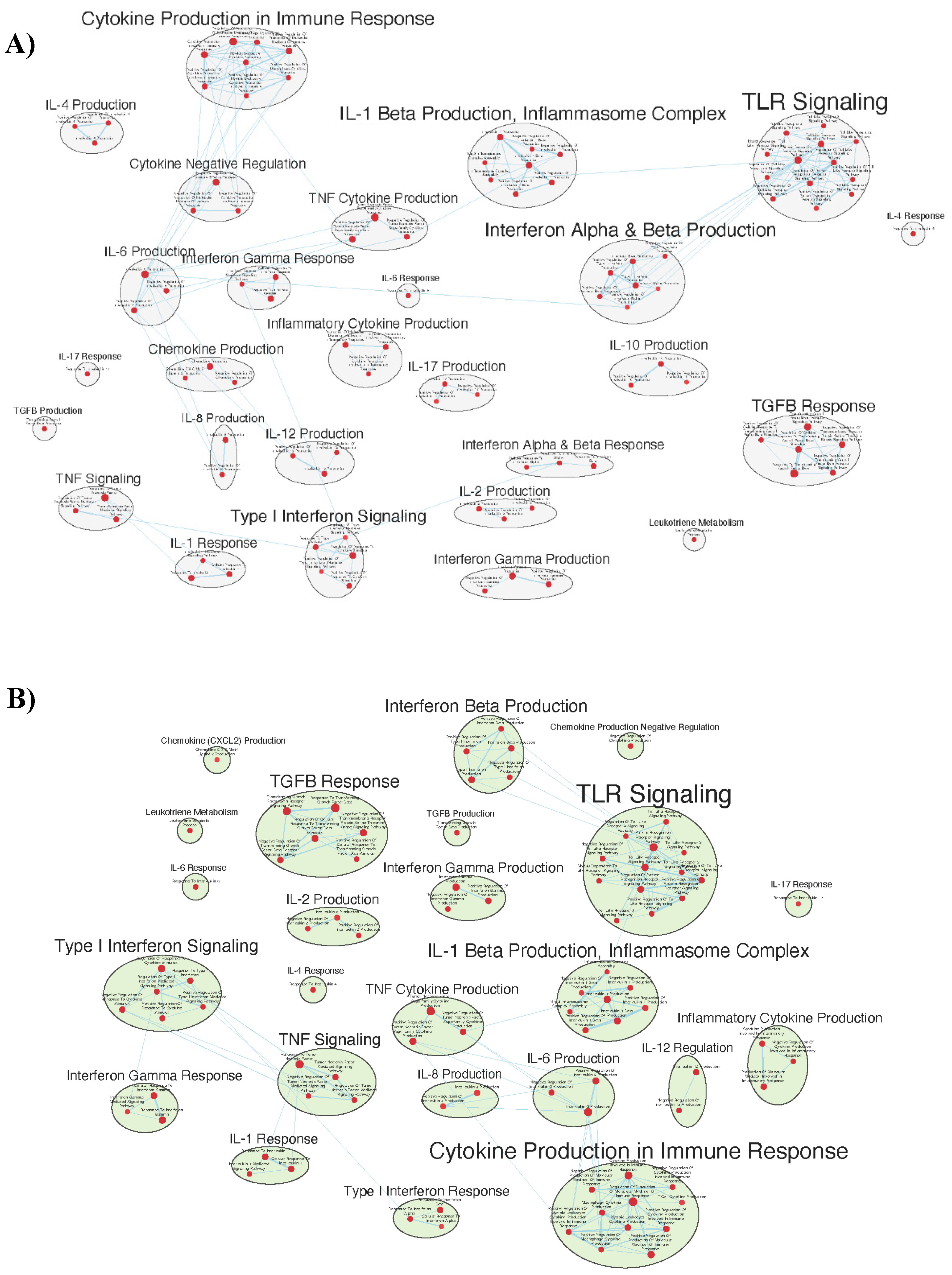
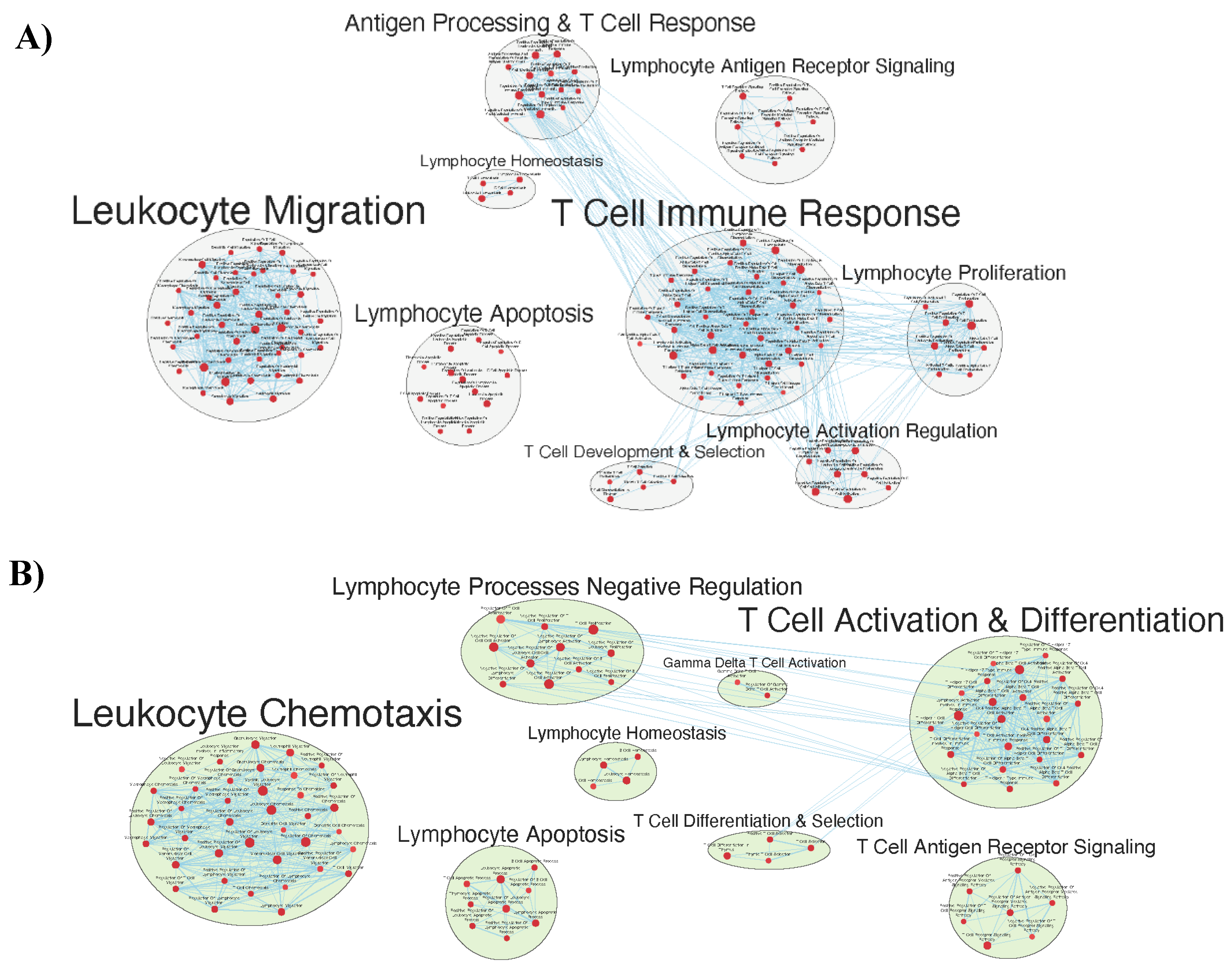
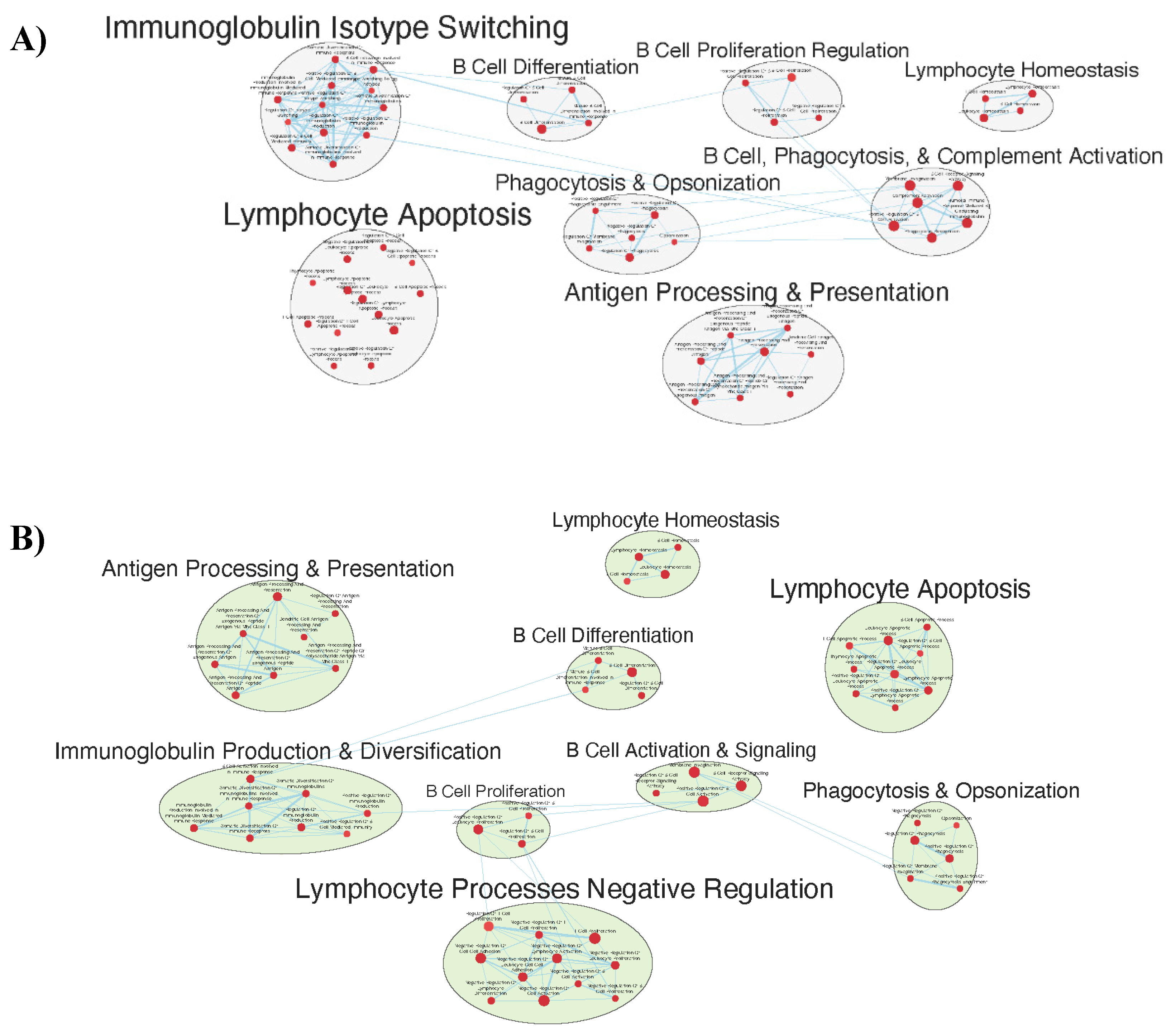
Immunologically Relevant Gene Sets Were Enriched in GAD85 Compared to NCK56
Similar Gene Sets Were Enriched in GAD85 and NCK56 when Compared to STI Control
Other Gene Sets that Were Enriched between the GAD85, NCK56, and STI Groups
Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lycke, N. Recent progress in mucosal vaccine development: potential and limitations. Nat. Rev. Immunol. 2012, 12, 592–605. [CrossRef]
- E. C. Lavelle, R. W. Ward, Mucosal vaccines — fortifying the frontiers. Nat Rev Immunol 22, 236–250 (2022).
- Lynn, D.J.; Benson, S.C.; Lynn, M.A.; Pulendran, B. Modulation of immune responses to vaccination by the microbiota: implications and potential mechanisms. Nat. Rev. Immunol. 2021, 22, 33–46. [CrossRef]
- LeCureux, J.S.; Dean, G.A. Lactobacillus Mucosal Vaccine Vectors: Immune Responses against Bacterial and Viral Antigens. mSphere 2018, 3, e00061-18. [CrossRef]
- Zimmermann, P.; Curtis, N. The influence of probiotics on vaccine responses – A systematic review. Vaccine 2018, 36, 207–213. [CrossRef]
- Mowat, A.M. To respond or not to respond — a personal perspective of intestinal tolerance. Nat. Rev. Immunol. 2018, 18, 405–415. [CrossRef]
- Joosse, M.E.; Nederlof, I.; Walker, L.S.K.; Samsom, J.N. Tipping the balance: inhibitory checkpoints in intestinal homeostasis. Mucosal Immunol. 2019, 12, 21–35. [CrossRef]
- Girardin, S.E.; Boneca, I.G.; Viala, J.; Chamaillard, M.; Labigne, A.; Thomas, G.; Philpott, D.J.; Sansonetti, P.J. Nod2 Is a General Sensor of Peptidoglycan through Muramyl Dipeptide (MDP) Detection. J. Biol. Chem. 2003, 278, 8869–8872. [CrossRef]
- Matsuguchi, T.; Takagi, A.; Matsuzaki, T.; Nagaoka, M.; Ishikawa, K.; Yokokura, T.; Yoshikai, Y. Lipoteichoic Acids fromLactobacillusStrains Elicit Strong Tumor Necrosis Factor Alpha-Inducing Activities in Macrophages through Toll-Like Receptor 2. Clin. Vaccine Immunol. 2003, 10, 259–266. [CrossRef]
- Zeuthen, L.H.; Fink, L.N.; Frøkiær, H. Toll-like receptor 2 and nucleotide-binding oligomerization domain-2 play divergent roles in the recognition of gut-derived lactobacilli and bifidobacteria in dendritic cells. Immunology 2008, 124, 489–502. [CrossRef]
- Fox, B.E.; Vilander, A.; Abdo, Z.; Dean, G.A. NOD2 signaling in CD11c + cells is critical for humoral immune responses during oral vaccination and maintaining the gut microbiome. Sci. Rep. 2022, 12, 1–14. [CrossRef]
- Kajikawa, A.; Zhang, L.; LaVoy, A.; Bumgardner, S.; Klaenhammer, T.R.; Dean, G.A. Mucosal Immunogenicity of Genetically Modified Lactobacillus acidophilus Expressing an HIV-1 Epitope within the Surface Layer Protein. PLOS ONE 2015, 10, e0141713. [CrossRef]
- Kajikawa, A.; Nordone, S.K.; Zhang, L.; Stoeker, L.L.; LaVoy, A.S.; Klaenhammer, T.R.; Dean, G.A. Dissimilar Properties of Two Recombinant Lactobacillus acidophilus Strains Displaying Salmonella FliC with Different Anchoring Motifs. Appl. Environ. Microbiol. 2011, 77, 6587–6596. [CrossRef]
- Gilfillan, D.; Vilander, A.C.; Pan, M.; Goh, Y.J.; O’flaherty, S.; Feng, N.; Fox, B.E.; Lang, C.; Greenberg, H.B.; Abdo, Z.; et al. Lactobacillus acidophilus Expressing Murine Rotavirus VP8 and Mucosal Adjuvants Induce Virus-Specific Immune Responses. Vaccines 2023, 11, 1774. [CrossRef]
- B. Pulendran, in Mucosal Vaccines, (Elsevier, 2020), pp. 753–772.
- Fourati, S.; Tomalin, L.E.; Mule, M.P.; Chawla, D.G.; Gerritsen, B.; Rychkov, D.; Henrich, E.; Miller, H.E.R.; Hagan, T.; Diray-Arce, J.; et al. Pan-vaccine analysis reveals innate immune endotypes predictive of antibody responses to vaccination. Nat. Immunol. 2022, 23, 1777–+. [CrossRef]
- S. Yanagihara, T. Kanaya, S. Fukuda, G. Nakato, M. Hanazato, X.-R. Wu, N. Yamamoto, H. Ohno, Uromodulin-SlpA binding dictates Lactobacillus acidophilus uptake by intestinal epithelial M cells. Int Immunol 29, 357–363 (2017).
- Moon, A.; Sun, Y.; Wang, Y.; Huang, J.; Khan, M.U.Z.; Qiu, H.-J. Lactic Acid Bacteria as Mucosal Immunity Enhancers and Antivirals through Oral Delivery. Appl. Microbiol. 2022, 2, 837–854. [CrossRef]
- Vlasova, A.N.; Kandasamy, S.; Chattha, K.S.; Rajashekara, G.; Saif, L.J. Comparison of probiotic lactobacilli and bifidobacteria effects, immune responses and rotavirus vaccines and infection in different host species. Veter- Immunol. Immunopathol. 2016, 172, 72–84. [CrossRef]
- van Baarlen, P.; Troost, F.; van der Meer, C.; Hooiveld, G.; Boekschoten, M.; Brummer, R.J.M.; Kleerebezem, M. Human mucosal in vivo transcriptome responses to three lactobacilli indicate how probiotics may modulate human cellular pathways. Proc. Natl. Acad. Sci. 2010, 108, 4562–4569. [CrossRef]
- Paturi, G.; Phillips, M.; Kailasapathy, K. Effect of Probiotic Strains Lactobacillus acidophilus LAFTI L10 and Lactobacillus paracasei LAFTI L26 on Systemic Immune Functions and Bacterial Translocation in Mice. J. Food Prot. 2008, 71, 796–801. [CrossRef]
- Medina, M.; Izquierdo, E.; Ennahar, S.; Sanz, Y. Differential immunomodulatory properties of Bifidobacterium logum strains: relevance to probiotic selection and clinical applications. Clin. Exp. Immunol. 2007, 150, 531–538. [CrossRef]
- Huang, Z.; Gong, L.; Jin, Y.; Stanton, C.; Ross, R.P.; Zhao, J.; Yang, B.; Chen, W. Different Effects of Different Lactobacillus acidophilus Strains on DSS-Induced Colitis. Int. J. Mol. Sci. 2022, 23, 14841. [CrossRef]
- Hagan, T.; Gerritsen, B.; Tomalin, L.E.; Fourati, S.; Mule, M.P.; Chawla, D.G.; Rychkov, D.; Henrich, E.; Miller, H.E.R.; Diray-Arce, J.; et al. Transcriptional atlas of the human immune response to 13 vaccines reveals a common predictor of vaccine-induced antibody responses. Nat. Immunol. 2022, 23, 1788–+. [CrossRef]
- B. Pulendran, S. Li, H. I. Nakaya, Systems Vaccinology. Immunity 33, 516–529 (2010).
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [CrossRef]
- Babraham Bioinformatics, FastQC A Quality Control tool for High Throughput Sequence Data (2018) (available at https://www.bioinformatics.babraham.ac.uk/projects/fastqc/).
- Ewels, P.; Magnusson, M.; Lundin, S.; Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 2016, 32, 3047–3048. [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [CrossRef]
- H. Li, Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv:1303.3997 [q-bio] (2013) (available at http://arxiv.org/abs/1303.3997).
- shilparaopradeep, Samtools guide: learning how to filter and manipulate with SAM/BAM filesMedium (2016) (available at https://medium.com/@shilparaopradeep/samtools-guide-learning-how-to-filter-and-manipulate-with-sam-bam-files-2c28b25d29e8).
- Kraken2 taxonomic sequence classification system (2018).
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [CrossRef]
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq—A Python framework to work with high-throughput sequencing data. Bioinformatics 2015, 31, 166–169. [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [CrossRef]
- H. Wickham, ggplot2: Elegant graphics for data analysis (Springer Science+Business Media, New York, NY, 2009).
- R Core Team, R: A Language and Environment for Statistical Computing (2023) (available at https://www.R-project.org).
- Wood, D.E.; Lu, J.; Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 2019, 20, 257. [CrossRef]
| Treatments | Male | Female | Total |
|---|---|---|---|
| GAD85 (rLA, expressing dual-antigens, VP8-1 and VP810AA, and adjuvants FimH and FliC) | 3 | 3 | 6 |
| NCK56 (wild-type LA, positive control) | 3 | 3 | 6 |
| STI (sham treatment, negative control) | 3 | 3 | 6 |
| Total | 9 | 9 | 18 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




