Submitted:
08 May 2024
Posted:
09 May 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. RNA Extraction, Reverse Transcription, Library Preparation and High-Throughput Sequencing
2.3. Complete Genome Assembly and Annotation of Novel Virus
2.4. Phylogenetic Analyses
3. Results
3.1. Mastadenovirus Detection
3.2. Complete Genome of Novel Mastadenovirus BatAdV/MOW15-Nn19
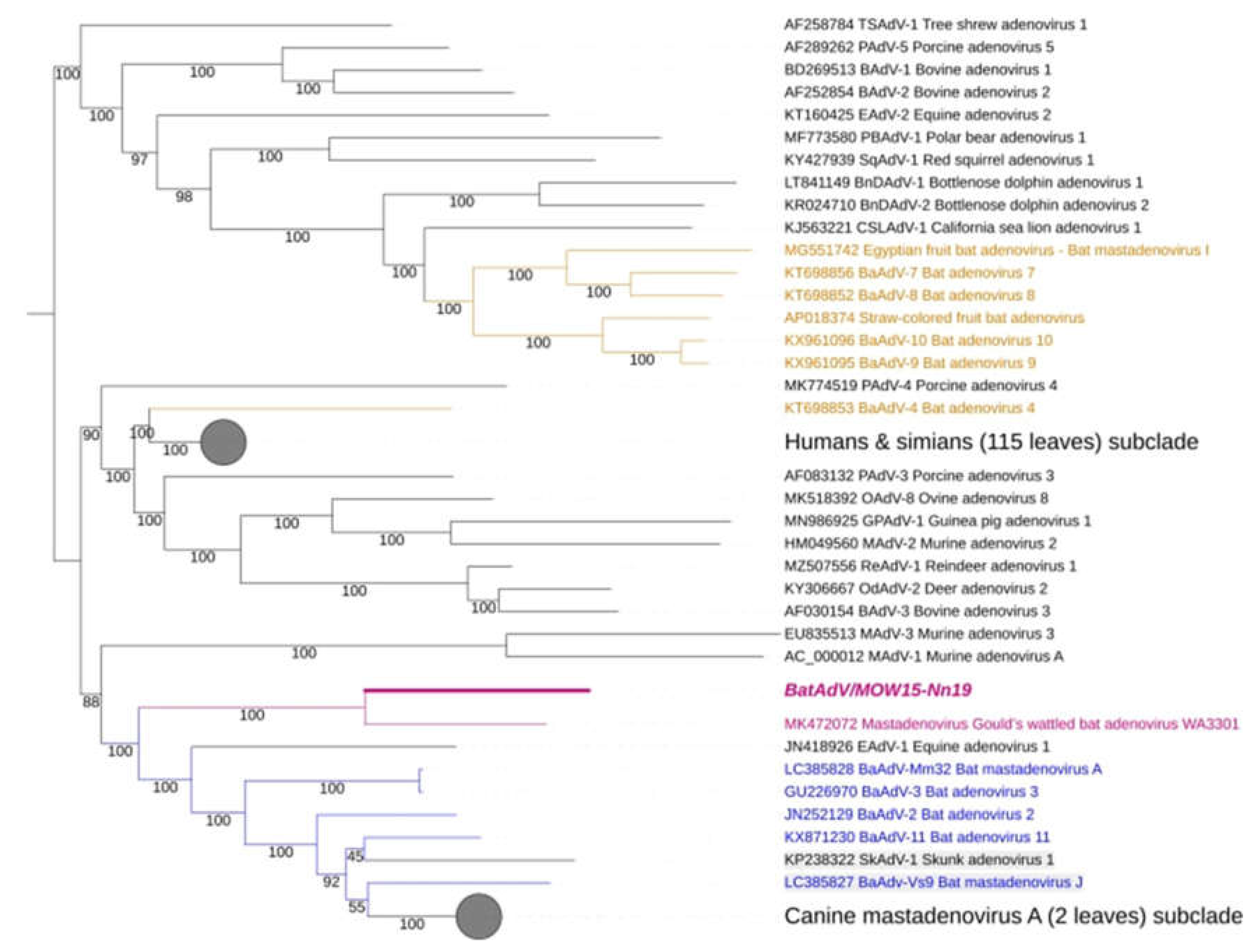
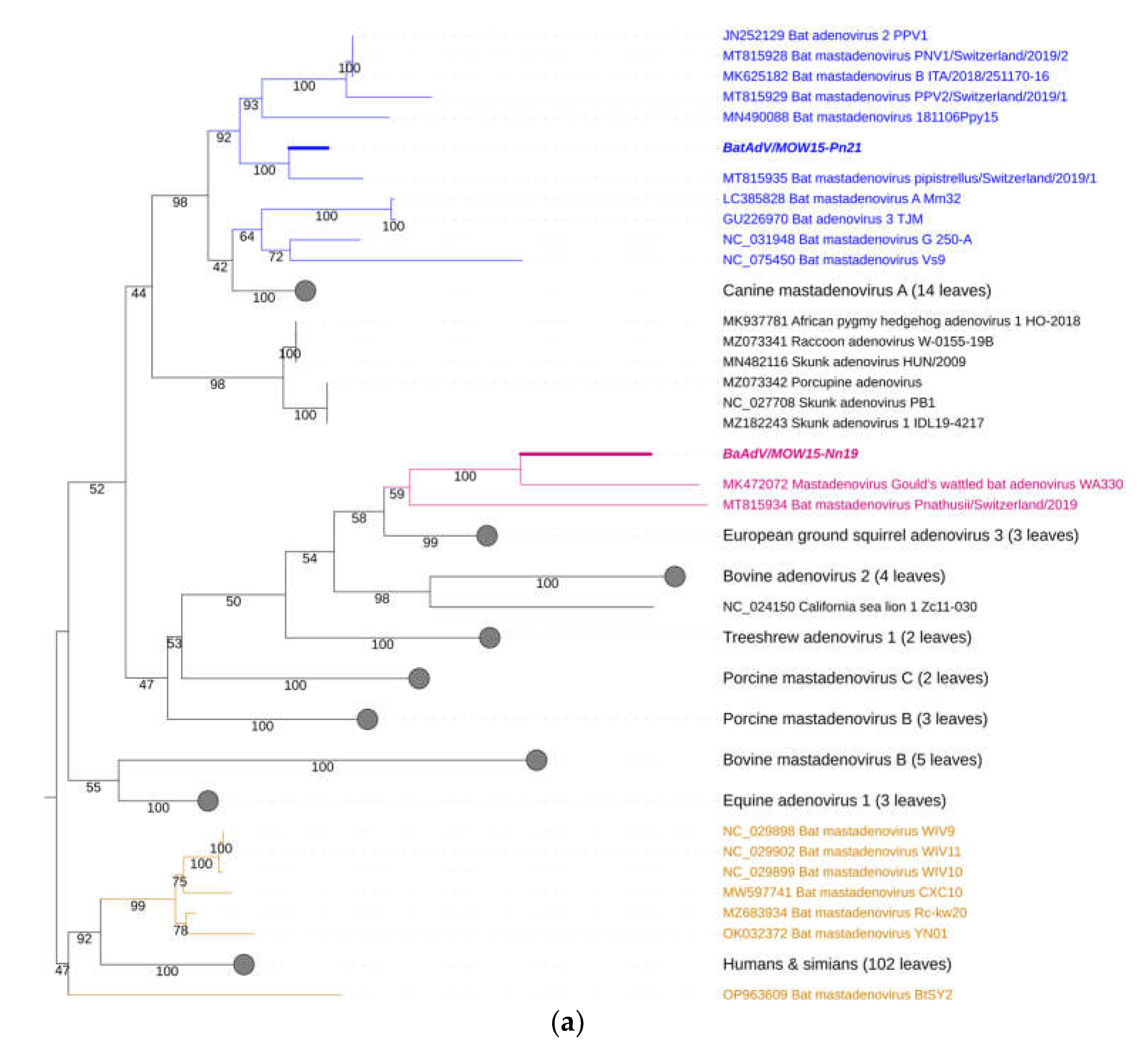
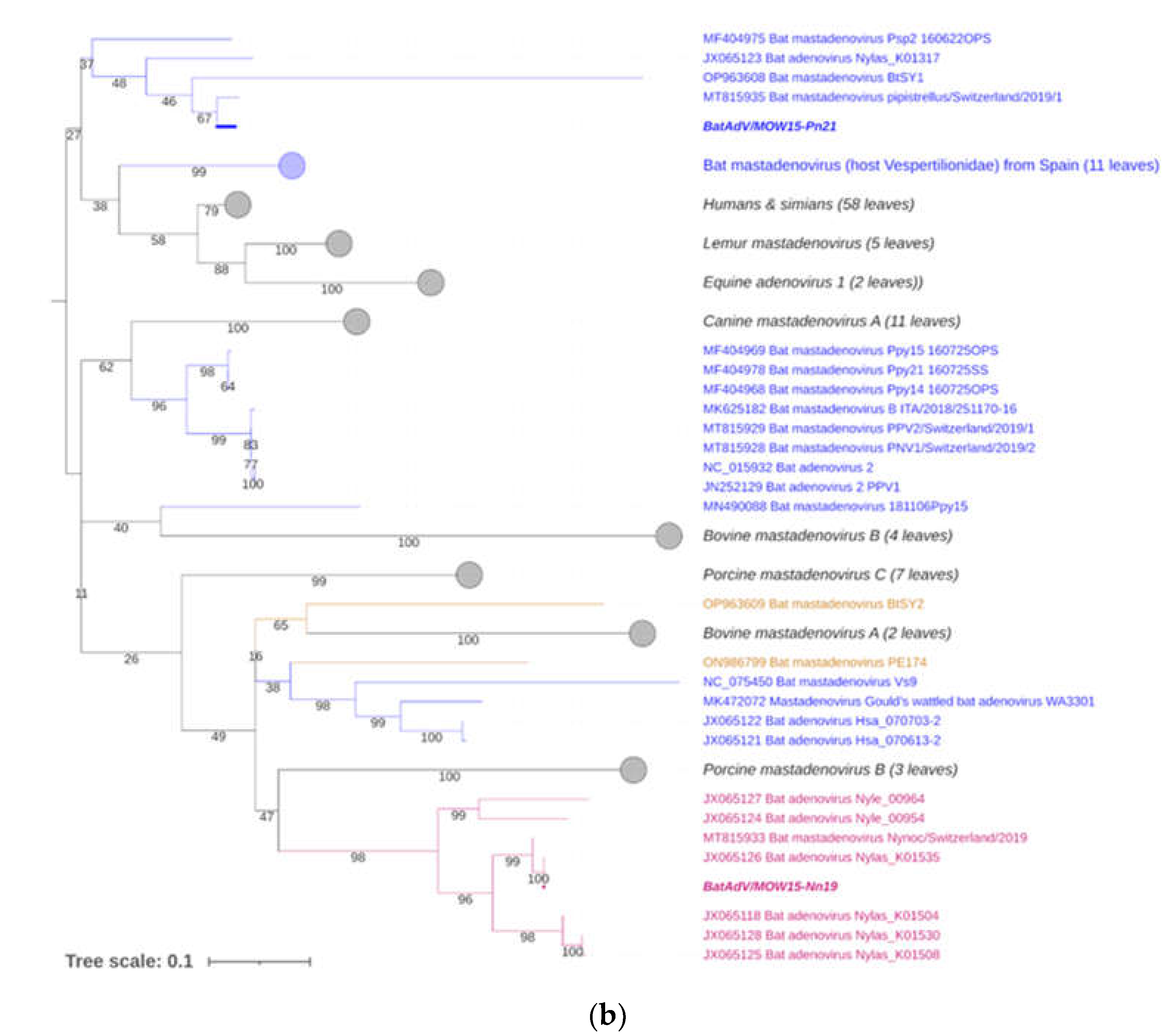
3.2. Phylogenetic Analysis for Bat Mastadenoviruses from Russia
4. Discussion
4.1. Prevalence of Mastadenoviruses in Bat Populations
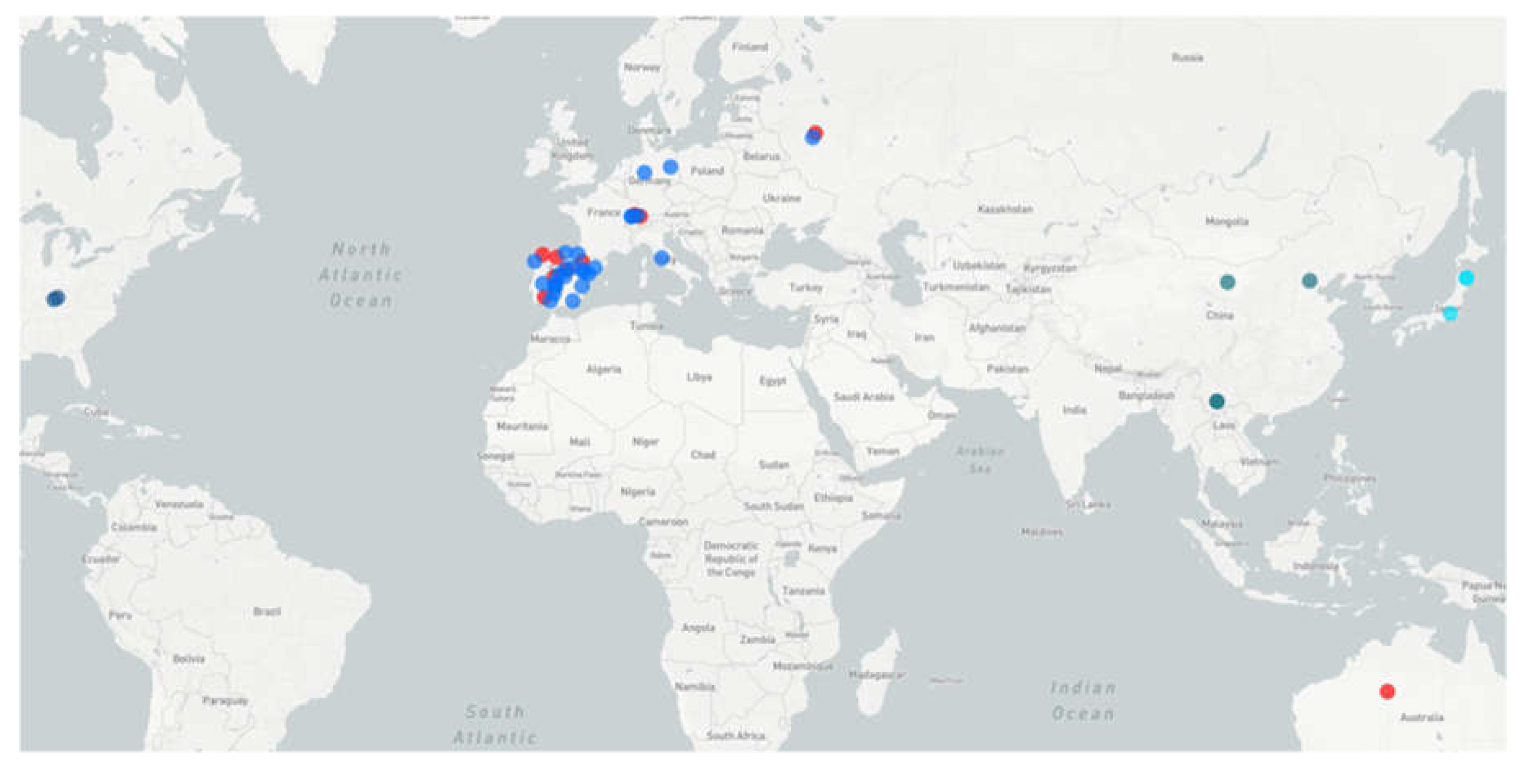
4.2. Diversity and Geographic Distribution of BatAdVs from European Bats
4.3. Could the Pipistrellus Bats Distribute Mastadenoviruses around the Old World until Australia?
4.4. Structural Features of the Genome of the Novel Mastadenovirus Species
5. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Balboni, A., Verin, R., Morandi, F., Poli, A., Prosperi, S., & Battilani, M. (2013). Molecular epidemiology of canine adenovirus type 1 and type 2 in free-ranging red foxes (Vulpes vulpes) in Italy. Veterinary Microbiology, 162(2–4). [CrossRef] [PubMed]
- Benko, M., Aoki, K., Arnberg, N., Davison, A. J., Echavarria, M., Hess, M., … Harrach, B. (2022). ICTV Virus Taxonomy Profile: Adenoviridae 2022. Journal of General Virology, 103(3). [CrossRef] [PubMed]
- Böszörményi, K. P., Podgorski, I. I., Vidovszky, M. Z., Sós, E., Benkő, M., & Harrach, B. (2020). Full genome sequence analysis of a novel adenovirus from a captive polar bear (Ursus maritimus). Virus Research, 277. [CrossRef] [PubMed]
- Chen, E. C., Yagi, S., Kelly, K. R., Mendoza, S. P., Maninger, N., Rosenthal, A., … Chiu, C. Y. (2011). Cross-species transmission of a novel adenovirus associated with a fulminant pneumonia outbreak in a new world monkey colony. PLoS Pathogens, 7(7). [CrossRef]
- Cholleti, H., de Jong, J., Blomström, A. L., & Berg, M. (2022). Characterization of Pipistrellus pygmaeus Bat Virome from Sweden. Viruses, 14(8). [CrossRef] [PubMed]
- Dacheux, L., Cervantes-Gonzalez, M., Guigon, G., Thiberge, J. M., Vandenbogaert, M., Maufrais, C., … Bourhy, H. (2014). A preliminary study of viral metagenomics of french bat species in contact with humans: Identification of new mammalian viruses. PLoS ONE, 9(1). [PubMed]
- Dehghan, S., Seto, J., Liu, E. B., Ismail, A. M., Madupu, R., Heim, A., … Seto, D. (2019). A Zoonotic Adenoviral Human Pathogen Emerged through Genomic Recombination among Human and Nonhuman Simian Hosts. Journal of Virology, 93(18). [CrossRef] [PubMed]
- Diakoudi, G., Lanave, G., Moreno, A., Chiapponi, C., Sozzi, E., Prosperi, A., … Lelli, D. (2019). Surveillance for adenoviruses in bats in Italy. Viruses, 11(6). [CrossRef] [PubMed]
- 9. Doerfler, W. (1996). Adenoviruses (4th ed., Vol. 67; Walter Doerfler & Samuel Baron, Eds.). Galveston (TX). Retrieved from https://pubmed.ncbi.nlm.nih.gov/21413345/.
- Drexler, J. F., Corman, V. M., Wegner, T., Tateno, A. F., Zerbinati, R. M., Gloza-Rausch, F., … Drosten, C. (2011). Amplification of emerging viruses in a bat colony. Emerging Infectious Diseases, 17(3). [CrossRef]
- Gallardo, J., Pérez-Illana, M., Martín-González, N., & Martín, C. S. (2021). Adenovirus structure: What is new? International Journal of Molecular Sciences, Vol. 22. [CrossRef]
- Hardmeier, I., Aeberhard, N., Qi, W., Schoenbaechler, K., Kraettli, H., Hatt, J. M., … Kubacki, J. (2021). Metagenomic analysis of fecal and tissue samples from 18 endemic bat species in Switzerland revealed a diverse virus composition including potentially zoonotic viruses. PLoS ONE, 16(6 June 2021).
- Harrach, B. (2014). Adenoviruses: General Features☆. In Reference Module in Biomedical Sciences. [CrossRef]
- Harvey, W. , Hutto, E. H., Chilton, J. A., Chamanza, R., Mysore, J. V., Parry, N. M. A., … Bradley, A. E. (2023). Infectious diseases of non-human primates. In Spontaneous Pathology of the Laboratory Non-human Primate. [CrossRef]
- Hou, J., Xu, J., Wang, B., Zhang, H., Yin, B., Li, G., … Wang, L. (2023). First identification of canine adenovirus 1 in mink and bioinformatics analysis of its 100 K protein. Frontiers in Microbiology, 14. [CrossRef] [PubMed]
- Iglesias-Caballero, M., Juste, J., Vázquez-Morón, S., Falcon, A., Aznar-Lopez, C., Ibáñez, C., … Casas, I. (2018). New adenovirus groups in western palaearctic bats. Viruses, 10(8). [CrossRef] [PubMed]
- Jánoska, M., Vidovszky, M., Molnár, V., Liptovszky, M., Harrach, B., & Benko, M. (2011). Novel adenoviruses and herpesviruses detected in bats. Veterinary Journal, 189(1). [CrossRef]
- Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., Von Haeseler, A., & Jermiin, L. S. (2017). ModelFinder: Fast model selection for accurate phylogenetic estimates. Nature Methods, 14(6). [CrossRef] [PubMed]
- Katayama, M., Murakami, S., Matsugo, H., Kamiki, H., Fujii, M., Takenaka-Uema, A., & Horimoto, T. (2022). Complete genome sequence of a novel bat mastadenovirus C strain isolated from Rhinolophus cornutus in Japan. Archives of Virology, 167(3). [CrossRef]
- Katoh, K., Rozewicki, J., & Yamada, K. D. (2018). MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Briefings in Bioinformatics, 20(4). [CrossRef]
- 21. Kitchener, D.J., Caputi, N., Jones, B. (1986). Revision of Australo-Papuan Pipistrellus and Falsistrellus (Microchiroptera: Vespertilionidae). Records of the Western Australian Museum, 12(4), 435–495.
- Kleinberger, T. (2015). Mechanisms of cancer cell killing by the adenovirus E4orf4 protein. Viruses, Vol. 7. [CrossRef]
- Kohl, C., Nitsche, A., & Kurth, A. (2021). Update on potentially zoonotic viruses of european bats. Vaccines, Vol. 9. [CrossRef]
- Kohl, C., Vidovszky, M. Z., Mühldorfer, K., Dabrowski, P. W., Radonić, A., Nitsche, A., … Harrach, B. (2012). Genome Analysis of Bat Adenovirus 2: Indications of Interspecies Transmission. Journal of Virology, 86(3). [CrossRef] [PubMed]
- Kosoltanapiwat, N., Tongshoob, J., Ampawong, S., Reamtong, O., Prasittichai, L., Yindee, M., … Boonnak, K. (2022). Simian adenoviruses: Molecular and serological survey in monkeys and humans in Thailand. One Health, 15. [CrossRef] [PubMed]
- Lack, J. B., & Van Den Bussche, R. A. (2010). Identifying the confounding factors in resolving phylogenetic relationships in Vespertilionidae. Journal of Mammalogy, 91(6). [CrossRef]
- Lagerveld, S., Poerink, B. J., & Geelhoed, S. C. V. (2021). Offshore occurrence of a migratory bat, pipistrellus nathusii, depends on seasonality and weather conditions. Animals, 11(12). [CrossRef] [PubMed]
- Letunic, I., & Bork, P. (2021). Interactive tree of life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Research, 49(W1). [CrossRef] [PubMed]
- Matsugo, H., Kitamura-Kobayashi, T., Kamiki, H., Ishida, H., Sekine, W., Takenaka-Uema, A., … Horimoto, T. (2021). A potential bat adenovirus-based oncolytic virus targeting canine cancers. Scientific Reports, 11(1). [CrossRef] [PubMed]
- Medkour, H., Amona, I., Akiana, J., Davoust, B., Bitam, I., Levasseur, A., … Mediannikov, O. (2020). Adenovirus infections in African humans and wild non-human primates: Great diversity and cross-species transmission. Viruses, 12(6). [CrossRef] [PubMed]
- Moratelli R, Burgin C, Cláudio V, Novaes R, López-Baucells A, Haslauer R (2019) Family Vespertilionidae (Vesper Bats): Bats. Lynx Edicions, Barcelona, pp 716-981.
- Minh, B. Q., Nguyen, M. A. T., & Von Haeseler, A. (2013). Ultrafast approximation for phylogenetic bootstrap. Molecular Biology and Evolution, 30(5). [CrossRef]
- Nguyen, L. T., Schmidt, H. A., Von Haeseler, A., & Minh, B. Q. (2015). IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Molecular Biology and Evolution, 32(1). [CrossRef]
- Nishimura, Y., Yoshida, T., Kuronishi, M., Uehara, H., Ogata, H., & Goto, S. (2017). ViPTree: The viral proteomic tree server. Bioinformatics, 33(15). [CrossRef] [PubMed]
- Ntumvi, N. F., Diffo, J. L. D., Tamoufe, U., Ndze, V. N., Takuo, J. M., Mouiche, M. M. M., … Lange, C. E. (2021). Evaluation of bat adenoviruses suggests co-evolution and host roosting behaviour as drivers for diversity. Microbial Genomics, 7(4). [CrossRef]
- Ogawa, H., Kajihara, M., Nao, N., Shigeno, A., Fujikura, D., Hang’Ombe, B. M., … Takada, A. (2017). Characterization of a novel bat adenovirus isolated from straw-colored fruit bat (Eidolon helvum). Viruses, 9(12). [CrossRef]
- Oleaga, A., Balseiro, A., Espí, A., & Royo, L. J. (2022). Wolf (Canis lupus) as canine adenovirus type 1 (CAdV-1) sentinel for the endangered cantabrian brown bear (Ursus arctos arctos). Transboundary and Emerging Diseases, 69(2). [CrossRef] [PubMed]
- Prada, D., Boyd, V., Baker, M. L., O’Dea, M., & Jackson, B. (2019). Viral diversity of microbats within the south west botanical province of western Australia. Viruses, 11(12). [CrossRef]
- Prjibelski, A., Antipov, D., Meleshko, D., Lapidus, A., & Korobeynikov, A. (2020). Using SPAdes De Novo Assembler. Current Protocols in Bioinformatics, 70(1). [CrossRef] [PubMed]
- Roy, S., Vandenberghe, L. H., Kryazhimskiy, S., Grant, R., Calcedo, R., Yuan, X., … Wilson, J. M. (2009). Isolation and characterization of adenoviruses persistently shed from the gastrointestinal tract of non-human primates. PLoS Pathogens, 5(7). [CrossRef]
- Sallard, E., Zhang, W., Aydin, M., Schröer, K., & Ehrhardt, A. (2023). The Adenovirus Vector Platform: Novel Insights into Rational Vector Design and Lessons Learned from the COVID-19 Vaccine. Viruses, Vol. 15. [CrossRef]
- Sanz MA, Almela EG, García-Moreno M, Marina AI, Carrasco L. 2019. A viral RNA motif involved in signaling the initiation of translation on non-AUG codons. RNA 25: 431–452. doi:10.1261/rna.068858.118. [CrossRef]
- Seemann, T. (2014). Prokka: Rapid prokaryotic genome annotation. Bioinformatics, 30(14). [CrossRef]
- Simmons, N.B. (2005). Order Chiroptera. In Wilson, D.E. & Reeder, D.M. (eds.) Mammal Species of the World, (Third edition). The Johns Hopkins University Press, Baltimore.
- Sonntag, M., Mühldorfer, K., Speck, S., Wibbelt, G., & Kurth, A. (2009). New adenovirus in bats, Germany. Emerging Infectious Diseases, 15(12). [CrossRef] [PubMed]
- Speranskaya, A. S., Artiushin, I. V., Samoilov, A. E., Korneenko, E. V., Khabudaev, K. V., Ilina, E. N., … Daszak, P. (2023). Identification and Genetic Characterization of MERS-Related Coronavirus Isolated from Nathusius’ Pipistrelle (Pipistrellus nathusii) near Zvenigorod (Moscow Region, Russia). International Journal of Environmental Research and Public Health, 20(4). [CrossRef] [PubMed]
- Tcherepanov, V., Ehlers, A., & Upton, C. (2006). Genome annotation transfer utility (GATU): Rapid annotation of viral genomes using a closely related reference genome. BMC Genomics, 7. [CrossRef] [PubMed]
- Vidovszky, M. Z., Kohl, C., Boldogh, S., Görföl, T., Wibbelt, G., Kurth, A., & Harrach, B. (2015). Random sampling of the Central European bat fauna reveals the existence of numerous hitherto unknown adenoviruses. Acta Veterinaria Hungarica, 63(4). [CrossRef] [PubMed]
- viralzone.expasy.org Mastadenovirus (taxid:10509). (n.d.). Retrieved January 25, 2024, from Mastadenovirus (taxid:10509) website: https://viralzone.expasy.org/183?outline=all_by_species.
- Wang, F., Zhu, R., Qian, Y., Sun, Y., Chen, D., Wang, F., … Zhao, L. (2023). The changed endemic pattern of human adenovirus from species B to C among pediatric patients under the pressure of non-pharmaceutical interventions against COVID-19 in Beijing, China. Virology Journal, 20(1). [CrossRef]
- Zhukova, S. S., Solovyeva, E. N., Artyushin, I. V., & Kruskop, S. V. (2022). Paraphyly of the Pipistrelles (Pipistrellus; Vespertilionidae) is Confirmed by the Analysis of the Nuclear Gene Markers. Doklady Biochemistry and Biophysics, 507(1). [CrossRef]
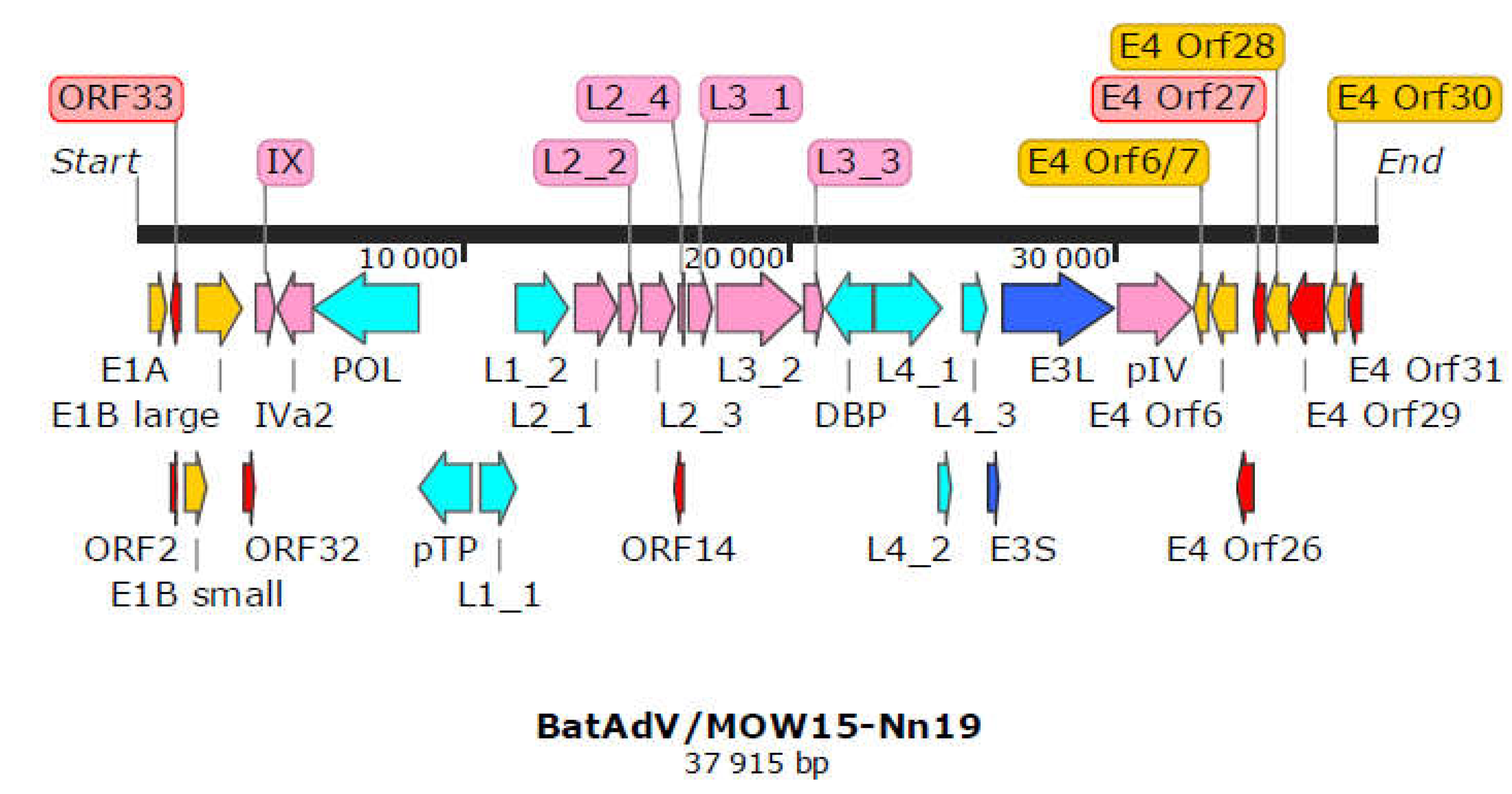
| Coordinates | Gene | Protein Description; Function, accordingly to ICTV | Transcription class | Nearest homolog (proteins) | |||
|---|---|---|---|---|---|---|---|
| Genbank ID | Identity | Query coverage | |||||
| 1 | 372-968 | E1A | R, E1A; Modulation of the host cell transcriptional machinery |
E1 | QGX41997* | 39.13% | 74% |
| 2 | 1034-1258 | ORF2 | Putative protein | unknown | - | - | |
| 3 | 1050-1289 complement |
ORF33 | Putative protein | unknown | - | - | |
| 4 | 1422-2150 | E1B 19K | R, E1B small; Modulation of the host cell transcriptional machinery |
QGX41989* | 39.92% | 96% | |
| 5 | 1811-3271 | E1B 55K | R, E1B Large; Modulation of the host cell transcriptional machinery |
QGX41980* | 47.00% | 99% | |
| 6 | 3271-3621 | ORF32 | Putative protein | Intermediate | unknown | - | - |
| 7 | 3602-4270 | IX | S (IX); Capsid minor protein |
QGX41995* | 53.55% | 66% | |
| 8 | 4274-5344 complement |
IVa2 | D, S (IVa2); Capsid minor protein |
QGX41983* | 86.52% | 99% | |
| 9 | 5371-8619 complement |
POL | D; DNA polymerase |
E2b | QGX41975* | 76.90% | 98% |
| 10 | 8619-10250 complement |
pTP | D; pTP, Preterminal protein; Important role in the initiation of viral DNA replication |
QGX41978* | 83.39% | 99% | |
| 11 | 10530-11684 | L1_1 | D; pP3 52k | L1 | QGX41986* | 73.42% | 99% |
| 12 | 11629-13278 | L1_2 | D, S (pIIIa); Pre-hexon-linking protein, phosphoprotein Capsid minor protein |
QGX41979* | 80.00% | 99% | |
| 13 | 13382-14776 | L2_1 | S (III); penton base; Capsid major protein |
L2 | QGX41981* | 84.70% | 99% |
| 14 | 14810-15349 | L2_2 | S (pVII), major core; Аssociated with the DNA and form the core within the virion | QGX41998* | 74.51% | 27% | |
| 15 | 15412-16509 | L2_3 | S (V), minor core; Аssociated with the DNA and form the core within the virion |
QGX41987* | 70.89% | 98% | |
| 16 | 16455-16754 complement | ORF14 | Putative protein | unknown | - | - | |
| 17 | 16585-16821 | L2_4 | S (pX), pre-core protein X; Аssociated with the DNA and form the core within the virion |
YP_009388318 | 61.19% | 84% | |
| 18 | 16908-17657 | L3_1 | S (pVI); Capsid minor protein |
L3 | QGX41988* | 72.31% | 99% |
| 19 | 17767-20466 | L3_2 | S (II), hexon; Capsid major protein |
QGX41976* | 83.70% | 99% | |
| 20 | 20468-21088 | L3_3 | D, S, protease; Peptidase_C5 | QGX41993* | 79.41% | 98% | |
| 21 | 21128-22564 complement | DBP | D, DBP; DNA binding protein |
E2a | QGX41982* | 58.44% | 99% |
| 22 | 22594-24696 | L4_1 | D, 100 kDa Shutoff | L4 | QGX41977* | 72.38% | 92% |
| 23 | 24542-25042 | L4_2 | D, R, phosphoprotein 2, pP2 | QGX41999* | 56.73% | 56% | |
| 24 | 25332-26099 | L4_3 | S (pVIII), hexon associated protein; Capsid minor protein |
QGX41992* | 85.15% | 89% | |
| 25 | 26083-26478 | E3S | H, 14,4 kDA protein | E3 | YP_010796290 | 30.65% | 80% |
| 26 | 26533-30003 | E3L | H, pE3L | QGX41974* | 29.21% | 99% | |
| 27 | 30059-32377 | pIV | S (IV), fiber; Capsid major protein |
L5 | QGX41984* | 33.24% | 56% |
| 28 | 32416-32874 complementt | E4 Orf6/7 | R, E4 protein; Modulation of the host cell transcriptional machinery |
E4 | AGT77890 | 44.59% | 36% |
| 29 | 32899-33717 complement | E4 Orf6 | R, E4 protein; Modulation of the host cell transcriptional machinery |
QGX41990* | 36.69% | 89% | |
| 30 | 33687-34226 | E4 Orf26 | Putative protein | unknown | - | - | |
| 31 | 34207-34572 | E4 Orf27 | Putative protein | unknown | - | - | |
| 32 | 34584-35285 | E4 Orf28 | R, Putative dUTPase | QGX41994* | 27.07% | 75% | |
| 33 | 35313-36422 | E4 Orf29 | ORF19, Putative protein | QGX41985* | 35.94% | 86% | |
| 34 | 36476-37060 | E4 Orf30 | R, Putative dUTPase | QGX41991* | 55.61% | 95% | |
| 35 | 37094-37558 | E4 Orf31 | Putative protein | unknown | - | - | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




