Submitted:
29 April 2024
Posted:
01 May 2024
You are already at the latest version
Abstract
Keywords:
Introduction
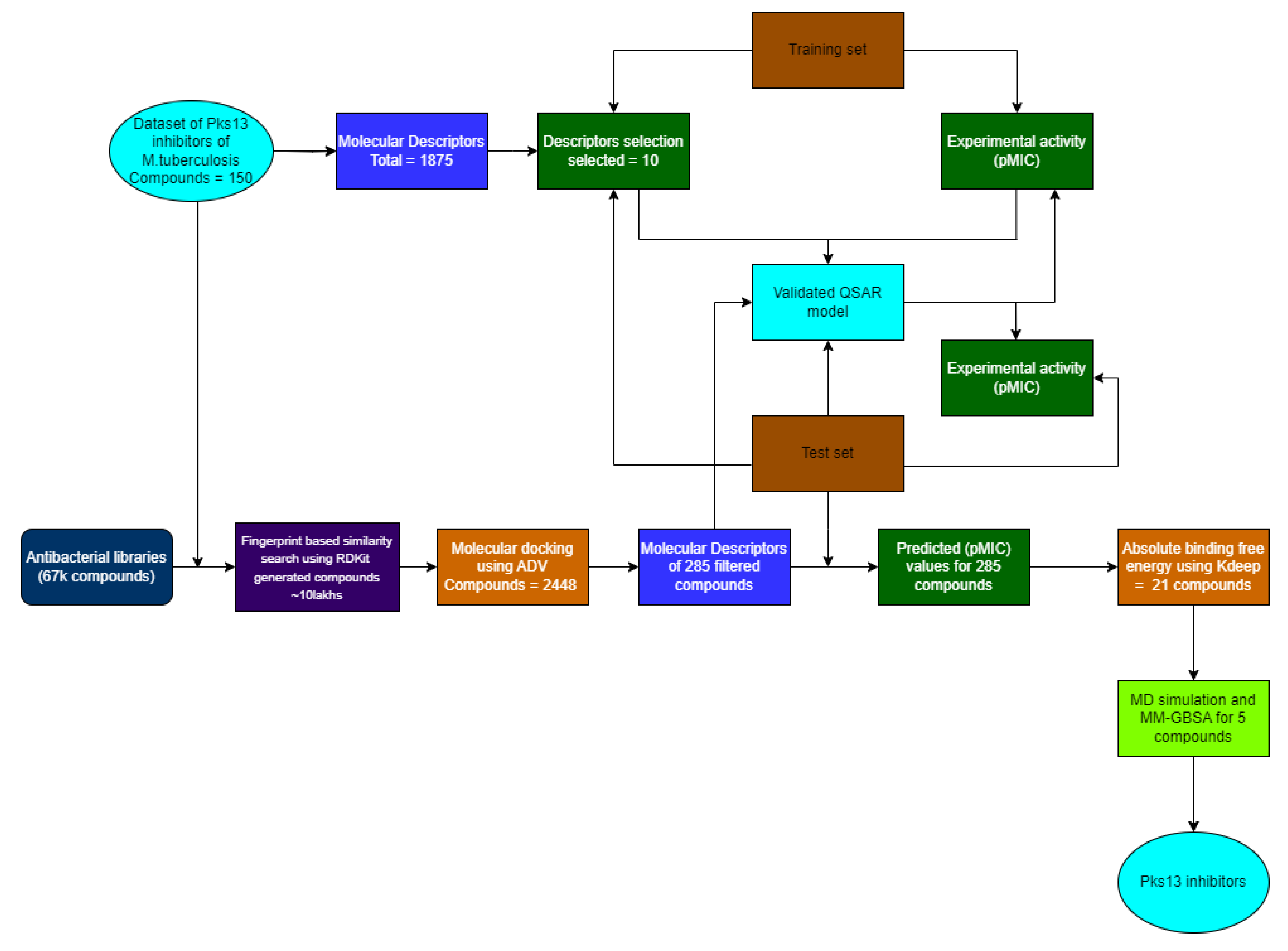
Materials & Methods
Dataset Construction
Molecular Descriptors Calculation
Data Pretreatment
QSAR Model Development and Validation
Protein Preparation of the Pks13-TE Domain
Preparation of Active Compound Library and Selection of Chemical Library Database
Fingerprint-Based Similarity Search Against Parent Antibacterial Chemical Library Database
Compounds Library Curation
Molecular Docking Studies
Biological Activity Prediction Using Validated QSAR Model
Absolute Binding Free Energy Calculations Employing KDeep
Molecular Dynamics (MD) Simulation Study
Molecular Mechanics with Generalised Born and Surface Area Solvation (MM-GBSA)
Binding Free Energy Calculation through MM-GBSA Approach
Results and Discussion
QSAR Modelling
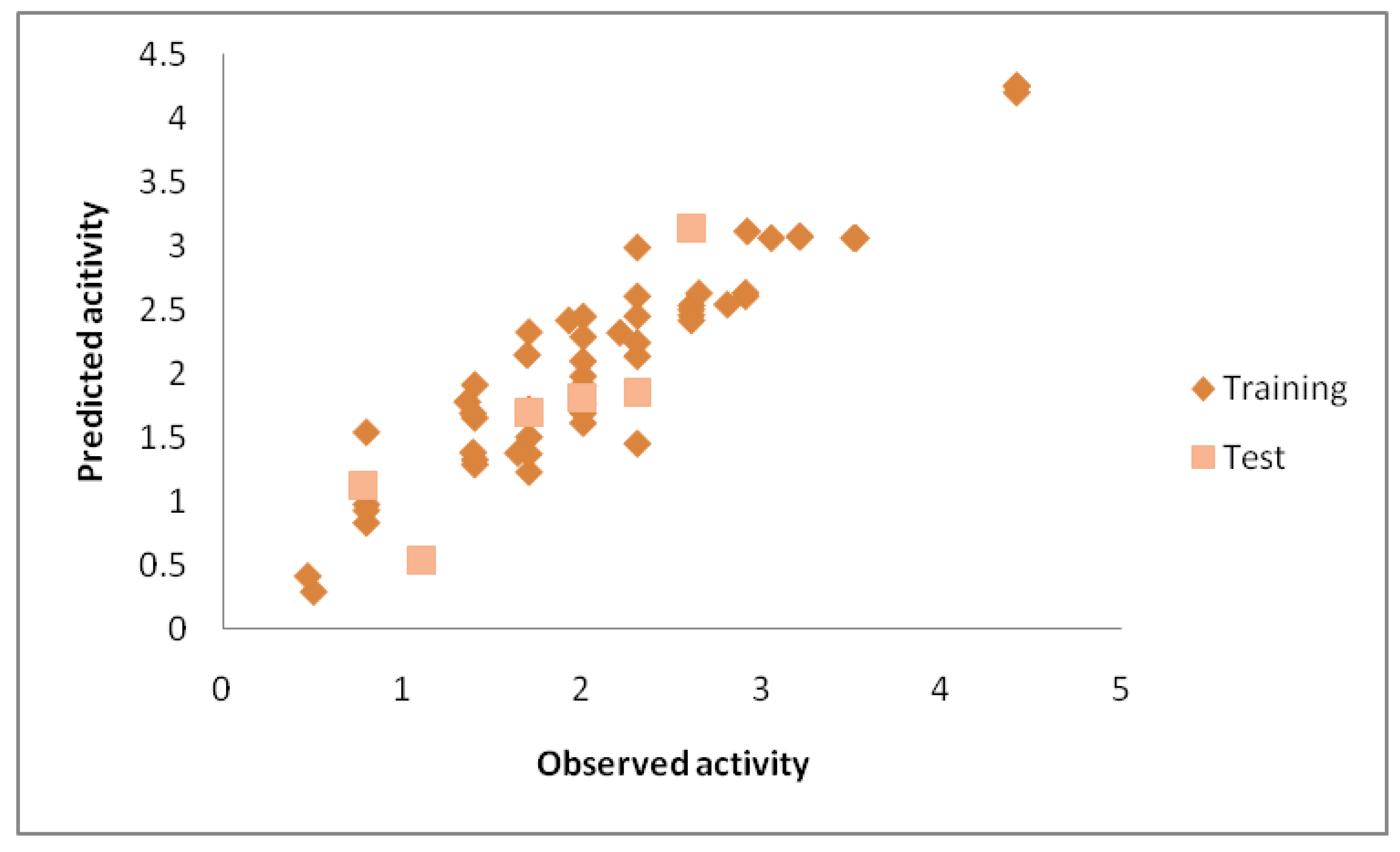
| S.No. | Descriptor | Type | Information |
|---|---|---|---|
| 1. | AATSC6c | 2D | Average centred Broto-Moreau autocorrelation - lag six / weighted by Sanderson electronegativities. |
| 2. | IC2 | 2D | Information content index (neighbourhood symmetry of 2-order) |
| 3. | ATSC1e | 2D | Centred Broto-Moreau autocorrelation - lag one / weighted by Sanderson electronegativities |
| 4. | ALogp2 | 2D | Square ofAlogP |
| 5. | ATSC4c | 2D | Centred Broto-Moreau autocorrelation - lag four / weighted by charges |
| 6. | SpMax5_Bhm | 2D | The largest absolute eigenvalue of Burden-modified matrix - n 5 / weighted by relative mass |
| 7. | VR3_Dzp | 2D | Logarithmic Randic-like eigenvector-based index from Barysz matrix / weighted by polarizabilities |
| 8. | MATS7s | 2D | Moran autocorrelation - lag seven / weighted by I-state |
| 9. | SpMax7_Bhm | 2D | The largest absolute eigenvalue of Burden-modified matrix - n 7 / weighted by relative mass |
| 10. | AATSC8p | 2D | Averaged-centred Broto-Moreau autocorrelation - lag eight / weighted by polarizability |
| Compounds | pMIC (observed) | pMIC (predicted) | Absolute Residual |
|---|---|---|---|
| 2 | 1.699 | 1.697 | 0.002 |
| 17 | 1.097 | 0.546 | 0.551 |
| 81 | 2.301 | 1.854 | 0.447 |
| 87 | 2.602 | 3.138 | 0.536 |
| 101 | 1.991 | 1.809 | 0.182 |
| 113 | 0.772 | 1.136 | 0.364 |
| Parameter | Observed | Threshold value |
|---|---|---|
| R2 (internal) | 0.879 | ≥0.6 |
| R2 pred | 0.752 | ≥0.5 |
| Q2 (LOO) | 0.820 | ≥0.5 |
| Scaled Average rm2 (LOO) | 0.753 | ≥0.5 |
| Scaled Average rm2 (test) | 0.600 | ≥0.5 |
| Scaled Delta rm2(LOO) | 0.081 | ≤0.2 |
| Scaled Delta rm2(test) | 0.185 | ≤0.2 |
| Mean Absolute Error (MAE) | 0.306 | ≤0.1×training set range |
Compounds Library Generation Using RDKit
Molecular Docking – AutoDock Vina Based Analysis
Biological Activity Prediction
Re-Evaluation of Screened Compounds by Calculating Absolute Binding Free Energy Estimation Using KDeep
| Compounds | Structure | IUPAC | Binding energy (kcal/mol) AutoDock Vina |
Absolute binding free energy (kcal/mol) |
|---|---|---|---|---|
| PKD1 | 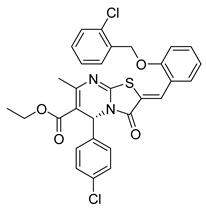 |
Ethyl (R,Z)-2-(2-((2-chlorobenzyl)oxy)benzylidene)-5-(4-chlorophenyl)-7-methyl-3-oxo-2,3-dihydro-5H-thiazolo[3,2-a]pyrimidine-6-carboxylate | -10.100 | -11.530 |
| PKD2 | 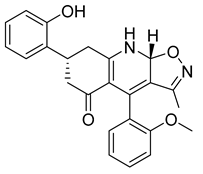 |
(7R,9aR)-7-(2-hydroxyphenyl)-4-(2-methoxyphenyl)-3-methyl-7,8,9,9a-tetrahydroisoxazolo[5,4-b]quinolin-5(6H)-one | -10.500 | -8.350 |
| PKD3 | 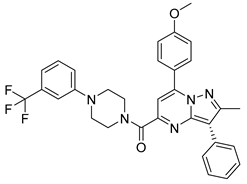 |
(7-(4-methoxyphenyl)-2-methyl-3-phenylpyrazolo[1,5-a]pyrimidin-5-yl)(4-(3-(trifluoromethyl)phenyl)piperazin-1-yl)methanone | -10.500 | -8.750 |
| PKD4 | 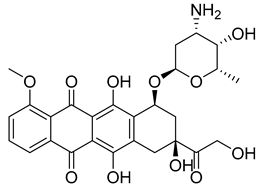 |
(8S,10S)-10-(((2R,4S,5S,6S)-4-amino-5-hydroxy-6-methyltetrahydro-2H-pyran-2-yl)oxy)-6,8,11-trihydroxy-8-(2-hydroxyacetyl)-1-methoxy-7,8,9,10-tetrahydrotetracene-5,12-dione | -10.200 | -10.250 |
| PKD5 | 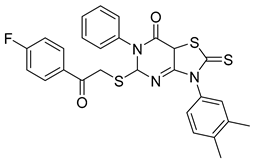 |
3-(3,4-dimethylphenyl)-5-((2-(4-fluorophenyl)-2-oxoethyl)thio)-6-phenyl-2-thioxo-2,3,5,7a-tetrahydrothiazolo[4,5-d]pyrimidin-7(6H)-one | -10.100 | -9.150 |
|
7IJ (Co-crystal) |
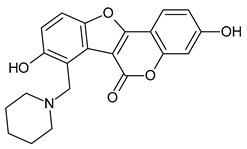 |
3,8-bis(oxidanyl)-7-(piperidin-1-ylmethyl)-[1]benzofuro[3,2-c]chromen-6-one | -12.100 | -9.900 |
| Compounds | Interactions | |||||
|---|---|---|---|---|---|---|
| Hydrogen bonds | Hydrophobic interactions | Pi-stacking | Halogen bonds | Salt bridges | Pi-cation | |
| PKD1 | His1664 | Pro1476, Trp1532, Tyr1637, Asn1640, Tyr1663, Ala1667, Phe1670, Ile1700 | Phe1670, Tyr1674 | Gln1633, Asp1644 | - | - |
| PKD2 | Ser1533, His1664, His1699 | Ile1643, Tyr1663, Ala1667, Ile1700 | Tyr1674 | - | - | - |
| PKD3 | His1664 | Trp1532, Tyr1582, Phe1585, Thr1589, Tyr1637, Asn1640, Ile1700 | Phe1670 | Tyr1637 | - | - |
| PKD4 | Gly1478, Ser1533, His1632, Gln1633, Ser1636, Asn1640, His1699 | Trp1532, Arg1578, Phe1585, Tyr1637, Asn1640, Phe1670, Ile1700 | Phe1670 | - | His1664 | - |
| PKD5 | Ser1533 | Ala1477, Phe1585, Tyr1637, Asn1640, Ile1643, Tyr1663, Ala1667 | Phe1670 | - | - | His1699 |
| 7IJ (Co-crystal) | Gln1633 | Arg1578, Tyr1582, Tyr1637, Asn1640, Ile1643, Tyr1663, Ala1667, Phe1670, Tyr1674 | Phe1670 | - | Asp1644 | - |
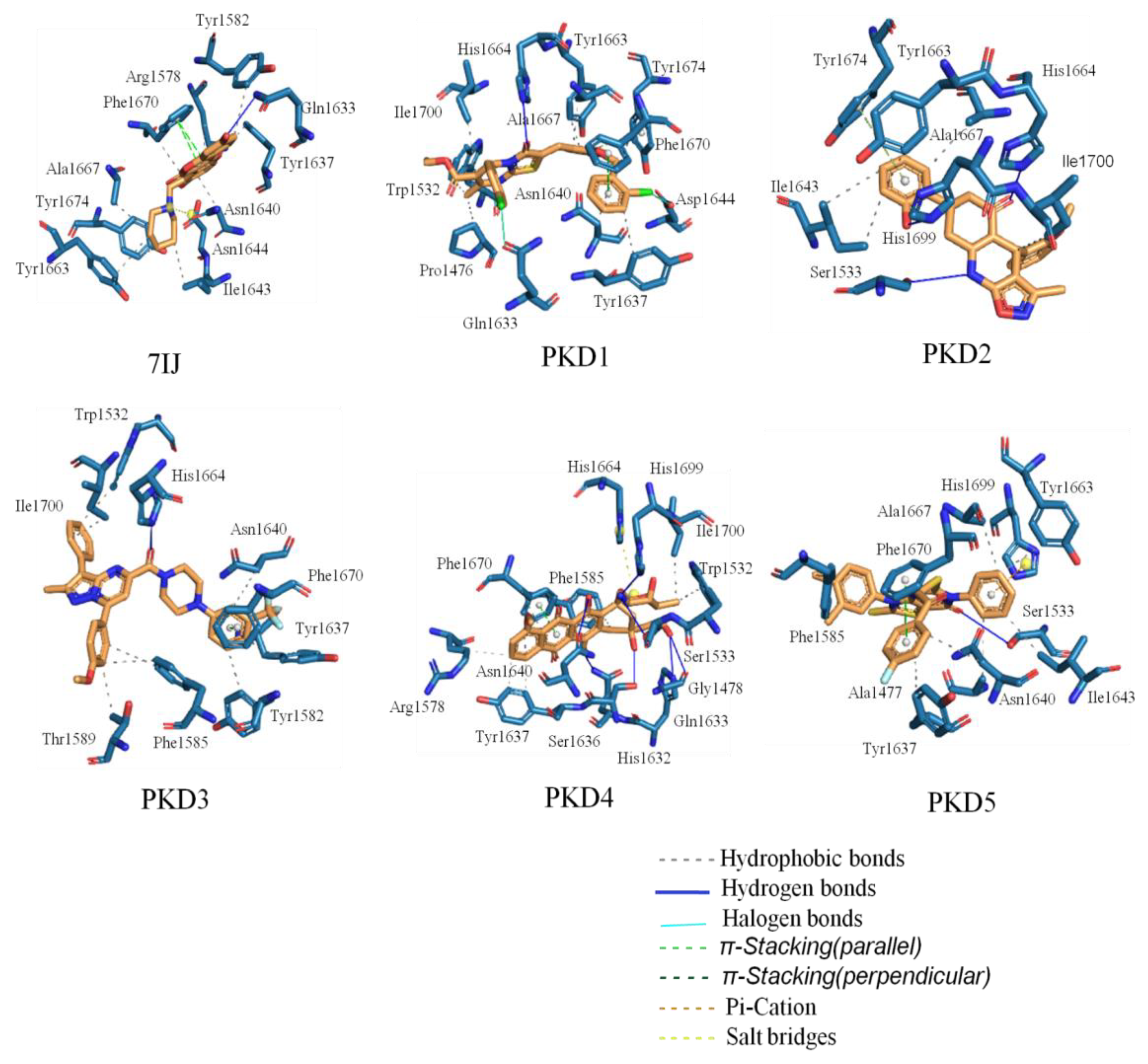
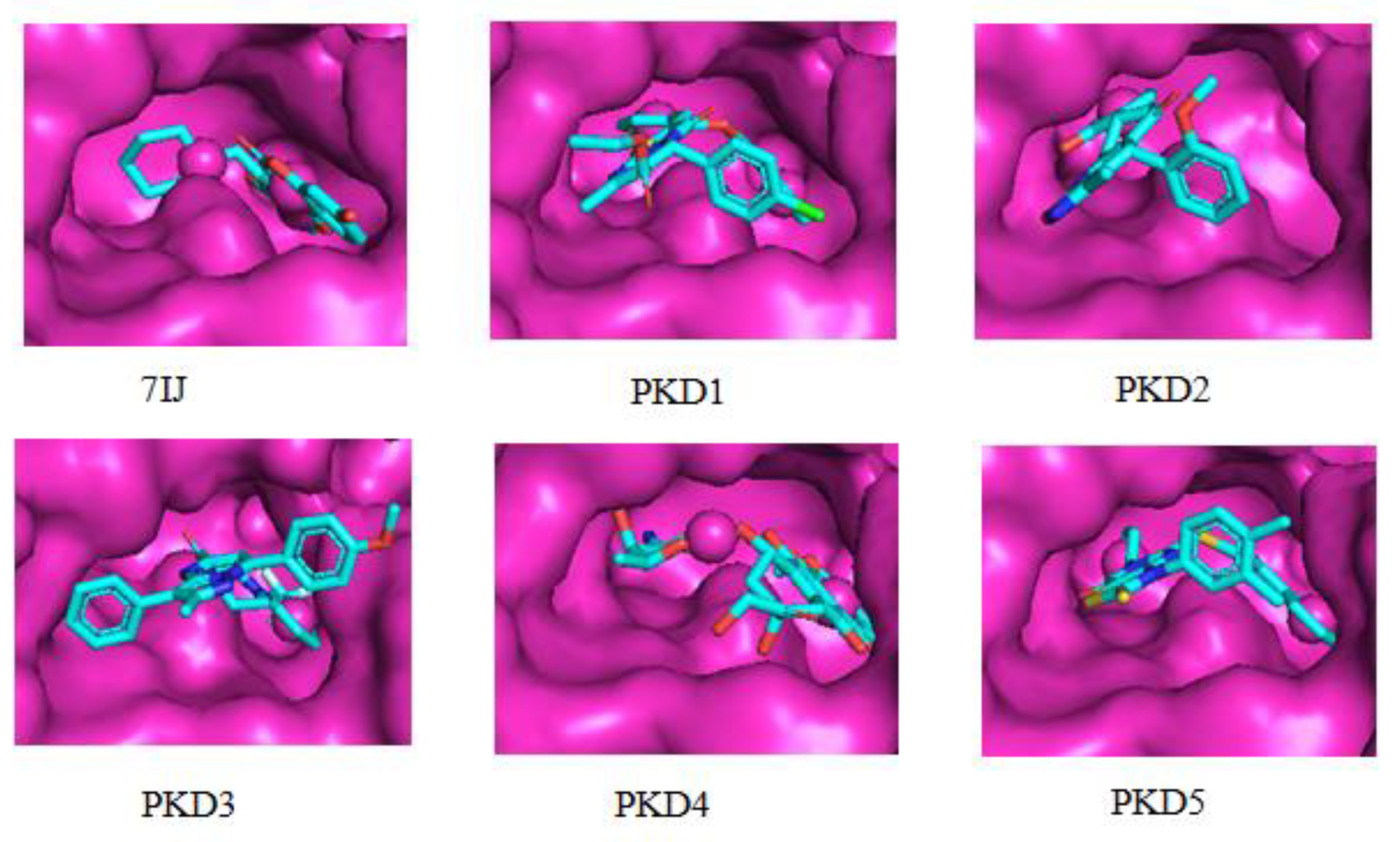
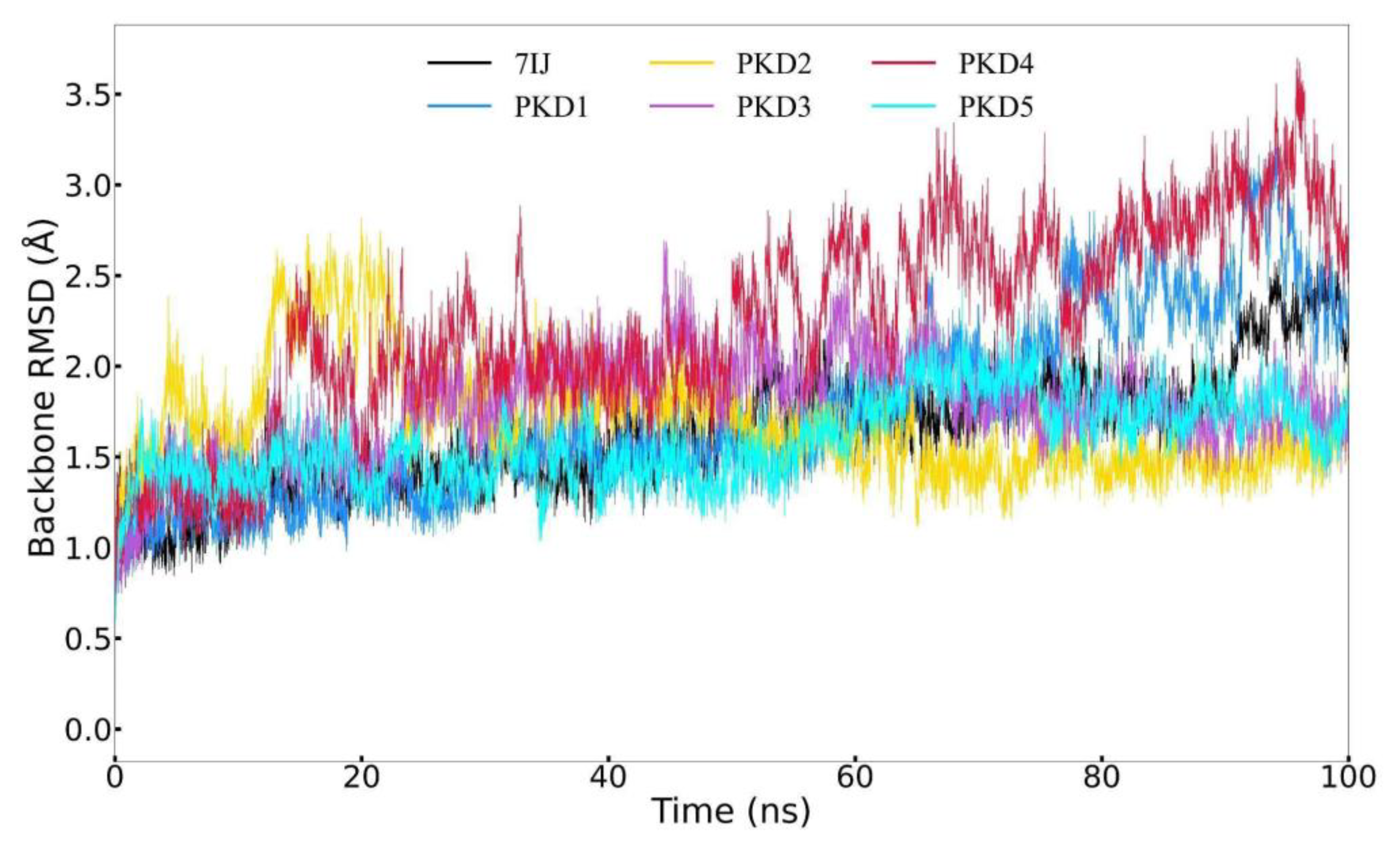
Molecular Dynamic Analysis
| Parameters | 7IJ(Co-crystal) | PKD1 | PKD2 | PKD3 | PKD4 | PKD5 | |
|---|---|---|---|---|---|---|---|
|
Backbone RMSD (Å) |
Average | 1.634 | 1.741 | 1.697 | 1.770 | 2.235 | 1.589 |
| Maximum | 2.588 | 3.300 | 2.815 | 2.691 | 3.697 | 2.436 | |
| Minimum | 0.005 | 0.004 | 0.005 | 0.005 | 0.005 | 0.004 | |
|
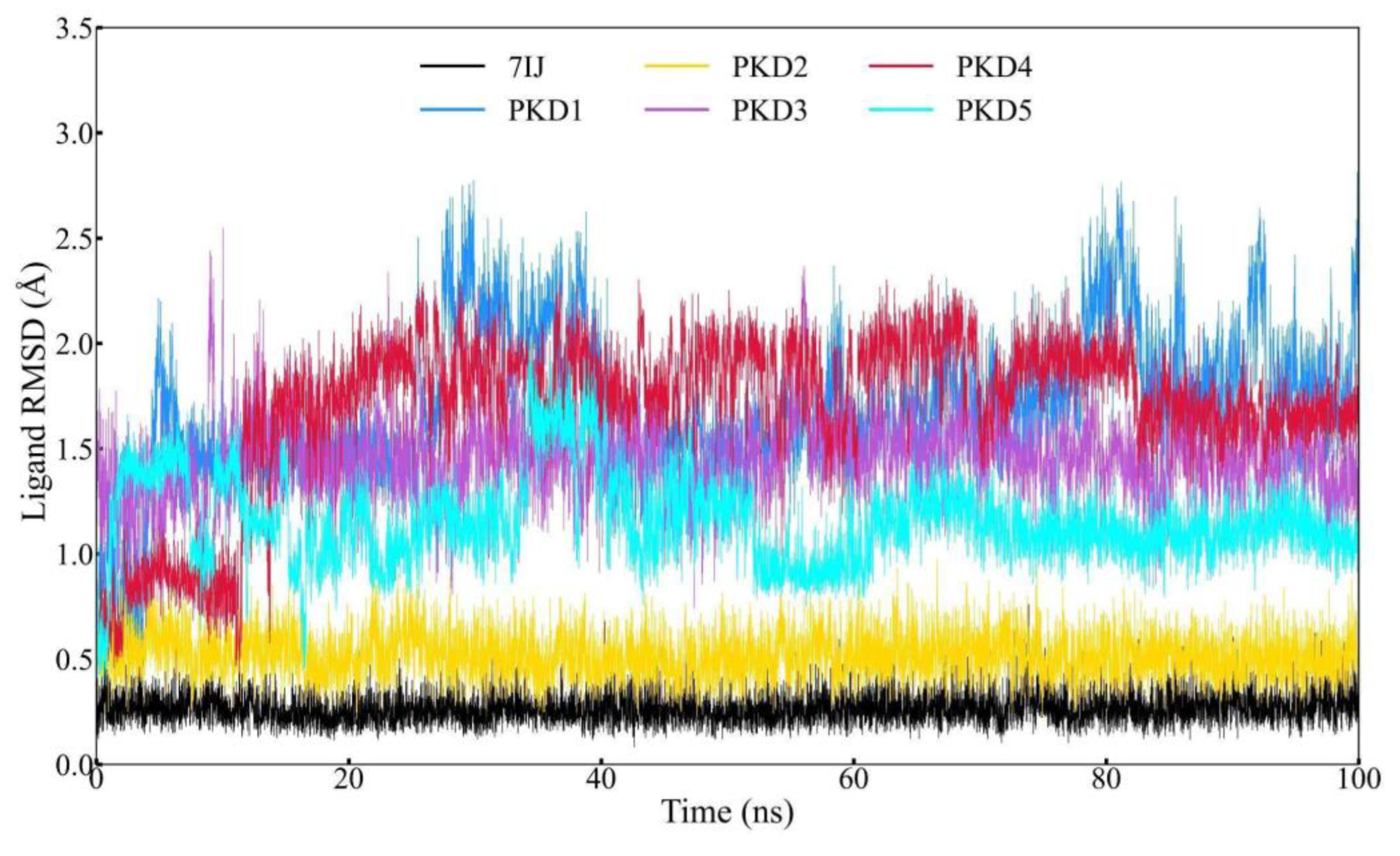
Ligand RMSD (Å) |
Average | 0.269 | 1.685 | 0.518 | 1.445 | 1.693 | 1.156 |
| Maximum | 0.759 | 2.810 | 0.972 | 2.546 | 2.366 | 2.040 | |
| Minimum | 0.004 | 0.004 | 0.004 | 0.004 | 0.004 | 0.004 | |
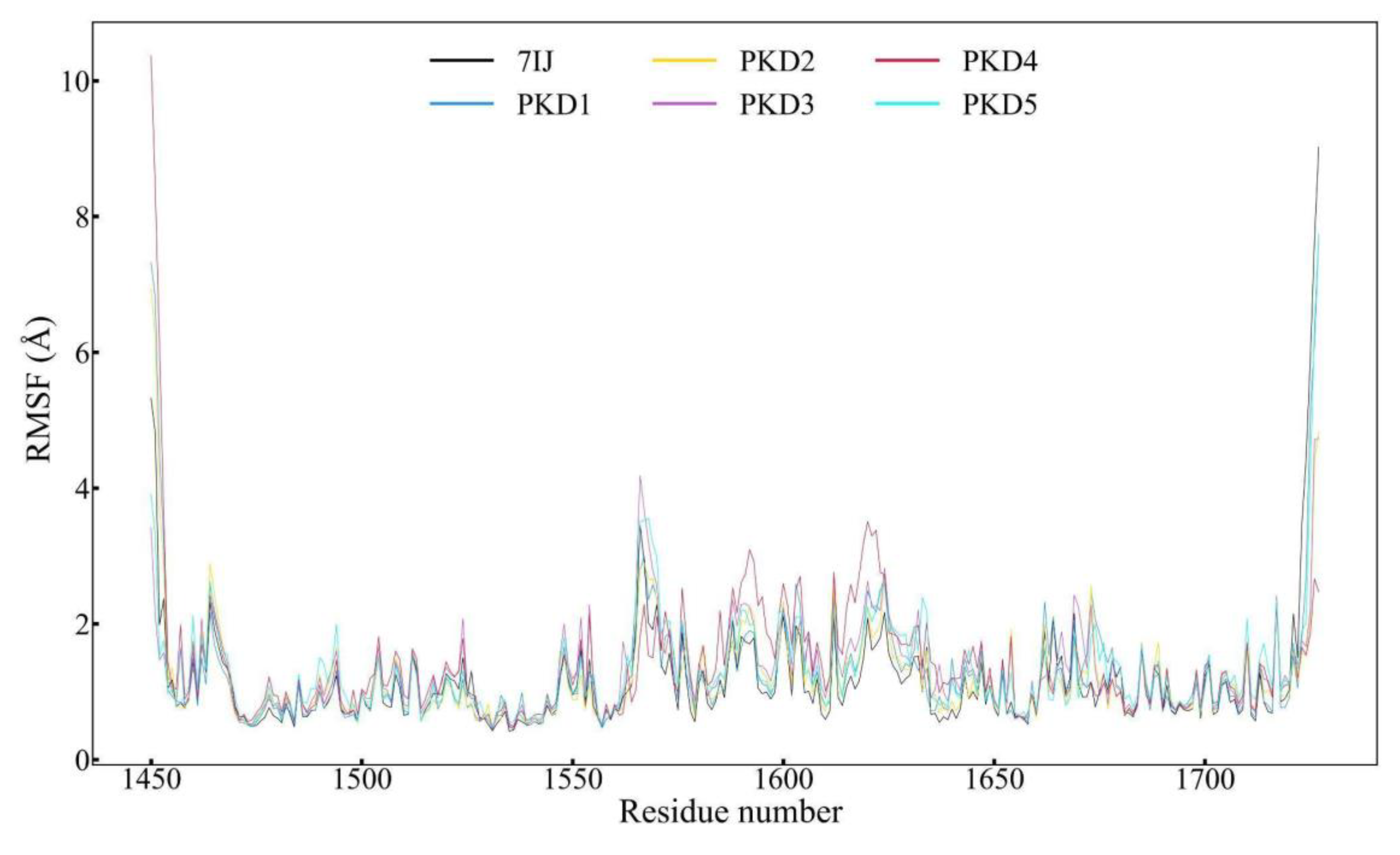
| RMSF (Å) | Average | 1.183 | 1.264 | 1.228 | 1.306 | 1.403 | 1.313 |
| Maximum | 9.020 | 7.703 | 6.931 | 4.734 | 10.365 | 7.740 | |
| Minimum | 0.419 | 0.459 | 0.444 | 0.442 | 0.467 | 0.477 | |
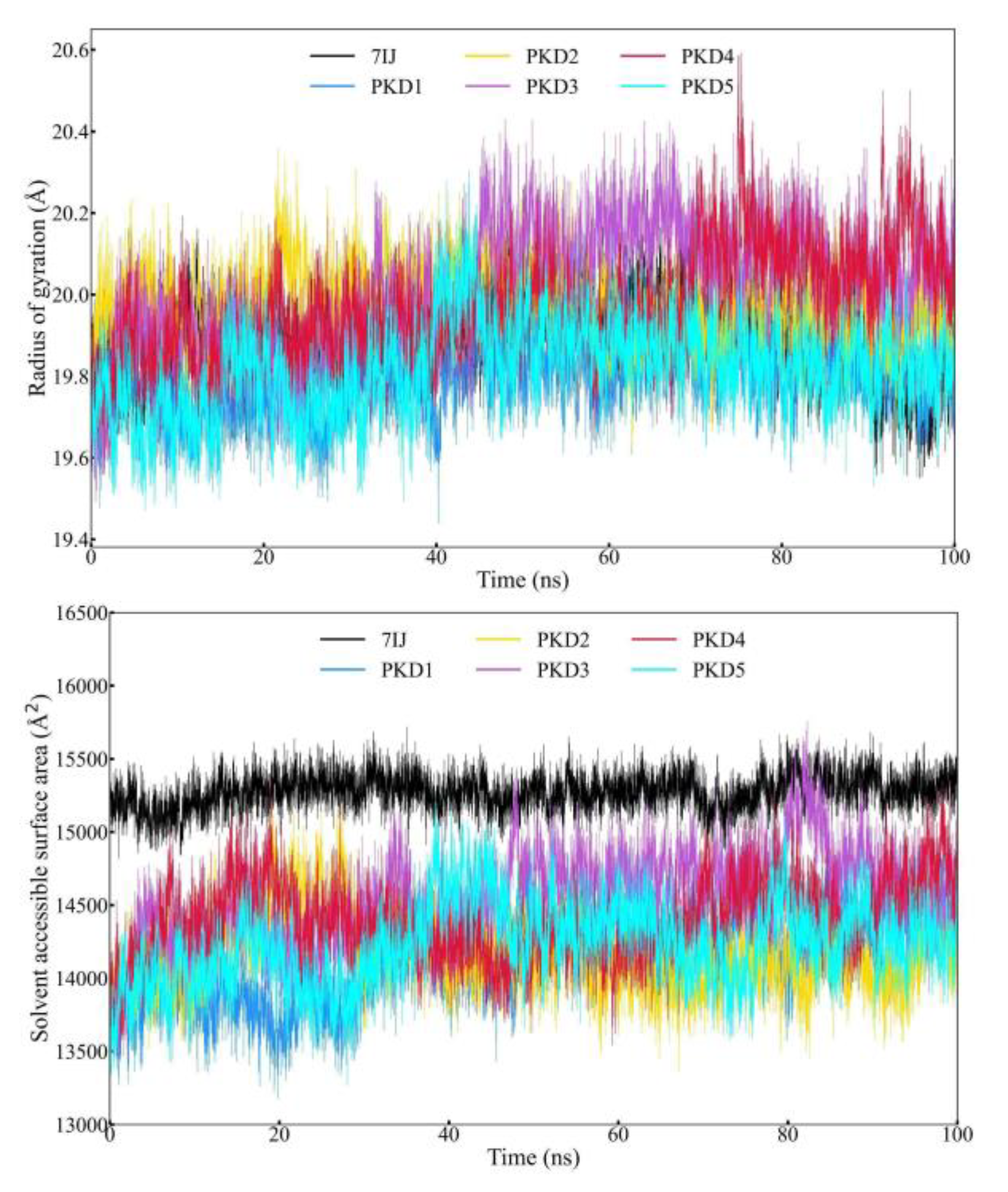
| Rg (Å) | Average | 19.887 | 19.807 | 19.976 | 20.018 | 19.970 | 19.819 |
| Maximum | 20.268 | 20.162 | 20.358 | 20.430 | 20.592 | 20.304 | |
| Minimum | 19.446 | 19.439 | 19.482 | 19.458 | 19.438 | 19.455 | |
| SASA (Å2) | Average | 15273.110 | 14162.140 | 14175.620 | 14555.830 | 14389.040 | 14228.800 |
| Maximum | 15718.000 | 15081.400 | 15204.400 | 15755.600 | 15404.000 | 15254.600 | |
| Minimum | 14799.100 | 13177.900 | 13279.800 | 13180.800 | 13261.500 | 13236.100 |
Root Mean Square Deviation (RMSD) Profile Analysis
Root Mean Square Fluctuation (RMSF) Profile Analysis
Radius of Gyration (Rg) and Solvent Accessible Surface Area (SASA) Profile Analysis
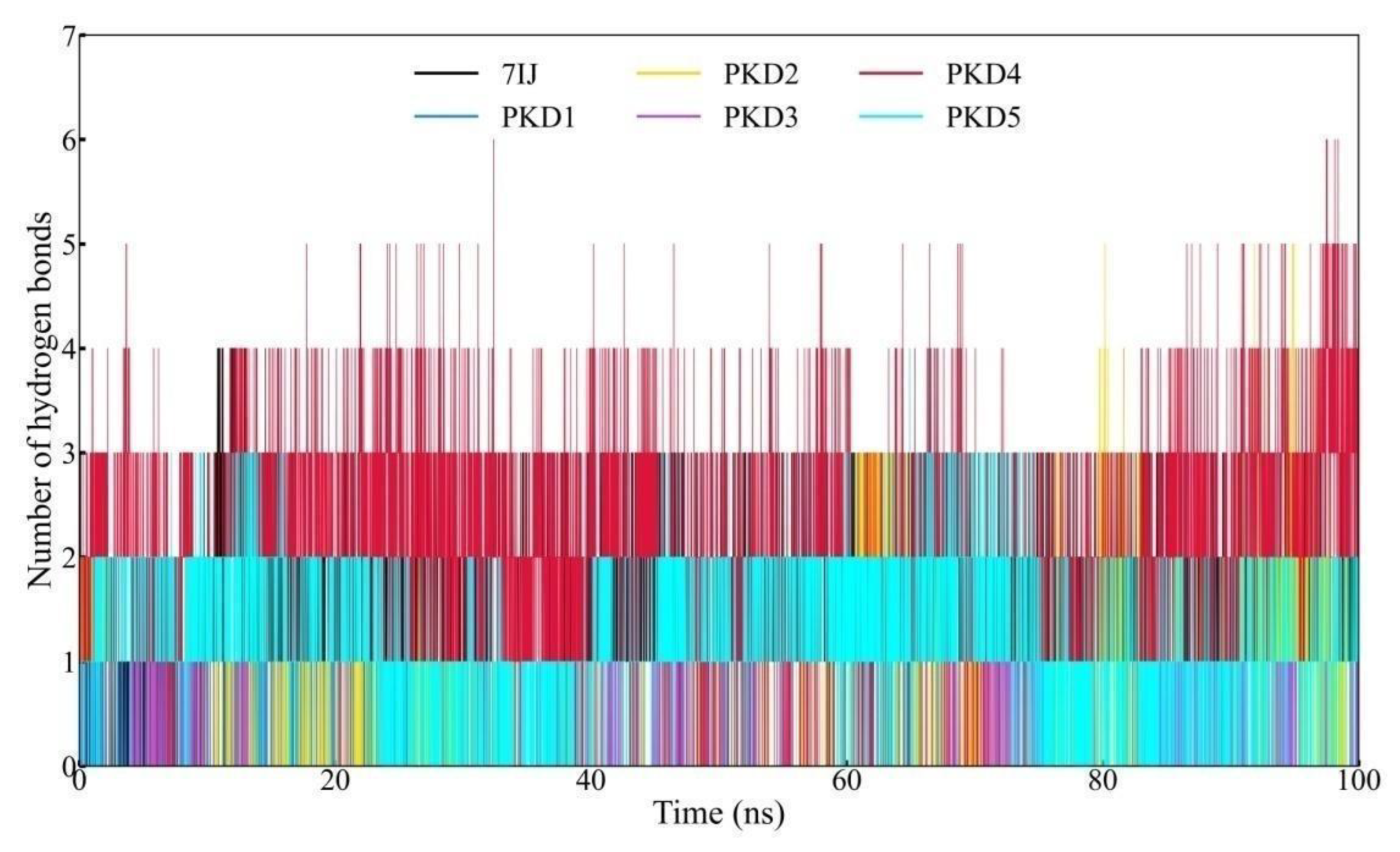
Hydrogen Bond (Hbond) Profile Analysis
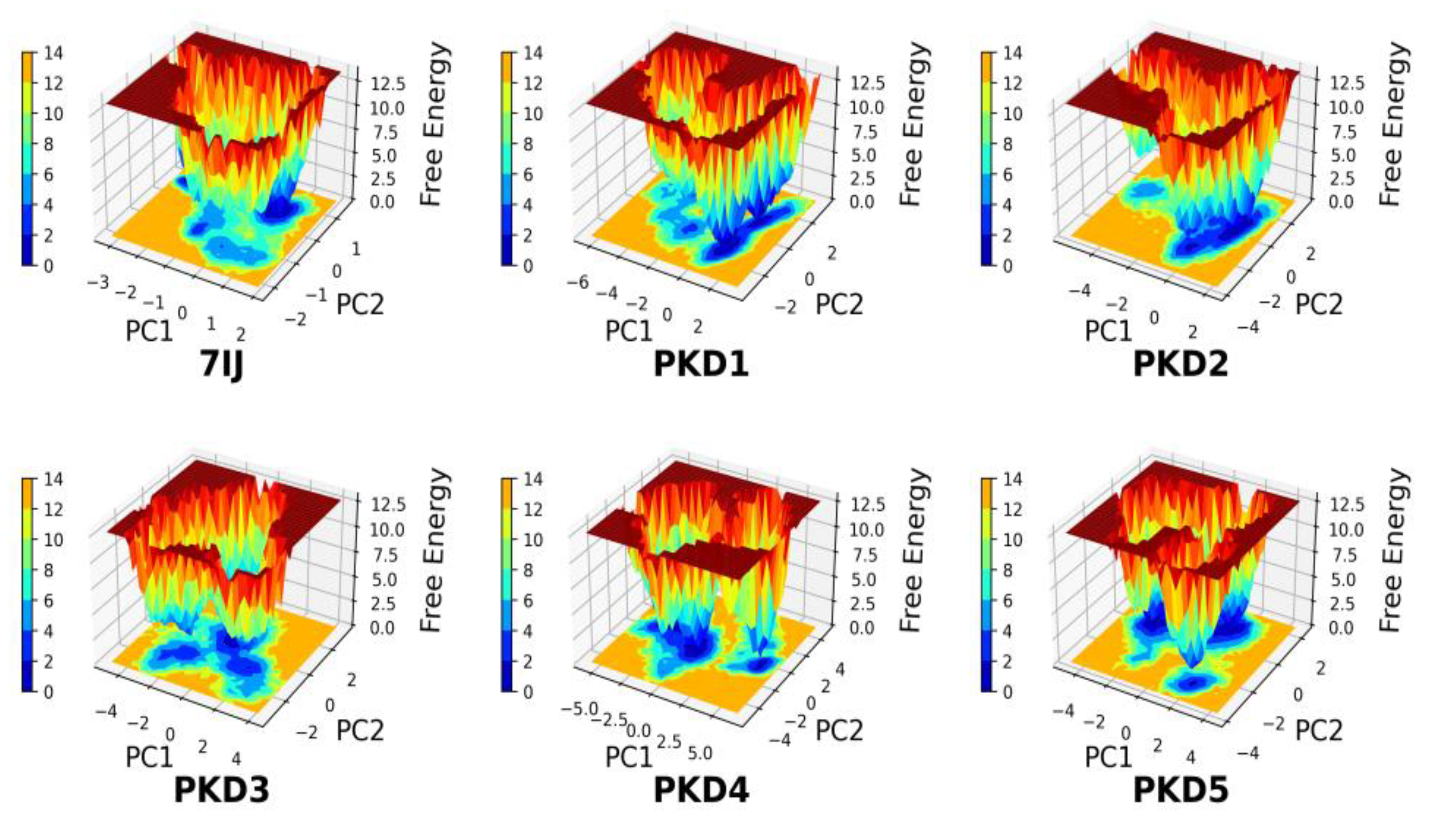
Free energy Landscape (FEL) Analysis
Dynamic Cross-Correlation Map (DCCM) Analyses
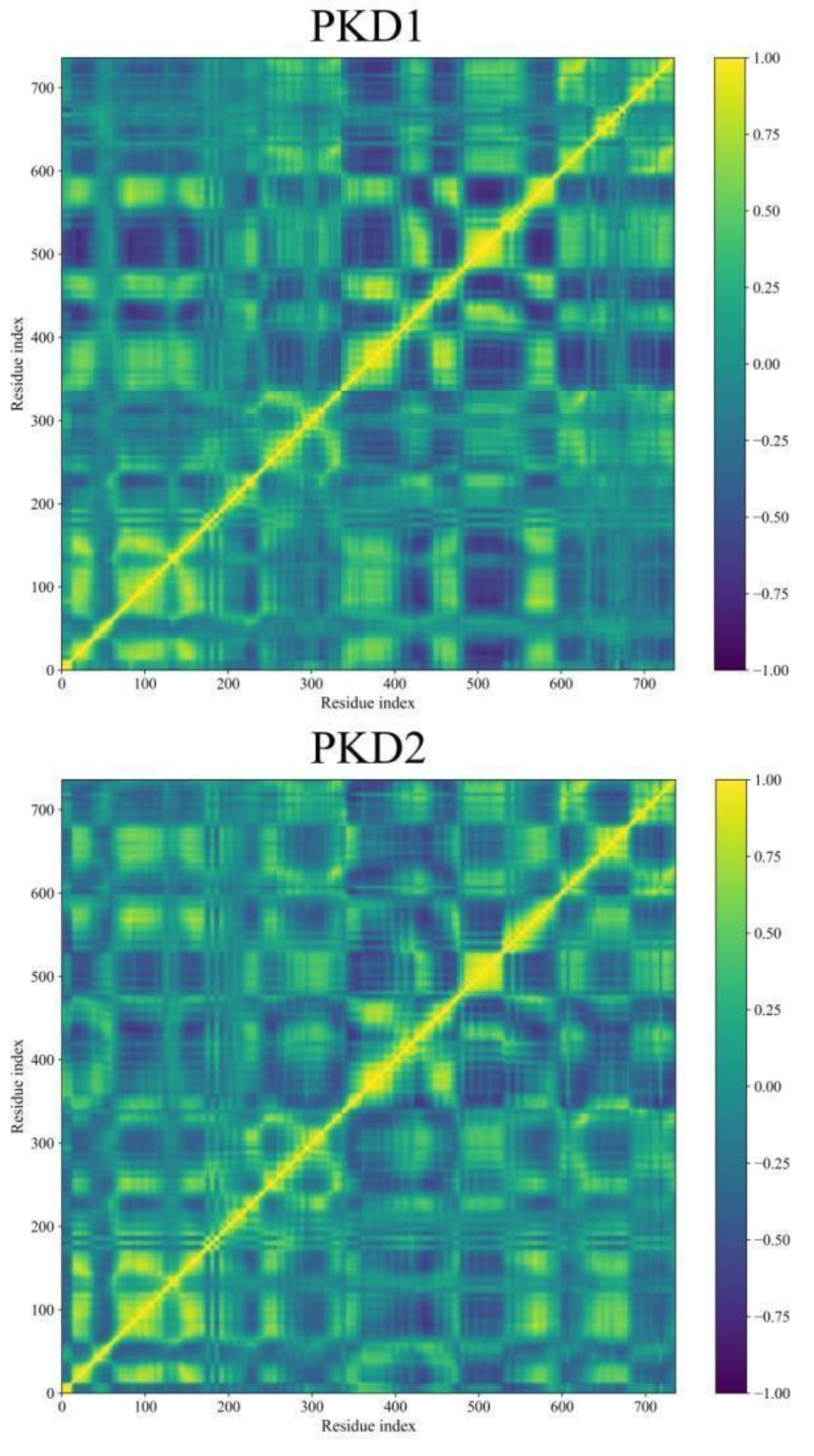
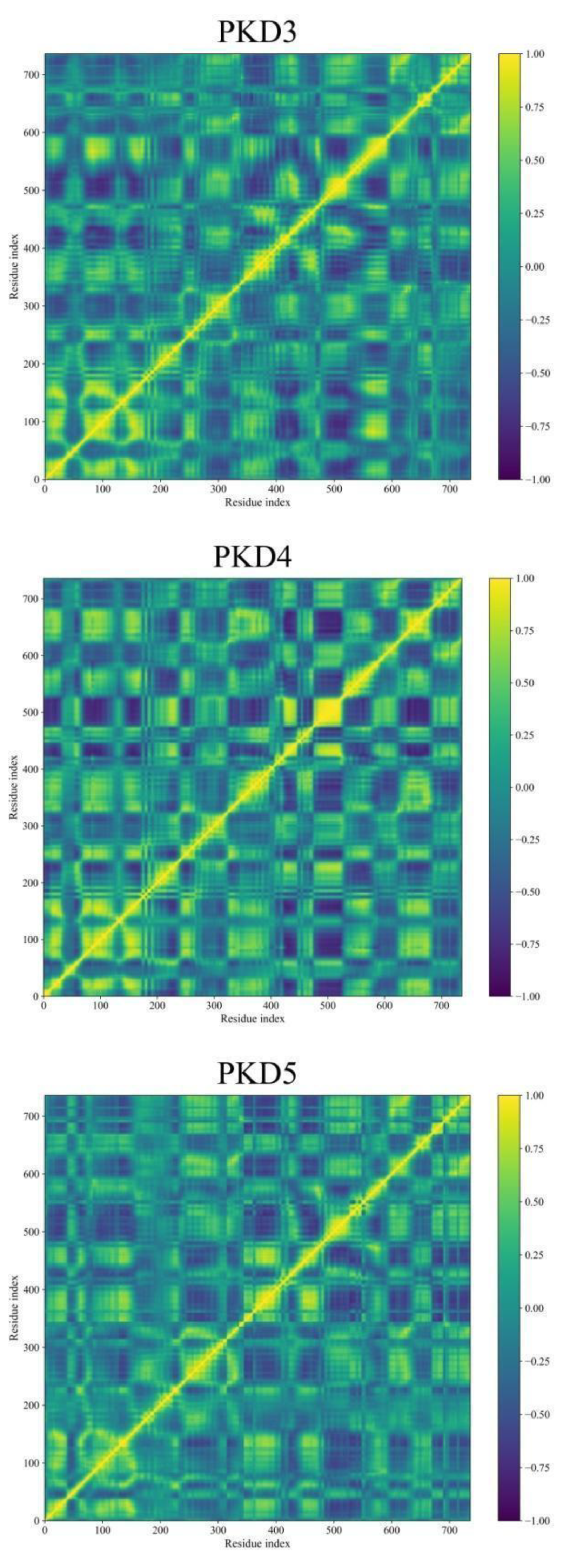
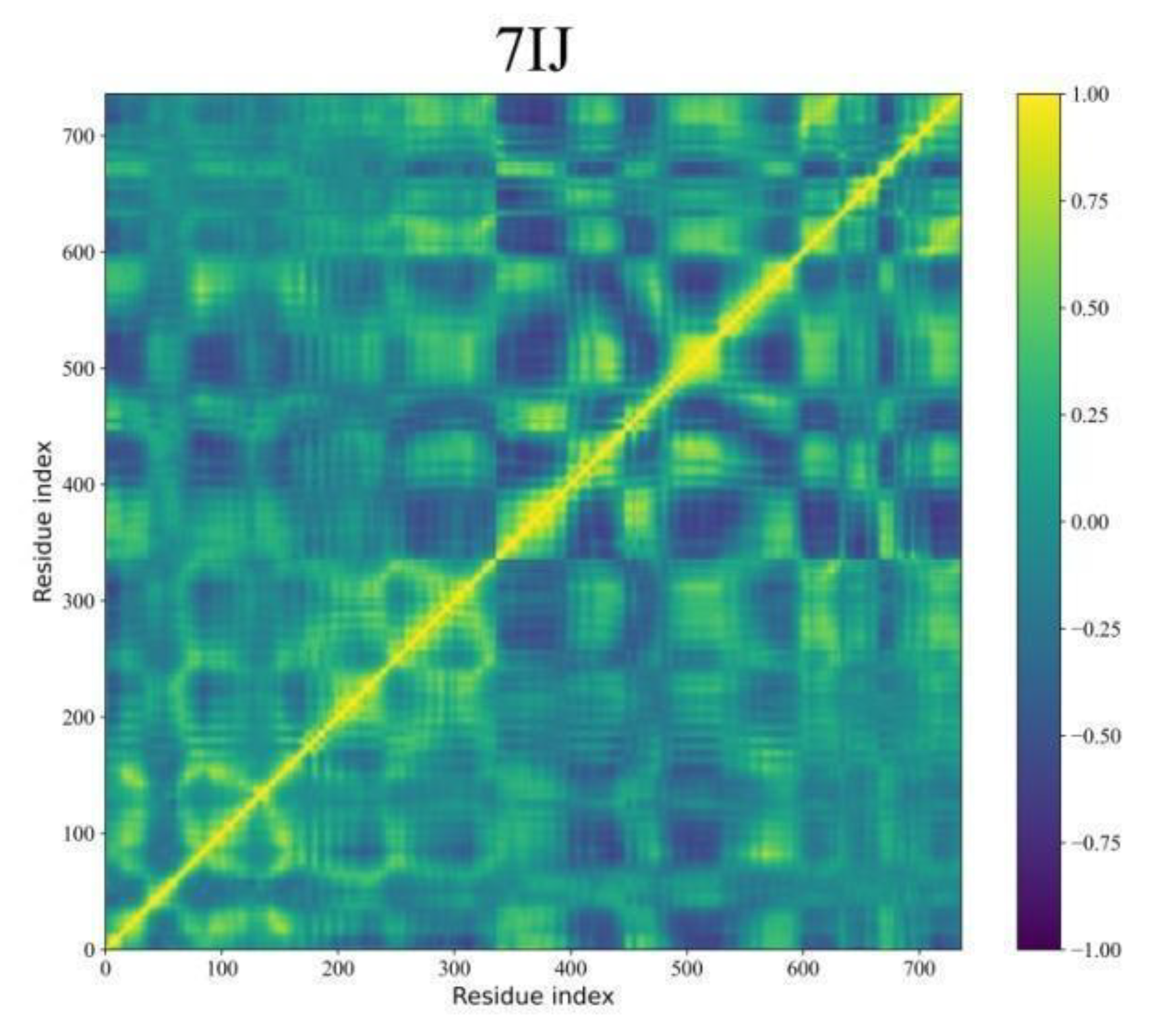
Binding Free Energy Analysis through MM-GBSA Approach
Conclusion
Conflict of Interest
Acknowledgment
Bibliography
- Global tuberculosis report 2023. https://www.who.int/publications/i/item/9789240083851 (accessed 2024-04-22).
- Dartois, V. A.; Rubin, E. J. Anti-Tuberculosis Treatment Strategies and Drug Development: Challenges and Priorities. Nature Reviews Microbiology 2022 20:11 2022, 20 (11), 685–701. [CrossRef]
- Farhat, M.; Cox, H.; Ghanem, M.; Denkinger, C. M.; Rodrigues, C.; Abd El Aziz, M. S.; Enkh-Amgalan, H.; Vambe, D.; Ugarte-Gil, C.; Furin, J.; Pai, M. Drug-Resistant Tuberculosis: A Persistent Global Health Concern. Nature Reviews Microbiology 2024 2024, 1–19. [CrossRef]
- Lee, J. Y. Diagnosis and Treatment of Extrapulmonary Tuberculosis. Tuberc Respir Dis (Seoul) 2015, 78 (2), 47–55. [CrossRef]
- Gavalda, S.; Bardou, F.; Laval, F.; Bon, C.; Malaga, W.; Chalut, C.; Guilhot, C.; Mourey, L.; Daffé, M.; Quémard, A. The Polyketide Synthase Pks13 Catalyzes a Novel Mechanism of Lipid Transfer in Mycobacteria. Chem Biol 2014, 21 (12), 1660–1669. [CrossRef]
- Wilson, R.; Kumar, P.; Parashar, V.; Vilchèze, C.; Veyron-Churlet, R.; Freundlich, J. S.; Barnes, S. W.; Walker, J. R.; Szymonifka, M. J.; Marchiano, E.; Shenai, S.; Colangeli, R.; Jacobs, W. R.; Neiditch, M. B.; Kremer, L.; Alland, D. Antituberculosis Thiophenes Define a Requirement for Pks13 in Mycolic Acid Biosynthesis. Nature Chemical Biology 2013 9:8 2013, 9 (8), 499–506. [CrossRef]
- Zhang, W.; Liu, L. ling; Lun, S.; Wang, S. S.; Xiao, S.; Gunosewoyo, H.; Yang, F.; Tang, J.; Bishai, W. R.; Yu, L. F. Design and Synthesis of Mycobacterial Pks13 Inhibitors: Conformationally Rigid Tetracyclic Molecules. Eur J Med Chem 2021, 213, 113202. [CrossRef]
- Kim, S. K.; Dickinson, M. S.; Finer-Moore, J.; Guan, Z.; Kaake, R. M.; Echeverria, I.; Chen, J.; Pulido, E. H.; Sali, A.; Krogan, N. J.; Rosenberg, O. S.; Stroud, R. M. Structure and Dynamics of the Essential Endogenous Mycobacterial Polyketide Synthase Pks13. Nature Structural & Molecular Biology 2023 30:3 2023, 30 (3), 296–308. [CrossRef]
- Chikhale, R. V; Eldesoky, E.; Kolpe, M. S.; Suryawanshi, V. S.; Patil, P. C.; Bhowmick, S. Identification of Mycobacterium Tuberculosis Transcriptional Repressor EthR Inhibitors: Shape-Based Search and Machine Learning Studies. 2024. [CrossRef]
- Chikhale, R. V.; Abdelghani, H. T. M.; Deka, H.; Pawar, A. D.; Patil, P. C.; Bhowmick, S. Machine Learning Assisted Methods for the Identification of Low Toxicity Inhibitors of Enoyl-Acyl Carrier Protein Reductase (InhA). Comput Biol Chem 2024, 110, 108034. [CrossRef]
- Liu, P.; Long, W. Current Mathematical Methods Used in QSAR/QSPR Studies. Int J Mol Sci 2009, 10 (5), 1978. [CrossRef]
- Lun, S.; Xiao, S.; Zhang, W.; Wang, S.; Gunosewoyo, H.; Yu, L. F.; Bishai, W. R. Therapeutic Potential of Coumestan Pks13 Inhibitors for Tuberculosis. Antimicrob Agents Chemother 2021, 65 (5). [CrossRef]
- Yap, C. W. PaDEL-Descriptor: An Open Source Software to Calculate Molecular Descriptors and Fingerprints. J Comput Chem 2011, 32 (7), 1466–1474. [CrossRef]
- Ambure, P.; Aher, R. B.; Gajewicz, A.; Puzyn, T.; Roy, K. “NanoBRIDGES” Software: Open Access Tools to Perform QSAR and Nano-QSAR Modeling. Chemometrics and Intelligent Laboratory Systems 2015, 147, 1–13. [CrossRef]
- Roy, K.; Mitra, I. On Various Metrics Used for Validation of Predictive QSAR Models with Applications in Virtual Screening and Focused Library Design. Comb Chem High Throughput Screen 2011, 14 (6), 450–474. [CrossRef]
- Roy, K.; Chakraborty, P.; Mitra, I.; Ojha, P. K.; Kar, S.; Das, R. N. Some Case Studies on Application of “Rm2” Metrics for Judging Quality of Quantitative Structure–Activity Relationship Predictions: Emphasis on Scaling of Response Data. J Comput Chem 2013, 34 (12), 1071–1082. [CrossRef]
- Zhang, W.; Lun, S.; Wang, S. S.; Cai, Y. P.; Yang, F.; Tang, J.; Bishai, W. R.; Yu, L. F. Structure-Based Optimization of Coumestan Derivatives as Polyketide Synthase 13-Thioesterase(Pks13-TE) Inhibitors with Improved HERG Profiles for Mycobacterium Tuberculosis Treatment. J Med Chem 2022, 65 (19), 13240–13252. [CrossRef]
- Compound Libraries/Screening Libraries for High Throughput/Content Screening|96-Well. https://www.selleckchem.com/screening-libraries.html (accessed 2024-04-22).
- Screening Libraries. https://www.chemdiv.com/catalog/screening-libraries/ (accessed 2024-04-22).
- Gaulton, A.; Bellis, L. J.; Bento, A. P.; Chambers, J.; Davies, M.; Hersey, A.; Light, Y.; McGlinchey, S.; Michalovich, D.; Al-Lazikani, B.; Overington, J. P. ChEMBL: A Large-Scale Bioactivity Database for Drug Discovery. Nucleic Acids Res 2012, 40 (D1), D1100–D1107. [CrossRef]
- Diversity Libraries - Enamine. https://enamine.net/compound-libraries/diversity-libraries (accessed 2024-04-22).
- Asinex.com - Antibacterial - Research Areas - Screening Libraries. https://www.asinex.com/antibacterial (accessed 2024-04-22).
- The RDKit Documentation — The RDKit 2024.03.1 documentation. https://rdkit.org/docs/index.html (accessed 2024-04-22).
- Ellingson, S. R.; Smith, J. C.; Baudry, J. VinaMPI: Facilitating Multiple Receptor High-Throughput Virtual Docking on High-Performance Computers. J Comput Chem 2013, 34 (25), 2212–2221. [CrossRef]
- Morris, G. M.; Ruth, H.; Lindstrom, W.; Sanner, M. F.; Belew, R. K.; Goodsell, D. S.; Olson, A. J. AutoDock4 and AutoDockTools4: Automated Docking with Selective Receptor Flexibility. J Comput Chem 2009, 30 (16), 2785. [CrossRef]
- O’Boyle, N. M.; Banck, M.; James, C. A.; Morley, C.; Vandermeersch, T.; Hutchison, G. R. Open Babel: An Open Chemical Toolbox. J Cheminform 2011, 3 (10), 1–14. [CrossRef]
- Jiménez, J.; Škalič, M.; Martínez-Rosell, G.; De Fabritiis, G. KDEEP: Protein-Ligand Absolute Binding Affinity Prediction via 3D-Convolutional Neural Networks. J Chem Inf Model 2018, 58 (2), 287–296. [CrossRef]
- Wang, R.; Fang, X.; Lu, Y.; Yang, C. Y.; Wang, S. The PDBbind Database: Methodologies and Updates. J Med Chem 2005, 48 (12), 4111–4119. [CrossRef]
- van Gunsteren, W. F.; Berendsen, H. J. C. Computer Simulation of Molecular Dynamics: Methodology, Applications, and Perspectives in Chemistry. Angewandte Chemie International Edition in English 1990, 29 (9), 992–1023. [CrossRef]
- Valdés-Tresanco, M. S.; Valdés-Tresanco, M. E.; Valiente, P. A.; Moreno, E. Gmx_MMPBSA: A New Tool to Perform End-State Free Energy Calculations with GROMACS. J Chem Theory Comput 2021, 17 (10), 6281–6291. [CrossRef]
| Compounds | ΔEVDW | ΔEELE | ΔGGB | ΔGSurf | ΔGgas | ΔGSol | ΔGbind |
|---|---|---|---|---|---|---|---|
| PKD1 | -50.430 (4.110) |
-8.620 (3.330) |
31.770 (3.610) |
-6.090 (0.430) |
-59.050 (6.110) |
25.680 (3.410) |
-33.370 (4.090) |
| PKD2 | -31.800 (3.100) |
-4.880 (4.230) |
24.500 (3.270) |
-4.050 (0.350) |
-36.680 (4.890) |
20.440 (3.140) |
-16.240 (2.670) |
| PKD3 | -47.160 (7.420) |
180.160 (9.180) |
-133.660 (10.920) |
-5.850 (0.850) |
133.000 (11.700) |
-139.520 (10.430) |
-6.520 (4.440) |
| PKD4 | -49.970 (4.750) |
-274.170 (19.210) |
298.300 (15.560) |
-6.690 (0.500) |
-324.140 (19.790) |
291.610 (15.350) |
-32.530 (6.950) |
| PKD5 | -55.830 (3.550) |
-24.150 (7.250) |
43.500 (4.210) |
-6.910 (0.350) |
-79.970 (7.150) |
36.600 (4.300) |
-43.380 (4.630) |
| 7IJ | -45.300 (3.810) |
-24.580 (6.330) |
38.690 (4.060) |
-5.560 (0.200) |
-69.870 (5.370) |
33.140 (4.090) |
-36.740 (3.500) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




