Submitted:
30 March 2024
Posted:
01 April 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
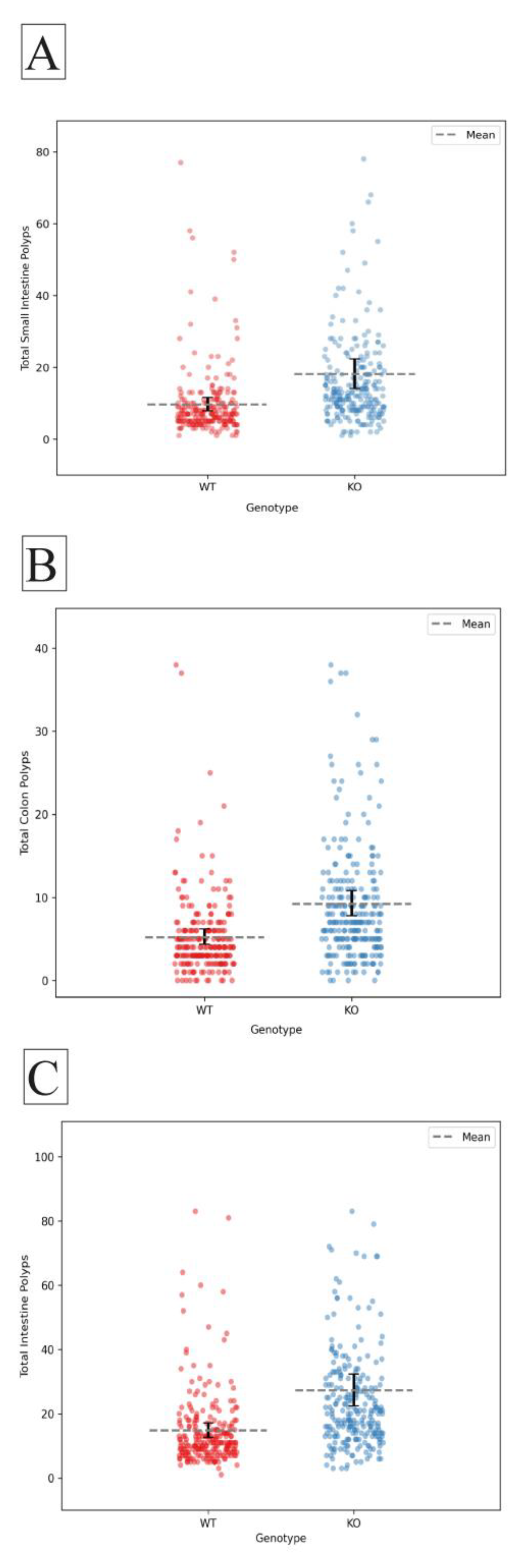
2.1. The Effect of Smad4 Heterozygous Kock Out in the General Population.
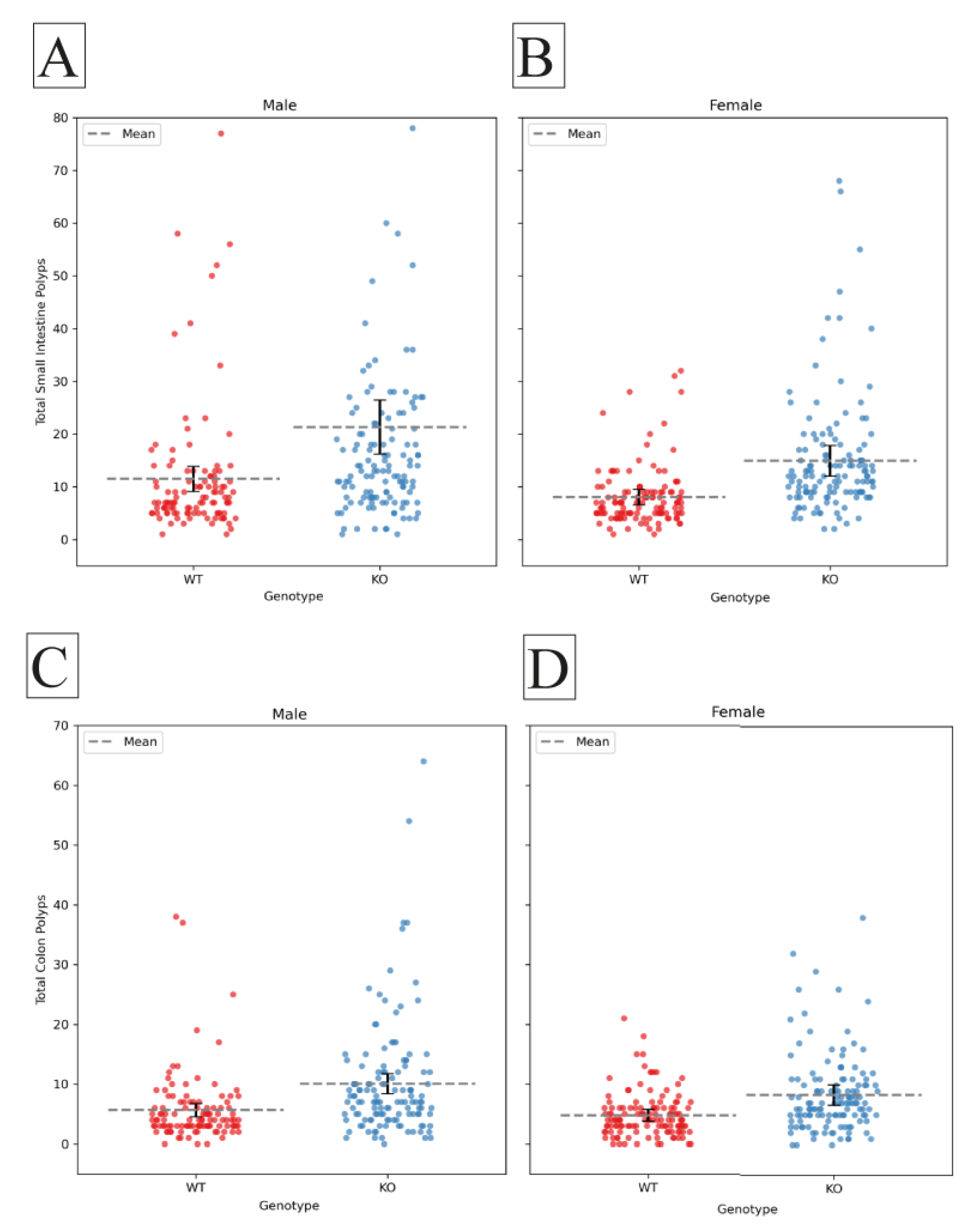
2.2. Sex effect
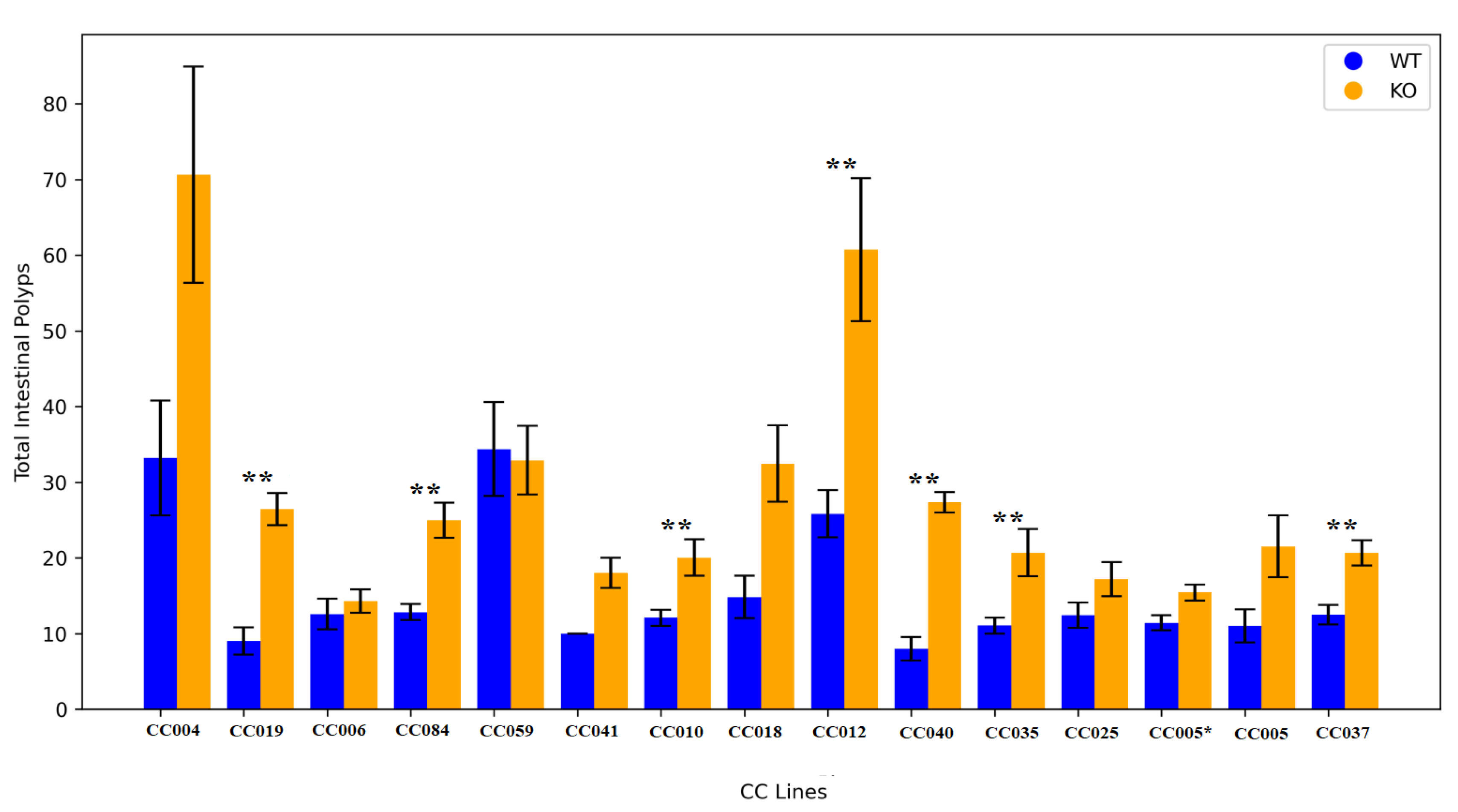
2.3. Line Genetic Effect
2.4. Multimorbidity Heatmaps of Polyp Counts, Body Weight
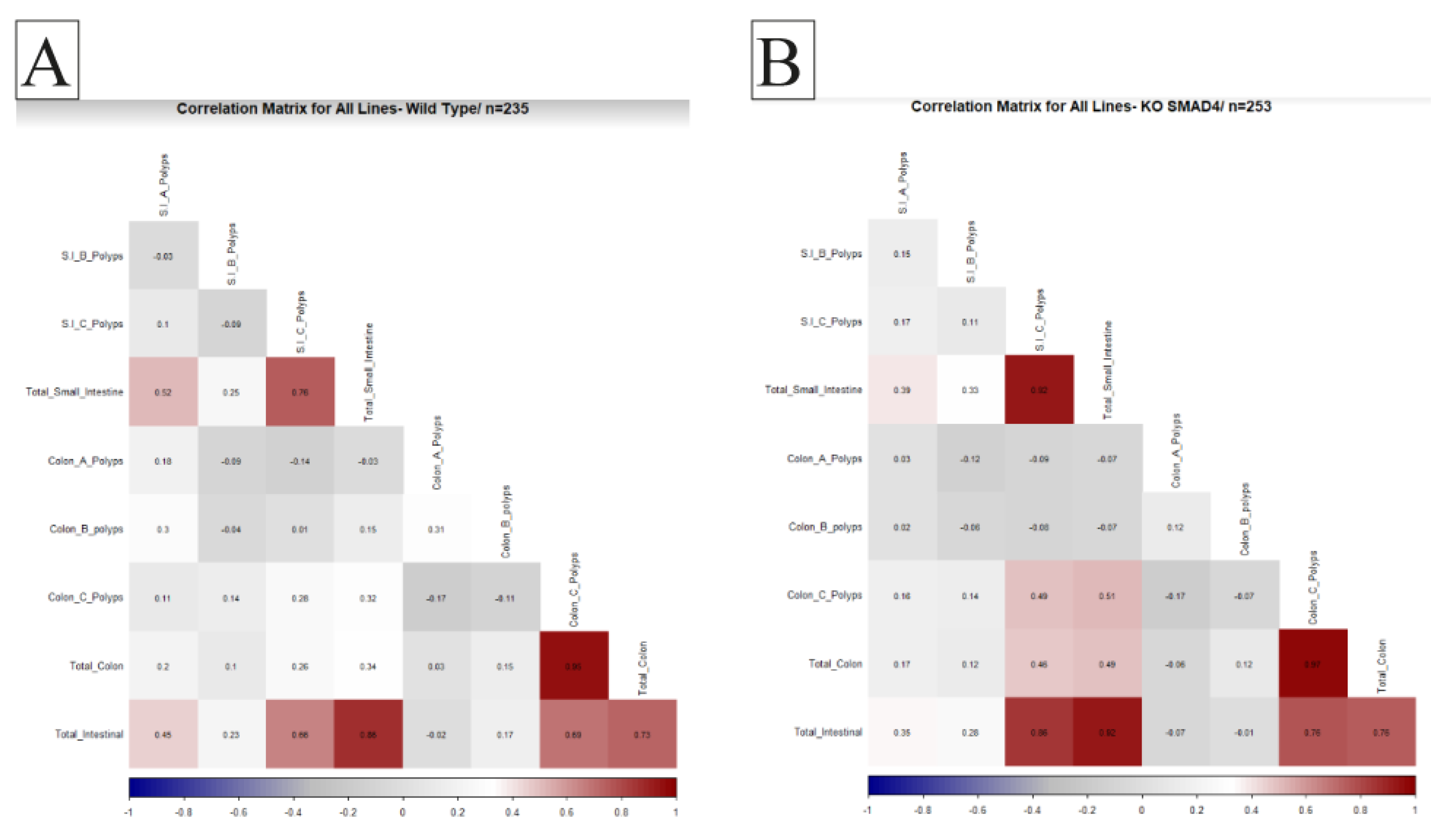
2.5. General Population Correlations
2.6. Sex Variation
2.7. Polyp Counts Correlations in Different Lines
2.8. Heritability
2.9. Machine Learning
3. Discussion
4. Materials and Methods
4.1. Ethical Aspects of the Project
4.2. Generation of F1 Crosses
4.3. Mouse Housing and Diet
4.4. Genomic DNA Extraction and Genotyping
4.5. Genotyping of F1 Mice
4.6. Tissue Collection
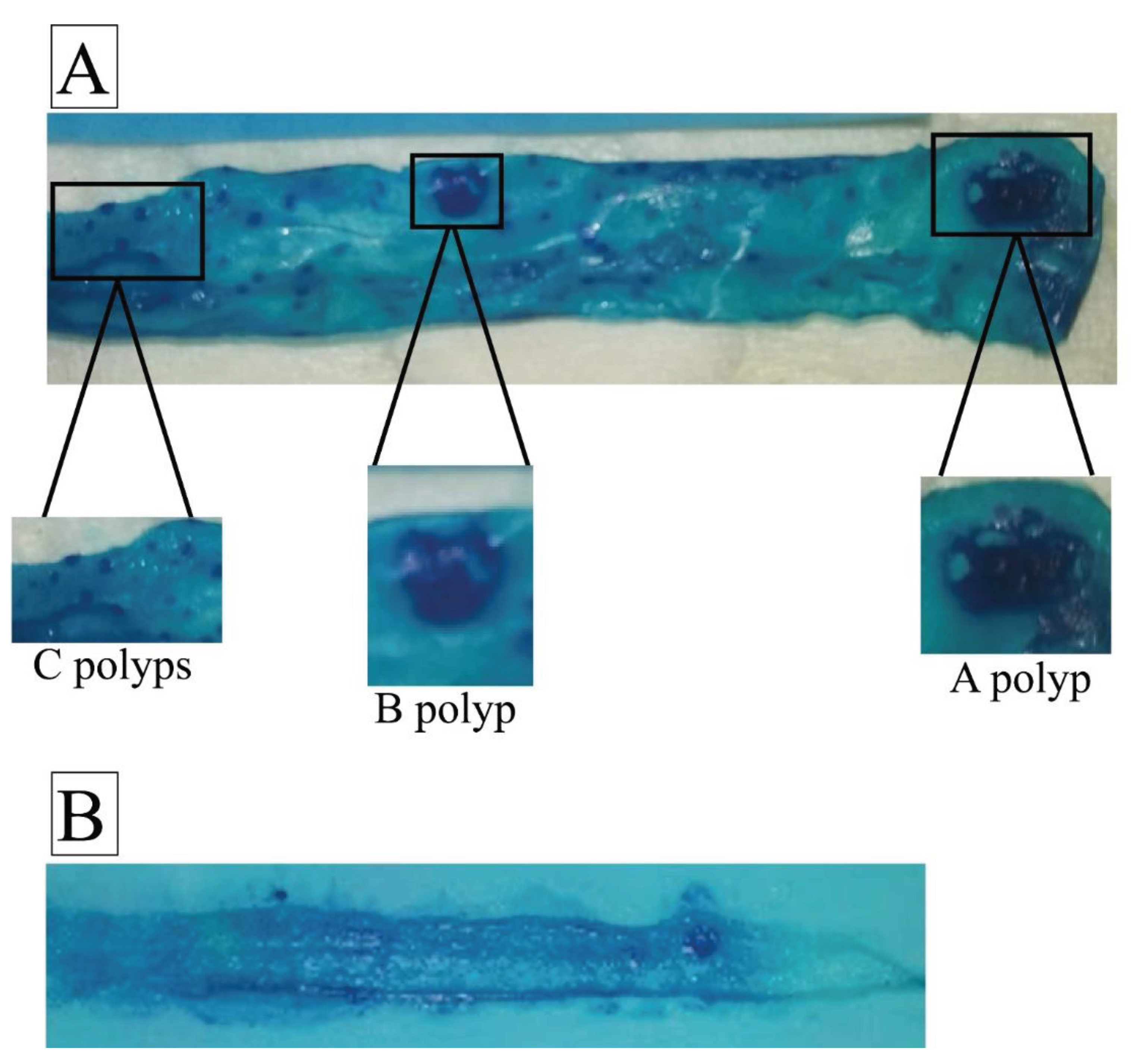
4.7. Intestine Whole Mounts Preparation and Assessing the Intestinal Polyp Counts
4.8. Estimation of the Heritability of The Assessed Phenotypes
4.9. Statistical Analysis
4.10. Machine Learning
4.11. Data Preprocessing
4.12. Model Evaluation
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Conflict of Interest
Ethics Statements
Informed Consent Statement
Data Availability Statement
References
- Van Hattem, W.A.; Brosens, L.A.A.; De Leng, W.W.J.; Morsink, F.H.; Lens, S.; Carvalho, R.; Giardiello, F.M.; Offerhaus, G.J.A. Large Genomic Deletions of SMAD4, BMPR1A and PTEN in Juvenile Polyposis. Gut 2008, 57, 623–627. [Google Scholar] [CrossRef]
- van Hattem, W.A.; Langeveld, D.; de Leng, W.W.J.; Morsink, F.H.; van Diest, P.J.; Iacobuzio-Donahue, C.A.; Giardiello, F.M.; Offerhaus, G.J.A.; Brosens, L.A.A. Histological Variations in Juvenile Polyp Phenotype Correlate with Genetic Defect Underlying Juvenile Polyposis. Am J Surg Pathol 2011, 35, 530. [Google Scholar] [CrossRef] [PubMed]
- Blatter, R.; Tschupp, B.; Aretz, S.; Bernstein, I.; Colas, C.; Evans, D.G.; Genuardi, M.; Hes, F.J.; Hüneburg, R.; Järvinen, H.; et al. Disease Expression in Juvenile Polyposis Syndrome: A Retrospective Survey on a Cohort of 221 European Patients and Comparison with a Literature-Derived Cohort of 473 SMAD4/BMPR1A Pathogenic Variant Carriers. Genetics in Medicine 2020 22:9 2020, 22, 1524–1532. [Google Scholar] [CrossRef] [PubMed]
- Brosens, L.A.A.; Langeveld, D.; van Hattem, W.A.; Giardiello, F.M.; Offerhaus, G.J.A. Juvenile Polyposis Syndrome. World Journal of Gastroenterology : WJG 2011, 17, 4839. [Google Scholar] [CrossRef] [PubMed]
- Malki, A.; Abu Elruz, R.; Gupta, I.; Allouch, A.; Vranic, S.; al Moustafa, A.-E. Molecular Sciences Molecular Mechanisms of Colon Cancer Progression and Metastasis: Recent Insights and Advancements. 2020. [CrossRef]
- Yamagishi, H.; Kuroda, H.; Imai, Y.; Hiraishi, H. Molecular Pathogenesis of Sporadic Colorectal Cancers. Chin J Cancer 2016, 35, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Jelsig, A.M.; Ousager, L.B.; Brusgaard, K.; Qvist, N. Juvenile Polyps in Denmark from 1995 to 2014. Dis Colon Rectum 2016, 59, 751–757. [Google Scholar] [CrossRef]
- Phen, C.; Rojas, I. Paediatric Polyposis Syndromes: Burden of Disease and Current Concepts. Curr Opin Pediatr 2021, 33, 509–514. [Google Scholar] [CrossRef] [PubMed]
- Latchford, A.R.; Neale, K.; Phillips, R.K.S.; Clark, S.K. Juvenile Polyposis Syndrome: A Study of Genotype, Phenotype, and Long-Term Outcome. Dis Colon Rectum 2012, 55, 1038–1043. [Google Scholar] [CrossRef] [PubMed]
- Aytac, E.; Sulu, B.; Heald, B.; O’Malley, M.; Laguardia, L.; Remzi, F.H.; Kalady, M.F.; Burke, C.A.; Church, J.M. Genotype-Defined Cancer Risk in Juvenile Polyposis Syndrome. British Journal of Surgery 2014, 102, 114–118. [Google Scholar] [CrossRef] [PubMed]
- Ishida, H.; Ishibashi, K.; Iwama, T. Malignant Tumors Associated with Juvenile Polyposis Syndrome in Japan. Surg Today 2018, 48, 253–263. [Google Scholar] [CrossRef] [PubMed]
- MacFarland, S.P.; Ebrahimzadeh, J.E.; Zelley, K.; Begum, L.; Bass, L.M.; Brand, R.E.; Dudley, B.; Fishman, D.S.; Ganzak, A.; Karloski, E.; et al. Phenotypic Differences in Juvenile Polyposis Syndrome with or without a Disease-Causing SMAD4/BMPR1A Variant. Cancer Prev Res (Phila) 2021, 14, 215. [Google Scholar] [CrossRef] [PubMed]
- Micolonghi, C.; Piane, M.; Germani, A.; Sadeghi, S.; Libi, F.; Savio, C.; Fabiani, M.; Mancini, R.; Ranieri, D.; Pizzuti, A.; et al. A New SMAD4 Splice Site Variant in a Three-Generation Italian Family with Juvenile Polyposis Syndrome. Diagnostics 2022, 12, 2684. [Google Scholar] [CrossRef] [PubMed]
- Samadder, N.J.; Baffy, N.; Giridhar, K. V.; Couch, F.J.; Riegert-Johnson, D. Hereditary Cancer Syndromes-A Primer on Diagnosis and Management, Part 2: Gastrointestinal Cancer Syndromes. Mayo Clin Proc 2019, 94, 1099–1116. [Google Scholar] [CrossRef] [PubMed]
- Vasen, H.F.A.; Tomlinson, I.; Castells, A. Clinical Management of Hereditary Colorectal Cancer Syndromes. Nature Reviews Gastroenterology & Hepatology 2015 12:2 2015, 12, 88–97. [Google Scholar] [CrossRef]
- Latchford, A.R.; Neale, K.; Phillips, R.K.S.; Clark, S.K. Juvenile Polyposis Syndrome: A Study of Genotype, Phenotype, and Long-Term Outcome. Dis Colon Rectum 2012, 55, 1038–1043. [Google Scholar] [CrossRef] [PubMed]
- Katz, L.H.; Gingold-Belfer, R.; Vainer, E.; Hegger, S.; Laish, I.; Derazne, E.; Weintraub, I.; Reznick-Levi, G.; Goldberg, Y.; Levi, Z.; et al. Phenotypic Diversity among Juvenile Polyposis Syndrome Patients from Different Ethnic Background. Hered Cancer Clin Pract 2022, 20, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Brodie, S.G.; Yang, X.; Im, Y.-H.; Parks, W.T.; Chen, L.; Zhou, Y.-X.; Weinstein, M.; Kim, S.-J.; Deng, C.-X. Haploid Loss of the Tumor Suppressor Smad4/Dpc4 Initiates Gastric Polyposis and Cancer in Mice. Oncogene 2000, 19, 1868–1874. [Google Scholar] [CrossRef] [PubMed]
- He, X.C.; Zhang, J.; Tong, W.G.; Tawfik, O.; Ross, J.; Scoville, D.H.; Tian, Q.; Zeng, X.; He, X.; Wiedemann, L.M.; et al. BMP Signaling Inhibits Intestinal Stem Cell Self-Renewal through Suppression of Wnt–β-Catenin Signaling. Nature Genetics 2004 36:10 2004, 36, 1117–1121. [Google Scholar] [CrossRef] [PubMed]
- Marsh Durban, V.; Jansen, M.; Davies, E.J.; Morsink, F.H.; Offerhaus, G.J.A.; Clarke, A.R. Epithelial-Specific Loss of PTEN Results in Colorectal Juvenile Polyp Formation and Invasive Cancer. Am J Pathol 2014, 184, 86–91. [Google Scholar] [CrossRef] [PubMed]
- Kaneda, A.; Feinberg, A.P. Loss of Imprinting of IGF2: A Common Epigenetic Modifier of Intestinal Tumor Risk. Cancer Res 2005, 65, 11236–11240. [Google Scholar] [CrossRef]
- Oshima, H.; Matsunaga, A.; Fujimura, T.; Tsukamoto, T.; Taketo, M.M.; Oshima, M. Carcinogenesis in Mouse Stomach by Simultaneous Activation of the Wnt Signaling and Prostaglandin E2 Pathway. Gastroenterology 2006, 131, 1086–1095. [Google Scholar] [CrossRef] [PubMed]
- Haramis, A.P.G.; Begthel, H.; Van Den Born, M.; Van Es, J.; Jonkheer, S.; Offerhaus, G.J.A.; Clevers, H. De Novo Crypt Formation and Juvenile Polyposis on BMP Inhibition in Mouse Intestine. Science (1979) 2004, 303, 1684–1686. [Google Scholar] [CrossRef] [PubMed]
- Büller, N.V.J.A.; Rosekrans, S.L.; Metcalfe, C.; Heijmans, J.; Van Dop, W.A.; Fessler, E.; Jansen, M.; Ahn, C.; Vermeulen, J.L.M.; Westendorp, B.F.; et al. Stromal Indian Hedgehog Signaling Is Required for Intestinal Adenoma Formation in Mice. Gastroenterology 2015, 148. [Google Scholar] [CrossRef] [PubMed]
- Van Es, J.H.; Clevers, H. Notch and Wnt Inhibitors as Potential New Drugs for Intestinal Neoplastic Disease. Trends Mol Med 2005, 11, 496–502. [Google Scholar] [CrossRef] [PubMed]
- Threadgill, D.W.; Churchill, G.A. Ten Years of the Collaborative Cross. Genetics 2012, 190, 291–294. [Google Scholar] [CrossRef] [PubMed]
- Nashef, A.; Qahaz, N.; El-Naaj, I.A.; Iraqi, F.A. Systems Genetics Analysis of Oral Squamous Cell Carcinoma Susceptibility Using the Mouse Model: Current Position and New Perspective. Mammalian Genome 2021, 32, 323–331. [Google Scholar] [CrossRef] [PubMed]
- Churchill, G.A.; Airey, D.C.; Allayee, H.; Angel, J.M.; Attie, A.D.; Beatty, J.; Beavis, W.D.; Belknap, J.K.; Bennett, B.; Berrettini, W.; et al. The Collaborative Cross, a Community Resource for the Genetic Analysis of Complex Traits. Nat Genet 2004, 36, 1133–1137. [Google Scholar] [CrossRef]
- Atamni, H.J.A.T.; Mott, R.; Soller, M.; Iraqi, F.A. High-Fat-Diet Induced Development of Increased Fasting Glucose Levels and Impaired Response to Intraperitoneal Glucose Challenge in the Collaborative Cross Mouse Genetic Reference Population. BMC Genet 2016, 17. [Google Scholar] [CrossRef] [PubMed]
- Dorman, A.; Binenbaum, I.; Abu-Toamih Atamni, H.J.; Chatziioannou, A.; Tomlinson, I.; Mott, R.; Iraqi, F.A. Genetic Mapping of Novel Modifiers for Apc Min Induced Intestinal Polyps’ Development Using the Genetic Architecture Power of the Collaborative Cross Mice. BMC Genomics 2021, 22. [Google Scholar] [CrossRef]
- Takaku, K.; Miyoshi, H.; Matsunaga, A.; Oshima, M.; Sasaki, N.; Taketo, M.M. Gastric and Duodenal Polyps in Smad4 (Dpc4) Knockout Mice. Cancer Res 1999, 59. [Google Scholar]
- Langeveld, D.; Van Hattem, W.A.; De Leng, W.W.J.; Morsink, F.H.; Ten Kate, F.J.W.; Giardiello, F.M.; Offerhaus, G.J.A.; Brosens, L.A.A. SMAD4 Immunohistochemistry Reflects Genetic Status in Juvenile Polyposis Syndrome. Clinical Cancer Research 2010, 16, 4126–4134. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Yang, X. Smad4-Mediated TGF-β Signaling in Tumorigenesis. Int J Biol Sci 2010, 6, 1. [Google Scholar] [CrossRef] [PubMed]
- Iraqi, F.A.; Churchill, G.; Mott, R. The Collaborative Cross, Developing a Resource for Mammalian Systems Genetics: A Status Report of the Wellcome Trust Cohort. Mammalian Genome 2008, 19. [Google Scholar] [CrossRef]
- Iraqi, F.A.; Mahajne, M.; Salaymah, Y.; Sandovski, H.; Tayem, H.; Vered, K.; Balmer, L.; Hall, M.; Manship, G.; Morahan, G.; et al. The Genome Architecture of the Collaborative Cross Mouse Genetic Reference Population. Genetics 2012, 190. [Google Scholar] [CrossRef]
- Truett, G.E.; Heeger, P.; Mynatt, R.L.; Truett, A.A.; Walker, J.A.; Warman, M.L. Preparation of PCR-Quality Mouse Genomic Dna with Hot Sodium Hydroxide and Tris (HotSHOT). Biotechniques 2000, 29. [Google Scholar] [CrossRef]
- Dorman, A.; Binenbaum, I.; Abu-Toamih Atamni, H.J.; Chatziioannou, A.; Tomlinson, I.; Mott, R.; Iraqi, F.A. Genetic Mapping of Novel Modifiers for Apc Min Induced Intestinal Polyps’ Development Using the Genetic Architecture Power of the Collaborative Cross Mice. BMC Genomics 2021, 22. [Google Scholar] [CrossRef] [PubMed]
- Rudling, R.; Hassan, A.B.; Kitau, J.; Mandir, N.; Goodlad, R.A. A Simple Device to Rapidly Prepare Whole Mounts of Murine Intestine. Cell Prolif 2006, 39. [Google Scholar] [CrossRef] [PubMed]
- Iraqi, F.A.; Athamni, H.; Dorman, A.; Salymah, Y.; Tomlinson, I.; Nashif, A.; Shusterman, A.; Weiss, E.; Houri-Haddad, Y.; Mott, R.; et al. Heritability and Coefficient of Genetic Variation Analyses of Phenotypic Traits Provide Strong Basis for High-Resolution QTL Mapping in the Collaborative Cross Mouse Genetic Reference Population. Mammalian Genome 2014, 25, 109–119. [Google Scholar] [CrossRef]
- Quinlan, J.R. Induction of Decision Trees. Mach Learn 1986, 1, 81–106. [Google Scholar] [CrossRef]
- Krzywinksi, M.; Altman, N. Points of Significance Classification and Regression Trees. Nat Methods 2017, 14. [Google Scholar] [CrossRef]
- Breiman, L. (Impo)Random Forests(Book). Mach Learn 2001. [Google Scholar]
- Zhao, X.; Zou, Q.; Liu, B.; Liu, X. Exploratory Predicting Protein Folding Model with Random Forest and Hybrid Features. Curr Proteomics 2015, 11. [Google Scholar] [CrossRef]
- Liao, Z.; Ju, Y.; Zou, Q. Prediction of G Protein-Coupled Receptors with SVM-Prot Features and Random Forest. Scientifica (Cairo) 2016, 2016. [Google Scholar] [CrossRef] [PubMed]
- Cawley, G.C.; Talbot, N.L.C. On Over-Fitting in Model Selection and Subsequent Selection Bias in Performance Evaluation. Journal of Machine Learning Research 2010, 11. [Google Scholar]


| Sex | Genotype | Trait | df between | df within | n | MS between | MS within | VG | H2 | Trait mean | CVg | Anova Sig |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Female | WT | SB1_C | 12 | 96 | 7.46 | 14.04 | 3.11 | 1.47 | 0.32 | 1.71 | 0.71 | 0.000 |
| Female | WT | SB2_A | 12 | 96 | 7.46 | 1.31 | 0.70 | 0.08 | 0.11 | 0.66 | 0.43 | 0.046 |
| Female | WT | SB2_B | 12 | 96 | 7.46 | 0.49 | 0.41 | 0.01 | 0.03 | 0.52 | 0.20 | 0.298 |
| Female | WT | SB3_A | 12 | 96 | 7.46 | 1.92 | 0.86 | 0.14 | 0.14 | 0.80 | 0.47 | 0.016 |
| Female | WT | SB3_C | 12 | 96 | 7.46 | 26.55 | 7.09 | 2.61 | 0.27 | 1.70 | 0.95 | 0.000 |
| Female | WT | S.I_A_Polyps | 12 | 96 | 7.46 | 7.21 | 3.55 | 0.49 | 0.12 | 2.04 | 0.34 | 0.029 |
| Female | WT | S.I_C_Polyps | 12 | 96 | 7.46 | 68.87 | 18.15 | 6.80 | 0.27 | 4.59 | 0.57 | 0.000 |
| Female | WT | Total_Small_Intestine | 12 | 96 | 7.46 | 96.44 | 26.12 | 9.42 | 0.27 | 8.48 | 0.36 | 0.000 |
| Female | WT | Colon_A_Polyps | 12 | 96 | 7.46 | 0.15 | 0.08 | 0.01 | 0.10 | 0.10 | 0.96 | 0.054 |
| Female | WT | Colon_B_polyps | 12 | 96 | 7.46 | 0.70 | 0.52 | 0.02 | 0.04 | 0.40 | 0.39 | 0.204 |
| Female | WT | Colon_C_Polyps | 12 | 96 | 7.46 | 50.97 | 9.60 | 5.54 | 0.37 | 4.64 | 0.51 | 0.000 |
| Female | WT | Total_Colon | 12 | 96 | 7.46 | 46.08 | 9.92 | 4.85 | 0.33 | 5.15 | 0.43 | 0.000 |
| Female | WT | Total_Intestinal | 12 | 96 | 7.46 | 191.35 | 43.68 | 19.79 | 0.31 | 13.62 | 0.33 | 0.000 |
| Female | KO | SB1_A | 13 | 100 | 7.21 | 0.96 | 0.85 | 0.02 | 0.02 | 0.87 | 0.14 | 0.340 |
| Female | KO | SB1_B | 13 | 100 | 7.21 | 0.77 | 0.74 | 0.00 | 0.01 | 0.92 | 0.07 | 0.417 |
| Female | KO | SB1_C | 13 | 100 | 7.21 | 43.26 | 29.78 | 1.87 | 0.06 | 3.37 | 0.41 | 0.149 |
| Female | KO | SB2_A | 13 | 100 | 7.21 | 0.93 | 0.84 | 0.01 | 0.02 | 1.00 | 0.11 | 0.358 |
| Female | KO | SB2_C | 13 | 100 | 7.21 | 16.22 | 7.22 | 1.25 | 0.15 | 2.39 | 0.47 | 0.013 |
| Female | KO | SB3_A | 13 | 100 | 7.21 | 2.14 | 1.00 | 0.16 | 0.14 | 1.20 | 0.33 | 0.018 |
| Female | KO | SB3_C | 13 | 100 | 7.21 | 118.77 | 36.60 | 11.39 | 0.24 | 3.96 | 0.85 | 0.000 |
| Female | KO | S.I_A_Polyps | 13 | 100 | 7.21 | 5.74 | 3.05 | 0.37 | 0.11 | 3.07 | 0.20 | 0.041 |
| Female | KO | S.I_B_Polyps | 13 | 100 | 7.21 | 3.35 | 3.00 | 0.05 | 0.02 | 2.57 | 0.09 | 0.355 |
| Female | KO | S.I_C_Polyps | 13 | 100 | 7.21 | 348.49 | 88.36 | 36.06 | 0.29 | 9.73 | 0.62 | 0.000 |
| Female | KO | Total_Small_Intestine | 13 | 100 | 7.21 | 394.98 | 92.78 | 41.89 | 0.31 | 15.37 | 0.42 | 0.000 |
| Female | KO | Colon_B_polyps | 13 | 100 | 7.21 | 1.90 | 0.88 | 0.14 | 0.14 | 0.62 | 0.60 | 0.017 |
| Female | KO | Colon_C_Polyps | 13 | 100 | 7.21 | 87.11 | 39.20 | 6.64 | 0.14 | 8.23 | 0.31 | 0.014 |
| Female | KO | Total_Colon | 13 | 100 | 7.21 | 81.18 | 38.90 | 5.86 | 0.13 | 8.96 | 0.27 | 0.021 |
| Female | KO | Total_Intestinal | 13 | 100 | 7.21 | 650.63 | 153.55 | 68.90 | 0.31 | 24.33 | 0.34 | 0.000 |
| Male | WT | SB1_A | 13 | 87 | 6.29 | 0.83 | 0.60 | 0.04 | 0.06 | 0.50 | 0.38 | 0.187 |
| Male | WT | SB1_C | 13 | 87 | 6.29 | 10.50 | 4.77 | 0.91 | 0.16 | 1.64 | 0.58 | 0.016 |
| Male | WT | SB2_A | 13 | 87 | 6.29 | 1.77 | 0.76 | 0.16 | 0.17 | 0.74 | 0.54 | 0.011 |
| Male | WT | SB2_C | 13 | 87 | 6.29 | 35.20 | 15.05 | 3.21 | 0.18 | 2.21 | 0.81 | 0.010 |
| Male | WT | SB3_B | 13 | 87 | 6.29 | 0.90 | 0.81 | 0.01 | 0.02 | 0.76 | 0.16 | 0.359 |
| Male | WT | SB3_C | 13 | 87 | 6.29 | 220.63 | 92.31 | 20.41 | 0.18 | 3.84 | 1.18 | 0.009 |
| Male | WT | S.I_A_Polyps | 13 | 87 | 6.29 | 5.49 | 3.48 | 0.32 | 0.08 | 1.98 | 0.29 | 0.107 |
| Male | WT | S.I_C_Polyps | 13 | 87 | 6.29 | 380.31 | 123.12 | 40.92 | 0.25 | 7.69 | 0.83 | 0.001 |
| Male | WT | Total_Small_Intestine | 13 | 87 | 6.29 | 445.20 | 122.28 | 51.37 | 0.30 | 11.67 | 0.61 | 0.000 |
| Male | WT | Colon_A_Polyps | 13 | 87 | 6.29 | 0.08 | 0.07 | 0.00 | 0.01 | 0.08 | 0.35 | 0.400 |
| Male | WT | Colon_B_polyps | 13 | 87 | 6.29 | 0.94 | 0.46 | 0.08 | 0.14 | 0.39 | 0.72 | 0.025 |
| Male | WT | Colon_C_Polyps | 13 | 87 | 6.29 | 72.84 | 28.17 | 7.11 | 0.20 | 5.38 | 0.50 | 0.004 |
| Male | WT | Total_Colon | 13 | 87 | 6.29 | 68.33 | 30.91 | 5.95 | 0.16 | 5.84 | 0.42 | 0.015 |
| Male | WT | Total_Intestinal | 13 | 87 | 6.29 | 699.96 | 172.11 | 83.98 | 0.33 | 17.51 | 0.52 | 0.000 |
| Male | KO | SB1_A | 13 | 95 | 6.86 | 2.07 | 1.12 | 0.14 | 0.11 | 1.19 | 0.31 | 0.046 |
| Two classes | ||||
| LDA | KNN | SVM | RF | |
| Accuracy | 0.67 | 0.55 | 0.69 | 0.64 |
| Kappa | 0.33 | 0.1 | 0.38 | 0.27 |
| Three classes | ||||
| Accuracy | 0.62 | 0.62 | 0.63 | 0.62 |
| Kappa | 0.1 | 0.07 | 0.0 | 0.1 |
| F1 (Samd4KO X CCxxx) | Sex | Total | ||||
|---|---|---|---|---|---|---|
| F | M | |||||
| CC037 | Genotype | WT | 17 | 15 | 32 | |
| KO | 14 | 19 | 33 | |||
| Total | 31 | 34 | 65 | |||
| CC004 | Genotype | WT | 4 | 5 | 9 | |
| KO | 11 | 8 | 19 | |||
| Total | 15 | 13 | 28 | |||
| CC040 | Genotype | WT | 1 | 2 | 3 | |
| KO | 1 | 2 | 3 | |||
| Total | 2 | 4 | 6 | |||
| CC005 | Genotype | WT | 4 | 4 | 8 | |
| KO | 3 | 3 | 6 | |||
| Total | 7 | 7 | 14 | |||
| CC019 | Genotype | WT | 4 | 3 | 7 | |
| KO | 12 | 4 | 16 | |||
| Total | 16 | 7 | 23 | |||
| CC006 | Genotype | WT | 7 | 5 | 12 | |
| KO | 10 | 7 | 17 | |||
| Total | 17 | 12 | 29 | |||
| CC084 | Genotype | WT | 12 | 12 | 24 | |
| KO | 13 | 11 | 24 | |||
| Total | 25 | 23 | 48 | |||
| CC059 | Genotype | WT | 4 | 9 | 13 | |
| KO | 4 | 6 | 10 | |||
| Total | 8 | 15 | 23 | |||
| CC041 | Genotype | WT | 2 | 0 | 2 | |
| KO | 2 | 0 | 2 | |||
| Total | 4 | 4 | ||||
| CC010 | Genotype | WT | 11 | 12 | 23 | |
| KO | 14 | 10 | 24 | |||
| Total | 25 | 22 | 47 | |||
| CC018 | Genotype | WT | 16 | 8 | 24 | |
| KO | 14 | 15 | 29 | |||
| Total | 30 | 23 | 53 | |||
| CC012 | Genotype | WT | 7 | 4 | 11 | |
| KO | 5 | 6 | 11 | |||
| Total | 12 | 10 | 22 | |||
| CC035 | Genotype | WT | 11 | 8 | 19 | |
| KO | 5 | 13 | 18 | |||
| Total | 16 | 21 | 37 | |||
| CC025 | Genotype | WT | 9 | 11 | 20 | |
| KO | 8 | 11 | 19 | |||
| Total | 17 | 22 | 39 | |||
| CC005* | Genotype | WT | 20 | 12 | 32 | |
| KO | 15 | 14 | 29 | |||
| Total | 35 | 26 | 61 | |||
| Total | Genotype | WT | 129 | 110 | 239 | |
| KO | 131 | 129 | 260 | |||
| Total | 260 | 239 | 499 | |||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




