Submitted:
31 March 2024
Posted:
01 April 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Clinical Samples
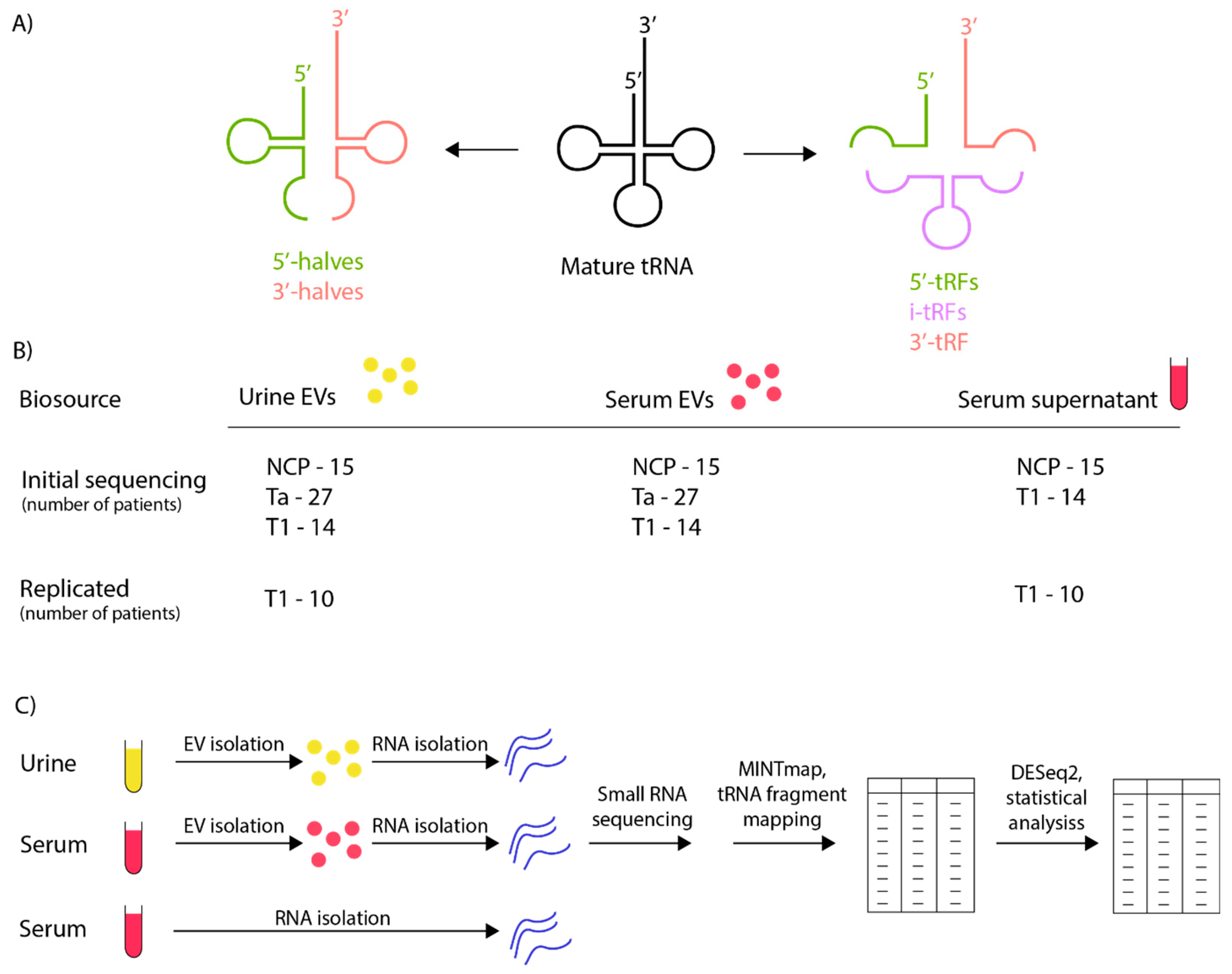
2.2. Workflow and Study Design
2.3. RNA isolation
2.4. Small RNA Sequencing
2.5. Statistical Analysis
2.6. BLAST Search
3. Results
3.1. Sample Characterisation
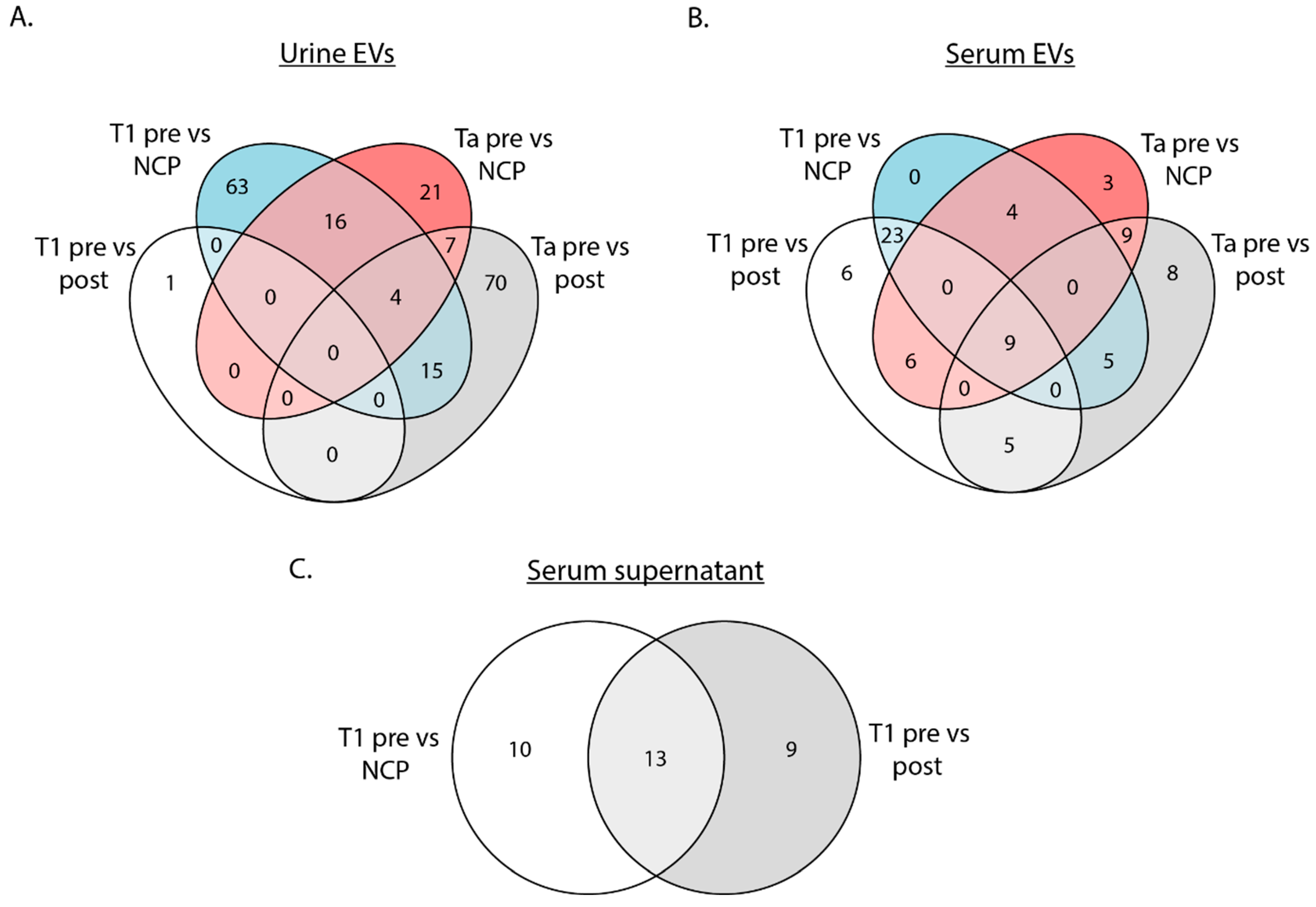
3.2. Differential Expression of tRFs in Urine EVs, Serum EVs and Serum Supernatant in the Initial Sequencing Run
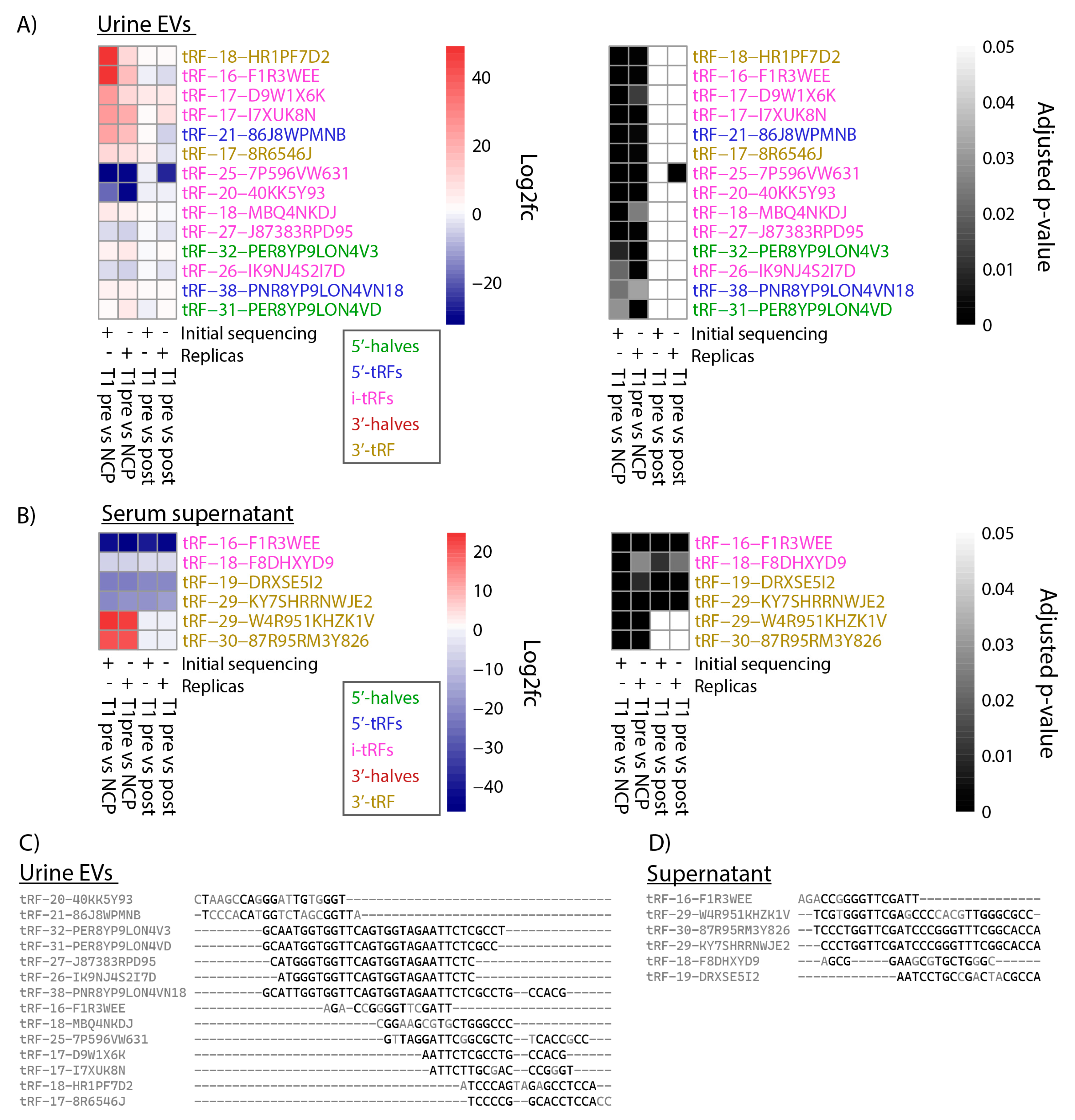
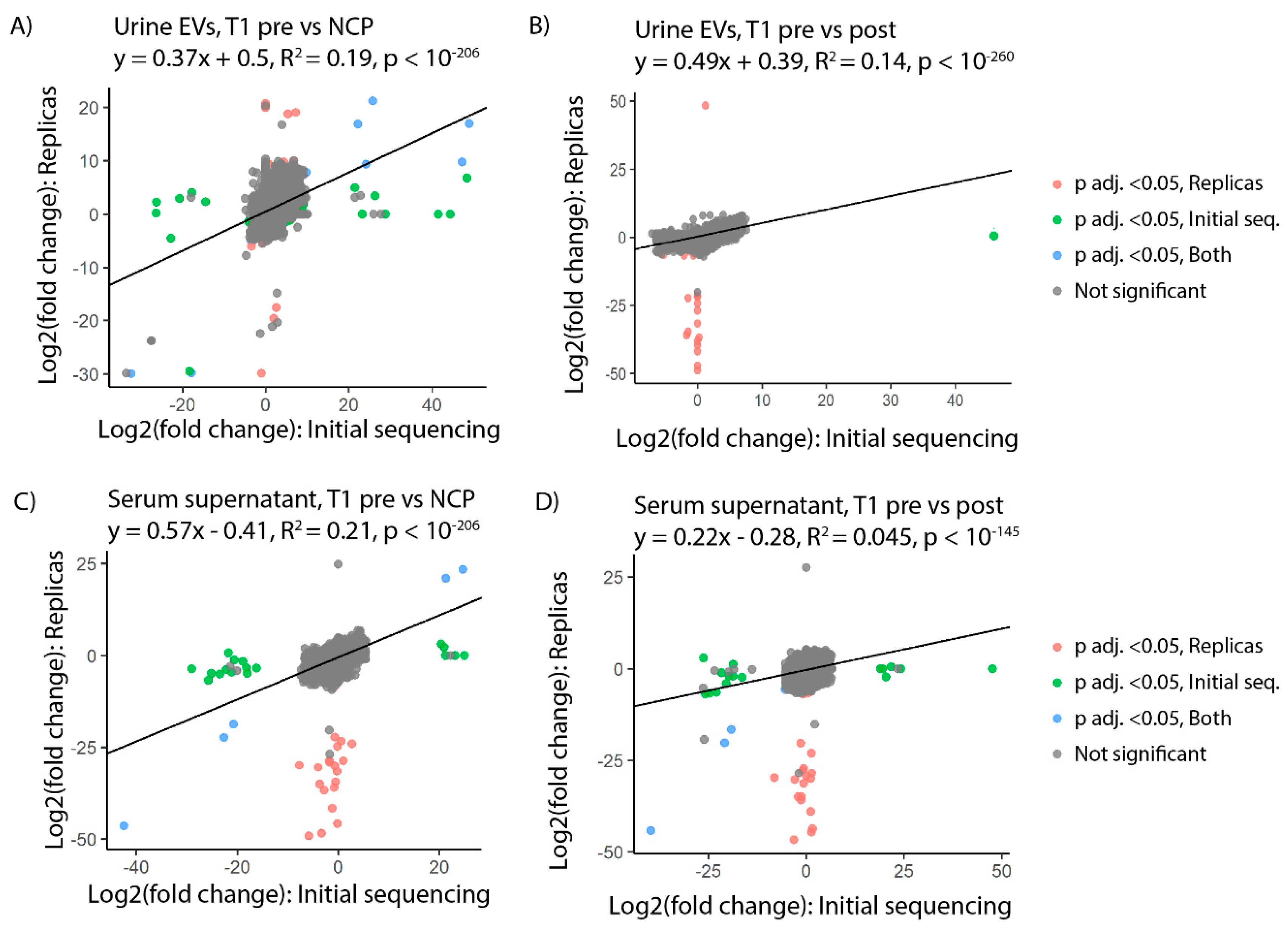
3.3. tRFs Are Confirmed as Differentially Expressed among Patients with Stage T1 Disease by Replica Sequencing
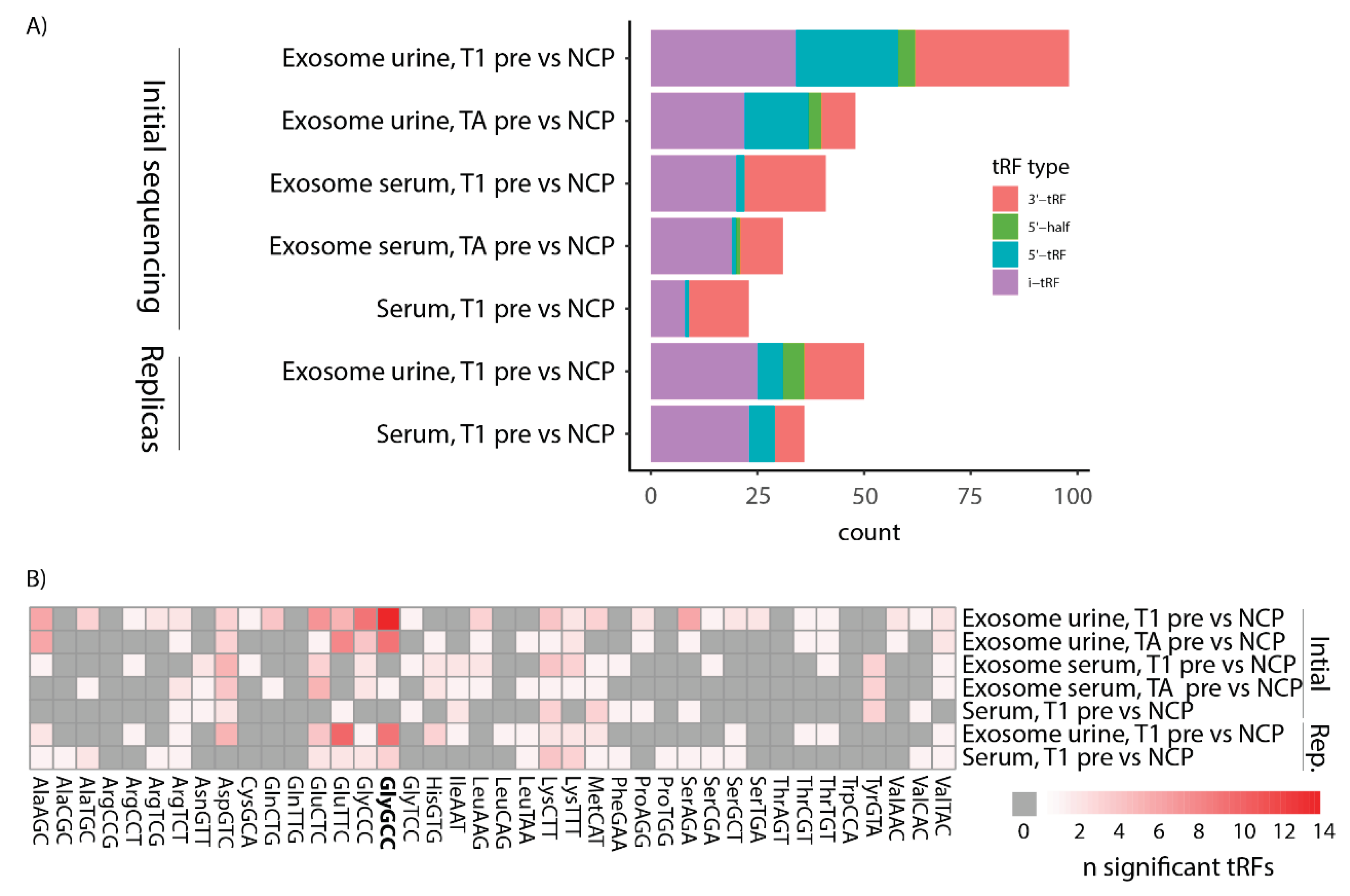
3.4. tRFs Differentially Expressed between Bladder Cancer and Non-Cancer Patients Are of Diverse Types and Originate from a Multitude of tRNAs
3.5. Assessment of Potential Role for tRFs in Biological Processes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Trail M, Cullen J, Fulton E, Clayton F, McGregor E, McWilliam F, et al. Evaluating the Safety of Performing Flexible Cystoscopy When Urinalysis Suggests Presence of “Infection”: Results of a Prospective Clinical Study in 2350 patients. Eur Urol Open Sci. 2021, 31, 28–36. [Google Scholar] [CrossRef] [PubMed]
- Toden S, Goel A. Non-coding RNAs as liquid biopsy biomarkers in cancer. Br J Cancer. 2022, 126, 351–360. [Google Scholar] [CrossRef] [PubMed]
- Hoagland MB, Stephenson ML, Scott JF, Hecht LI, Zamecnik PC. A soluble ribonucleic acid intermediate in protein synthesis. J Biol Chem. 1958, 231, 241–257. [Google Scholar] [CrossRef]
- Speer J, Gehrke CW, Kuo KC, Waalkes TP, Borek E. tRNA breakdown products as markers for cancer. Cancer. 1979, 44, 2120–2123. [Google Scholar] [CrossRef]
- Waalkes TP, Gehrke CW, Zumwalt RW, Chang SY, Lakings DB, Tormey DC, et al. The urinary excretion of nucleosides of ribonucleic acid by patients with advanced cancer. Cancer. 1975, 36, 390–398. [Google Scholar] [CrossRef]
- Lee YS, Shibata Y, Malhotra A, Dutta A. A novel class of small RNAs: tRNA-derived RNA fragments (tRFs). Genes Dev. 2009, 23, 2639–2649. [Google Scholar] [CrossRef]
- Weng Q, Wang Y, Xie Y, Yu X, Zhang S, Ge J, et al. Extracellular vesicles-associated tRNA-derived fragments (tRFs): biogenesis, biological functions, and their role as potential biomarkers in human diseases. J Mol Med (Berl). 2022, 100, 679–695. [Google Scholar] [CrossRef] [PubMed]
- Wilson B, Dutta A. Function and Therapeutic Implications of tRNA Derived Small RNAs. Front Mol Biosci. 2022, 9, 888424. [Google Scholar]
- Kumar P, Kuscu C, Dutta A. Biogenesis and Function of Transfer RNA-Related Fragments (tRFs). Trends Biochem Sci. 2016, 41, 679–689. [Google Scholar] [CrossRef]
- Zhang M, Li F, Wang J, He W, Li Y, Li H, et al. tRNA-derived fragment tRF-03357 promotes cell proliferation, migration and invasion in high-grade serous ovarian cancer. Onco Targets Ther. 2019, 12, 6371–6383. [Google Scholar] [CrossRef]
- Mao M, Chen W, Huang X, Ye D. Role of tRNA-derived small RNAs(tsRNAs) in the diagnosis and treatment of malignant tumours. Cell Commun Signal. 2023, 21, 178. [Google Scholar] [CrossRef] [PubMed]
- Maute RL, Schneider C, Sumazin P, Holmes A, Califano A, Basso K, Dalla-Favera R. tRNA-derived microRNA modulates proliferation and the DNA damage response and is down-regulated in B cell lymphoma. Proc Natl Acad Sci U S A. 2013, 110, 1404–1409. [Google Scholar] [CrossRef] [PubMed]
- Fu M, Gu J, Wang M, Zhang J, Chen Y, Jiang P, et al. Emerging roles of tRNA-derived fragments in cancer. Mol Cancer. 2023, 22, 30. [Google Scholar] [CrossRef] [PubMed]
- Endres, L. tRNA modification and cancer: potential for therapeutic prevention and intervention. Futire Medicinal Cemistry. 2019, 8. [Google Scholar] [CrossRef]
- Keam SP, Hutvagner G. tRNA-Derived Fragments (tRFs): Emerging New Roles for an Ancient RNA in the Regulation of Gene Expression. Life (Basel). 2015, 5, 1638–1651. [Google Scholar]
- Xie Y, Yao L, Yu X, Ruan Y, Li Z, Guo J. Action mechanisms and research methods of tRNA-derived small RNAs. Signal Transduct Target Ther. 2020, 5, 109. [Google Scholar] [CrossRef] [PubMed]
- Su Z, Monshaugen I, Klungland A, Ougland R, Dutta A. Characterization of novel small non-coding RNAs and their modifications in bladder cancer using an updated small RNA-seq workflow. Front Mol Biosci. 2022, 9, 887686. [Google Scholar]
- van Niel G, D’Angelo G, Raposo G. Shedding light on the cell biology of extracellular vesicles. Nat Rev Mol Cell Biol. 2018, 19, 213–228. [Google Scholar] [CrossRef] [PubMed]
- Kowal J, Tkach M, Thery C. Biogenesis and secretion of exosomes. Curr Opin Cell Biol. 2014, 29, 116–125. [Google Scholar] [CrossRef]
- Cocucci E, Racchetti G, Meldolesi J. Shedding microvesicles: artefacts no more. Trends Cell Biol. 2009, 19, 43–51. [Google Scholar] [CrossRef]
- Maas SLN, Breakefield XO, Weaver AM. Extracellular Vesicles: Unique Intercellular Delivery Vehicles. Trends Cell Biol. 2017, 27, 172–188. [Google Scholar] [CrossRef]
- Pitt JM, Kroemer G, Zitvogel L. Extracellular vesicles: masters of intercellular communication and potential clinical interventions. J Clin Invest. 2016, 126, 1139–1143. [Google Scholar] [CrossRef] [PubMed]
- Bebelman MP, Janssen E, Pegtel DM, Crudden C. The forces driving cancer extracellular vesicle secretion. Neoplasia. 2021, 23, 149–157. [Google Scholar] [CrossRef] [PubMed]
- Lane RE, Korbie D, Hill MM, Trau M. Extracellular vesicles as circulating cancer biomarkers: opportunities and challenges. Clin Transl Med. 2018, 7, 14. [Google Scholar] [CrossRef] [PubMed]
- Zhu L, Li J, Gong Y, Wu Q, Tan S, Sun D, et al. Exosomal tRNA-derived small RNA as a promising biomarker for cancer diagnosis. Mol Cancer. 2019, 18, 74. [Google Scholar] [CrossRef]
- Lin, C. tRFs as Potential Exosome tRNA-Derived Fragment Biomarkers for Gastric Carcinoma. Clin Lab. 2020, 1. [Google Scholar] [CrossRef]
- Guzman N, Agarwal K, Asthagiri D, Yu L, Saji M, Ringel MD, Paulaitis ME. Breast Cancer-Specific miR Signature Unique to Extracellular Vesicles Includes “microRNA-like” tRNA Fragments. Mol Cancer Res. 2015, 13, 891–901. [Google Scholar] [CrossRef]
- Koi Y, Tsutani Y, Nishiyama Y, Ueda D, Ibuki Y, Sasada S, et al. Predicting the presence of breast cancer using circulating small RNAs, including those in the extracellular vesicles. Cancer Sci. 2020, 111, 2104–2115. [Google Scholar] [CrossRef]
- Armstrong DA, Green BB, Seigne JD, Schned AR, Marsit CJ. MicroRNA molecular profiling from matched tumor and bio-fluids in bladder cancer. Mol Cancer. 2015, 14, 194. [Google Scholar] [CrossRef]
- Stromme O, Heck KA, Brede G, Lindholm HT, Otterlei M, Arum CJ. Differentially Expressed Extracellular Vesicle-Contained microRNAs before and after Transurethral Resection of Bladder Tumors. Curr Issues Mol Biol. 2021, 43, 286–300. [Google Scholar] [CrossRef]
- Loher P, Telonis AG, Rigoutsos I. MINTmap: fast and exhaustive profiling of nuclear and mitochondrial tRNA fragments from short RNA-seq data. Sci Rep. 2017, 7, 41184. [Google Scholar] [CrossRef] [PubMed]
- Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550.
- Srinivasan S, Yeri A, Cheah PS, Chung A, Danielson K, De Hoff P, et al. Small RNA Sequencing across Diverse Biofluids Identifies Optimal Methods for exRNA Isolation. Cell. 2019, 177, 446–462. [Google Scholar] [CrossRef] [PubMed]
- Chan C, Pham P, Dedon PC, Begley TJ. Lifestyle modifications: coordinating the tRNA epitranscriptome with codon bias to adapt translation during stress responses. Genome Biol. 2018, 19, 228. [Google Scholar]
- Hanahan, D. Hallmarks of Cancer: New Dimensions. Cancer Discov. 2022, 12, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Djingo, D. Mammalian-wide interspersed repeat (MIR)-derived enhancers and the regulation of human gene expression. Mobile DNA. 2014, 5. [Google Scholar]
- Maraver A, Fernandez-Marcos PJ, Cash TP, Mendez-Pertuz M, Duenas M, Maietta P, et al. NOTCH pathway inactivation promotes bladder cancer progression. J Clin Invest. 2015, 125, 824–830. [Google Scholar] [CrossRef]
- Ozata DM, Gainetdinov I, Zoch A, O’Carroll D, Zamore PD. PIWI-interacting RNAs: small RNAs with big functions. Nat Rev Genet. 2019, 20, 89–108. [Google Scholar] [CrossRef]

| Phrase | n | tRF names |
|---|---|---|
| FCGR2A | 6 | tRF-38-PNR8YP9LON4VN18, tRF-32-PER8YP9LON4V3, tRF-31-PER8YP9LON4VD, tRF-27-J87383RPD95, tRF-26-IK9NJ4S2I7D, tRF-17-D9W1X6K |
| GPD2 | 6 | tRF-38-PNR8YP9LON4VN18, tRF-32-PER8YP9LON4V3, tRF-31-PER8YP9LON4VD, tRF-27-J87383RPD95, tRF-26-IK9NJ4S2I7D, tRF-17-D9W1X6K |
| H3K4me1 hESC enhancer | 11 | tRF-38-PNR8YP9LON4VN18, tRF-32-PER8YP9LON4V3, tRF-31-PER8YP9LON4VD, tRF-27-J87383RPD95, tRF-26-IK9NJ4S2I7D, tRF-25-7P596VW631, tRF-20-40KK5Y93, tRF-17-I7XUK8N, tRF-17-D9W1X6K, tRF-17-8R6546J, tRF-16-F1R3WEE |
| HES7 | 6 | tRF-38-PNR8YP9LON4VN18, tRF-32-PER8YP9LON4V3, tRF-31-PER8YP9LON4VD, tRF-27-J87383RPD95, tRF-26-IK9NJ4S2I7D, tRF-17-D9W1X6K |
| HSPA6 | 6 | tRF-38-PNR8YP9LON4VN18, tRF-32-PER8YP9LON4V3, tRF-31-PER8YP9LON4VD, tRF-27-J87383RPD95, tRF-26-IK9NJ4S2I7D, tRF-17-D9W1X6K |
| lncRNA | 7 | tRF-27-J87383RPD95, tRF-26-IK9NJ4S2I7D, tRF-25-7P596VW631, tRF-21-86J8WPMNB, tRF-18-MBQ4NKDJ, tRF-17-D9W1X6K, tRF-16-F1R3WEE |
| LTA4 | 1 | tRF-16-F1R3WEE |
| piRNA | 3 | tRF-27-J87383RPD95, tRF-26-IK9NJ4S2I7D, tRF-18-MBQ4NKDJ |
| regulatory element | 7 | tRF-38-PNR8YP9LON4VN18, tRF-32-PER8YP9LON4V3, tRF-31-PER8YP9LON4VD, tRF-27-J87383RPD95, tRF-26-IK9NJ4S2I7D, tRF-18-MBQ4NKDJ, tRF-17-D9W1X6K |
| tRNA | 12 | tRF-38-PNR8YP9LON4VN18, tRF-32-PER8YP9LON4V3, tRF-31-PER8YP9LON4VD, tRF-27-J87383RPD95, tRF-26-IK9NJ4S2I7D, tRF-25-7P596VW631, tRF-21-86J8WPMNB, tRF-20-40KK5Y93, tRF-18-MBQ4NKDJ, tRF-17-I7XUK8N, tRF-17-D9W1X6K, tRF-16-F1R3WEE |
| VAC14 | 6 | tRF-38-PNR8YP9LON4VN18, tRF-32-PER8YP9LON4V3, tRF-31-PER8YP9LON4VD, tRF-27-J87383RPD95, tRF-26-IK9NJ4S2I7D, tRF-17-D9W1X6K |
| Phrase | n | tRF names |
|---|---|---|
| H3K4me1 hESC enhancer | 2 | tRF-29-W4R951KHZK1V, tRF-16-F1R3WEE |
| lncRNA | 3 | tRF-29-W4R951KHZK1V, tRF-18-F8DHXYD9, tRF-16-F1R3WEE |
| LTA4 | 1 | tRF-16-F1R3WEE |
| piRNA | 3 | tRF-30-87R95RM3Y826, tRF-29-KY7SHRRNWJE2, tRF-18-F8DHXYD9 |
| regulatory element | 1 | tRF-18-F8DHXYD9 |
| tRNA | 5 | tRF-30-87R95RM3Y826, tRF-29-KY7SHRRNWJE2, tRF-19-DRXSE5I2, tRF-18-F8DHXYD9, tRF-16-F1R3WEE |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




