Submitted:
28 March 2024
Posted:
29 March 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
- We employed the U-Net model as the backbone for our performance analysis, incorporating various transfer learning models for comparison. The top five transfer learning models we explored are ResECA-U-Net, ResNet34, Efficient-NetB0, EfficientNetB1, and EfficientNetB2—all exhibiting commendable performance and leveraging the ECA-Net architecture.
- Our research introduced a U-Net model in computer vision, specifically designed to enhance local features for segmentation tasks. The ResECA-U-Net model proposed in this work was applied to the UW-Madison GI tract image segmentation dataset.
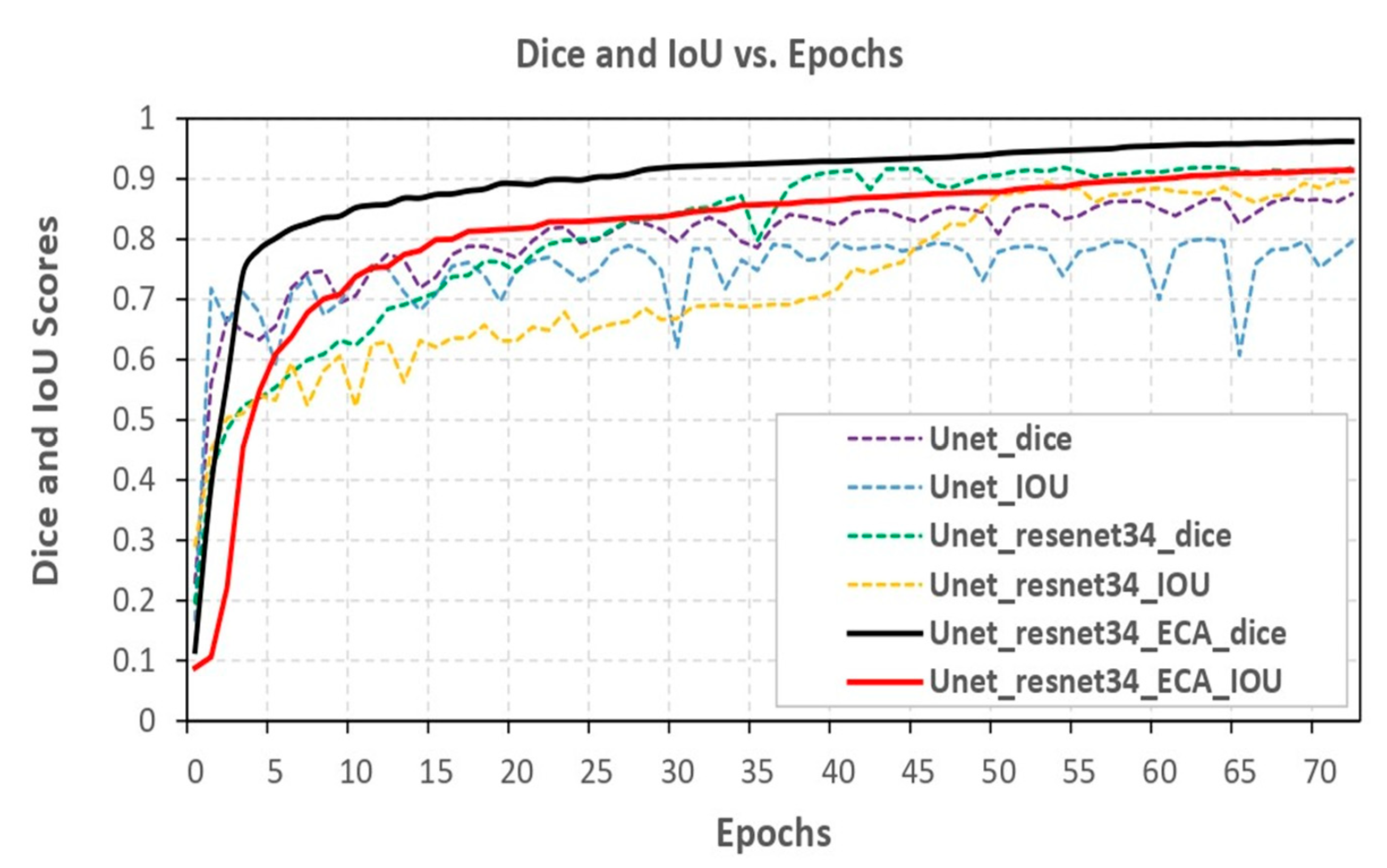
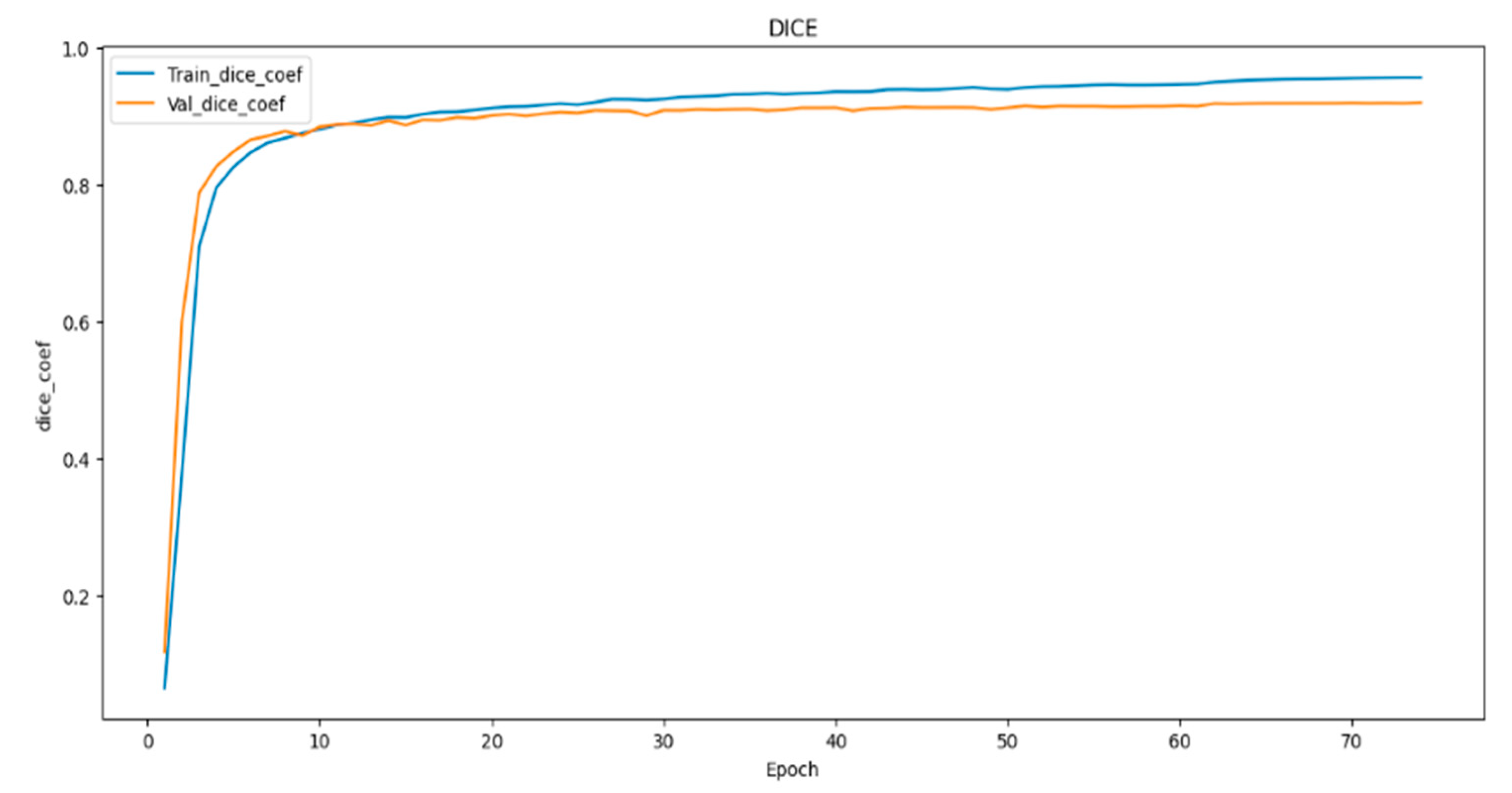
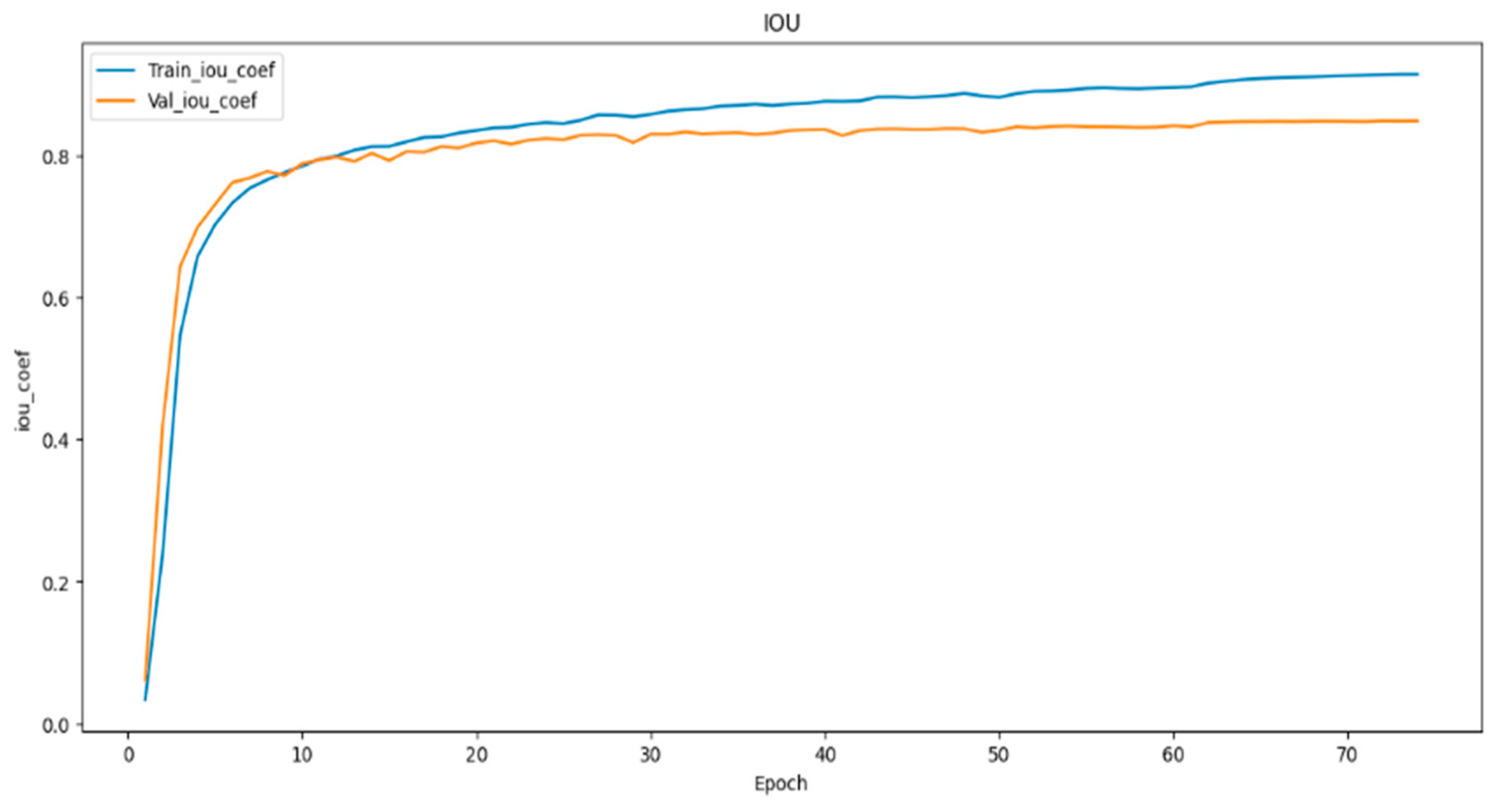
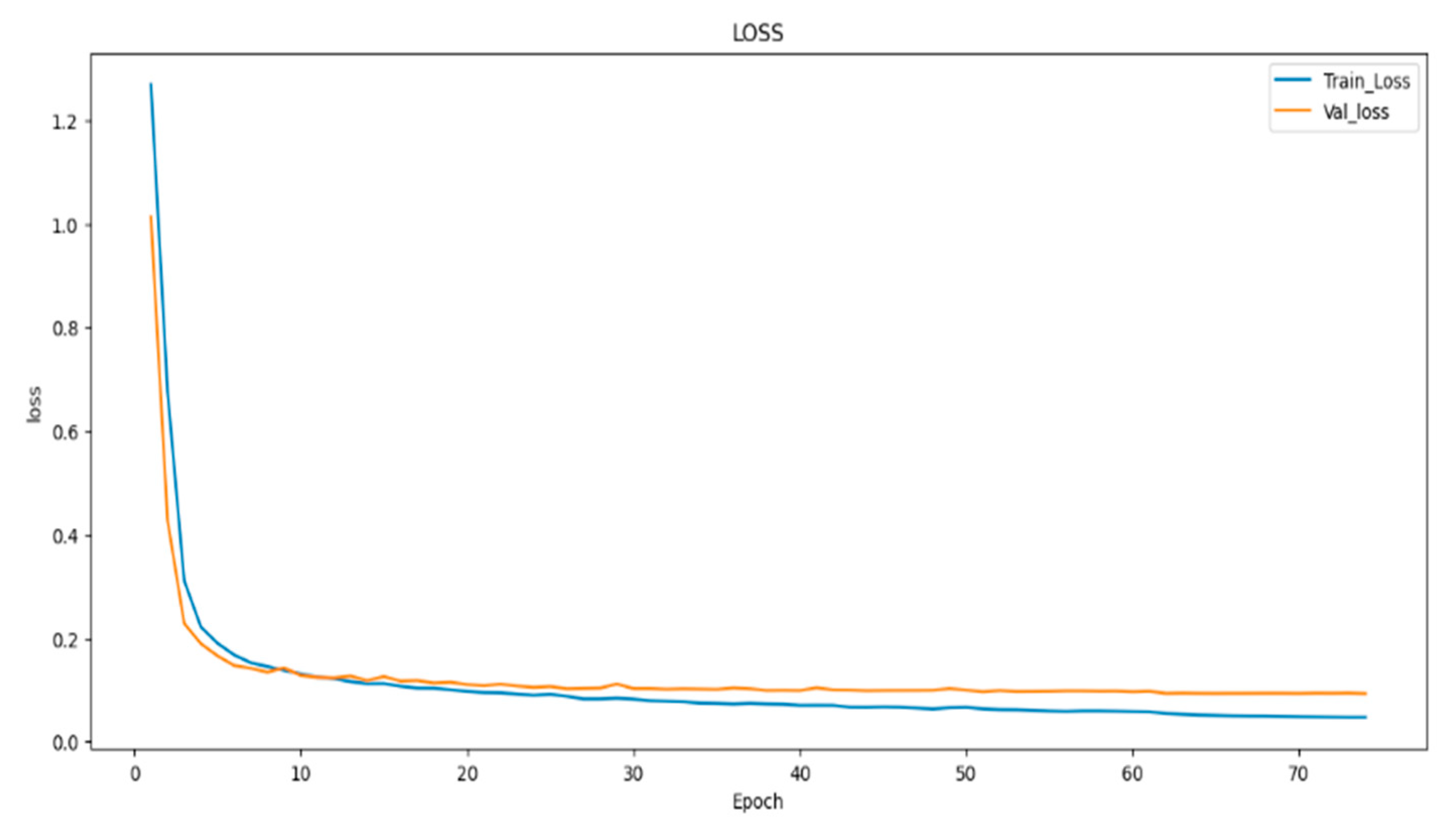
- To evaluate the performance of the models, we employed metrics such as dice coefficient, Intersection over Union (IoU), and model loss. These metrics offered a comprehensive assessment of the proposed models’ effectiveness in image segmentation.
2. Related Work
3. Methodology
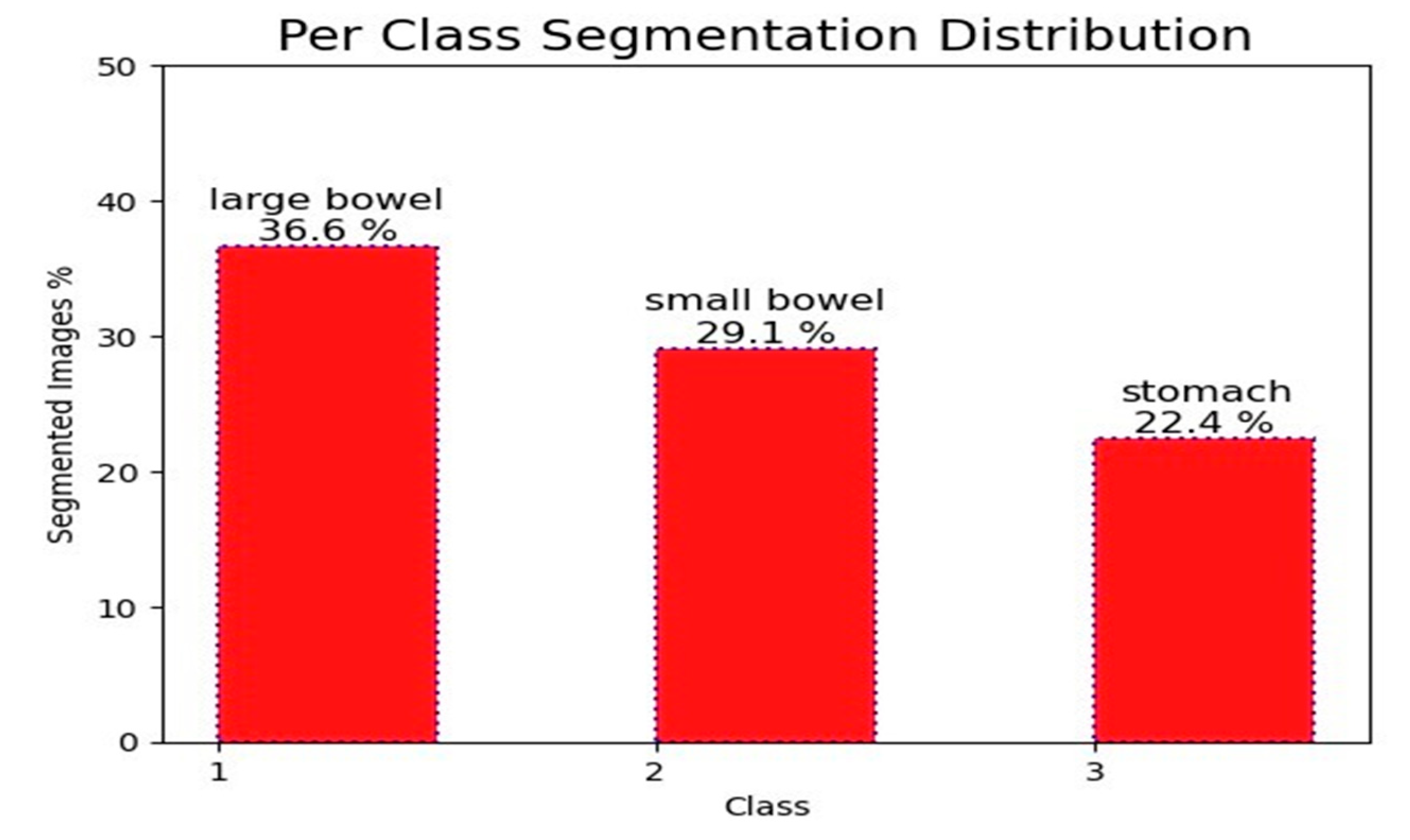
3.1. Data Set
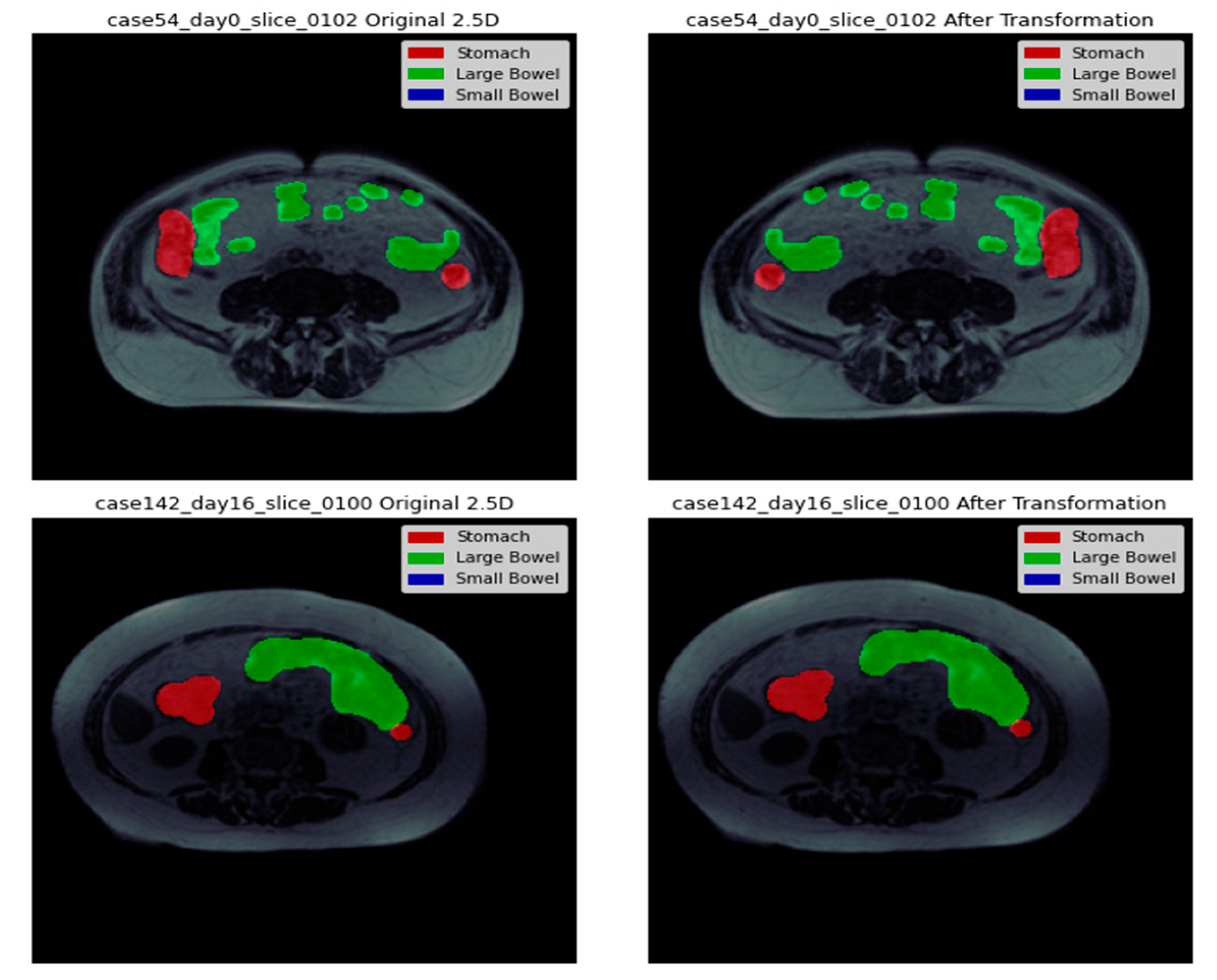
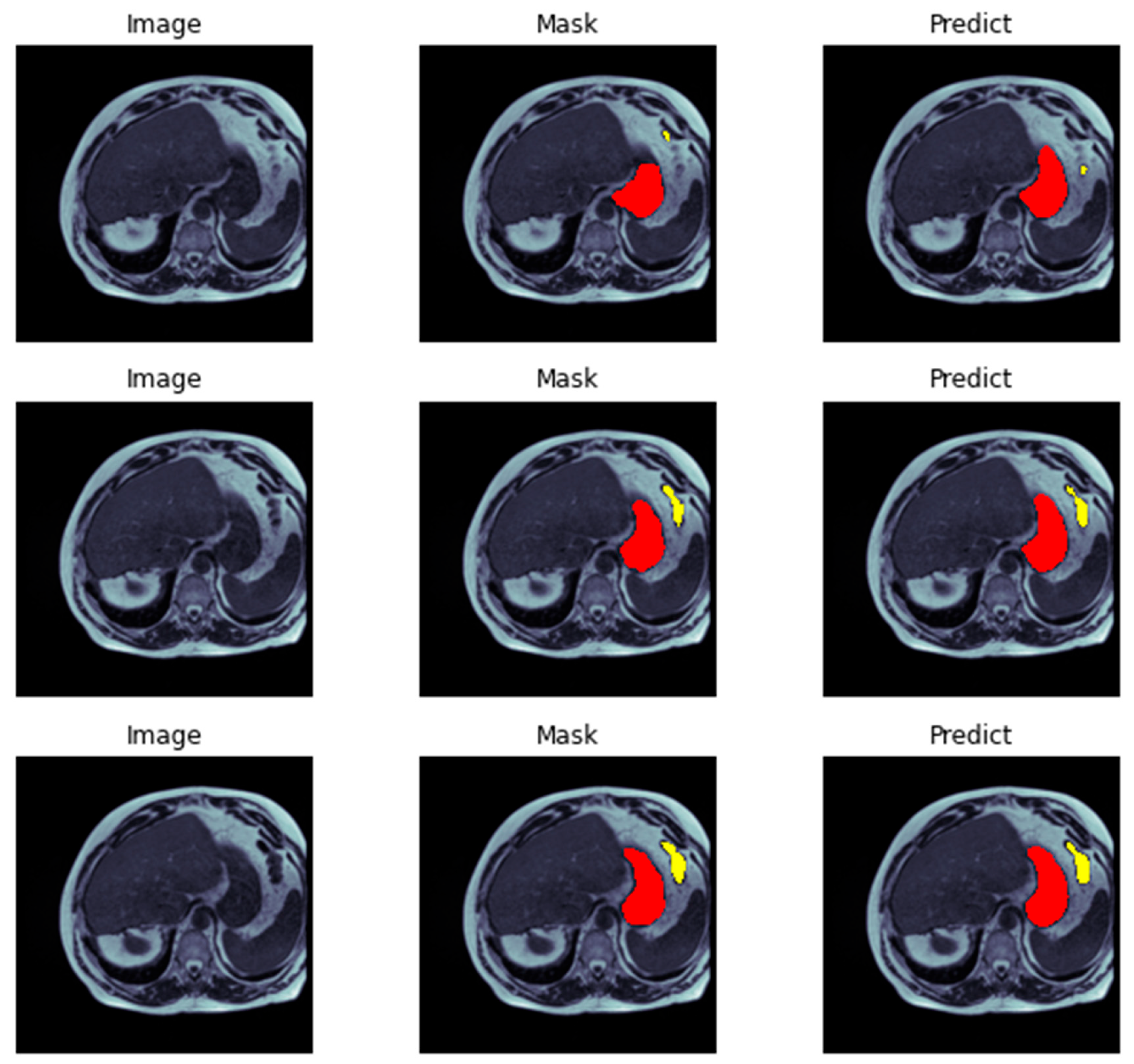
3.4. Data Visualization
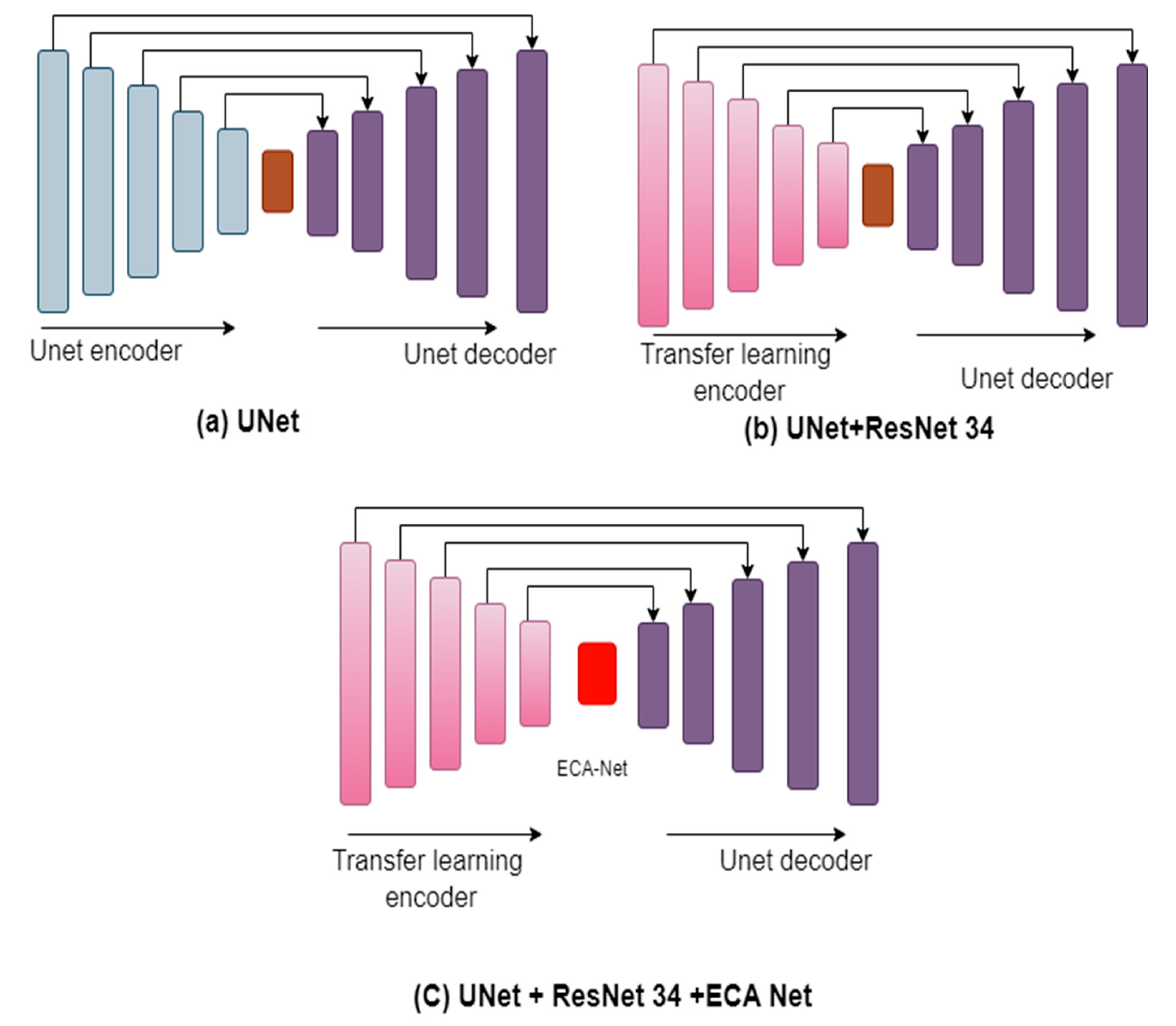
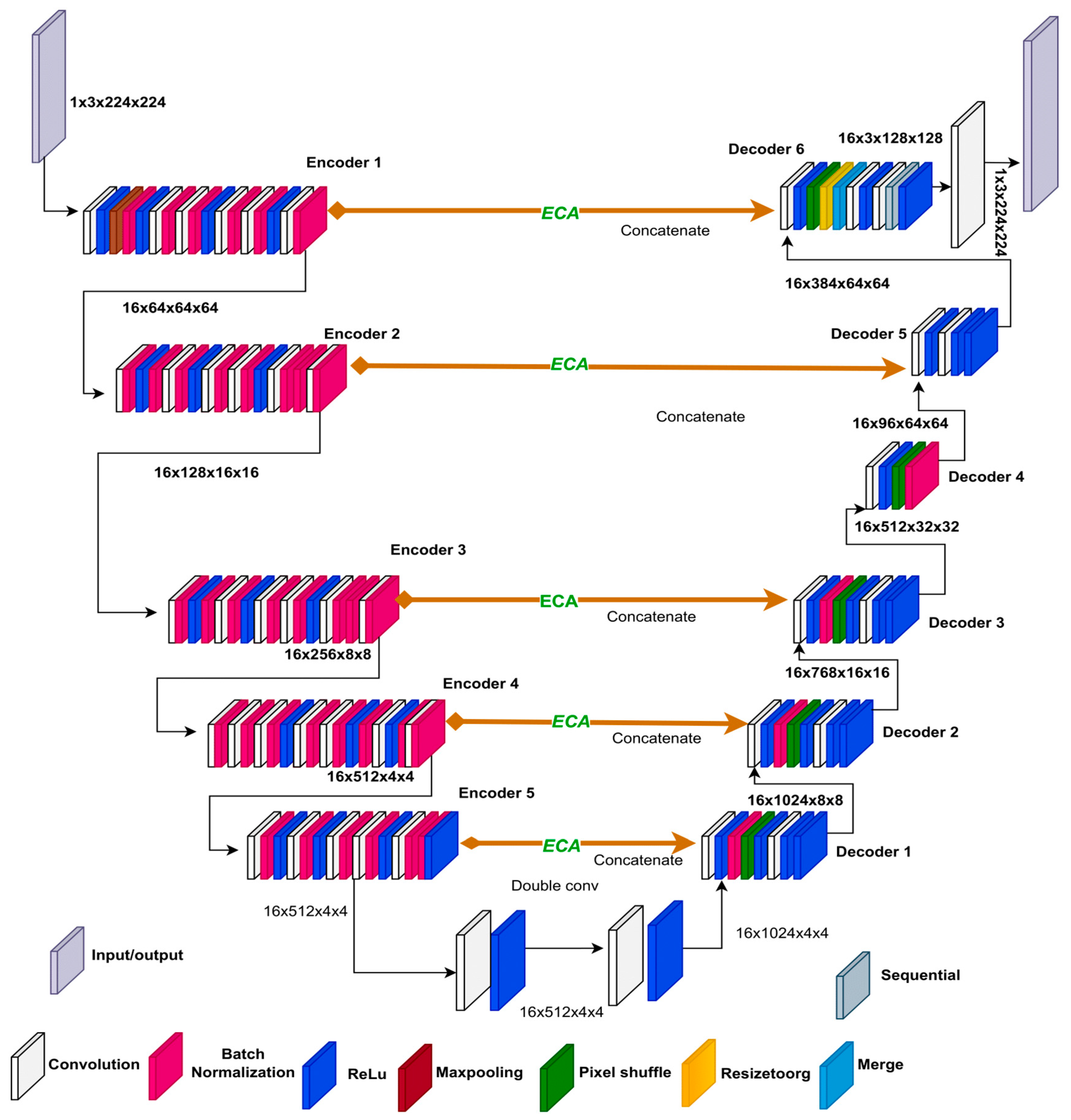
3.5. ResECA-U-Net
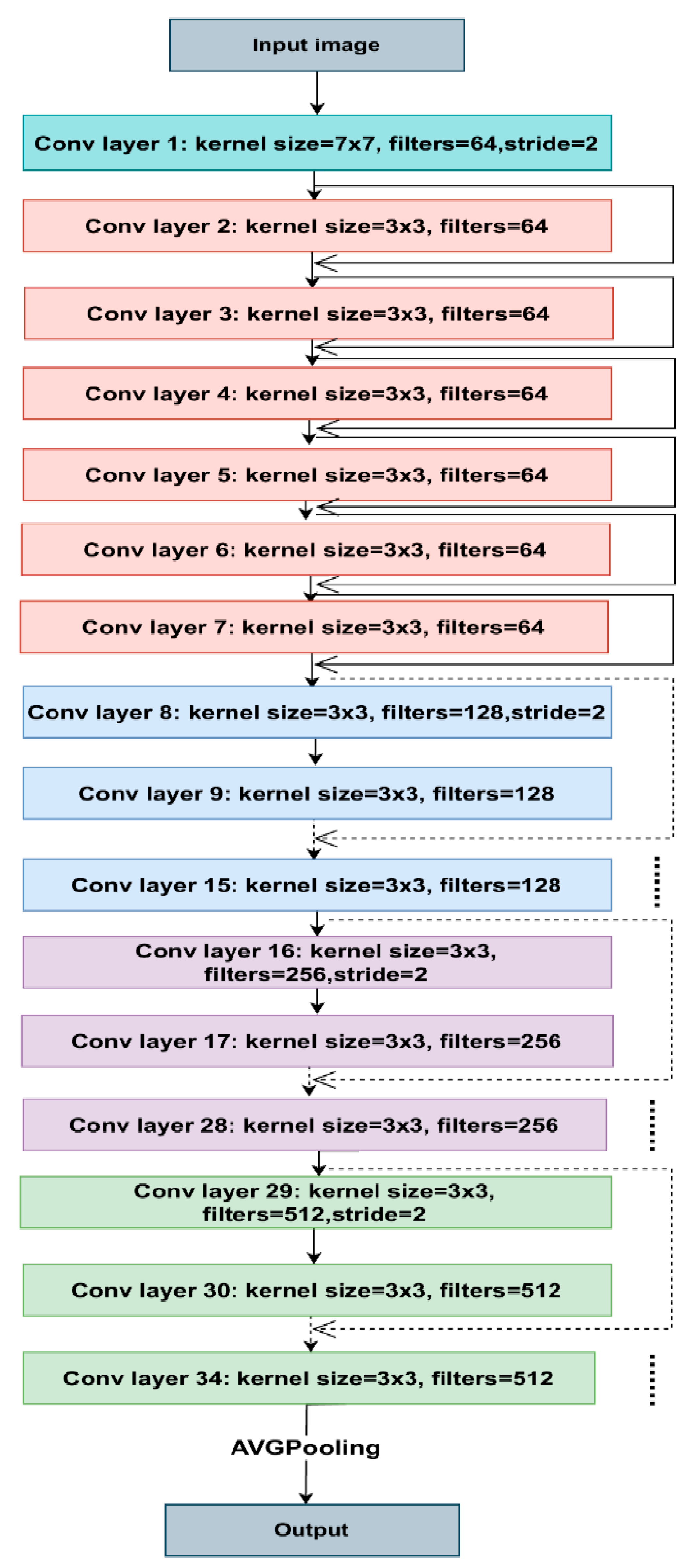
3.6. Residual Module Basic Block
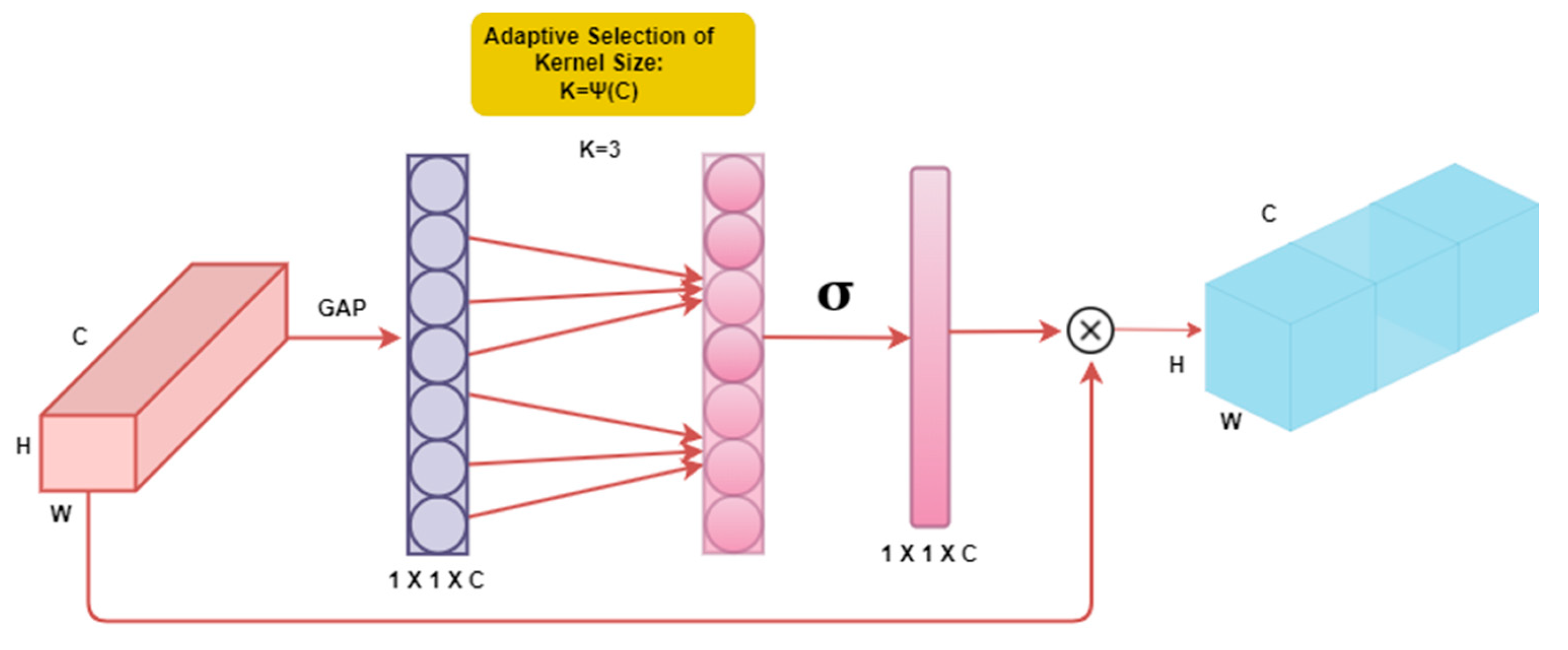
3.7. Efficient Channel Attention (ECA-Net)
3.8. Transfer Learning
3.9. Loss Variations
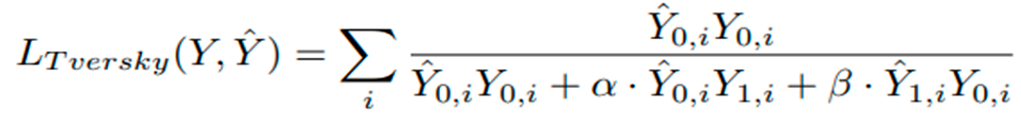
3.10. Diversity-Promoting Ensemble
1.11. Diversity-Promoting Ensemble Creation Algorithm
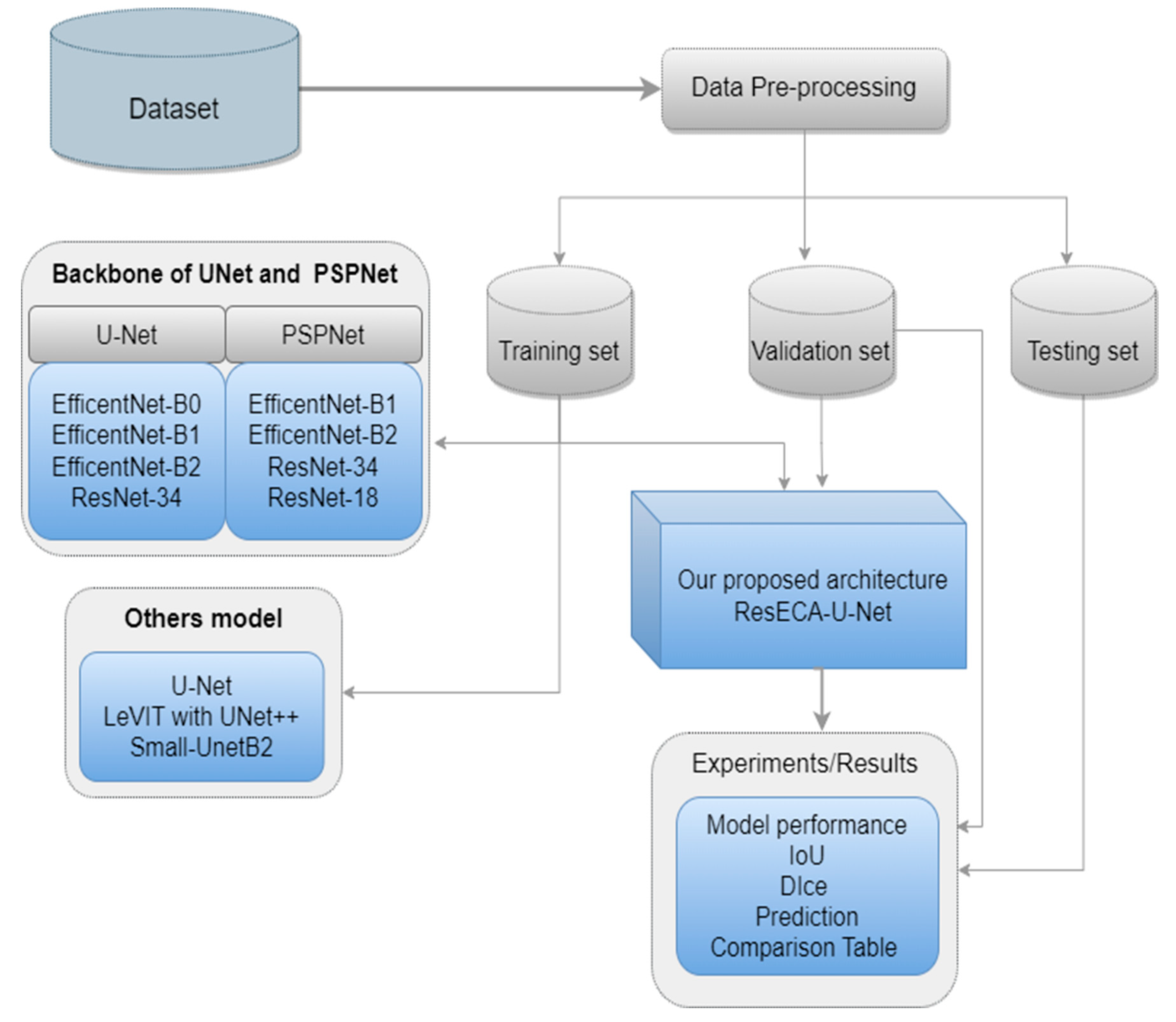
4. Experiments/Results
4.1. Result
4.2. Discussion
5. Conclusion
6. Future Work
Funding
Data Availability Statement
Conflicts of Interest
References
- Rawla, P.; Barsouk, A. Epidemiology of gastric cancer: global trends, risk factors and prevention. Gastroenterology Review/Przeglad Gastroenterologiczny 2019, 14(1), 26–38. [Google Scholar] [CrossRef] [PubMed]
- Alam MJ, Zaman S, Shill PC, Kar S, Hakim MA. Automated Gastrointestinal Tract Image Segmentation Of Cancer Patient Using LeVit-UNet To Automate Radiotherapy. In: 2023 International Conference on Electrical, Computer and Communication Engineering (ECCE). IEEE; 2023. p. 1–5.
- Jaffray DA, Gospodarowicz MK. Radiation therapy for cancer. Cancer: disease control priorities. 2015;3:239–248.
- Heavey PM, Rowland IR. Gastrointestinal cancer. Best Practice & Research Clinical Gastroenterology. 2004;18(2):323–336.
- Lagendijk JJ, Raaymakers BW, Van Vulpen M. The magnetic resonance imaging–linac system. In: Seminars in radiation oncology. vol. 24. Elsevier; 2014. p. 207–209.
- Chou A, Li W, Roman E. GI Tract Image Segmentation with U-Net and Mask R-CNN.
- Sharma M. Automated GI tract segmentation using deep learning. arXiv preprint arXiv:220611048. 2022.
- Chia B, Gu H, Lui N. Gastro-Intestinal Tract Segmentation Using Multi-Task Learning;
- Khan MA, Khan MA, Ahmed F, Mittal M, Goyal LM, Hemanth DJ, et al. Gastrointestinal diseases segmentation and classification based on duo-deep architectures. Pattern Recognition Letters. 2020;131:193–204.
- Guggari S, Srivastava BC, Kumar V, Harshita H, Farande V, Kulkarni U, et al. RU-Net: A Novel Approach for Gastro-Intestinal Tract Image Segmentation Using Convolutional Neural Network. In: International Conference on Applied Machine Learning and Data Analytics. Springer; 2022. p. 131–141.
- Wang S, Cong Y, Zhu H, Chen X, Qu L, Fan H, et al. Multi-scale context-guided deep network for automated lesion segmentation with endoscopy images of gastrointestinal tract. IEEE Journal of Biomedical and Health Informatics. 2020;25(2):514–525.
- Sharma N, Gupta S, Koundal D, Alyami S, Alshahrani H, Asiri Y, et al. U-Net model with transfer learning model as a backbone for segmentation of gastrointestinal tract. Bioengineering. 2023;10(1):119.
- Holzmann GJ. State compression in SPIN: Recursive indexing and compression training runs. In: Proceedings of third international Spin workshop; 1997.
- Olaf Ronneberger TB Philipp Fischer. U-Net: Convolutional Networks for Biomedical Image Segmentation; 2015. https://arxiv.org/abs/1505.04597.
- Wang Q, Wu B, Zhu P, Li P, Zuo W, Hu Q. ECA-Net: Efficient channel attention for deep convolutional neural networks. In: Proceedings of the IEEE/CVF conference on computer vision and pattern recognition; 2020. p. 11534–11542.
- Bian J, Liu Y. Dual Channel Attention Networks. In: Journal of Physics: Conference Series. vol. 1642. IOP Publishing; 2020. p. 012004.
- Garc’ıa-Pedrajas N, Herv´as-Mart´ınez C, Ortiz-Boyer D. Cooperative coevolution of artificial neural network ensembles for pattern classification. IEEE transactions on evolutionary computation. 2005;9(3):271–302.
- Rusu AA, Rabinowitz NC, Desjardins G, Soyer H, Kirkpatrick J, Kavukcuoglu K, et al. Progressive neural networks. arXiv preprint arXiv:160604671. 2016.
- Kirst C, Skriabine S, Vieites-Prado A, Topilko T, Bertin P, Gerschenfeld G, et al. Mapping the fine-scale organization and plasticity of the brain vasculature. Cell. 2020;180(4):780–795.
- Pacheco M, Oliva GA, Rajbahadur GK, Hassan AE. Is my transaction done yet? An empirical study of transaction processing times in the Ethereum Blockchain Platform. ACM Transactions on Software Engineering and Methodology. 2022.
- Mashhadi PS, Nowaczyk S, Pashami S. Parallel orthogonal deep neural network. Neural Networks. 2021;140:167–183.
- Hu L, Cao J, Xu G, Cao L, Gu Z, Zhu C. Personalized recommendation via cross-domain triadic factorization. In: Proceedings of the 22nd international conference on World Wide Web; 2013. p. 595–606.
- Fern´andez-Mart´ınez F, Luna-Jim´enez C, Kleinlein R, Griol D, Callejas Z, Montero JM. Fine-tuning BERT models for intent recognition using a frequency cut-off strategy for domain-specific vocabulary extension. Applied Sciences. 2022;12(3):1610.
- Georgescu MI, Ionescu RT, Miron AI. Diversity-Promoting Ensemble for Medical Image Segmentation. In: Proceedings of the 38th ACM/SIGAPP Symposium on Applied Computing; 2023. p. 599–606.
- Jiang W, Chen Y, Wen S, Zheng L, Jin H. PDAS: Improving network pruning based on Progressive Differentiable Architecture Search for DNNs. Future Generation Computer Systems. 2023;146:98–113.
- Chou, A., Li, W. and Roman, E., 2022. GI tract image segmentation with U-Net and mask R-CNN. Image Segmentation with U-Net and Mask R-CNN. Available online: http://cs231n. stanford. edu/reports/2022/pdfs/164. pdf (accessed on 4 June 2023).
- Sai, M.J. and Punn, N.S., 2023. LWU-Net approach for Efficient Gastro-Intestinal Tract Image Segmentation in Resource-Constrained Environments. medRxiv, pp.2023-12.
- Guggari, S., Srivastava, B.C., Kumar, V., Harshita, H., Farande, V., Kulkarni, U. and Meena, S.M., 2022, December. RU-Net: A Novel Approach for Gastro-Intestinal Tract Image Segmentation Using Convolutional Neural Network. In International Conference on Applied Machine Learning and Data Analytics (pp. 131-141). Cham: Springer Nature Switzerland.
- Zhou, H., Lou, Y., Xiong, J., Wang, Y. and Liu, Y., 2023. Improvement of Deep Learning Model for Gastrointestinal Tract Segmentation Surgery. Frontiers in Computing and Intelligent Systems, 6(1), pp.103-106.
- Sharma, N., Gupta, S., Rajab, A., Elmagzoub, M.A., Rajab, K. and Shaikh, A., 2023. Semantic Segmentation of Gastrointestinal Tract in MRI Scans Using PSPNet Model With ResNet34 Feature Encoding Network. IEEE Access, 11, pp.132532-132543.
- Sharma, N., Gupta, S., Koundal, D., Alyami, S., Alshahrani, H., Asiri, Y. and Shaikh, A., 2023. U-Net model with transfer learning model as a backbone for segmentation of gastrointestinal tract. Bioengineering, 10(1), p.119.
- Zhou, H., Lou, Y., Xiong, J., Wang, Y. and Liu, Y., 2023. Improvement of Deep Learning Model for Gastrointestinal Tract Segmentation Surgery. Frontiers in Computing and Intelligent Systems, 6(1), pp.103-106.
- Zhou, H., Lou, Y., Xiong, J., Wang, Y. and Liu, Y., 2023. Improvement of Deep Learning Model for Gastrointestinal Tract Segmentation Surgery. Frontiers in Computing and Intelligent Systems, 6(1), pp.103-106.
- Elgayar, S.M., Hamad, S. and El-Horbaty, E.S.M., 2023. Revolutionizing Medical Imaging through Deep Learning Techniques: An Overview. International Journal of Intelligent Computing and Information Sciences, 23(3), pp.59-72.
- Zhang, C., Xu, J., Tang, R., Yang, J., Wang, W., Yu, X. and Shi, S., 2023. Novel research and future prospects of artificial intelligence in cancer diagnosis and treatment. Journal of Hematology & Oncology, 16(1), p.114.
| Transfer-learning | Dice (%) | IoU (%) | Valid dice (%) | Valid IoU (%) |
|---|---|---|---|---|
| EfficientNet-B0 | 91.11 | 88.39 | 88.75 | 87.92 |
| EfficientNet-B1 | 91.86 | 88.60 | 91.72 | 88.03 |
| EfficientNet-B2 | 91.75 | 88.48 | 91.16 | 87.51 |
| ResNet-34 | 91.92 | 89.57 | 87.83 | 87.46 |
| ResECA-U-Net (proposed) | 96.27 | 91.48 | 92.03 | 87.90 |
| Transfer-learning | Dice (%) | IoU (%) | Valid dice (%) | Valid IoU (%) |
|---|---|---|---|---|
| RestNet-34 | 89.75 | 90.78 | 82.8 | 82.72 |
| EfficientNet-B1 | 88.57 | 87.12 | 80.52 | 79.66 |
| EfficientNet-B2 | 89.84 | 89.96 | 85.52 | 84.88 |
| ResNet-18 | 92.25 | 93.15 | 80.08 | 80.88 |
| ResECA-U-Net (proposed) | 96.27 | 91.48 | 92.03 | 87.90 |
| Model | Dice (%) | IoU (%) |
|---|---|---|
| U-Net with EfficientNet-B1 | 91.30 | 88.60 |
| LeViT with U-Net++ | 79.50 | 72.80 |
| SmallUnet-B2 | 83.14 | 83.14 |
| U-Net | 88.54 | 88.19 |
| Proposed model (ResECA-U-Net) | 96.27 | 91.47 |
| Ref. | Dataset type | Model | score |
| [26] | UW-Madison GI Tract Image Segmentation | R-CNN U-Net |
Dice score- 73% Dice score- 51% |
| [28] | UW-Madison GI Tract Image Segmentation | ResNet34 - U-Net | Dice score-90.49% |
| [27] | UW-Madison GI Tract Image Segmentation | Light Weight U-Net U-Net with ResNet34 |
Dice score- 77.91% || IoU score- 82.69% Dice score- 77.91% || IoU score- 82.69% |
| [29] | UW-Madison Carbone Cancer Center | Unet2.5D model | Dice score- 84.8% |
| [30] | UW Madison GI tract dataset | PSPNet Model With ResNet34 | Dice score- 88.44% |
| [31] | U-Net | Dice score- 88.54% || IoU score- 88.19% |
|
| (Proposed Model) | UW-Madison GI Tract Image Segmentation | ResECA-U-Net | Dice score- 96.27 || IoU score- 91.47 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




