Submitted:
27 March 2024
Posted:
28 March 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Dataset
2.2. Statistical Analysis
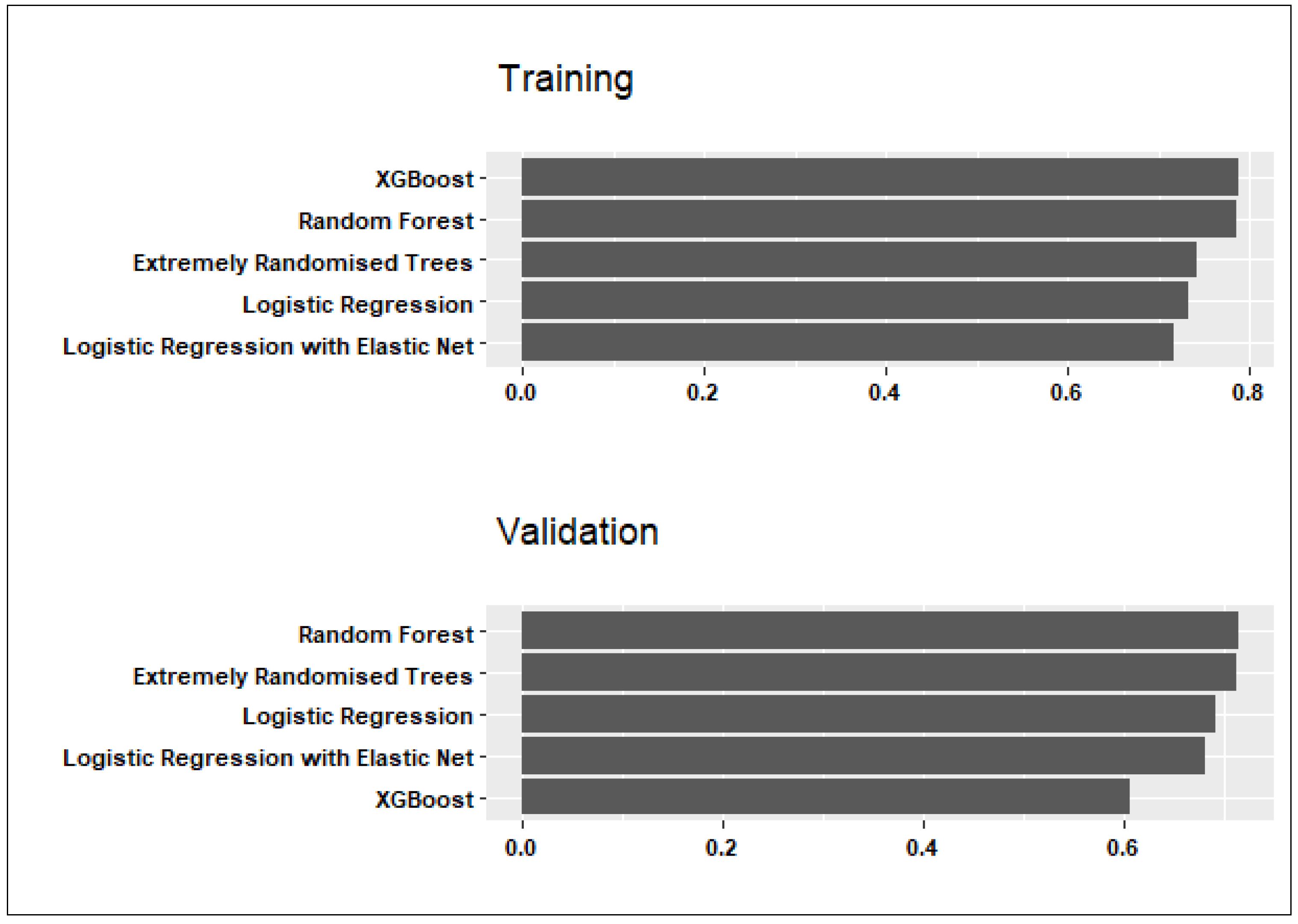
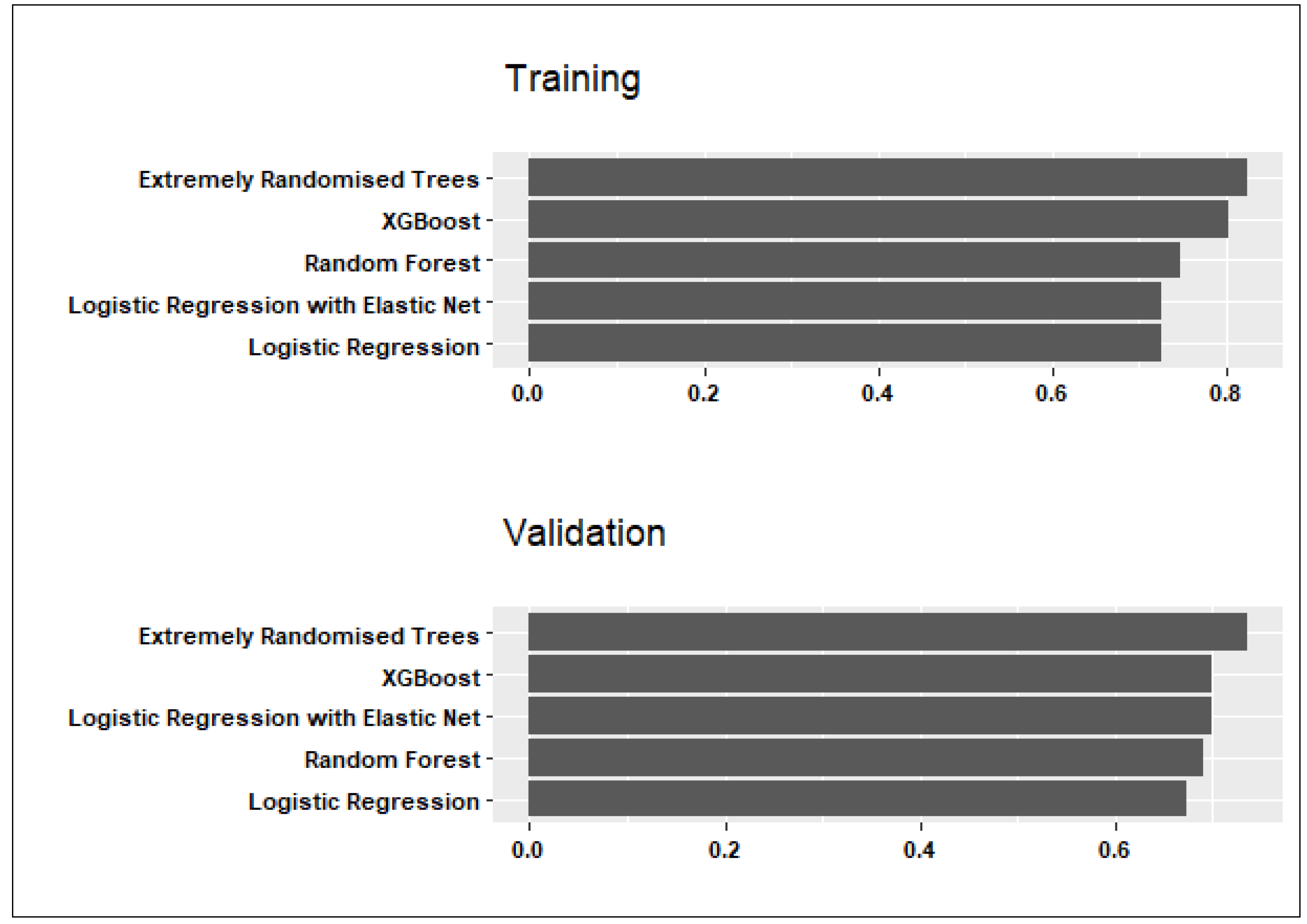
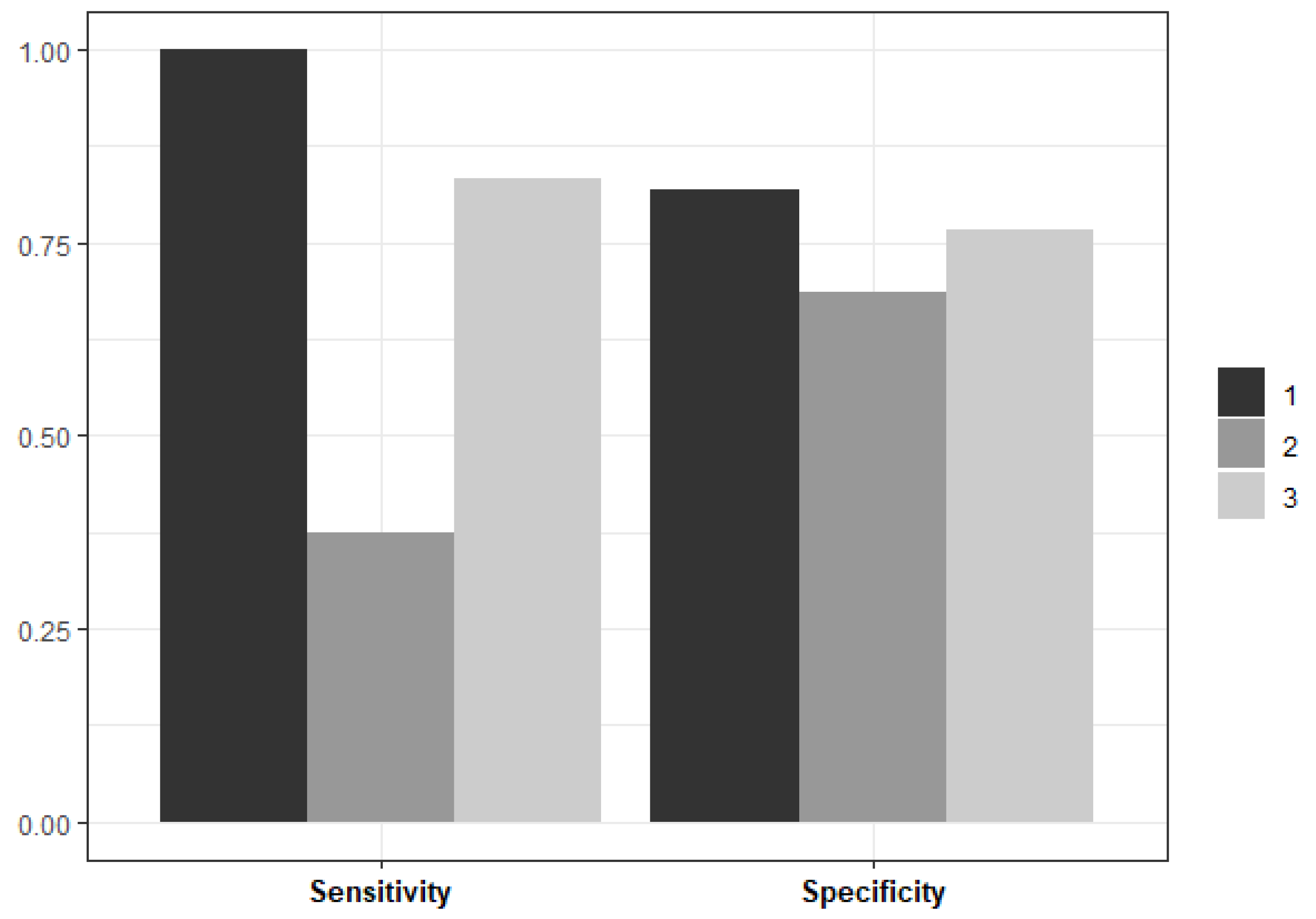
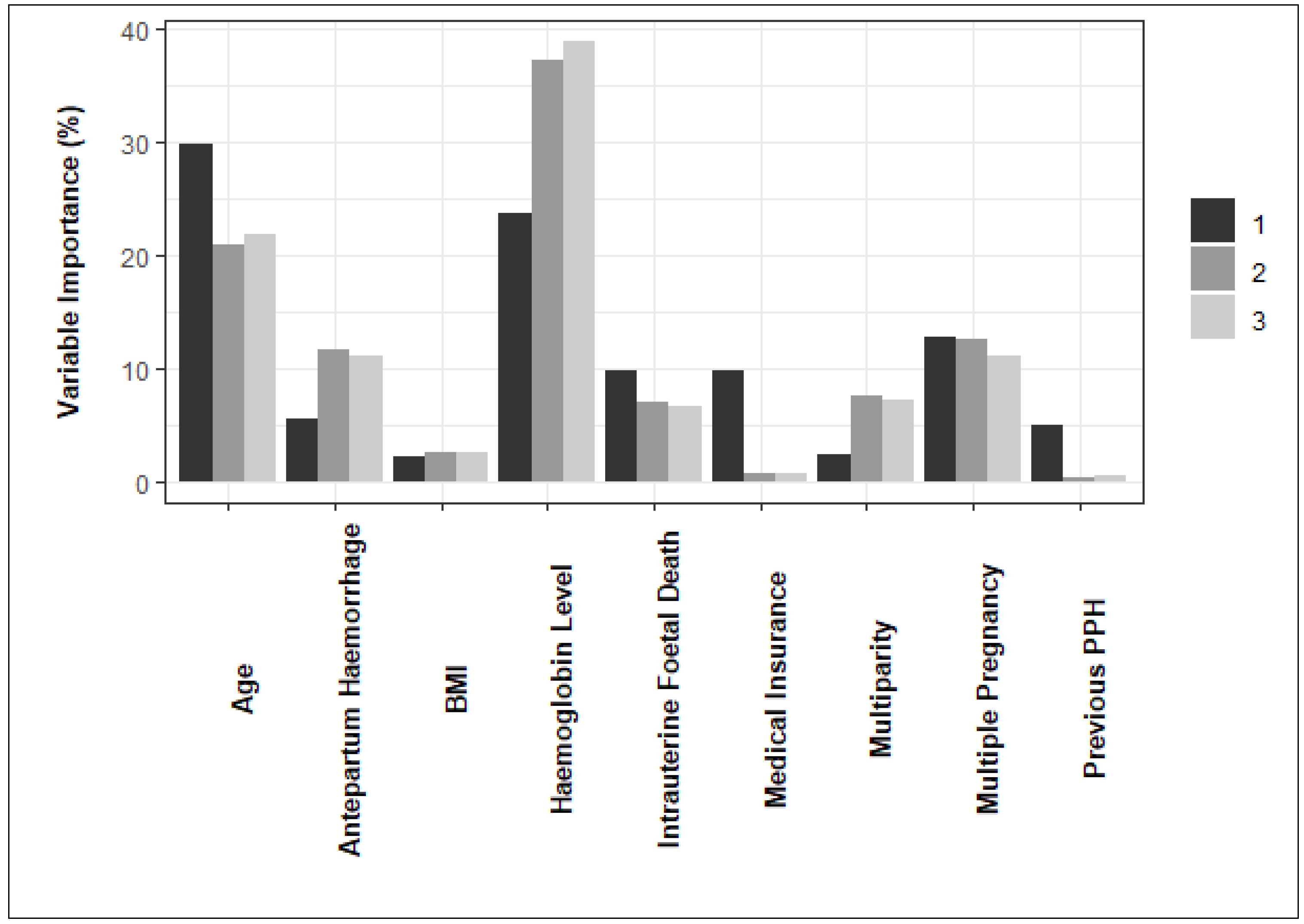
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hamm, R.F.; Wang, E.; Romanos, A.; O'Rourke, K.; Srinivas, S.K. Implementation of Quantification of Blood Loss Does Not Improve Prediction of Hemoglobin Drop in Deliveries with Average Blood Loss. Am. J. Perinatol. 2018, 35, 134–139. [Google Scholar] [CrossRef] [PubMed]
- Committee on Practice Bulletins-Obstetrics. Practice bulletin no. 183: Postpartum haemorrhage. Obstet. Gynecol. 2017, 130, e168–e186. [CrossRef]
- Knight, M.; Callaghan, W.M.; Berg, C.; et al. Trends in postpartum haemorrhage in high resource countries: A review and recommendations from the International postpartum haemorrhage Collaborative group. BMC Pregnancy Childbirth 2009, 9, 55. [Google Scholar] [CrossRef] [PubMed]
- Say, L.; Chou, D.; Gemmill, A.; et al. Global causes of maternal death: A WHO systematic analysis. Lancet Glob. Health 2014, 2, e323–e333. [Google Scholar] [CrossRef] [PubMed]
- Boujarzadeh, B.; Ranjbar, A.; Banihashemi, F.; Mehrnoush, V.; Darsareh, F.; Saffari, M. Machine learning approach to predict postpartum haemorrhage: A systematic review protocol. BMJ Open 2023, 13, e067661. [Google Scholar] [CrossRef]
- Creanga, A.A.; Berg, C.J.; Ko, J.Y.; et al. Maternal mortality and morbidity in the United States: Where are we now? J. Women's Health 2014, 23, 3–9. [Google Scholar] [CrossRef] [PubMed]
- Chandraharan, E.; Krishna, A. Diagnosis and management of postpartum haemorrhage. BMJ 2017, 358, j3875. [Google Scholar] [CrossRef]
- Bazirete, O.; Nzayirambaho, M.; Umubyeyi, A.; Karangwa, I.; Evans, M. Risk factors for postpartum haemorrhage in the Northern Province of Rwanda: A case control study. PLoS ONE 2022, 17, e0263731. [Google Scholar] [CrossRef]
- Mesfin, S.; Dheresa, M.; Fage, S.G.; Tura, A.K. Assessment of postpartum hemorrhage in a university hospital in Eastern Ethiopia: A cross-sectional study. Int. J. Women's Health 2021, 663–669. [Google Scholar] [CrossRef]
- Ononge, S.; et al. Incidence and risk factors for postpartum hemorrhage in Uganda. Reprod. Health 2016, 13, 1. [Google Scholar] [CrossRef]
- Boulesteix, A.L.; Schmid, M. Machine learning versus statistical modeling. Biometrical J. 2014, 56, 588–593. [Google Scholar] [CrossRef] [PubMed]
- Christodoulou, E.; Ma, J.; Collins, G.S.; Steyerberg, E.W.; Verbakel, J.Y.; Van Calster, B. A systematic review shows no performance benefit of machine learning over logistic regression for clinical prediction models. J. Clin. Epidemiol. 2019, 110, 12–22. [Google Scholar] [CrossRef] [PubMed]
- Venkatesh, K.K.; Strauss, R.A.; Grotegut, C.; et al. Machine learning and statistical models to predict postpartum hemorrhage. Obstet. Gynecol. 2020, 135, 935. [Google Scholar] [CrossRef]
- Zhou, P.; Hu, X.; Li, P.; Wu, X. Online feature selection for high-dimensional class-imbalanced data. Knowl.-Based Syst. 2017, 136, 187–199. [Google Scholar] [CrossRef]
- Zou, H.; Hastie, T. Regularization and variable selection via the elastic net. J. R. Stat. Soc. Ser. B Stat. Methodol. 2005, 67, 301–320. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Natekin, A.; Knoll, A. Gradient boosting machines, a tutorial. Front. Neurorobot. 2013, 7, 21. [Google Scholar] [CrossRef] [PubMed]
- Geurts, P.; Ernst, D.; Wehenkel, L. Extremely randomized trees. Mach. Learn. 2006, 63, 3–42. [Google Scholar] [CrossRef]
- Chen, T.; He, T.; Benesty, M.; et al. Xgboost: Extreme gradient boosting. R package version 2015, 0.4-2, 1–4. [CrossRef]
- Mehrnoush, V.; Ranjbar, A.; Farashah, M.V.; Darsareh, F.; Shekari, M.; Jahromi, M.S. Prediction of postpartum hemorrhage using traditional statistical analysis and a machine learning approach. AJOG Glob. Rep. 2023, 3, 100185. [Google Scholar] [CrossRef]
- Søreide, K. Receiver-operating characteristic curve analysis in diagnostic, prognostic and predictive biomarker research. J. Clin. Pathol. 2009, 62, 1–5. [Google Scholar] [CrossRef] [PubMed]
- RStudio Team. RStudio: Integrated Development for R. RStudio, PBC. 2021. Available online: https://www.rstudio.com/.
- Kuhn, M.; Wickham, H. Tidymodels: A collection of packages for modeling and machine learning using tidyverse principles. 2020. Available online: https://www.tidymodels.org.
- Kuhn, M. caret: Classification and Regression Training. R package version 6.0-88. 2022. Available online: https://CRAN.R-project.org/package=caret.
- Wright, M.N.; Wager, S.; Probst, P.; Wright, M.M.N. Package ‘ranger’. Version 0.11.2. 2019. Available online: https://CRAN.R-project.org/package=ranger.
- Khalilia, M.; Chakraborty, S.; Popescu, M. Predicting disease risks from highly imbalanced data using random forest. BMC Med. Inform. Decis. Mak. 2011, 11, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Ghiasi, M.M.; Zendehboudi, S. Application of decision tree-based ensemble learning in the classification of breast cancer. Comput. Biol. Med. 2021, 128, 104089. [Google Scholar] [CrossRef] [PubMed]

| Data Partition | Number of random features | Minimum terminal node size |
|---|---|---|
| 1 | 2 | 20 |
| 2 | 2 | 25 |
| 3 | 2 | 20 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




