Submitted:
06 March 2024
Posted:
07 March 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
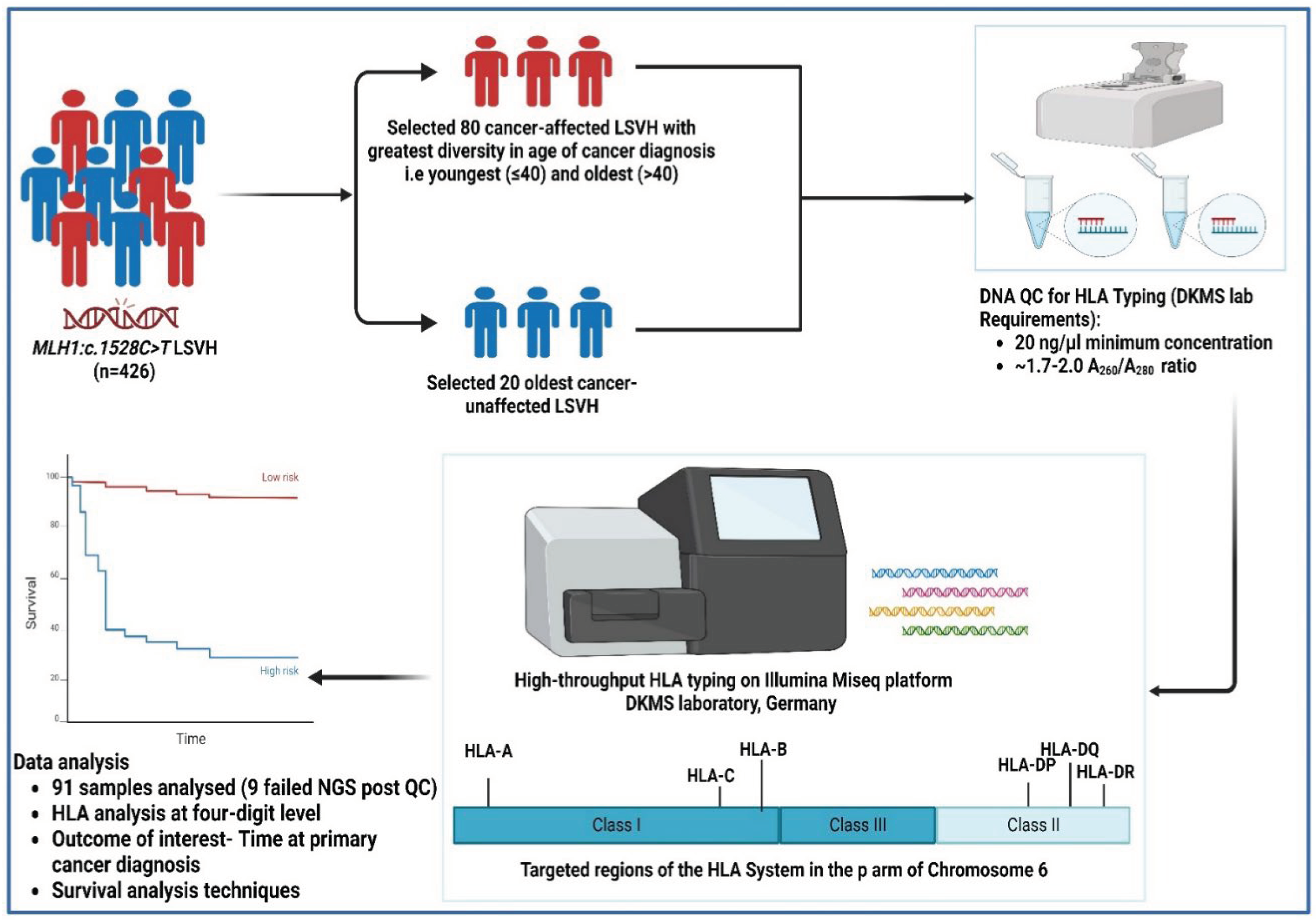
2.1. Patients
2.2. DNA Samples
2.3. HLA Typing
2.4. Statistical Analysis
3. Results
Overall Demographic and Clinical Characteristics of the Patients
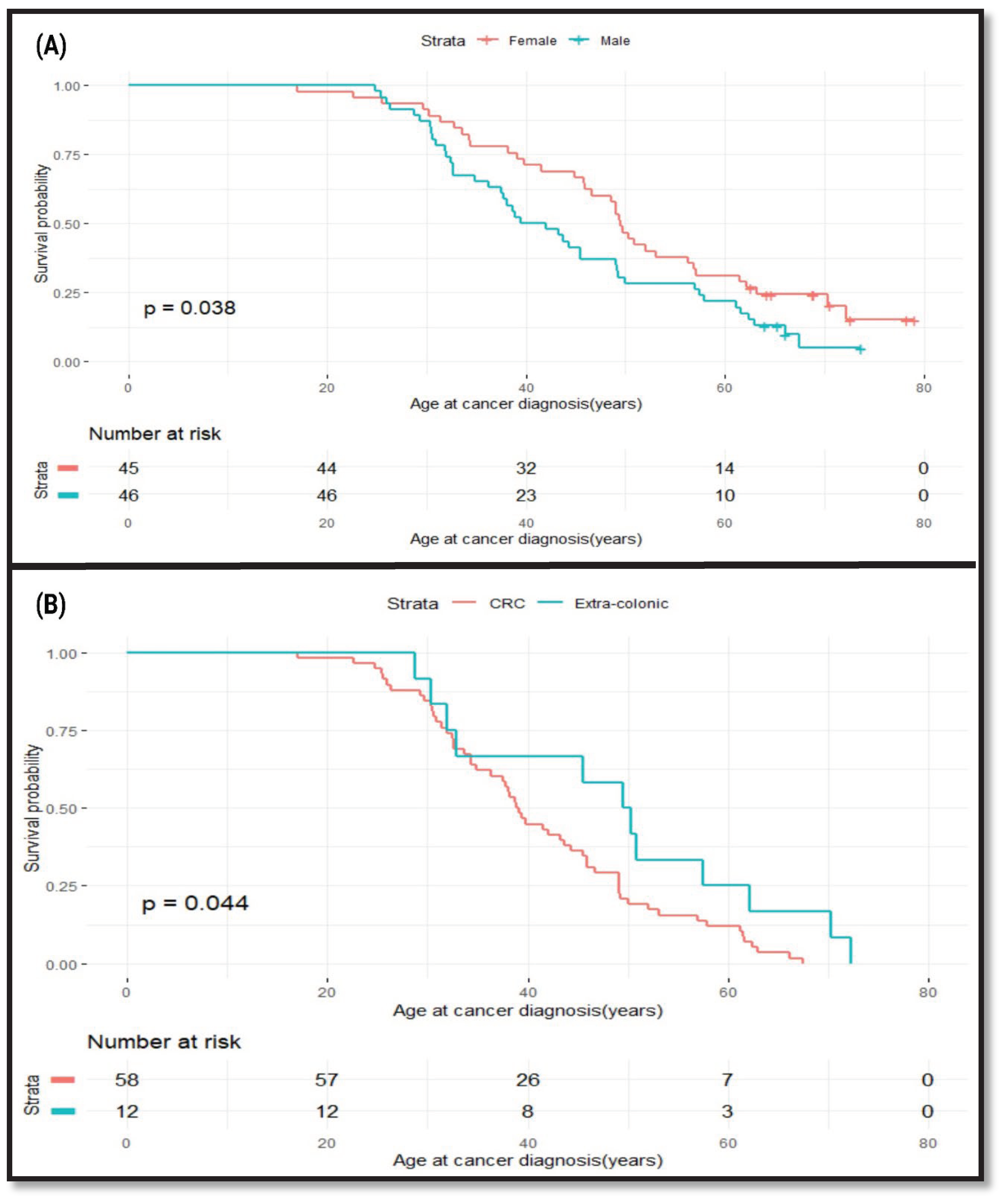
Effects of Gender and Cancer Site on Age at Cancer Diagnosis in LSVH
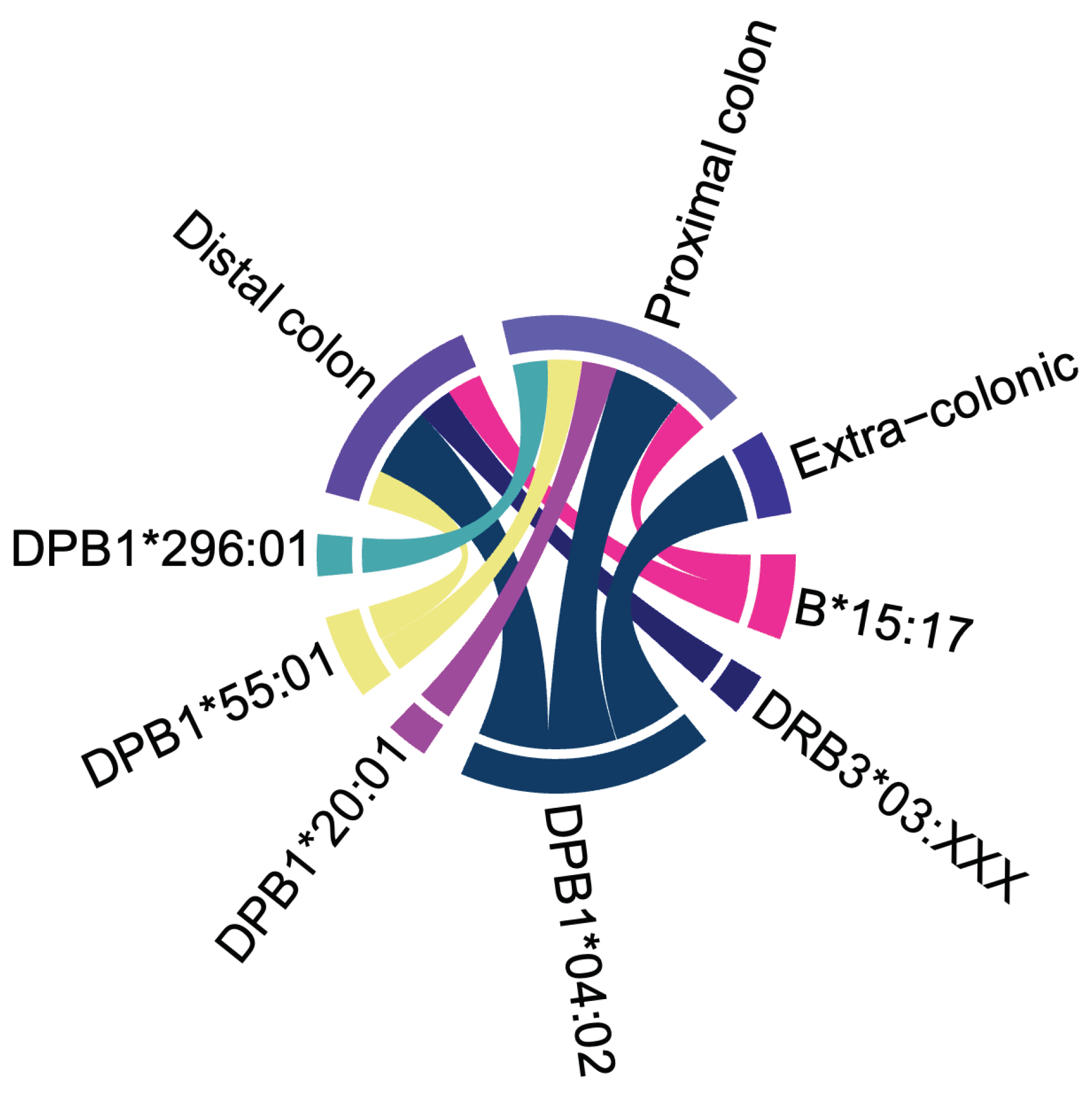
Effects of HLA Alleles on the Age at Cancer Diagnosis in LSVH
Effects of HLA Alleles on the Age at CRC Diagnosis in LSVH
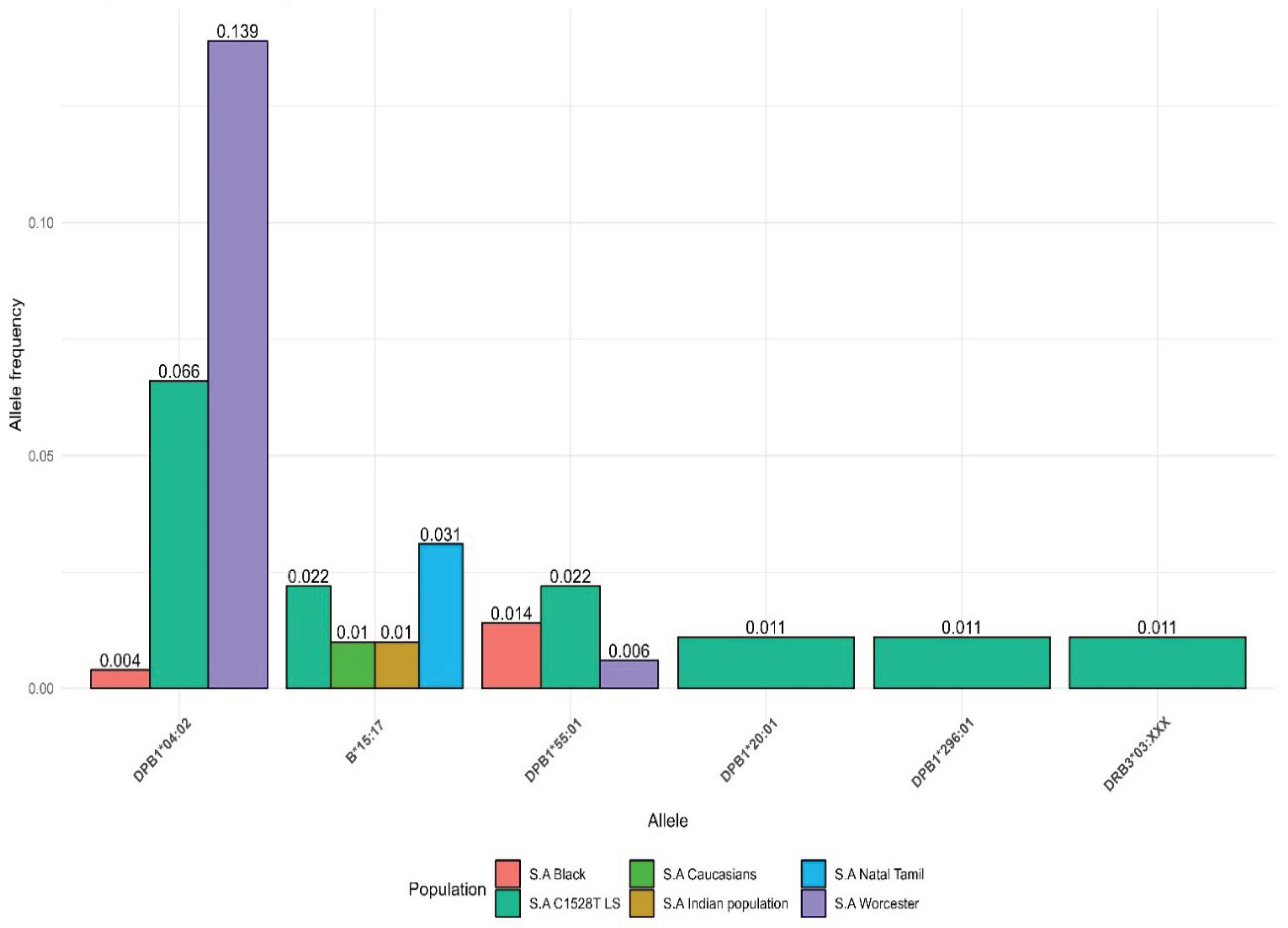
Different HLA Allele Frequencies between LSVH and the Previously Studied South African General Populations
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lynch HT, Krush AJ. Cancer family “G” revisited: 1895-1970. Cancer 1971, 27, 1505–1511. [CrossRef]
- Lynch HT, Lynch PM, Lanspa SJ, Snyder CL, Lynch JF, Boland CR. Review of the Lynch syndrome: history, molecular genetics, screening, differential diagnosis, and medicolegal ramifications. Clin. Genet. 2009, 76, 1–18. [CrossRef]
- Lynch HT, de la Chapelle A. Genetic susceptibility to non-polyposis colorectal cancer. J. Med. Genet. 1999, 36, 801–818.
- Talseth-Palmer BA, Wijnen JT, Grice DM, Scott RJ. Genetic modifiers of cancer risk in Lynch syndrome: a review. Fam. Cancer 2013, 12, 207–216. [CrossRef] [PubMed]
- Bucksch K, Zachariae S, Aretz S, Buttner R, Holinski-Feder E, Holzapfel S, et al. Cancer risks in Lynch syndrome, Lynch-like syndrome, and familial colorectal cancer type X: a prospective cohort study. BMC Cancer 2020, 20, 460. [CrossRef]
- Dominguez-Valentin M, Sampson JR, Seppala TT, Ten Broeke SW, Plazzer JP, Nakken S, et al. Cancer risks by gene, age, and gender in 6350 carriers of pathogenic mismatch repair variants: findings from the Prospective Lynch Syndrome Database. Genet Med. 2020, 22, 15–25. [CrossRef]
- Ahadova A, Seppala TT, Engel C, Gallon R, Burn J, Holinski-Feder E, et al. The “unnatural” history of colorectal cancer in Lynch syndrome: Lessons from colonoscopy surveillance. Int. J. Cancer 2021, 148, 800–811. [CrossRef] [PubMed]
- Felix R, Bodmer W, Fearnhead NS, van der Merwe L, Goldberg P, Ramesar RS. GSTM1 and GSTT1 polymorphisms as modifiers of age at diagnosis of hereditary nonpolyposis colorectal cancer (HNPCC) in a homogeneous cohort of individuals carrying a single predisposing mutation. Mutat. Res. 2006, 602, 175–181. [CrossRef]
- Stupart DA, Goldberg PA, Algar U, Ramesar R. Cancer risk in a cohort of subjects carrying a single mismatch repair gene mutation. Fam. Cancer 2009, 8, 519–523. [CrossRef]
- Ballhausen A, Przybilla MJ, Jendrusch M, Haupt S, Pfaffendorf E, Seidler F, et al. The shared frameshift mutation landscape of microsatellite-unstable cancers suggests immunoediting during tumor evolution. Nat. Commun. 2020, 11, 4740. [CrossRef]
- Ahadova A, Witt J, Haupt S, Gallon R, Huneburg R, Nattermann J, et al. Is HLA type a possible cancer risk modifier in Lynch syndrome? Int. J. Cancer. 2023, 152, 2024–2031. [CrossRef]
- Holoshitz, J. The quest for better understanding of HLA-disease association: scenes from a road less travelled by. Discov. Med. 2013, 16, 93–101. [Google Scholar] [PubMed]
- Klein J, Sato A. The HLA system. Second of two parts. N. Engl. J. Med. 2000, 343, 782–786. [CrossRef]
- Li XC, Raghavan M. Structure and function of major histocompatibility complex class I antigens. Curr. Opin. Organ Transplant. 2010, 15, 499–504. [CrossRef]
- Jurtz V, Paul S, Andreatta M, Marcatili P, Peters B, Nielsen M. NetMHCpan-4.0: Improved Peptide-MHC Class I Interaction Predictions Integrating Eluted Ligand and Peptide Binding Affinity Data. J. Immunol. 2017, 199, 3360–3368. [CrossRef] [PubMed]
- Falk K, Rötzschke O, Stevanović S, Jung G, Rammensee HG. Allele-specific motifs revealed by sequencing of self-peptides eluted from MHC molecules. Nature 1991, 351, 290–296. [CrossRef]
- Bjorkman PJ, Saper MA, Samraoui B, Bennett WS, Strominger JL, Wiley DC. The foreign antigen binding site and T cell recognition regions of class I histocompatibility antigens. Nature 1987, 329, 512–518. [CrossRef] [PubMed]
- Wysocki T, Olesińska M, Paradowska-Gorycka A. Current Understanding of an Emerging Role of HLA-DRB1 Gene in Rheumatoid Arthritis–From Research to Clinical Practice. Cells 2020, 9. [CrossRef]
- Klein J, Sato A. The HLA system. First of two parts. N. Engl. J. Med. 2000, 343, 702–709. [CrossRef]
- Chowell D, Morris LGT, Grigg CM, Weber JK, Samstein RM, Makarov V, et al. Patient HLA class I genotype influences cancer response to checkpoint blockade immunotherapy. Science 2018, 359, 582–587. [CrossRef]
- Naranbhai V, Viard M, Dean M, Groha S, Braun DA, Labaki C, et al. HLA-A*03 and response to immune checkpoint blockade in cancer: an epidemiological biomarker study. Lancet Oncol. 2022, 23, 172–184. [CrossRef]
- Li H, Liu D, Li X. HLA-DPB1 and Epstein-Barr virus gp42 protein jointly contribute to the development of Hodgkin lymphoma. Transl. Cancer Res. 2020, 9, 4424–4432. [CrossRef]
- Liu Z, Hildesheim A. Association Between Human Leukocyte Antigen Class I and II Diversity and Non-virus-associated Solid Tumors. Frontiers in Genetics 2021, 12. [CrossRef]
- Liu Z, Huang CJ, Huang YH, Pan MH, Lee MH, Yu KJ, et al. HLA Zygosity Increases Risk of Hepatitis B Virus-Associated Hepatocellular Carcinoma. J. Infect. Dis. 2021, 224, 1796–1805. [CrossRef]
- Hirata I, Murano M, Ishiguro T, Toshina K, Wang FY, Katsu K. HLA genotype and development of gastric cancer in patients with Helicobacter pylori infection. Hepatogastroenterology 2007, 54, 990–994.
- Chambuso R, Ramesar R, Kaambo E, Denny L, Passmore JA, Williamson AL, et al. Human Leukocyte Antigen (HLA) Class II -DRB1 and -DQB1 Alleles and the Association with Cervical Cancer in HIV/HPV Co-Infected Women in South Africa. J. Cancer 2019, 10, 2145–2152. [CrossRef]
- Albrecht V, Zweiniger C, Surendranath V, Lang K, Schofl G, Dahl A, et al. Dual redundant sequencing strategy: Full-length gene characterisation of 1056 novel and confirmatory HLA alleles. HLA 2017, 90, 79–87. [CrossRef]
- Listgarten J, Brumme Z, Kadie C, Xiaojiang G, Walker B, Carrington M, et al. Statistical resolution of ambiguous HLA typing data. PLoS Comput. Biol. 2008, 4, e1000016. [CrossRef]
- Katharina Imkeller [cre a. immunotation: Tools for working with diverse immune genes. R package version 1.8.0. 2023. Available online: https://bioconductor.org/packages/immunotation/.
- Cohen SA, Leininger A. The genetic basis of Lynch syndrome and its implications for clinical practice and risk management. Appl. Clin. Genet. 2014, 7, 147–158. [CrossRef]
- Schneider R, Schneider C, Jakobeit C, Furst A, Moslein G. Gender-Specific Aspects of Lynch Syndrome and Familial Adenomatous Polyposis. Viszeralmedizin 2014, 30, 82–88. [CrossRef]
- Vasen HFA, Moslein G, Alonso A, Bernstein I, Bertario L, Blanco I, et al. Guidelines for the clinical management of Lynch syndrome (hereditary non-polyposis cancer). Journal of Medical Genetics 2007, 44, 353–362. [CrossRef] [PubMed]
- Dominguez-Valentin M, Haupt S, Seppälä TT, Sampson JR, Sunde L, Bernstein I, et al. Mortality by age, gene and gender in carriers of pathogenic mismatch repair gene variants receiving surveillance for early cancer diagnosis and treatment: a report from the prospective Lynch syndrome database. EClinicalMedicine 2023, 58, 101909. [CrossRef]
- Bonadona V, Bonaïti B, Olschwang S, Grandjouan S, Huiart L, Longy M, et al. Cancer risks associated with germline mutations in MLH1, MSH2, and MSH6 genes in Lynch syndrome. Jama 2011, 305, 2304–2310. [CrossRef] [PubMed]
- Barker DJ, Maccari G, Georgiou X, Cooper MA, Flicek P, Robinson J, et al. The IPD-IMGT/HLA Database. Nucleic Acids Res. 2023, 51, D1053-D60. [CrossRef]
- Tshabalala M, Mellet J, Vather K, Nelson D, Mohamed F, Christoffels A, et al. High Resolution HLA ∼A, ∼B, ∼C, ∼DRB1, ∼DQA1, and ∼DQB1 Diversity in South African Populations. Front. Genet. 2023, 13, 711944. [CrossRef]
- Janse van Rensburg WJ, de Kock A, Bester C, Kloppers JF. HLA major allele group frequencies in a diverse population of the Free State Province, South Africa. Heliyon 2021, 7, e06850. [CrossRef] [PubMed]
- Lombard Z, Brune AE, Hoal EG, Babb C, Van Helden PD, Epplen JT, et al. HLA class II disease associations in southern Africa. Tissue Antigens 2006, 67, 97–110. [CrossRef]
- Yurgelun MB, Hampel H. Recent Advances in Lynch Syndrome: Diagnosis, Treatment, and Cancer Prevention. Am Soc Clin Oncol Educ Book 2018, 38, 101–109. [CrossRef]
- Seppala TT, Dominguez-Valentin M, Crosbie EJ, Engel C, Aretz S, Macrae F, et al. Uptake of hysterectomy and bilateral salpingo-oophorectomy in carriers of pathogenic mismatch repair variants: a Prospective Lynch Syndrome Database report. Eur. J. Cancer 2021, 148, 124–133. [CrossRef]
- Williams HB, Turner TR, Cambridge CA, Marsh SGE, Mayor NP. The novel HLA-DRB1*03:01:01:05 and -DPB1*04:02:01:21 alleles identified in patients with acute leukemia. HLA 2022, 99, 650–652. [CrossRef]
- Xin YN, Lin ZH, Jiang XJ, Zhan SH, Dong QJ, Wang Q, et al. Specific HLA-DQB1 alleles associated with risk for development of hepatocellular carcinoma: a meta-analysis. World J. Gastroenterol. 2011, 17, 2248–2254. [CrossRef]
- Kohno T, Kunitoh H, Mimaki S, Shiraishi K, Kuchiba A, Yamamoto S, et al. Contribution of the TP53, OGG1, CHRNA3, and HLA-DQA1 genes to the risk for lung squamous cell carcinoma. J. Thorac. Oncol. 2011, 6, 813–817. [CrossRef] [PubMed]
- Jackow CM, McHam JB, Friss A, Alvear J, Reveille JR, Duvic M. HLA-DR5 and DQB1*03 class II alleles are associated with cutaneous T-cell lymphoma. J. Invest. Dermatol. 1996, 107, 373–376. [CrossRef] [PubMed]
- Wu MS, Hsieh RP, Huang SP, Chang YT, Lin MT, Chang MC, et al. Association of HLA-DQB1*0301 and HLA-DQB1*0602 with different subtypes of gastric cancer in Taiwan. Jpn. J. Cancer Res. 2002, 93, 404–410. [CrossRef] [PubMed]
- Magnusson PKE, Enroth H, Eriksson I, Held M, Nyren O, Engstrand L, et al. Gastric cancer and human leukocyte antigen: distinct DQ and DR alleles are associated with development of gastric cancer and infection by Helicobacter pylori. Cancer Res. 2001, 61, 2684–2689.
- Cheng L, Guo Y, Zhan S, Xia P. Association between HLA-DP Gene Polymorphisms and Cervical Cancer Risk: A Meta-Analysis. Biomed. Res. Int. 2018, 2018, 7301595. [CrossRef]
- Shi Y, Li L, Hu Z, Li S, Wang S, Liu J, et al. A genome-wide association study identifies two new cervical cancer susceptibility loci at 4q12 and 17q12. Nat. Genet. 2013, 45, 918–922. [CrossRef] [PubMed]
- Wu Y, Liu B, Lin W, Xu Y, Li L, Zhang Y, et al. Human leukocyte antigen class II alleles and risk of cervical cancer in China. Hum. Immunol. 2007, 68, 192–200. [CrossRef]
- Liang J, Xu A, Xie Y, Awonuga AO, Lin Z. Some but not all of HLA-II alleles are associated with cervical cancer in Chinese women. Cancer Genet Cytogenet. 2008, 187, 95–100. [CrossRef]
- Rivera-Pirela SE, Echeverría M, Salcedo P, Márquez G, Carrillo Z, Parra Y, et al. [HLA DRB1*, DQB1*, DPA1*, and DPB1* and their association with the pathogenesis of leukemia in the population of Venezuela]. Rev. Alerg. Mex. 2016, 63, 237–251. [CrossRef]
- Sivapalan L, Anagnostou V. Genetic variation in antigen presentation and cancer immunotherapy. Immunity 2022, 55, 3–6. [CrossRef] [PubMed]
- Seliger B, Kloor M, Ferrone S. HLA class II antigen-processing pathway in tumors: Molecular defects and clinical relevance. Oncoimmunology 2017, 6, e1171447. [CrossRef] [PubMed]
- Leo PJ, Madeleine MM, Wang S, Schwartz SM, Newell F, Pettersson-Kymmer U, et al. Defining the genetic susceptibility to cervical neoplasia-A genome-wide association study. PLoS Genet. 2017, 13, e1006866. [CrossRef]
- Ekanayake Weeramange C, Shu D, Tang KD, Batra J, Ladwa R, Kenny L, et al. Analysis of human leukocyte antigen associations in human papillomavirus-positive and -negative head and neck cancer: Comparison with cervical cancer. Cancer 2022, 128, 1937–1947. [CrossRef] [PubMed]
- Augusto DG, Murdolo LD, Chatzileontiadou DSM, Sabatino JJ, Jr., Yusufali T, Peyser ND, et al. A common allele of HLA is associated with asymptomatic SARS-CoV-2 infection. Nature 2023, 620, 128–136. [CrossRef] [PubMed]
- Pastor DM, Schlom J. Immunology of Lynch Syndrome. Curr. Oncol. Rep. 2021, 23, 96. [CrossRef] [PubMed]
- Lynch HT, Drescher KM, de la Chapelle A. Immunology and the Lynch syndrome. Gastroenterology 2008, 134, 1246–1249. [CrossRef]
- Chambuso R, Kaambo E, Rebello G, Ramesar R. Correspondence on “Cancer risks by gene, age, and gender in 6350 carriers of pathogenic mismatch repair variants: findings from the Prospective Lynch Syndrome Database” by Dominguez-Valentin et al. Genet Med. 2022, 24, 1148–1150. [CrossRef]
- Burn J, Bishop DT, Chapman PD, Elliott F, Bertario L, Dunlop MG, et al. A randomized placebo-controlled prevention trial of aspirin and/or resistant starch in young people with familial adenomatous polyposis. Cancer Prev Res (Phila) 2011, 4, 655–665. [CrossRef]
- Pande M, Amos CI, Osterwisch DR, Chen J, Lynch PM, Broaddus R, et al. Genetic variation in genes for the xenobiotic-metabolizing enzymes CYP1A1, EPHX1, GSTM1, GSTT1, and GSTP1 and susceptibility to colorectal cancer in Lynch syndrome. Cancer Epidemiol Biomarkers Prev. 2008, 17, 2393–2401. [CrossRef]
- Campbell PT, Edwards L, McLaughlin JR, Green J, Younghusband HB, Woods MO. Cytochrome P450 17A1 and catechol O-methyltransferase polymorphisms and age at Lynch syndrome colon cancer onset in Newfoundland. Clin. Cancer Res. 2007, 13, 3783–3788. [CrossRef]
- Hitchins MP, Lynch HT. Dawning of the epigenetic era in hereditary cancer. Clin. Genet. 2014, 85, 413–416. [CrossRef] [PubMed]
- Hitchins MP, Rapkins RW, Kwok CT, Srivastava S, Wong JJ, Khachigian LM, et al. Dominantly inherited constitutional epigenetic silencing of MLH1 in a cancer-affected family is linked to a single nucleotide variant within the 5’UTR. Cancer Cell 2011, 20, 200–213. [CrossRef] [PubMed]
- Watson P, Ashwathnarayan R, Lynch HT, Roy HK. Tobacco use and increased colorectal cancer risk in patients with hereditary nonpolyposis colorectal cancer (Lynch syndrome). Arch. Intern. Med. 2004, 164, 2429–2431. [CrossRef] [PubMed]
- Pande M, Lynch PM, Hopper JL, Jenkins MA, Gallinger S, Haile RW, et al. Smoking and colorectal cancer in Lynch syndrome: results from the Colon Cancer Family Registry and the University of Texas M.D. Anderson Cancer Center. Clin. Cancer Res. 2010, 16, 1331–1339. [CrossRef]
- Dashti SG, Buchanan DD, Jayasekara H, Ait Ouakrim D, Clendenning M, Rosty C, et al. Alcohol Consumption and the Risk of Colorectal Cancer for Mismatch Repair Gene Mutation Carriers. Cancer Epidemiol Biomarkers Prev. 2017, 26, 366–375. [CrossRef]
- Botma A, Vasen HF, van Duijnhoven FJ, Kleibeuker JH, Nagengast FM, Kampman E. Dietary patterns and colorectal adenomas in Lynch syndrome: the GEOLynch cohort study. Cancer 2013, 119, 512–521. [CrossRef]
| Variable |
Cancer unaffected-controls (LS carriers) (N=13) |
Cancer affected -cases (N=78) |
Total subjects (N=91) |
*p-value |
|
| Early-diagnosis (≤40) (N=35) | Late-diagnosis (>40) (N=43) | ||||
| Gender | |||||
| Male | 4 (30.8%) | 23 (65.7%) | 19 (44.2%) | 46 | 0.057 |
| Female | 9 (69.2%) | 12 (34.3%) | 24 (55.8%) | 45 | 0.057 |
| *Age at diagnosis (years) | |||||
| Mean (SD) | 69.2 (5.44) | 31.9 (5.24) | 53.1 (8.31) | ||
| Median [Min, Max] | 68.8 [62.7, 79.1] | 32.0 [17.0- 39.4] | 50.0 [39.8-72.3] | ||
| Tumour site | |||||
| Proximal colon | NA | 24 | 21 | 45 | 0.080 |
| Distal colon | NA | 5 | 5 | 10 | 0.726 |
| Rectum | NA | 2 | 1 | 3 | 0.441 |
| Endometrium | NA | 0 | 8 | 8 | N/A |
| Breast | NA | 0 | 3 | 3 | N/A |
| Ovary | NA | 1 | 0 | 1 | N/A |
| Small intestine | NA | 2 | 3 | 5 | 0.818 |
| Bladder | NA | 1 | 0 | 1 | N/A |
| Kidney | NA | 0 | 1 | 1 | N/A |
| Skin | NA | 0 | 1 | 1 | N/A |
|
*p-value for the difference between cancer cases with early-diagnosis (≤40) and late-diagnosis (>40) calculated using a Z-score test for two proportions with a two-tailed significance level of 0.05. *Age at cancer diagnosis only for individuals with cancer and age at censoring for unaffected mutation carriers | |||||

| Variable |
Cancer unaffected-controls (LS carriers) (N=13) |
Cancer affected -cases (N=78) |
Total subjects (N=91) |
*p-value |
|
| Early-diagnosis (≤40) (N=35) | Late-diagnosis (>40) (N=43) | ||||
| Gender | |||||
| Male | 4 (30.8%) | 23 (65.7%) | 19 (44.2%) | 46 | 0.057 |
| Female | 9 (69.2%) | 12 (34.3%) | 24 (55.8%) | 45 | 0.057 |
| *Age at diagnosis (years) | |||||
| Mean (SD) | 69.2 (5.44) | 31.9 (5.24) | 53.1 (8.31) | ||
| Median [Min, Max] | 68.8 [62.7, 79.1] | 32.0 [17.0- 39.4] | 50.0 [39.8-72.3] | ||
| Tumour site | |||||
| Proximal colon | NA | 24 | 21 | 45 | 0.080 |
| Distal colon | NA | 5 | 5 | 10 | 0.726 |
| Rectum | NA | 2 | 1 | 3 | 0.441 |
| Endometrium | NA | 0 | 8 | 8 | N/A |
| Breast | NA | 0 | 3 | 3 | N/A |
| Ovary | NA | 1 | 0 | 1 | N/A |
| Small intestine | NA | 2 | 3 | 5 | 0.818 |
| Bladder | NA | 1 | 0 | 1 | N/A |
| Kidney | NA | 0 | 1 | 1 | N/A |
| Skin | NA | 0 | 1 | 1 | N/A |
| *p-value for the difference between cancer cases with early-diagnosis (≤40) and late-diagnosis (>40) calculated using a Z-score test for two proportions with a two-tailed significance level of 0.05. *Age at cancer diagnosis only for individuals with cancer and age at censoring for unaffected mutation carriers | |||||
| HLA Locus | HLA allele | Unadjusted | Gender Adjusted |
Affected: Unaffected ratio |
Mean age (range) for cancer affected | ||||
| HR (95% CI) | p-value | q-value | HR (95% CI) | P-value | q-value | ||||
| HLA-B |
B*15:17 |
94.65 (16.05- 558.11) | <0.001 | <0.001 | 87.64 (14.94-514.03) | <0.001 | <0.001 | 2:0 | 21(17, 25) |
| HLA-DRB3 |
DRB3*03:XXX |
247.15 (14.16-4314.89) | <0.001 | <0.001 | 301.04 (17.19-5270.18) | <0.001 | <0.001 | 1:0 | 17 |
| HLA-DPB1 |
DPB1*04:02 |
2.92 (1.22-7.02) | 0.011 | 0.045 | 3.56 (1.46-8.65) | 0.005 | 0.029 | 6:0 | 37 (25-50) |
|
DPB1*20:01 |
20.67 (2.60- 164.55) | 0.004 | 0.040 | 16.88 (2.11-134.75) | 0.006 | 0.006 | 1:0 | 26 | |
|
DPB1*296:01 |
12.97 (1.68-100.14) | 0.014 | 0.079 | 18.05 (2.31-141.30) | 0.006 | 0.029 | 1:0 | 30 | |
|
DPB1*55:01 |
74.13 (1.68-100.14) | <0.001 | <0.001 | 64.22 (12.26-366.25) | <0.001 | <0.001 | 2:0 | 21 (17,25) | |
| HLA allele | Gender Adjusted-HR | P-value | q-value | 95% CI | Affected: Unaffected ratio | Mean age (range) for cancer affected |
|
B*15:17 |
71.59 | <0.001 | <0.001 | 11.68-438.65 | 2:0 | 21(17, 25) |
| DRB3*03:XXX | 264.83 | <0.001 | 0.001 | 14.96- 4689.40 | 1:0 | 17 |
|
DPB1*55:01 |
54.05 | <0.001 | <0.001 | 11.16-319.47 | 2:0 | 21 (17,25) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




