Submitted:
05 March 2024
Posted:
05 March 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results and Discussion
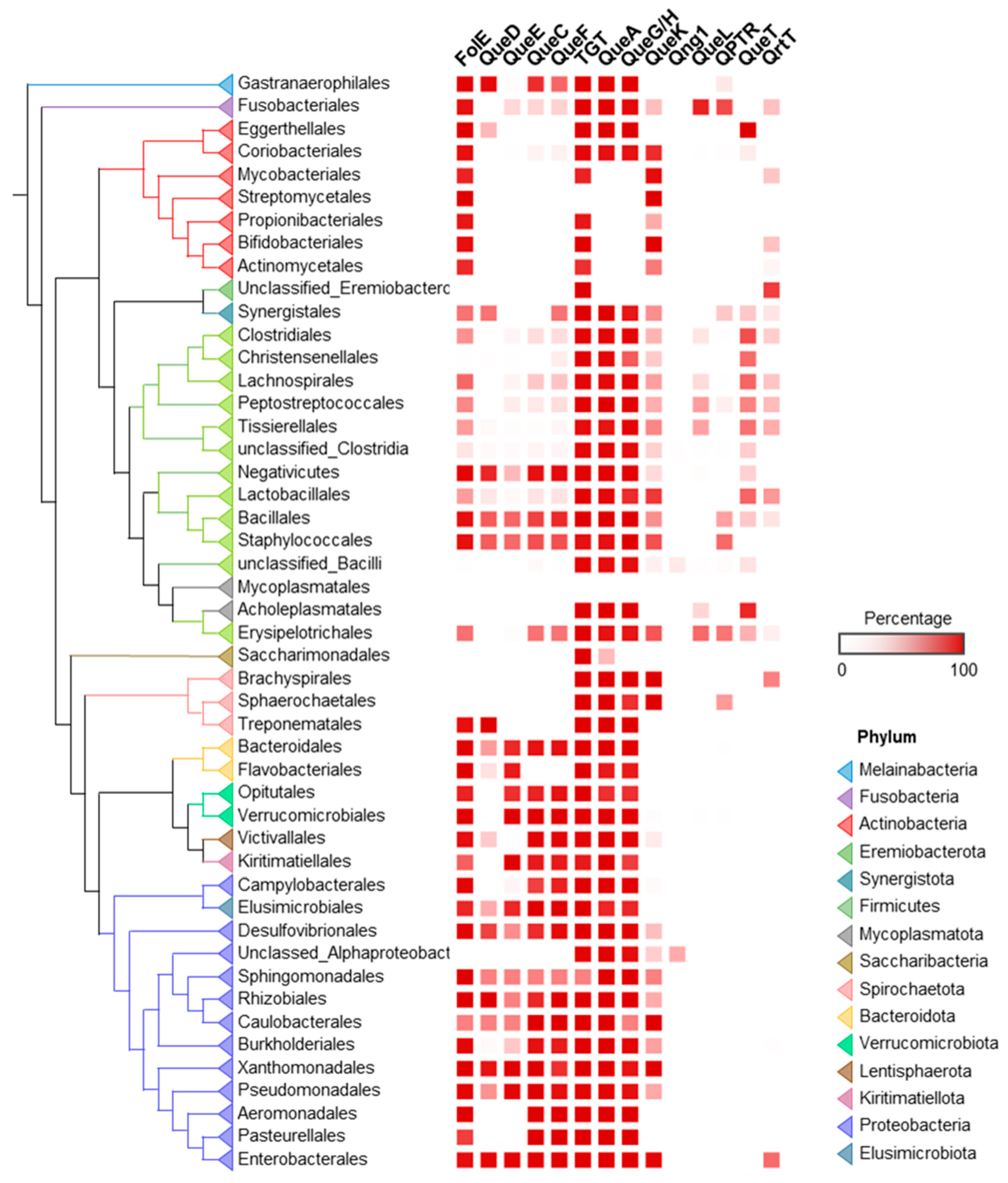
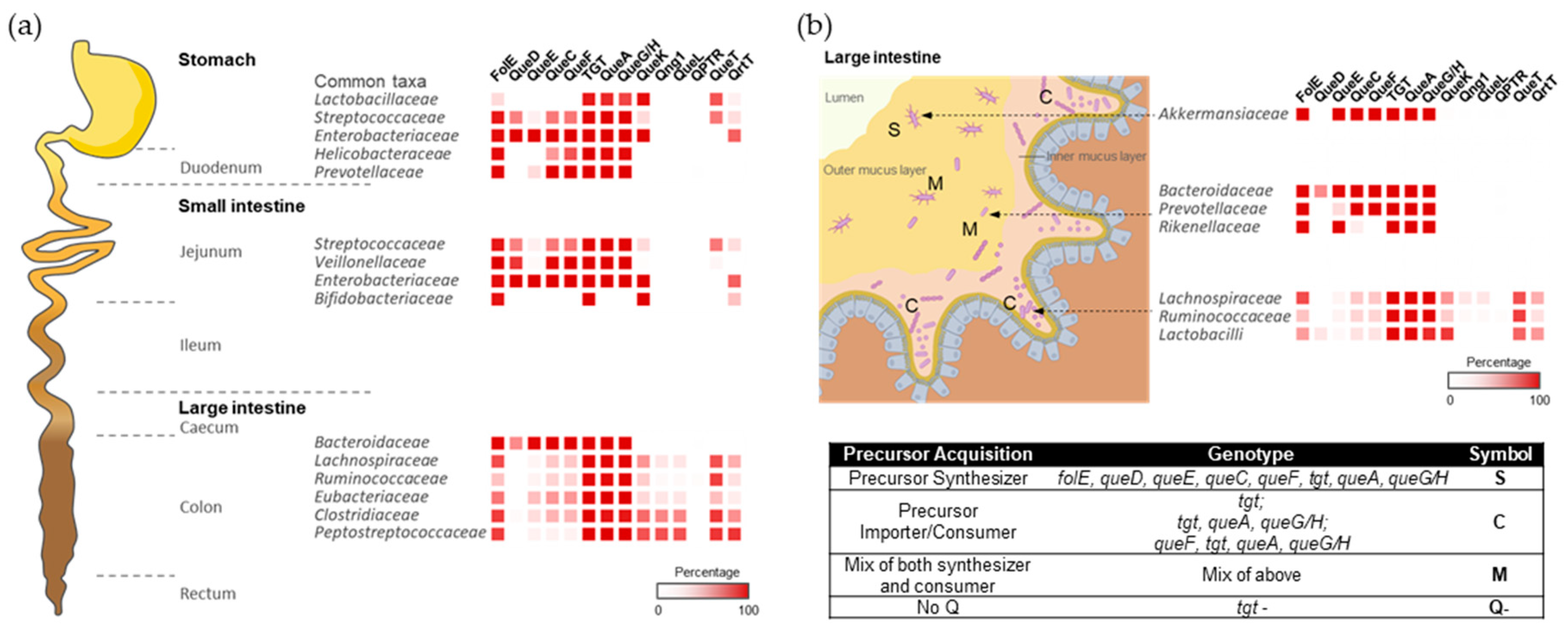
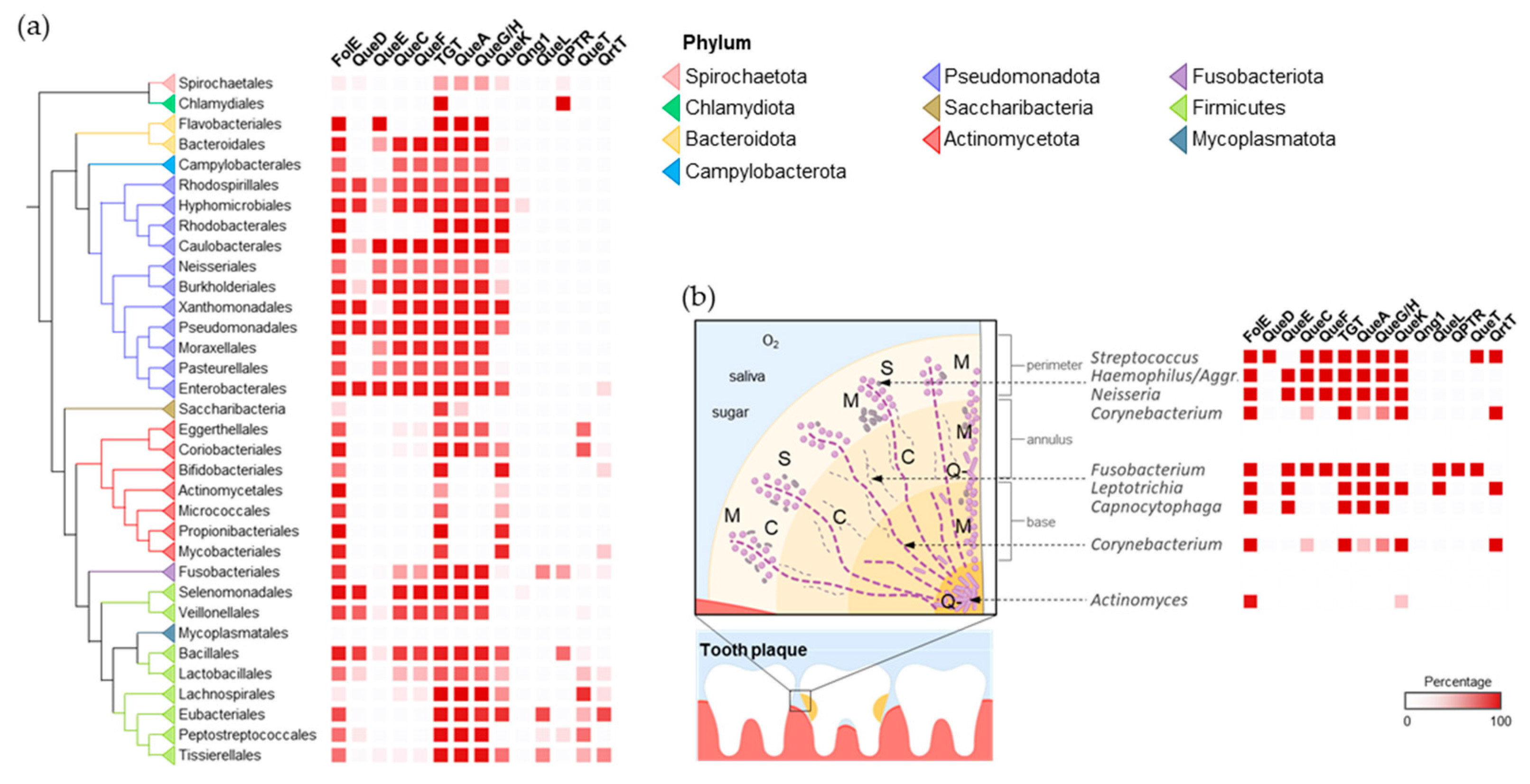
2.1. Q Makers and Users Are Spatially Distributed in the Human Gut and Oral Microbiomes
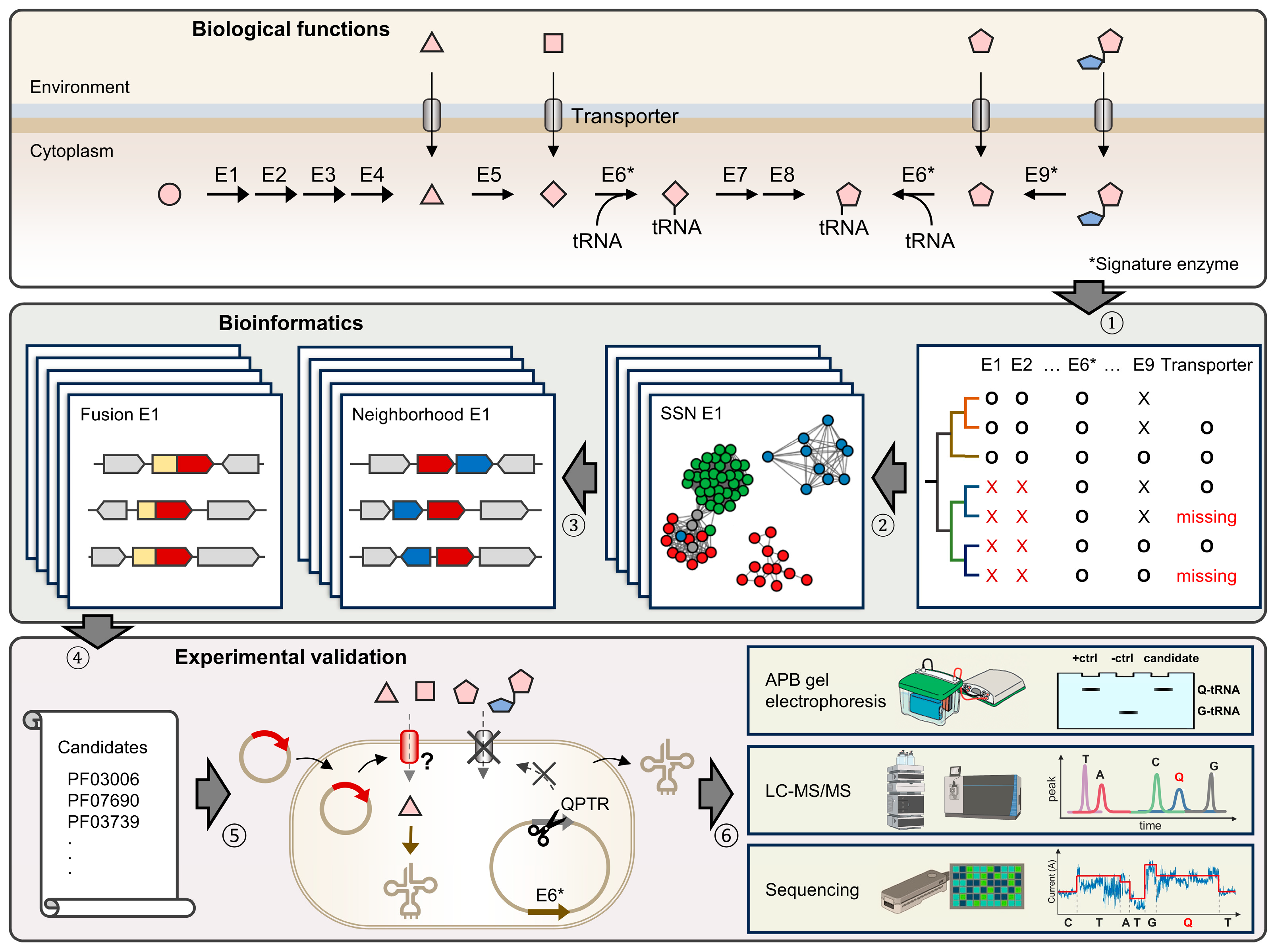
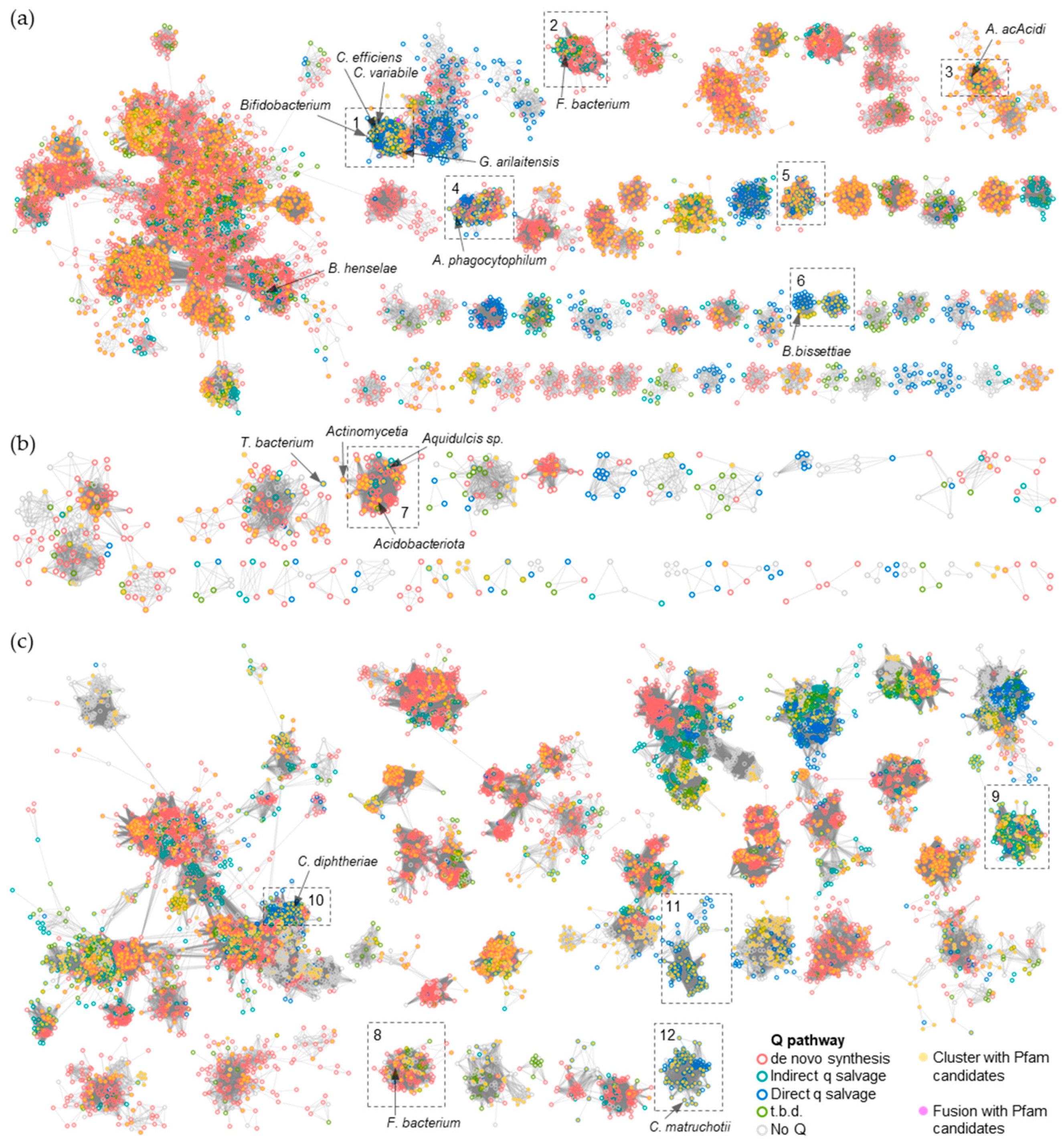
2.2. Identification of Q Precursor Transporter Candidates through Comparative Genomics
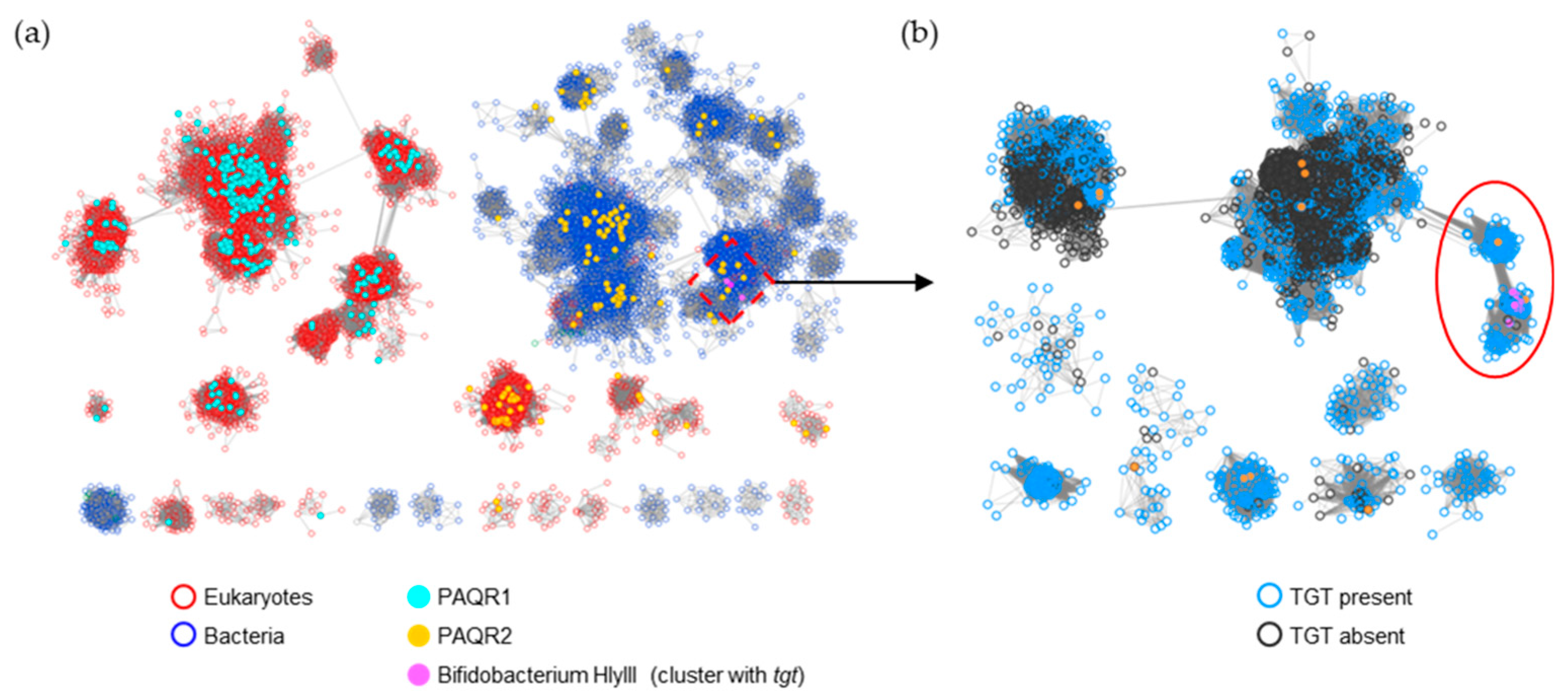
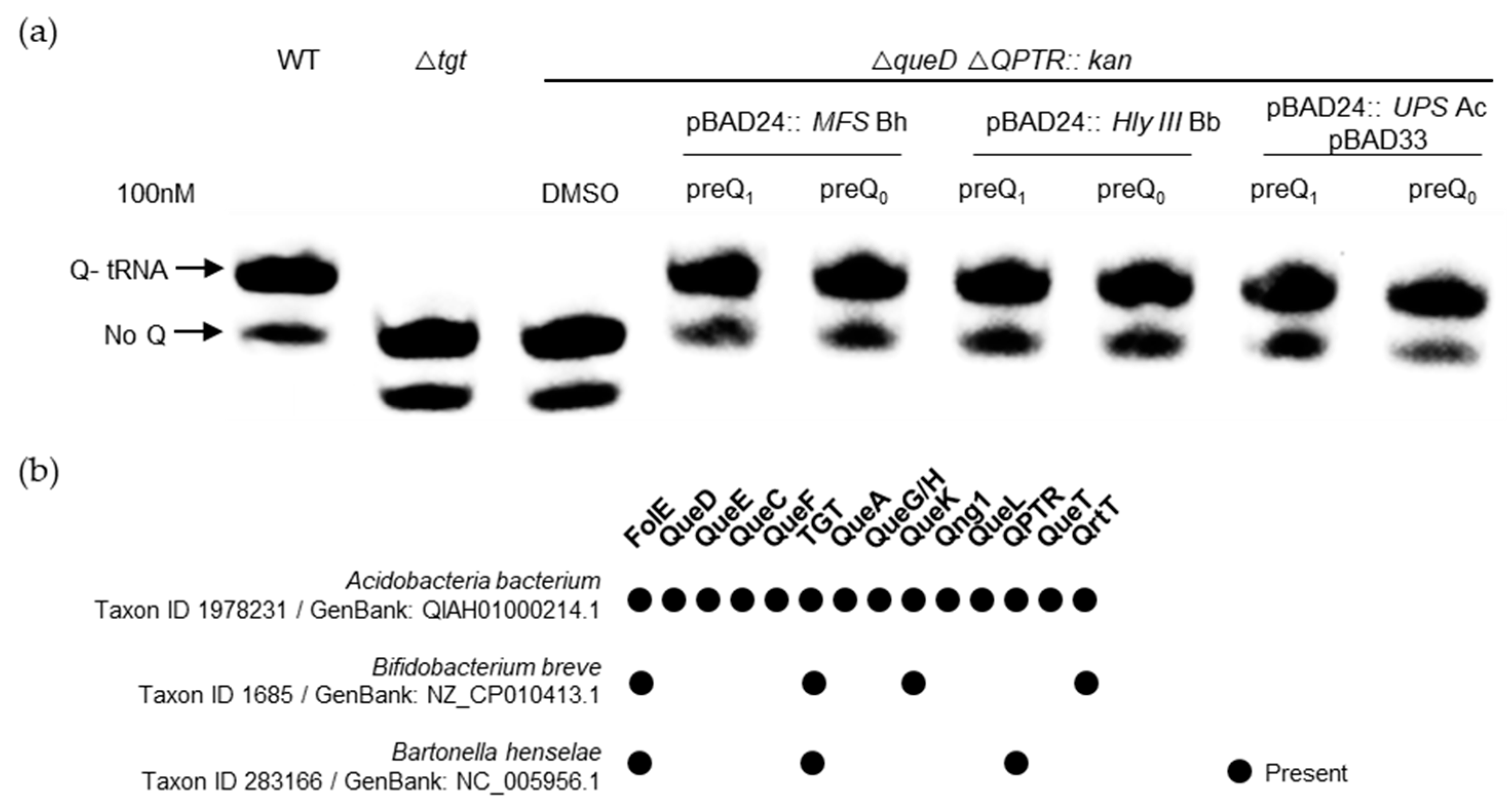
2.3. Members of Three Transporters Superfamilies Can Evolve to Transport PreQ0 and PreQ1
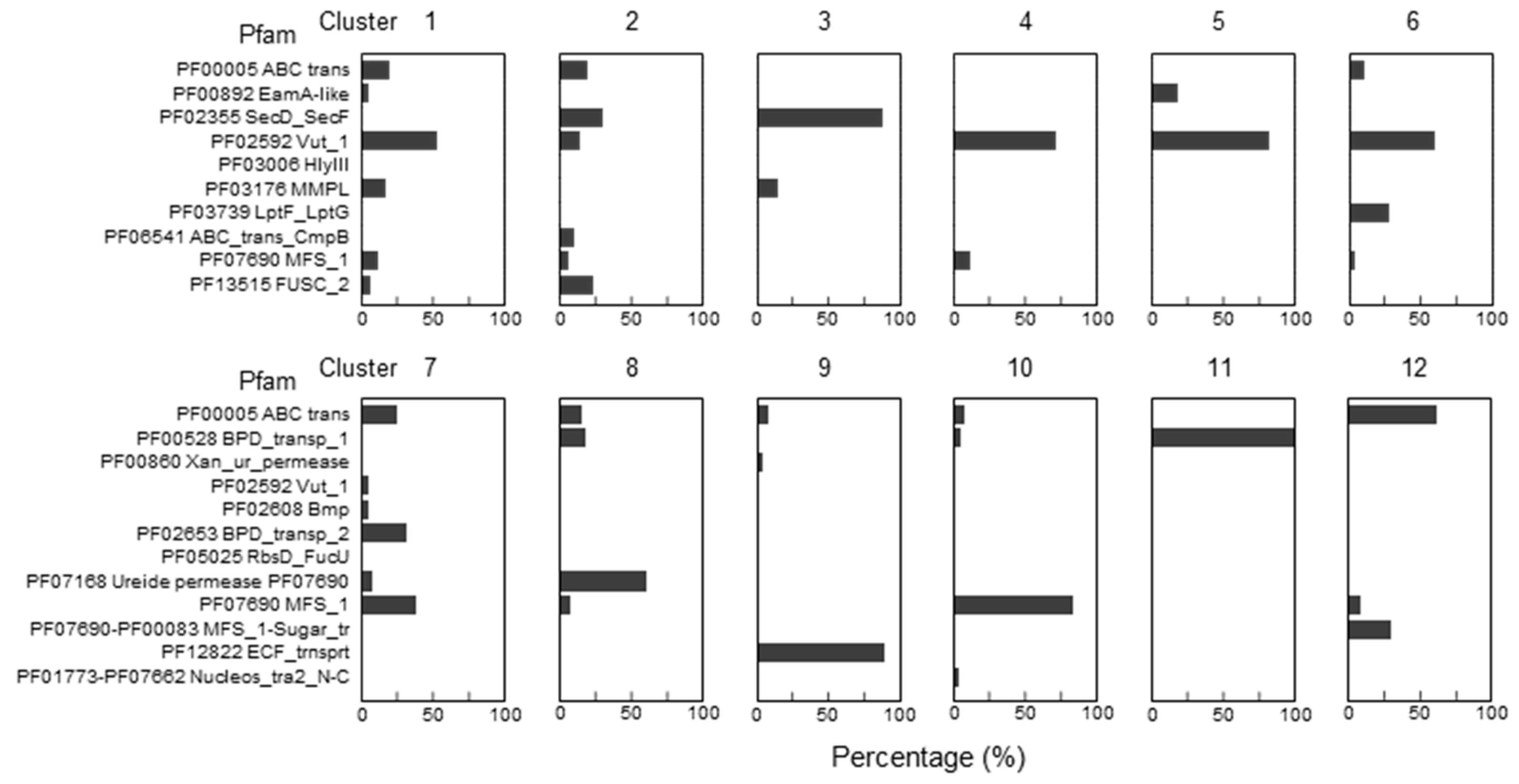
2.4. Tentative Identification of PreQ1 Transporters Subgroups in Three Transporters Superfamilies
3. Conclusions
4. Materials and Methods
4.1. Human Gut Microbiome Analysis
4.2. Human Oral Microbiome Analysis
4.3. Comparative Genomics and Sequence Similarity Networks (SSNs)
4.4. Strains, Media, and Growth Conditions
4.3. Construction of E. coli Strains and Plasmids
4.4. Q detection Assay
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- de Crécy-Lagard V, Hutinet G, Cediel-Becerra JD, Yuan Y, Zallot R, Chevrette MG, et al. Biosynthesis and function of 7-deazaguanine derivatives in bacteria and phages. Microbiol Mol Biol Rev. 2024, 0, e00199-23. [CrossRef]
- Yuan Y, Zallot R, Grove TL, Payan DJ, Martin-Verstraete I, Šepić S, et al. Discovery of novel bacterial queuine salvage enzymes and pathways in human pathogens. Proc Natl Acad Sci U S A. 2019, 116, 19126–19135. [Google Scholar] [CrossRef]
- Hung SH, Elliott GI, Ramkumar TR, Burtnyak L, McGrenaghan CJ, Alkuzweny S, et al. Structural basis of Qng1-mediated salvage of the micronutrient queuine from queuosine-5′-monophosphate as the biological substrate. Nucleic Acids Res. 2023, 51, 935–951. [Google Scholar] [CrossRef]
- Zallot R, Yuan Y, de Crécy-Lagard V. The Escherichia coli COG1738 member YhhQ is involved in 7-cyanodeazaguanine (preQ₀) Transport. Biomolecules. 2017, 7, 12. [Google Scholar]
- Ames, BN. Prolonging healthy aging: Longevity vitamins and proteins. Proc Natl Acad Sci U S A. 2018, 115, 10836. [Google Scholar] [CrossRef]
- Varghese S, Cotter M, Chevot F, Fergus C, Cunningham C, Mills KH, et al. In vivo modification of tRNA with an artificial nucleobase leads to full disease remission in an animal model of multiple sclerosis. Nucleic Acids Res. 2017, 45, 2029–2039.
- Cirzi C, Dyckow J, Legrand C, Schott J, Guo W, Perez Hernandez D, et al. Queuosine-tRNA promotes sex-dependent learning and memory formation by maintaining codon-biased translation elongation speed. EMBO J. 2023, 42, 112507. [Google Scholar] [CrossRef]
- Skolnick SD, Greig NH. Microbes and monoamines: potential neuropsychiatric consequences of dysbiosis. Trends Neurosci. 2019, 42, 151–163. [Google Scholar] [CrossRef]
- Rashad S, Al-Mesitef S, Mousa A, Zhou Y, Ando D, Sun G, et al. Translational response to mitochondrial stresses is orchestrated by tRNA modifications. bioRxiv. 2024 Jan 1;2024.02.14.580389.
- Díaz-Rullo J, González-Pastor JE. tRNA queuosine modification is involved in biofilm formation and virulence in bacteria. Nucleic Acids Res. 2023, 51, 9821–9837. [Google Scholar] [CrossRef] [PubMed]
- Magnúsdóttir S, Ravcheev D, de Crécy-Lagard V, Thiele I. Systematic genome assessment of B-vitamin biosynthesis suggests co-operation among gut microbes. Front Genet. 2015, 6, 148. [Google Scholar]
- Rodionov DA, Arzamasov AA, Khoroshkin MS, Iablokov SN, Leyn SA, Peterson SN, et al. Micronutrient requirements and sharing capabilities of the human gut microbiome. Front Microbiol. 2019, 10, 1316. [Google Scholar]
- Yan F, Xiang S, Shi L, Zhu X. Synthesis of queuine by colonic gut microbiome via cross-feeding. Food Front. 2023, 1–14.
- Kesh K, Mendez R, Mateo-Victoriano B, Garrido VT, Durden B, Gupta VK, et al. Obesity enriches for tumor protective microbial metabolites and treatment refractory cells to confer therapy resistance in PDAC. Gut Microbes. 2022, 14, 2096328. [Google Scholar] [CrossRef]
- Varriale L, Coretti L, Dipineto L, Green BD, Pace A, Lembo F, et al. An outdoor access period improves chicken cecal microbiota and potentially increases micronutrient biosynthesis. Front Vet Sci. 2022, 9, 904522. [Google Scholar] [CrossRef]
- Mark Welch JL, Rossetti BJ, Rieken CW, Dewhirst FE, Borisy GG. Biogeography of a human oral microbiome at the micron scale. Proc Natl Acad Sci U S A. 2016, 113, E791–800. [Google Scholar]
- McCallum G, Tropini C. The gut microbiota and its biogeography. Nat Rev Microbiol. 2024, 22, 105–118. [Google Scholar] [CrossRef] [PubMed]
- Mondragón-Palomino O, Poceviciute R, Lignell A, Griffiths JA, Takko H, Ismagilov RF. Three-dimensional imaging for the quantification of spatial patterns in microbiota of the intestinal mucosa. Proc Natl Acad Sci U S A 2022, 119, e2118483119. [Google Scholar] [CrossRef]
- Derrien M, Van Baarlen P, Hooiveld G, Norin E, Müller M, de Vos WM. Modulation of mucosal immune response, tolerance, and proliferation in mice colonized by the mucin-degrader Akkermansia muciniphila. Front Microbiol. 2011, 2, 166. [Google Scholar]
- Yasuda K, Oh K, Ren B, Tickle TL, Franzosa EA, Wachtman LM, et al. Biogeography of the intestinal mucosal and lumenal microbiome in the Rhesus macaque. Cell Host Microbe. 2015, 17, 385–391. [Google Scholar] [CrossRef] [PubMed]
- Bowen WH, Burne RA, Wu H, Koo H. Oral biofilms: pathogens, matrix, and polymicrobial interactions in microenvironments. Trends Microbiol. 2018, 26, 229–242. [Google Scholar] [CrossRef] [PubMed]
- Sangha JS, Barrett P, Curtis TP, Métris A, Jakubovics NS, Ofiteru ID. Effects of glucose and lactate on Streptococcus mutans abundance in a novel multispecies oral biofilm model. Microbiol Spectr. 2024, 0, e03713–23. [Google Scholar]
- Pereira FC, Berry D. Microbial nutrient niches in the gut. Environ Microbiol. 2017, 19, 1366–1378. [Google Scholar] [CrossRef] [PubMed]
- Gorkiewicz G, Moschen A. Gut microbiome: a new player in gastrointestinal disease. Virchows Archiv. 2018, 472, 159–172. [Google Scholar] [CrossRef] [PubMed]
- Clarke G, Sandhu K V. , Griffin BT, Dinan TG, Cryan JF, Hyland NP. Gut reactions: breaking down xenobiotic–microbiome interactions. Pharmacol Rev. 2019, 71, 198–224. [Google Scholar] [CrossRef] [PubMed]
- Henry CS, Lerma-Ortiz C, Gerdes SY, Mullen JD, Colasanti R, Zhukov A, et al. Systematic identification and analysis of frequent gene fusion events in metabolic pathways. BMC Genomics. 2016, 17, 473. [Google Scholar]
- Zallot R, Oberg N, Gerlt JA. The EFI web resource for genomic enzymology tools: leveraging protein, genome, and metagenome databases to discover novel enzymes and metabolic pathways. Biochemistry. 2019, 58, 4169–4182. [Google Scholar] [CrossRef]
- Quaiyum S, Yuan Y, Sun. Giuanxin, Ratnayakec RMM, Hutinet G, Dedon PC, et al. Queuosine salvage in Bartonella henselae Houston 1: a unique evolutionary path". BioRxiv. 2023;2023.12.05.570228.
- Pei J, Millay DP, Olson EN, Grishin N V. CREST - a large and diverse superfamily of putative transmembrane hydrolases. Biol Direct. 2011, 6, 37. [Google Scholar]
- Desimone M, Catoni E, Ludewig U, Hilpert M, Schneider A, Kunze R, et al. A novel superfamily of transporters for allantoin and other oxo derivatives of nitrogen heterocyclic compounds in Arabidopsis. Plant Cell. 2002, 14, 847–856. [Google Scholar] [CrossRef]
- Schmidt A, Baumann N, Schwarzkopf A, Frommer WB, Desimone M. Comparative studies on ureide permeases in Arabidopsis thaliana and analysis of two alternative splice variants of AtUPS5. Planta. 2006, 224, 1329–1340. [Google Scholar] [CrossRef]
- Schmidt A, Su YH, Kunze R, Warner S, Hewitt M, Slocum RD, et al. UPS1 and UPS2 from Arabidopsis mediate high affinity transport of uracil and 5-fluorouracil. J Biol Chem. 2004, 279, 44817–44824. [Google Scholar] [CrossRef]
- Sauve S, Williamson J, Polasa A, Moradi M. Ins and outs of rocker switch mechanism in major facilitator superfamily of transporters. Membranes (Basel). 2023, 13, 462. [Google Scholar] [CrossRef]
- Quistgaard EM, Löw C, Guettou F, Nordlund P. Understanding transport by the major facilitator superfamily (MFS): structures pave the way. Nat Rev Mol Cell Biol. 2016, 17, 123–132. [Google Scholar] [CrossRef]
- Donia MS, Fischbach MA. Small molecules from the human microbiota. Science (1979). 2015, 349, 1254766.
- Krypotou E, Evangelidis T, Bobonis J, Pittis AA, Gabaldón T, Scazzocchio C, et al. Origin, diversification and substrate specificity in the family of NCS1/FUR transporters. Mol Microbiol. 2015, 96, 927–950. [Google Scholar] [CrossRef]
- Jørgensen ME, Xu D, Crocoll C, Ernst HA, Ramírez D, Motawia MS, et al. Origin and evolution of transporter substrate specificity within the NPF family. Elife 2017, 6, e19466. [CrossRef]
- Kourkoulou A, Pittis AA, Diallinas G. Evolution of substrate specificity in the Nucleobase-Ascorbate Transporter (NAT) protein family. Microb Cell. 2018, 5, 280–292. [Google Scholar] [CrossRef] [PubMed]
- Gournas C, Athanasopoulos A, Sophianopoulou V. On the evolution of specificity in members of the yeast amino acid transporter family as parts of specific metabolic pathways. Int J Mol Sci. 2018, 19, 1398. [Google Scholar] [CrossRef] [PubMed]
- Teichmann L, Chen C, Hoffmann T, Smits SHJ, Schmitt L, Bremer E. From substrate specificity to promiscuity: hybrid ABC transporters for osmoprotectants. Mol Microbiol. 2017, 104, 761–780. [Google Scholar] [CrossRef]
- Saier MH, Reddy VS, Moreno-Hagelsieb G, Hendargo KJ, Zhang Y, Iddamsetty V, et al. The Transporter Classification Database (TCDB): 2021 update. Nucleic Acids Res. 2021, 49, D461–7. [Google Scholar] [CrossRef] [PubMed]
- Cerna-Vargas JP, Sánchez-Romera B, Matilla MA, Ortega Á, Krell T. Sensing preferences for prokaryotic solute binding protein families. Microb Biotechnol. 2023, 16, 1823–1833. [Google Scholar] [CrossRef] [PubMed]
- Stanchev LD, Møller-Hansen I, Lojko P, Rocha C, Borodina I. Screening of Saccharomyces cerevisiae metabolite transporters by 13C isotope substrate labeling. Front Microbiol. 2023, 14, 1286597. [Google Scholar]
- Pochini L, Galluccio M. Heterologous (Over) Expression of human SoLute Carrier (SLC) in yeast: a well-recognized tool for human transporter function/structure studies. Life. 2022, 12, 1206. [Google Scholar] [CrossRef] [PubMed]
- Fernández, Rico-Jiménez, Ortega, Daddaoua, García García, Martín-Mora, et al. Determination of ligand profiles for Pseudomonas aeruginosa Solute Binding Proteins. Int J Mol Sci. 2019, 20, 5156. [Google Scholar] [CrossRef] [PubMed]
- Elbourne LDH, Tetu SG, Hassan KA, Paulsen IT. TransportDB 2.0: a database for exploring membrane transporters in sequenced genomes from all domains of life. Nucleic Acids Res. 2017, 45, D320–4. [Google Scholar] [CrossRef] [PubMed]
- Poyet M, Groussin M, Gibbons SM, Avila-Pacheco J, Jiang X, Kearney SM, et al. A library of human gut bacterial isolates paired with longitudinal multiomics data enables mechanistic microbiome research. Nat Med. 2019, 25, 1442–1452. [Google Scholar] [CrossRef] [PubMed]
- Groussin M, Poyet M, Sistiaga A, Kearney SM, Moniz K, Noel M, et al. Elevated rates of horizontal gene transfer in the industrialized human microbiome. Cell. 2021, 184, 2053–2067. [Google Scholar] [CrossRef]
- Almeida A, Nayfach S, Boland M, Strozzi F, Beracochea M, Shi ZJ, et al. A unified catalog of 204,938 reference genomes from the human gut microbiome. Nat Biotechnol. 2021, 39, 105–114. [Google Scholar] [CrossRef]
- Buchfink B, Xie C, Huson DH. Fast and sensitive protein alignment using DIAMOND. Nat Methods. 2015, 12, 59–60. [Google Scholar] [CrossRef]
- Edgar, RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Criscuolo A, Gribaldo S. BMGE (Block Mapping and Gathering with Entropy): a new software for selection of phylogenetic informative regions from multiple sequence alignments. BMC Evol Biol 2010, 10, 210. [Google Scholar]
- Gouy M, Guindon S, Gascuel O. SeaView Version 4: A multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 2010, 27, 221–224. [Google Scholar] [CrossRef] [PubMed]
- Price MN, Dehal PS, Arkin AP. FastTree 2 – Approximately maximum-likelihood trees for large alignments. PLoS One. 2010, 5, e9490. [Google Scholar]
- Letunic I, Bork P. Interactive tree of life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef]
- Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Marchler-Bauer A, Derbyshire MK, Gonzales NR, Lu S, Chitsaz F, Geer LY, et al. CDD: NCBI’s conserved domain database. Nucleic Acids Res. 2015, 43, D222–6. [Google Scholar] [CrossRef] [PubMed]
- Bateman A, Martin MJ, Orchard S, Magrane M, Ahmad S, Alpi E, et al. UniProt: the Universal Protein Knowledgebase in 2023. Nucleic Acids Res. 2023, 51, D523–31. [Google Scholar] [CrossRef]
- Paysan-Lafosse T, Blum M, Chuguransky S, Grego T, Pinto BL, Salazar GA, et al. InterPro in 2022. Nucleic Acids Res. 2023, 51, D418–27. [Google Scholar] [CrossRef]
- Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Harrison KJ, de Crécy-Lagard V, Zallot R. Gene Graphics: A genomic neighborhood data visualization web application. Bioinformatics. 2018, 34, 1406–1408. [Google Scholar] [CrossRef]
- Guzman LM, Belin D, Carson MJ, Beckwith J. Tight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J Bacteriol. 1995, 177, 4121–4130. [Google Scholar] [CrossRef] [PubMed]
- Green R, Rogers EJ. Transformation of chemically competent E. coli. In Methods in Enzymology; Academic Press Inc., 2013; pp. 329–336.

| Pfam | Description | Experimentally tested candidates (species and accession number) |
| PF07690 | Major Facilitator Superfamily (MFS_1) | Bartonella henselae (A0A0H3LX18) (Bh_MFS) |
| PF03176 | MMPL family (MMPL) | Corynebacterium propoinquum (0A2W5NLC3) |
| PF03006 | Haemolysin-III related (Hly_III) | Brevibacterium breve (A0A0M3T8W5) (Bb_HlyII) |
| PF13515 | Fusaric acid resistance protein-like (FUSC_2) |
Not tested |
| PF00892 | EamA-like transporter family (EanA) | Not tested |
| PF02355 | Protein export membrane protein (SecD/SecF) |
Not tested |
| PF03739 | Lipopolysaccharide export system permease (LptF/LptG) |
Winogradskyella sp. (A0A024FC69) Chryseobacterium piperi (A0A086BN18) Bacteroides (Phocaeicola) dorei (A0A076J562) |
| PF06541 | Putative ABC-transporter type IV (ABC_trans_CmpB) | Not tested |
| PF07168 | Ureide permease (UPS) | Acidobacteriota bacterium (A0A2V9U0M9) (Ac_UPS) |
| PF00005 | ABC transporter (ABC_tran) | Not tested |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




