Submitted:
06 February 2024
Posted:
06 February 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results and Discussion
3. Materials and Methods
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Boison, D.; Yegutkin, G.G. Adenosine Metabolism: Emerging Concepts for Cancer Therapy. Cancer Cell 2019, 36, 582–596. [Google Scholar] [CrossRef]
- Dunwiddie, T.V.; Masino, S.A. The Role and Regulation of Adenosine in the Central Nervous System. Annu. Rev. Neurosci. 2001, 24, 31–55. [Google Scholar] [CrossRef] [PubMed]
- Fredholm, BB Adenosine, an Endogenous Distress Signal, Modulates Tissue Damage and Repair. Cell Death Differ 2007, 14, 1315–1323. [CrossRef] [PubMed]
- Garcia-Gil, M.; Camici, M.; Allegrini, S.; Pesi, R.; Tozzi, M.G. Metabolic Aspects of Adenosine Functions in the Brain. Front. Pharmacol. 2021, 12, 672182. [Google Scholar] [CrossRef] [PubMed]
- Guieu, R.; Deharo, J.-C.; Maille, B.; Crotti, L.; Torresani, E.; Brignole, M.; Parati, G. Adenosine and the Cardiovascular System: The Good and the Bad. JCM 2020, 9, 1366. [Google Scholar] [CrossRef] [PubMed]
- Haskó, G.; Antonioli, L.; Cronstein, B.N. Adenosine Metabolism, Immunity and Joint Health. Biochemical Pharmacology 2018, 151, 307–313. [Google Scholar] [CrossRef] [PubMed]
- Ohta, A. A Metabolic Immune Checkpoint: Adenosine in Tumor Microenvironment. Front. Immunol. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Yegutkin, G.G.; Boison, D. ATP and Adenosine Metabolism in Cancer: Exploitation for Therapeutic Gain. Pharmacol Rev 2022, 74, 799–824. [Google Scholar] [CrossRef] [PubMed]
- Headrick, J.P.; Ashton, K.J.; Rose’Meyer, R.B.; Peart, J.N. Cardiovascular Adenosine Receptors: Expression, Actions and Interactions. Pharmacology & Therapeutics 2013, 140, 92–111. [Google Scholar] [CrossRef]
- Gaudry, M.; Vairo, D.; Marlinge, M.; Gaubert, M.; Guiol, C.; Mottola, G.; Gariboldi, V.; Deharo, P.; Sadrin, S.; Maixent, J.M.; et al. Adenosine and Its Receptors: An Expected Tool for the Diagnosis and Treatment of Coronary Artery and Ischemic Heart Diseases. IJMS 2020, 21, 5321. [Google Scholar] [CrossRef]
- Guieu, R.; Deharo, J.-C.; Ruf, J.; Mottola, G.; Kipson, N.; Bruzzese, L.; Gerolami, V.; Franceschi, F.; Ungar, A.; Tomaino, M.; et al. Adenosine and Clinical Forms of Neurally-Mediated Syncope. Journal of the American College of Cardiology 2015, 66, 204–205. [Google Scholar] [CrossRef]
- Altstidl, J.M.; Gaede, L.; Troebs, M.; Marwan, M.; Achenbach, S. Side Effects and Major Adverse Cardiac Events Caused by Fractional Flow Reserve Measurement: A Systematic Review and Meta-Analysis of 12,215 Patients. European Heart Journal 2022, 43, ehac5442022. [Google Scholar] [CrossRef]
- Borea, P.A.; Gessi, S.; Merighi, S.; Vincenzi, F.; Varani, K. Pathological Overproduction: The Bad Side of Adenosine. British J Pharmacology 2017, 174, 1945–1960. [Google Scholar] [CrossRef] [PubMed]
- Handbook of Advanced Chromatography—Mass Spectrometry Techniques; Holčapek, M. , Byrdwell, W.C., Eds.; Elsevier, Academic Press, AOCS Press: London, 2017; ISBN 978-0-12-811732-3. [Google Scholar]
- Virgiliou, C.; Fragakis, N.; Sotiriadou, M.; Vassilikos, V.; Gerou, S.; Theodoridis, G.; Gika, H. HILIC-MS/MS Analysis of Adenosine in Patient Blood. Separations 2021, 8, 222. [Google Scholar] [CrossRef]
- González-Domínguez, R.; Jáuregui, O.; Queipo-Ortuño, M.I.; Andrés-Lacueva, C. Characterization of the Human Exposome by a Comprehensive and Quantitative Large-Scale Multianalyte Metabolomics Platform. Anal. Chem. 2020, 92, 13767–13775. [Google Scholar] [CrossRef] [PubMed]
- Capogrossi, M.C. ; Holdiness, MR; Israili, Z. H. Determination of Adenosine in Normal Human Plasma and Serum by High-Performance Liquid Chromatography. Journal of Chromatography B: Biomedical Sciences and Applications 1982, 227, 168–173. [Google Scholar] [CrossRef]
- Ontyd, J.; Schrader, J. Measurement of Adenosine, Inosine, and Hypoxanthine in Human Plasma. Journal of Chromatography B: Biomedical Sciences and Applications 1984, 307, 404–409. [Google Scholar] [CrossRef] [PubMed]
- Thane Eells, J.; Spector, R. Purine and Pyrimidine Base and Nucleoside Concentrations in Human Cerebrospinal Fluid and Plasma. Neurochem Res 1983, 8, 1451–1457. [Google Scholar] [CrossRef] [PubMed]
- Kaufmann, I.; Hoelzl, A.; Schliephake, F.; Hummel, T.; Chouker, A.; Lysenko, L.; Peter, K.; Thiel, M. EFFECTS OF ADENOSINE ON FUNCTIONS OF POLYMORPHONUCLEAR LEUKOCYTES FROM PATIENTS WITH SEPTIC SHOCK. Shock 2007, 27, 25–31. [Google Scholar] [CrossRef] [PubMed]
- Tan, A.; Boudreau, N.; Lévesque, A. Internal Standards for Quantitative LC-MS Bioanalysis. In LC-MS in Drug Bioanalysis; Xu, Q.A., Madden, TL, Eds.; Springer US: Boston, MA, 2012; pp. 1–32. ISBN 978-1-4614-3828-1. [Google Scholar]
- Bergeron, A.; Furtado, M.; Garofolo, F. Importance of Using Highly Pure Internal Standards for Successful Liquid Chromatography/Tandem Mass Spectrometric Bioanalytical Assays. Rapid Commun Mass Spectrom 2009, 23, 1287–1297. [Google Scholar] [CrossRef]
- Traut, T.W. Physiological Concentrations of Purines and Pyrimidines. Mol Cell Biochem 1994, 140, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Human Metabolome Database: Showing Metabocard for Adenosine (HMDB0000050) Available online:. Available online: https://hmdb.ca/metabolites/HMDB0000050 (accessed on 10 January 2024).
- Human Metabolome Database: Showing Metabocard for Inosine (HMDB0000195) Available online:. Available online: https://hmdb.ca/metabolites/HMDB0000195 (accessed on 10 January 2024).
- Human Metabolome Database: Showing Metabocard for Hypoxanthine (HMDB0000157) Available online:. Available online: https://hmdb.ca/metabolites/HMDB0000157 (accessed on 10 January 2024).
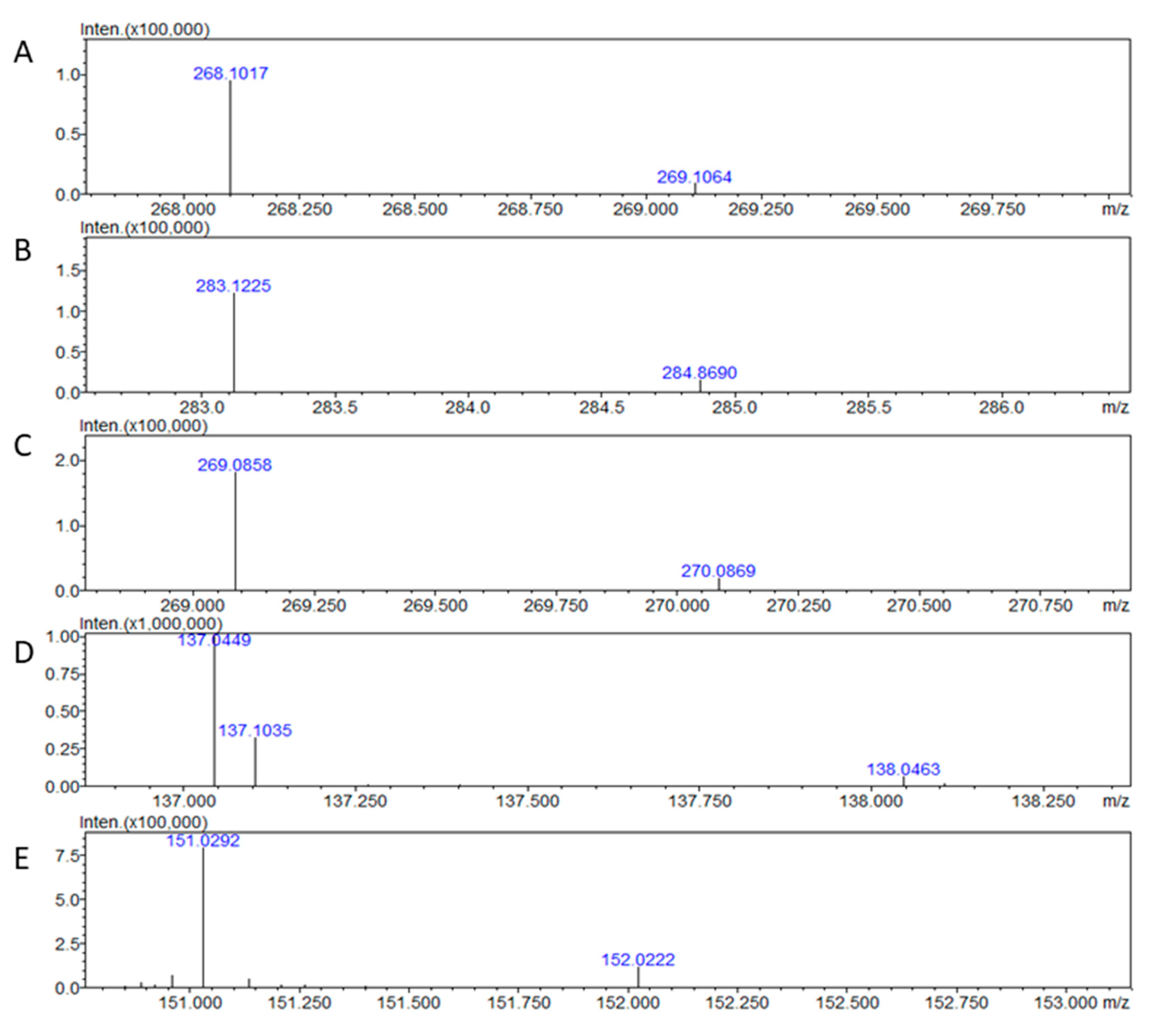
| Compound | m/z | AUP | Precision |
|---|---|---|---|
| 106 a.u. | % | ||
| 12C10,14N5-Adenosine | 268.1016 ±0.0004 | 31.3 ±5.4 | 17.4 |
| 13C10,15N5-Adenosine | 283.1212 ±0.0008 | 30.5 ±5.5 | 18.0 |
| Ratio | - | 1.10 ±0.01 | 0.6 |
| Column | Retention time | Concentration range |
Linearity (R2) | LOQ | Accuracy | |
|---|---|---|---|---|---|---|
| min | nM | w/o IS | w IS1 | nM | % | |
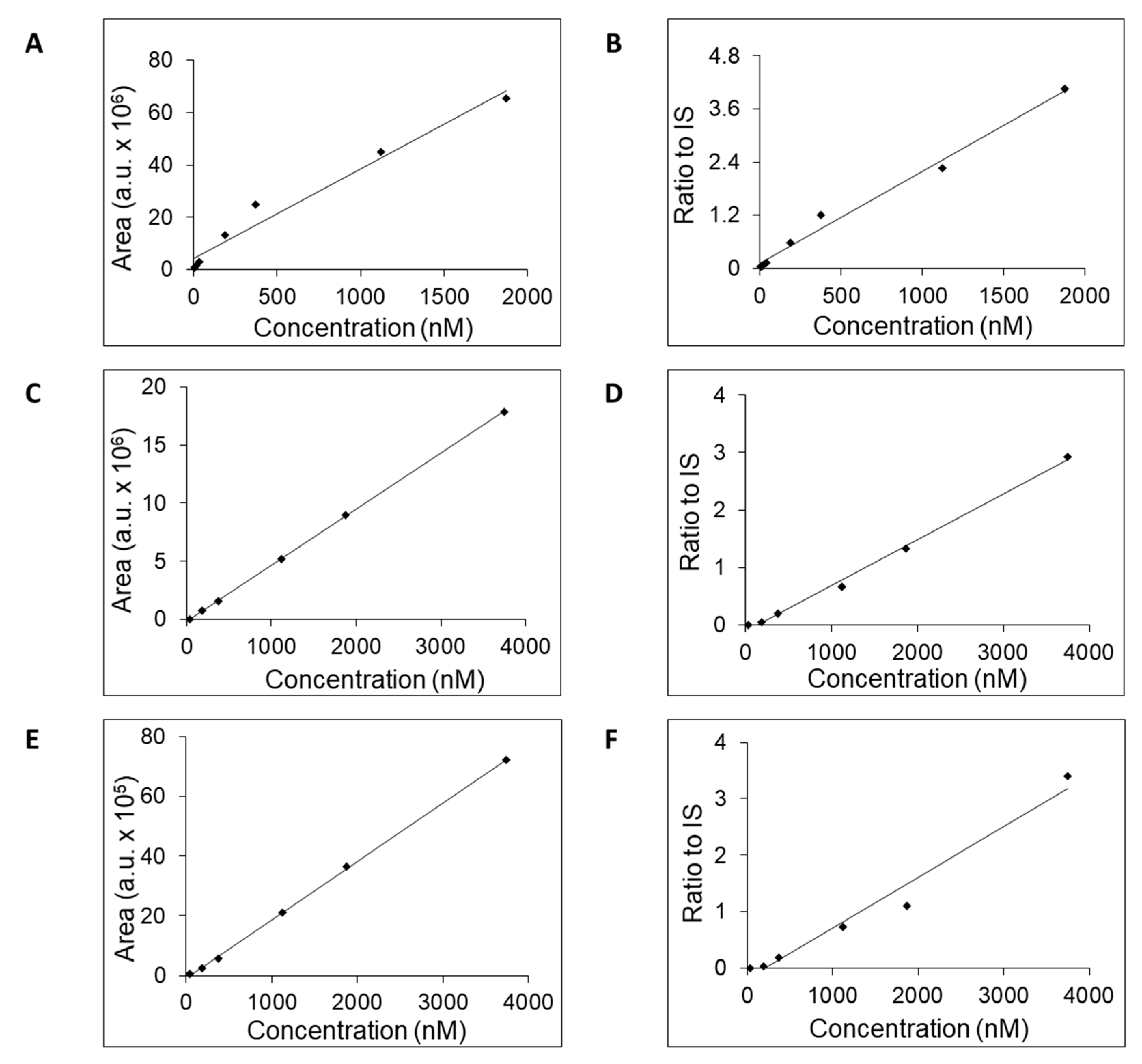
| SeQuant® ZIC®-cHILIC | 5.5 | 3.75–1872 | 0.9702 | 0.9882 | 3.75 | 103.9 |
| SeQuant® ZIC®-pHILIC | 4.9 | 37.45–3745 | 0.9998 | 0.9955 | 37.45 | 125.3 |
| YMC-Triart Diol-HILIC | 5.5 | 37.45–3745 | 0.9996 | 0.9716 | 37.45 | 117.4 |
| Compound | Retention time | Concentration range | Linearity (R2) | LOQ | |
|---|---|---|---|---|---|
| min | nM | w/o IS | w IS | nM | |
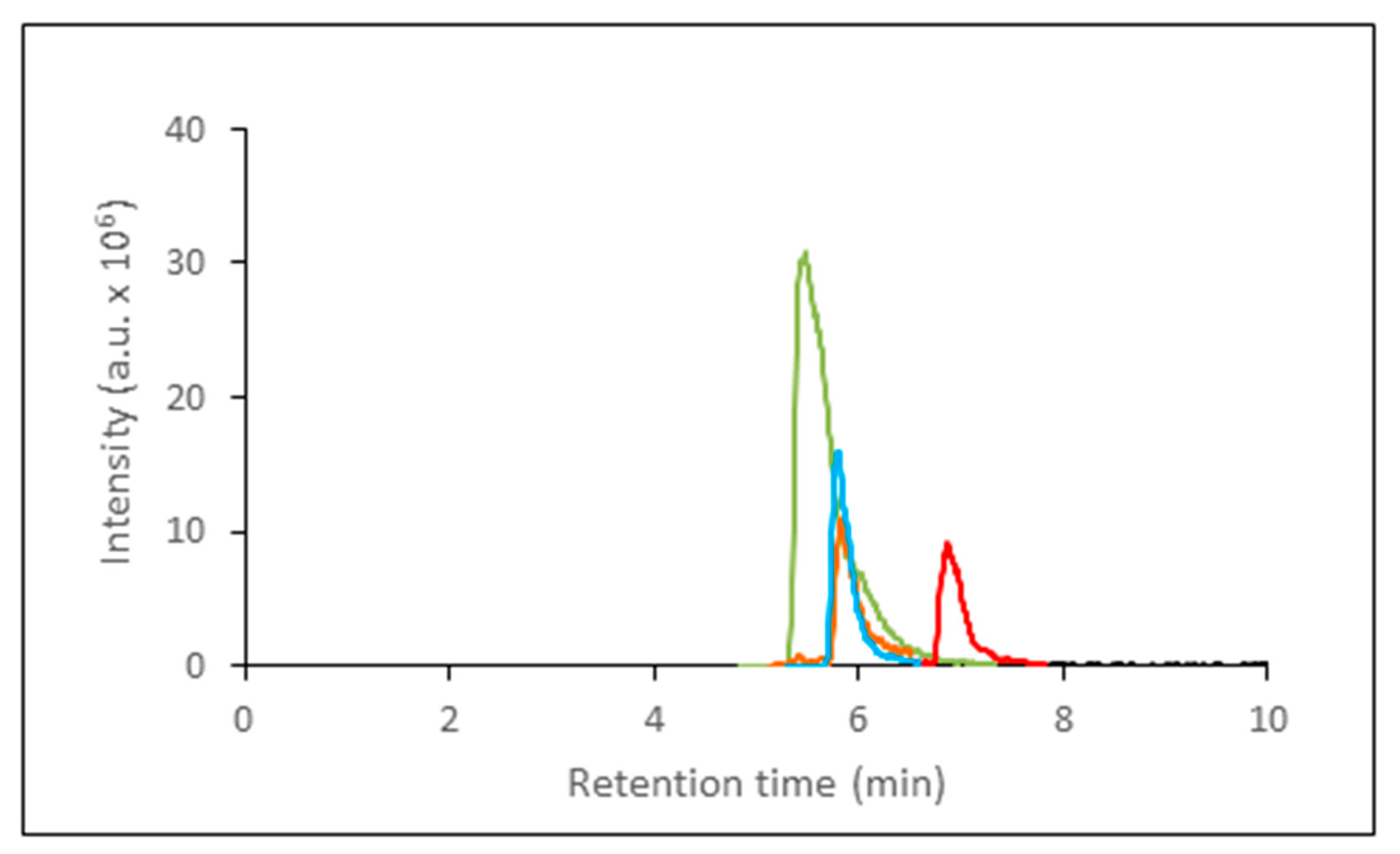
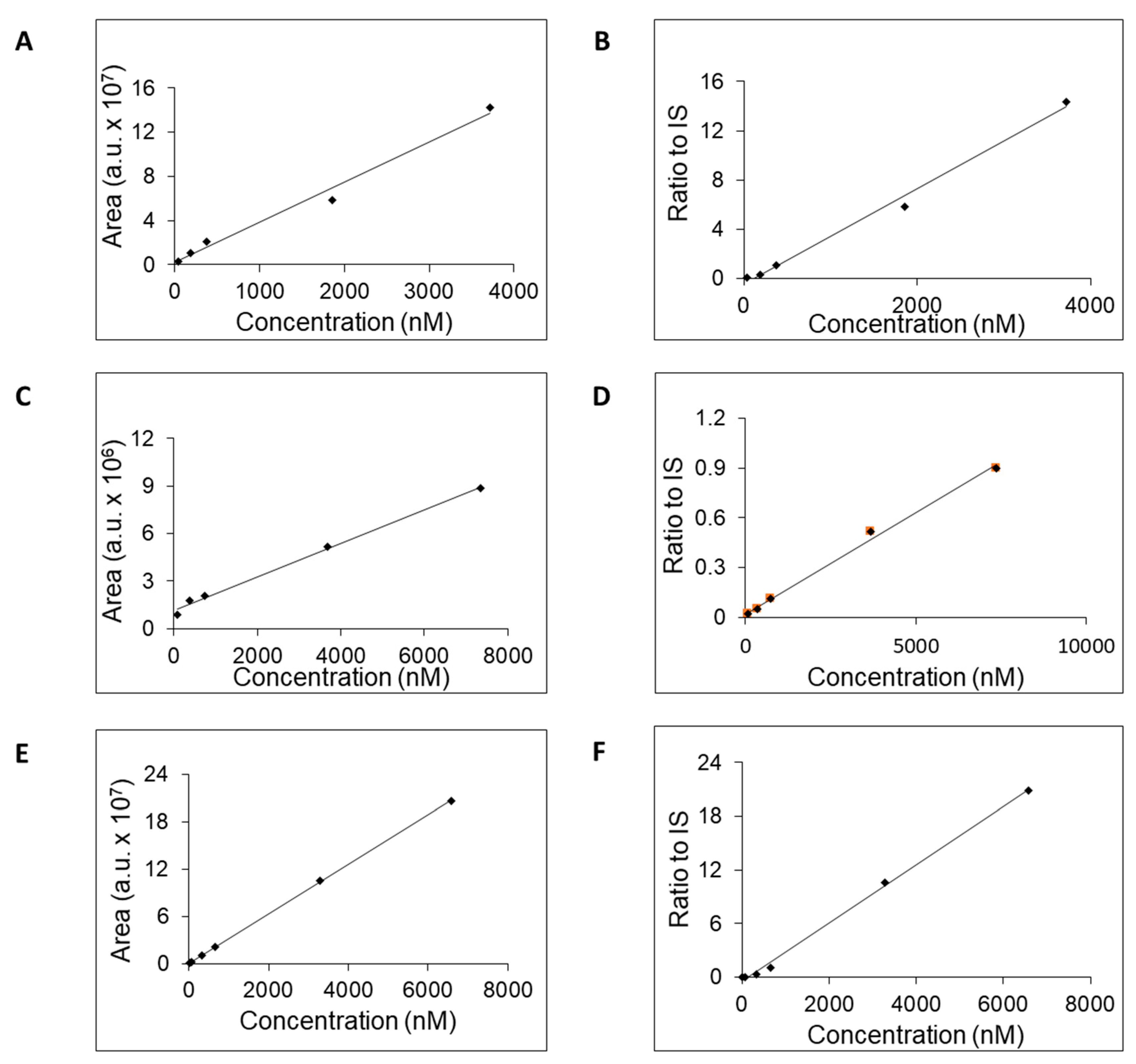
| Inosine | 6.9 | 37.2–3718 | 0.9858 | 0.9922 | 37.2 |
| Hypoxanthine | 5.8 | 73.5–7353 | 0.9951 | 0.9946 | 73.5 |
| Xanthine | 5.8 | 6.5–6579 | 0.9998 | 0.9977 | 6.5 |
| Adenosine | 5.5 | 3.75–1872 | 0.9702 | 0.9882 | 3.75 |
| Samples | [Adenosine] (µM) |
[Inosine] (µM) |
[Xanthine] (µM) |
|---|---|---|---|
| sample 1 | 0.92 | 1.66 | < LOQ |
| sample 2 | 4.33 | 8.18 | 70.42 |
| sample 3 | 0.13 | 1.15 | 1.38 |
| sample 4 | 0.36 | 1.29 | < LOQ |
| sample 5 | 2.78 | 3.66 | < LOQ |
| Average ±SD | 1.7 ±1.8 | 3.2 ±3.0 | ND. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




