Submitted:
16 January 2024
Posted:
17 January 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Isolation and culture of the strains
2.2. Sequencing and assembly of the phosphobacteria genomes
2.3. Genome annotation and comparisons
3. Results and Discussion
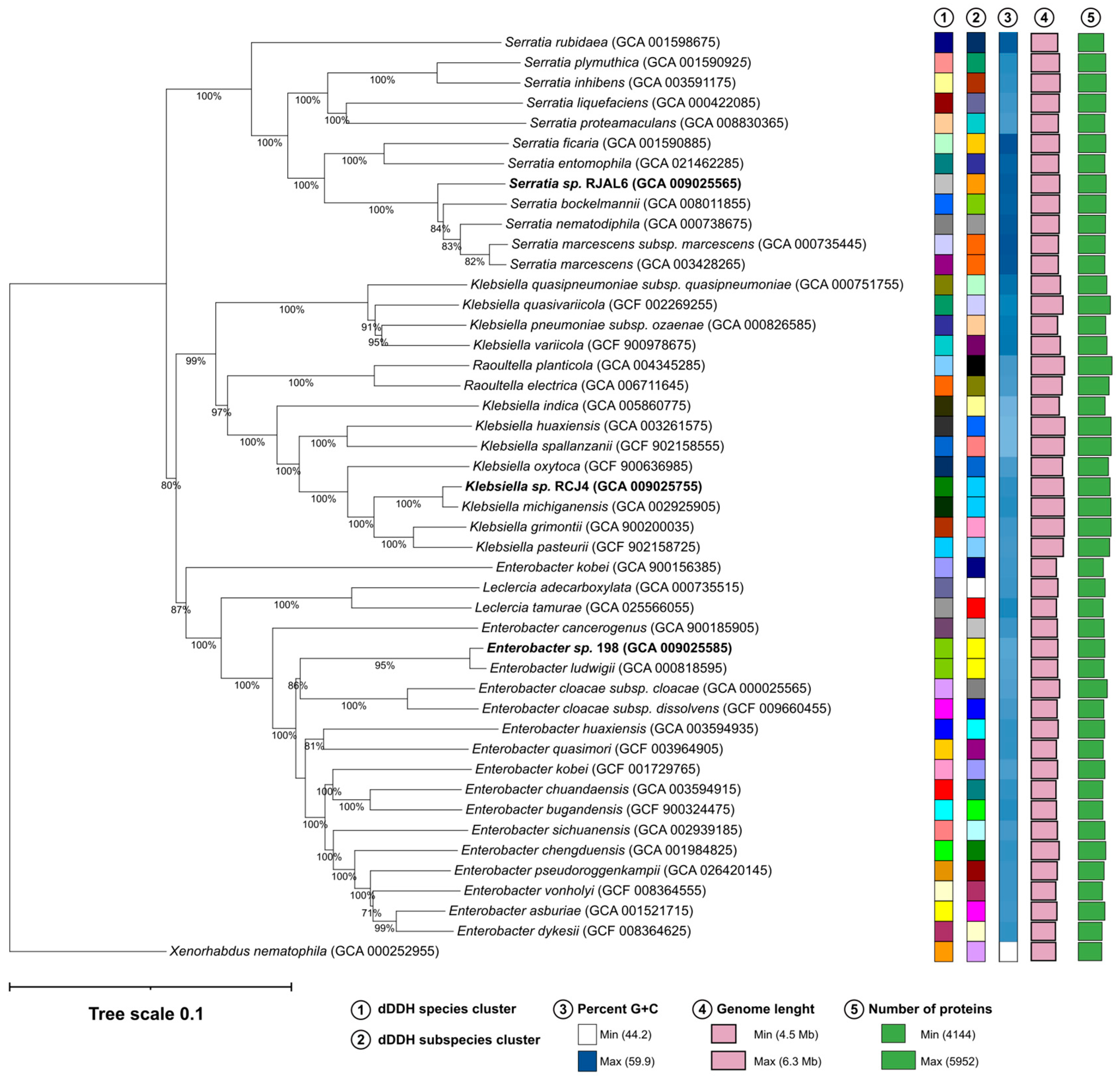
3.1. Genome based taxonomy
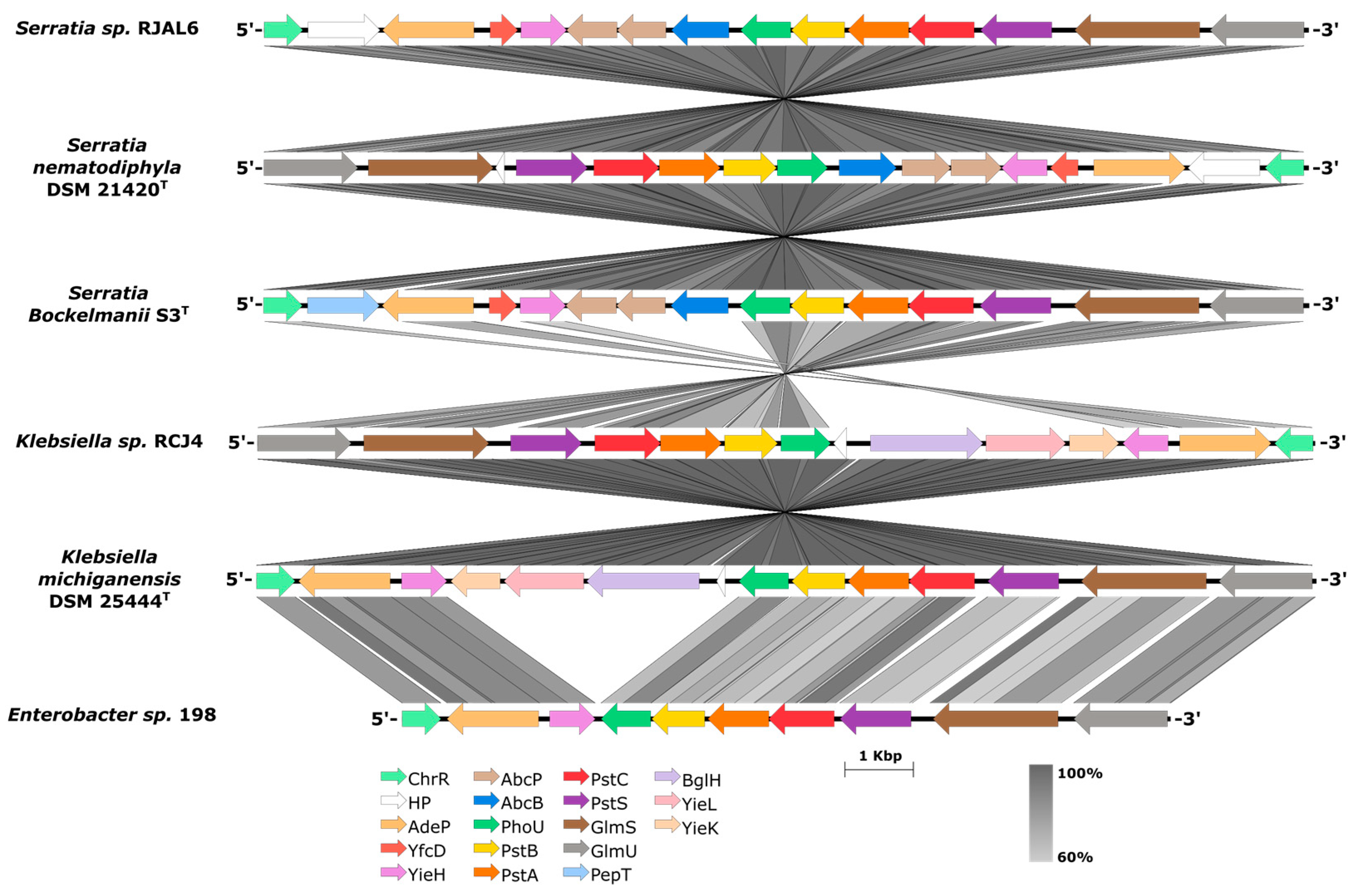
3.2. Phosphate uptake regulation genes
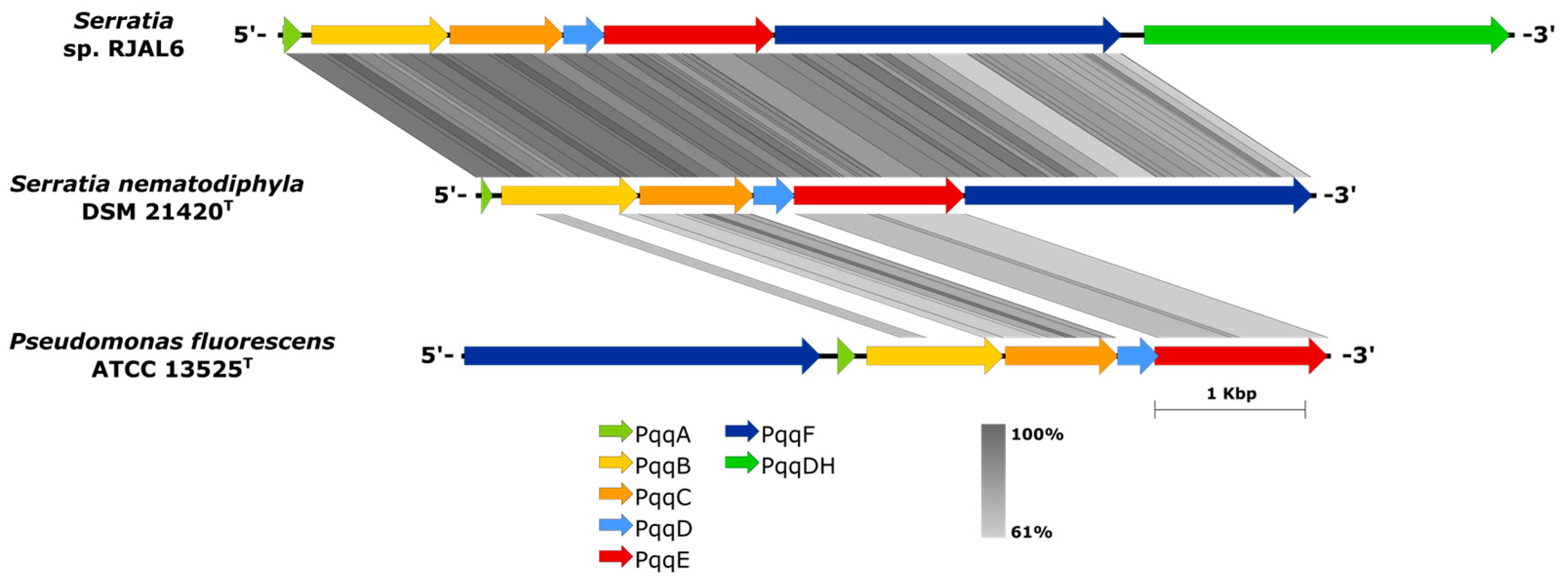
3.3. Gluconic acid production (Pqq genes)
3.4. Tryptophan-dependant auxin biosynthesis
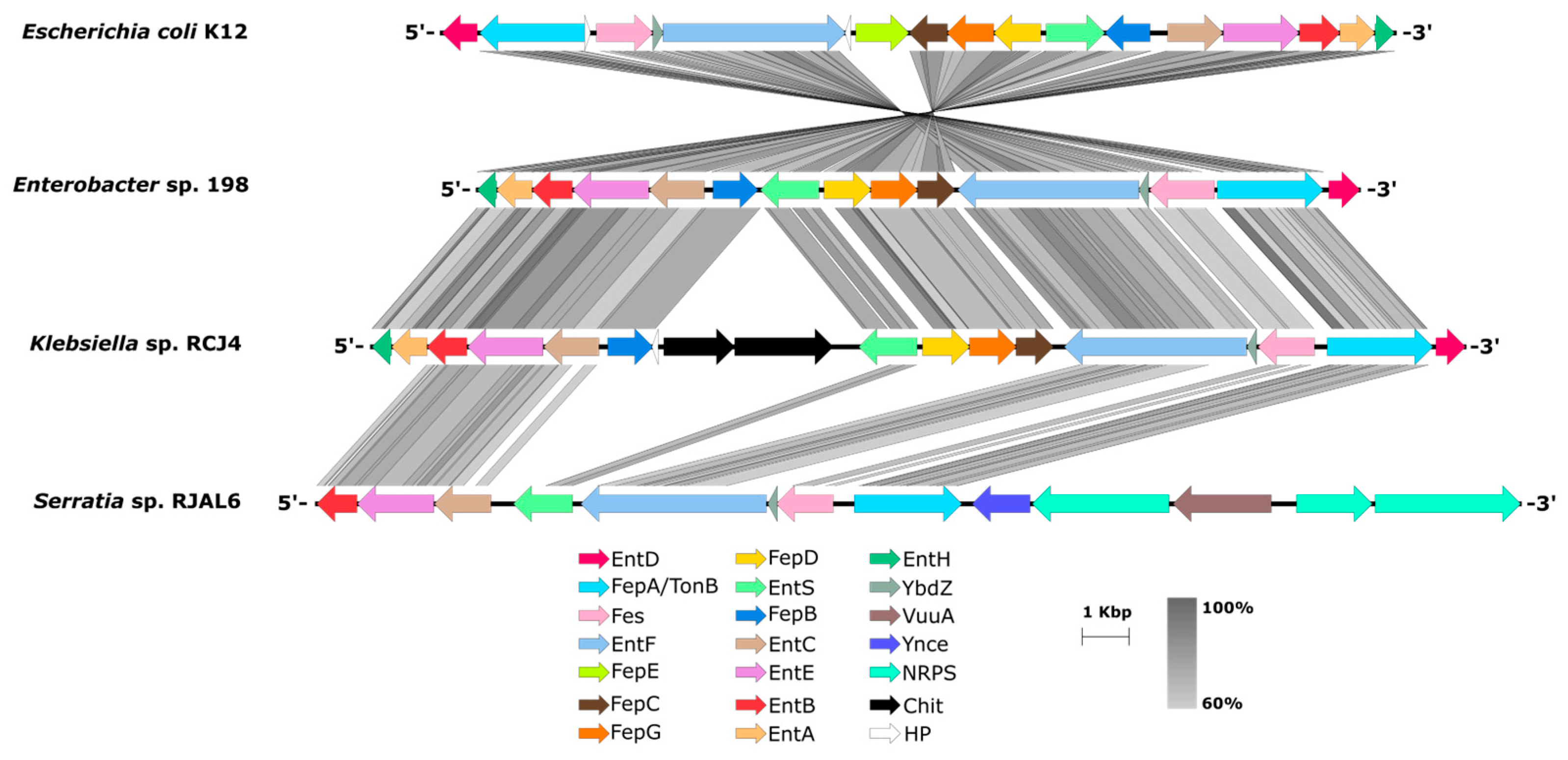
3.5. Secondary metabolites
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Barra, P. J., Inostroza, N. G., Acuña, J. J., Mora, M. L., Crowley, D. E., Jorquera, M. A. (2016). Formulation of bacterial consortia from avocado (Persea americana Mill.) and their effect on growth, biomass and superoxide dismutase activity of wheat seedlings under salt stress. Appl. Soil Ecol. 102, 80–91. [CrossRef]
- Barra, P. J., Inostroza, N. G., Mora, M. L., Crowley, D. E., Jorquera, M. A. (2017). Bacterial consortia inoculation mitigates the water shortage and salt stress in an avocado (Persea americana Mill.) nursery. Appl. Soil Ecol. 111, 39–47. [CrossRef]
- Barra, P. J., Pontigo, S., Delgado, M., Parra–Almuna, L., Duran, P., Valentine, A. J., Jorquera, M. L., Mora, M. L. (2019). Phosphobacteria inoculation enhances the benefit of P–fertilization on Lolium perenne in soils contrasting in P–availability. Soil Biol. Biochem. 136, 107516. [CrossRef]
- Inostroza, N.G., Barra, P.J., Wick, L.Y., Mora, M.L., Jorquera, M.A. (2017). Effect of rhizobacterial consortia from undisturbed arid- and agro-ecosystems on wheat growth under different conditions. Lett. Appl. Microbiol. 64, 158–163. [CrossRef]
- Glick, B. R. (2012). Plant growth-promoting bacteria: mechanisms and applications. Scientifica, 2012, 963401. [CrossRef]
- Ali, S. and Glick, B. R. (2019). Plant–bacterial interactions in management of plant growth under abiotic stresses. In New and Future Developments in Microbial Biotechnology and Bioengineering, 21–45. Amsterdam, the Netherlands: Elsevier. [CrossRef]
- Singh, J. S., and Gupta, V. K. (2018). Soil microbial biomass: A key soil driver in management of ecosystem functioning. Sci. Total Environ., 634, 497–500. [CrossRef]
- Castagno, L. N., Sannazzaro, A. I., Gonzalez, M. E., Pieckenstain, F. L., Estrella, M. J. (2021). Phospkhobacteria as key actors to overcome phosphorus deficiency in plants. Ann Appl Biol. 2021; 178: 256– 267. [CrossRef]
- Viruel, E., Lucca, M. E., Siñeriz, F. (2011). Plant growth promotion traits of phosphobacteria isolated from Puna, Argentina. Archives of microbiology, 193(7), 489–496. [CrossRef]
- Mora, M. L., Demanet, R., Acuña J., Viscardi, S., Jorquera, M., Rengel, Z., Durán, P. (2017). Aluminum-tolerant bacteria improve the plant growth and phosphorus content in ryegrass grown in a volcanic soil amended with cattle dung manure. Appl. Soil Ecol. 115, 19–26. [CrossRef]
- Barra, P. J., Viscardi, S., Jorquera, M. A., Duran, P. A., Valentine, A. J., Mora, M. L. (2018). Understanding the Strategies to Overcome Phosphorus-Deficiency and Aluminum-Toxicity by Ryegrass Endophytic and Rhizosphere Phosphobacteria. Frontiers in microbiology, 9, 1155. [CrossRef]
- Jackman, S. D., Vandervalk, B. P., Mohamadi, H., Chu, J., Yeo, S., Hammond, S. A., Jahesh, G., Khan, H., Coombe, L., Warren, R. L., Birol, I. (2017). ABySS 2.0: resource-efficient assembly of large genomes using a Bloom filter. Genome research, 27(5), 768–777. [CrossRef]
- Aziz, R.K., Devoid, S., Disz, T., Edwards, R.A., Henry, C.S., Olsen, G.J., Olson., R., Overbeek, R., Parello, B., Pusch, G.D., Stevens, R.L., Vonstein, V., Xia, F. (2012). SEED Servers: High-Performance Access to the SEED Genomes, Annotations, and Metabolic Models. Plos one. 7(10), e48053. [CrossRef]
- Yoon, S. H., Ha, S. M., Lim, J., Kwon, S., Chun, J. (2017a). A largescale evaluation of algorithms to calculate average nucleotide identity. Antonie van Leeuwenhoek. 110, 1281–1286. [CrossRef]
- Yoon, S. H., Ha, S. M., Kwon, S., Lim, J., Kim, Y., Seo, H., and Chun, J. (2017b). Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. International journal of systematic and evolutionary microbiology, 67(5), 1613–1617. [CrossRef]
- Meier-Kolthoff, J.P., Auch, A.F., Klenk, H-P., Göker, M. (2013). Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform. 14, 60. [CrossRef]
- Auch, A. F., Henz, S. R., Holland, B. R., Göker, M. (2006). Genome BLAST distance phylogenies inferred from whole plastid and whole mitochondrion genome sequences. BMC Bioinform.7, 350. [CrossRef]
- Meier-Kolthoff, J.P. and Göker, M. (2019). TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat Commun. 10, 2182. [CrossRef]
- Berriman, M., and Rutherford, K. M. (2003). Viewing and annotating sequence data with Artemis. Brief. Bioinform. 4, 124–132. [CrossRef]
- Bibb, M. J., Findlay, P. R., Johnson, M. W. (1984). The relationship between base composition and codon usage in bacterial genes and its use for the simple and reliable identification of protein-coding sequences. Gene 30, 157–166. [CrossRef]
- Meier-Kolthoff, J.P., Hahnke, R.L., Petersen, J., Scheuner, C., Michael, V., Fiebig, A., Rohde, C., Rohde, M., Fartmann, B., Goodwin, L.A., Chertkov, O., Reddy, T., Pati, A., Ivanova, N.N., Markowitz, V., Kyrpides, N.C., Woyke, T., Göker, M., Klenk, H-P. (2014). Complete genome sequence of DSM 30083T, the type strain (U5/41T) of Escherichia coli, and a proposal for delineating subspecies in microbial taxonomy. Stand Genomic Sci. 9, 2. [CrossRef]
- Richter, M. and Rosselló-Móra, R. (2009). Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl. Acad. Sci. U.S.A., 106(45), 19126–19131. [CrossRef]
- Rao, N. N., and Torriani, A. (1990). Molecular aspects of phosphate transport in Escherichia coli. Mol. Microbiol., 4(7), 1083–1090. [CrossRef]
- Aguena, M., Yagil, E. Spira, B. (2002). Transcriptional analysis of the pst operon of Escherichia coli. Mol. Genet. Genom., 268(4), 518–524. [CrossRef]
- Gardner, S. G., McCleary, W. R. (2019). Control of the phoBR Regulon in Escherichia coli. EcoSal Plus, 8(2). [CrossRef]
- Puehringer, S., Metlitzky, M. Schwarzenbacher, R. (2008). The pyrroloquinoline quinone biosynthesis pathway revisited: a structural approach. BMC biochemistry, 9, 8. [CrossRef]
- Klinman, J. P., and Bonnot, F. (2014). Intrigues and intricacies of the biosynthetic pathways for the enzymatic quinocofactors: PQQ, TTQ, CTQ, TPQ, and LTQ. Chem. Rev., 114(8), 4343–4365. [CrossRef]
- Chen, Y. P., Rekha, P. D., Arun, A. B., Shen, F. T., Lai, W. A., Young, C. C. (2006). Phosphate solubilizing bacteria from subtropical soil and their tricalcium phosphate solubilizing abilities. Appl. Soil Ecol., 34, 33–41. [CrossRef]
- Prabhu, N., Borkar, S., Garg, S. (2019). Phosphate solubilization by microorganisms. In Advances in Biological Science Research, 161–176. Amsterdam, the Netherlands: Elsevier. [CrossRef]
- Kogl, F. and Kostermans, D. (1934). Hetero-auxin als Stoffwechselprodukt niederer pflanzlicher Organismen. Isolierung aus Hefe.13. Mitteilung uber pflanzliche Wachstumsstoffe. Hoppe-Seyler´s Zeitschrift fur physiologische Chemie, 228:113-121. [CrossRef]
- Morffy, N. and Strader, L. C. (2020). Old Town Roads: routes of auxin biosynthesis across kingdoms. Curr. Opin. Plant Biol., 55, 21–27. [CrossRef]
- Kagan, J., Sharon, I., Beja, O., Kuhn, J. C. (2008). The tryptophan pathway genes of the Sargasso Sea metagenome: new operon structures and the prevalence of non-operon organization. Genome Biol., 9(1), R20. [CrossRef]
- Khan, A. R., Park, G. S., Asaf, S., Hong, S. J., Jung, B. K., Shin, J. H. (2017). Complete genome analysis of Serratia marcescens RSC-14: A plant growth-promoting bacterium that alleviates cadmium stress in host plants. PloS one, 12(2), e0171534. [CrossRef]
- Chen, T.-Y., Chen, J., Tang, Y., Zhou, J., Guo, Y., Chang, W.-C. (2021), Current Understanding toward Isonitrile Group Biosynthesis and Mechanism. Chin. J. Chem., 39: 463-472. [CrossRef]
- Brady, S. F., and Clardy, J. (2005). Cloning and heterologous expression of isocyanide biosynthetic genes from environmental DNA. Angewandte Chemie (International ed. in English), 44(43), 7063–7065. [CrossRef]
- Crawford, J. M., Kontnik, R., Clardy, J. (2010). Regulating alternative lifestyles in entomopathogenic bacteria. Curr. Biol. 20(1), 69–74. [CrossRef]
- Reitz, Z. L., Sandy, M., Butler, A. (2017). Biosynthetic considerations of triscatechol siderophores framed on serine and threonine macrolactone scaffolds. Metallomics, 9(7), 824–839. [CrossRef]
- Rusnak, F., Faraci, W. S., Walsh, C. T. (1989). Subcloning, expression, and purification of the enterobactin biosynthetic enzyme 2,3-dihydroxybenzoate-AMP ligase: demonstration of enzyme-bound (2,3-dihydroxybenzoyl) adenylate product. Biochemistry, 28(17), 6827–6835. [CrossRef]
- Zane, H. K., Naka, H., Rosconi, F., Sandy, M., Haygood, M. G., Butler, A. (2014). Biosynthesis of amphi-enterobactin siderophores by Vibrio harveyi BAA-1116: identification of a bifunctional nonribosomal peptide synthetase condensation domain. J. Am. Chem. Soc. 136(15), 5615–5618. [CrossRef]
- Leduc, D., Battesti, A., Bouveret, E. (2007). The hotdog thioesterase EntH (YbdB) plays a role in vivo in optimal enterobactin biosynthesis by interacting with the ArCP domain of EntB. Journal of bacteriology, 189(19), 7112–7126. [CrossRef]
- Furrer, J. L., Sanders, D. N., Hook-Barnard, I. G., McIntosh, M. A. (2002). Export of the siderophore enterobactin in Escherichia coli: involvement of a 43 kDa membrane exporter. Mol. Microbiol., 44(5), 1225–1234. [CrossRef]
- Chenault, S. S., and Earhart, C. F. (1991). Organization of genes encoding membrane proteins of the Escherichia coli ferrienterobactin permease. Mol. Microbiol., 5(6), 1405–1413. [CrossRef]
- Caza, M., Garénaux, A., Lépine, F., Dozois, C. M. (2015). Catecholate siderophore esterases Fes, IroD and IroE are required for salmochelins secretion following utilization, but only IroD contributes to virulence of extra-intestinal pathogenic Escherichia coli. Mol. Microbiol., 97(4), 717–732. [CrossRef]
- Barra, P. J., Duran, P., Delgado, M., Viscardi, S., Claverol, S., Larama, G., Dumont, M., Mora, M. L. (2023). Proteomic response to phosphorus deficiency and aluminum stress of three aluminum-tolerant phosphobacteria isolated from acidic soils. iScience, 26(10), 107910. [CrossRef]
- Schalk, I. J., Hannauer, M., Braud, A. (2011). New roles for bacterial siderophores in metal transport and tolerance. Environ. Microbiol., 13(11), 2844–2854. [CrossRef]
- Hu, X., and Boyer, G. L. (1996). Siderophore-Mediated Aluminum Uptake by Bacillus megaterium ATCC 19213. Appl. Environ. Microbiol., 62(11), 4044–4048. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



