Submitted:
09 January 2024
Posted:
10 January 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
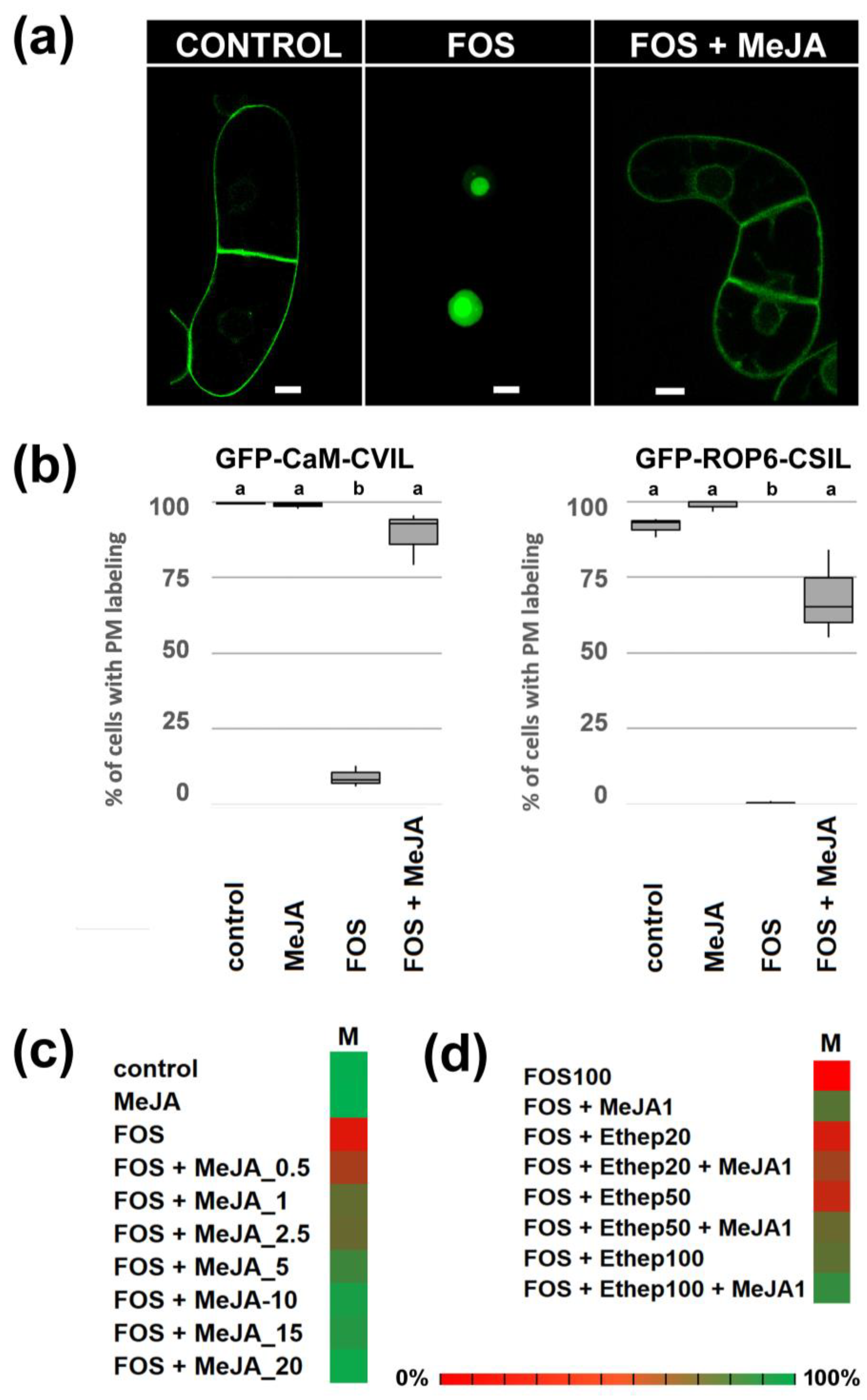
2.1. Me-JA Promotes Membrane Localization of Prenylated MEP-Derived CaaL-Box Proteins in the Presence of Fosmidomycin
2.2. PFT Modifies GFP-CaM-CVIL in the Presence of Methyl-Jasmonate
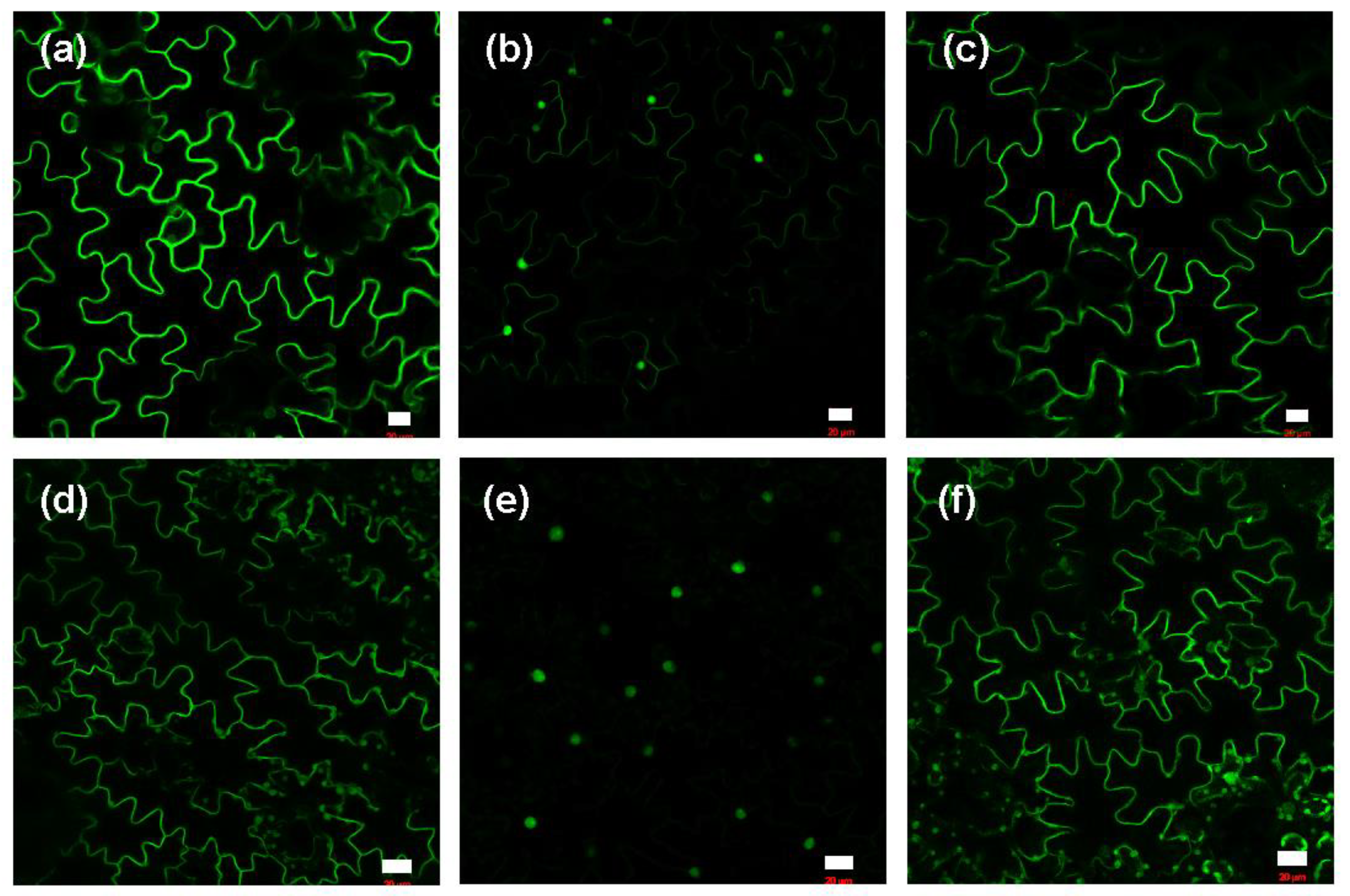
2.3. Me-JA Endorses Protein Prenylation Capacity in Tobacco
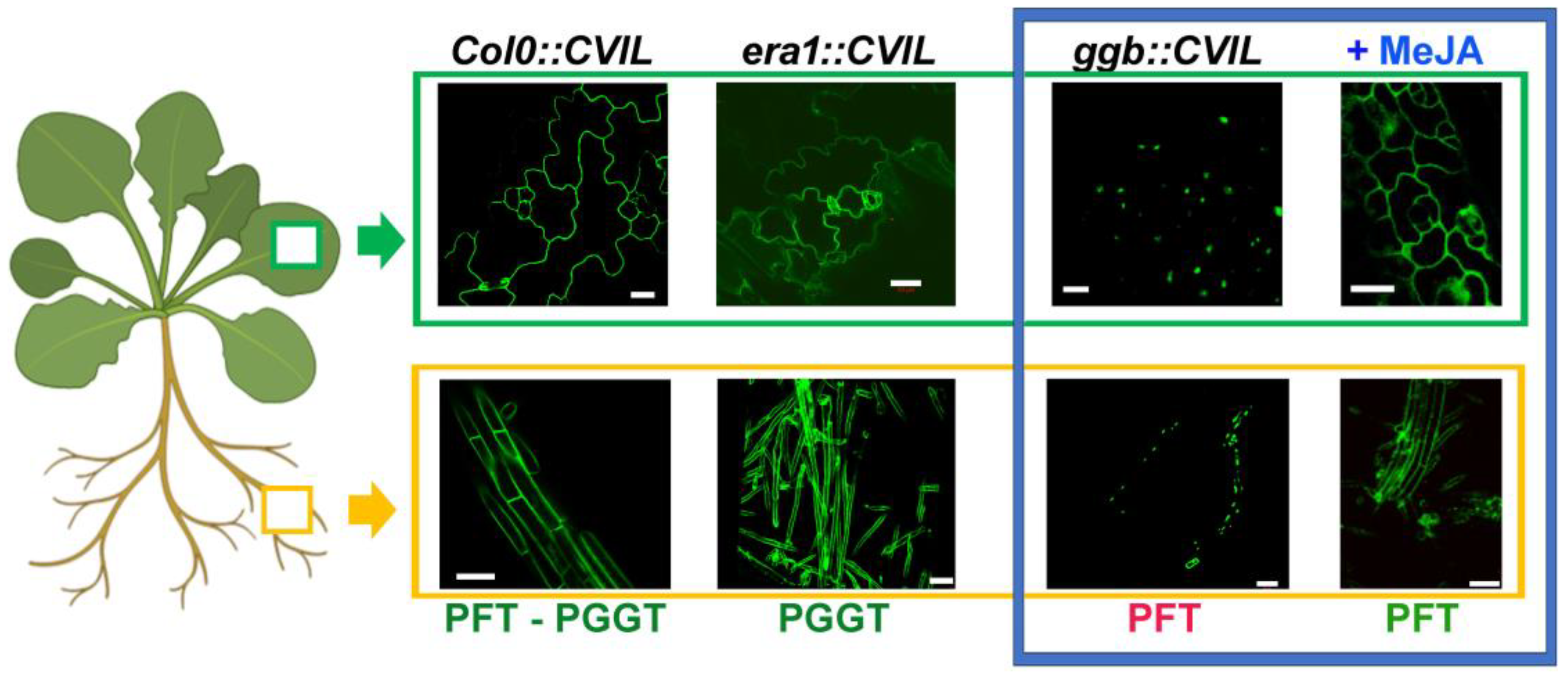
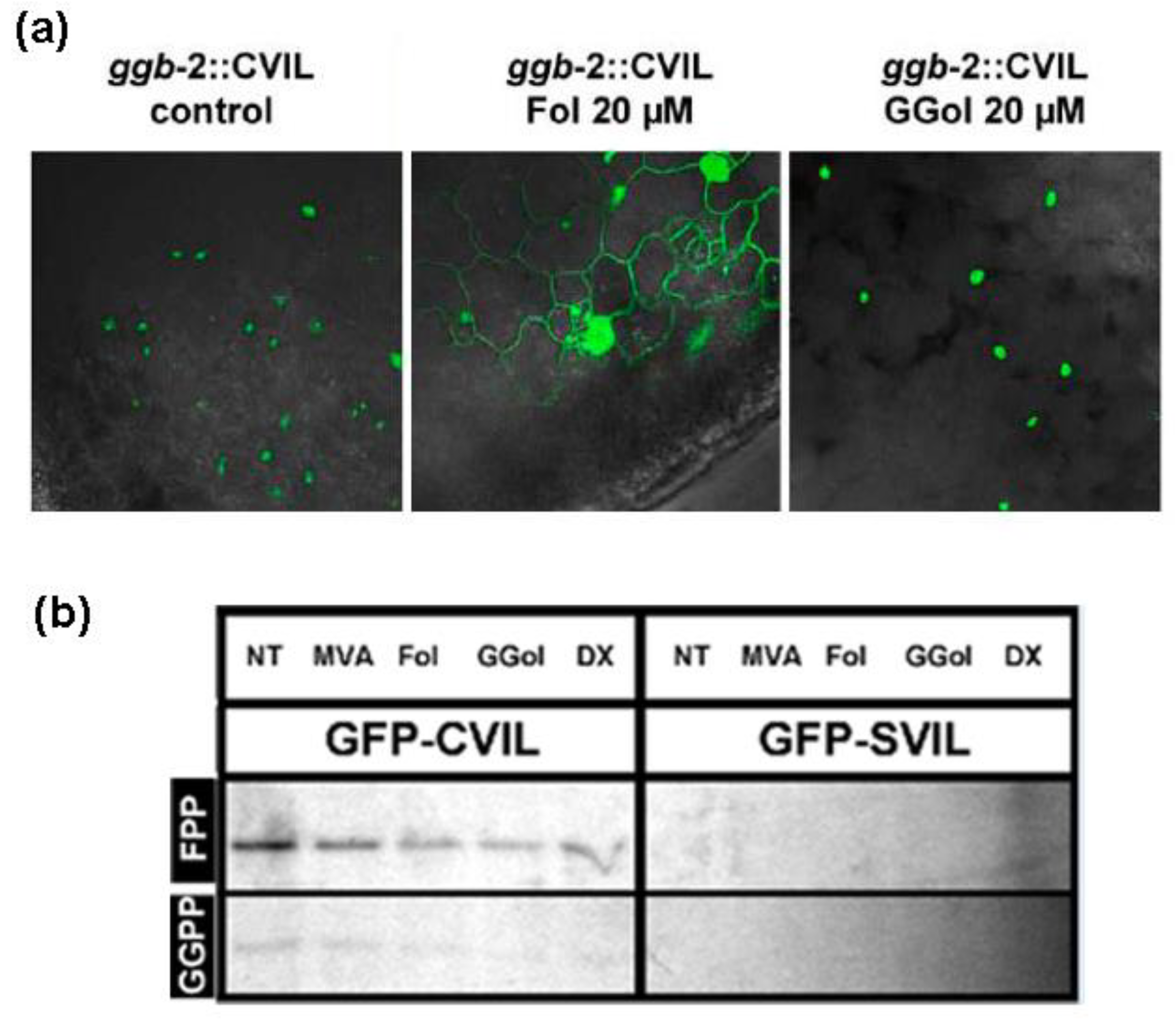
2.4. GFP-CaM-CVIL Prenylation under Restrictive Conditions Is Correlated with Mobilization of the MVA Pathway
3. Discussion
3.1. Modulation of PFT Substrate Specificities by Me-JA
3.2. MVA as a Sensor for Cross-Talk between MVA and MEP Pathway?
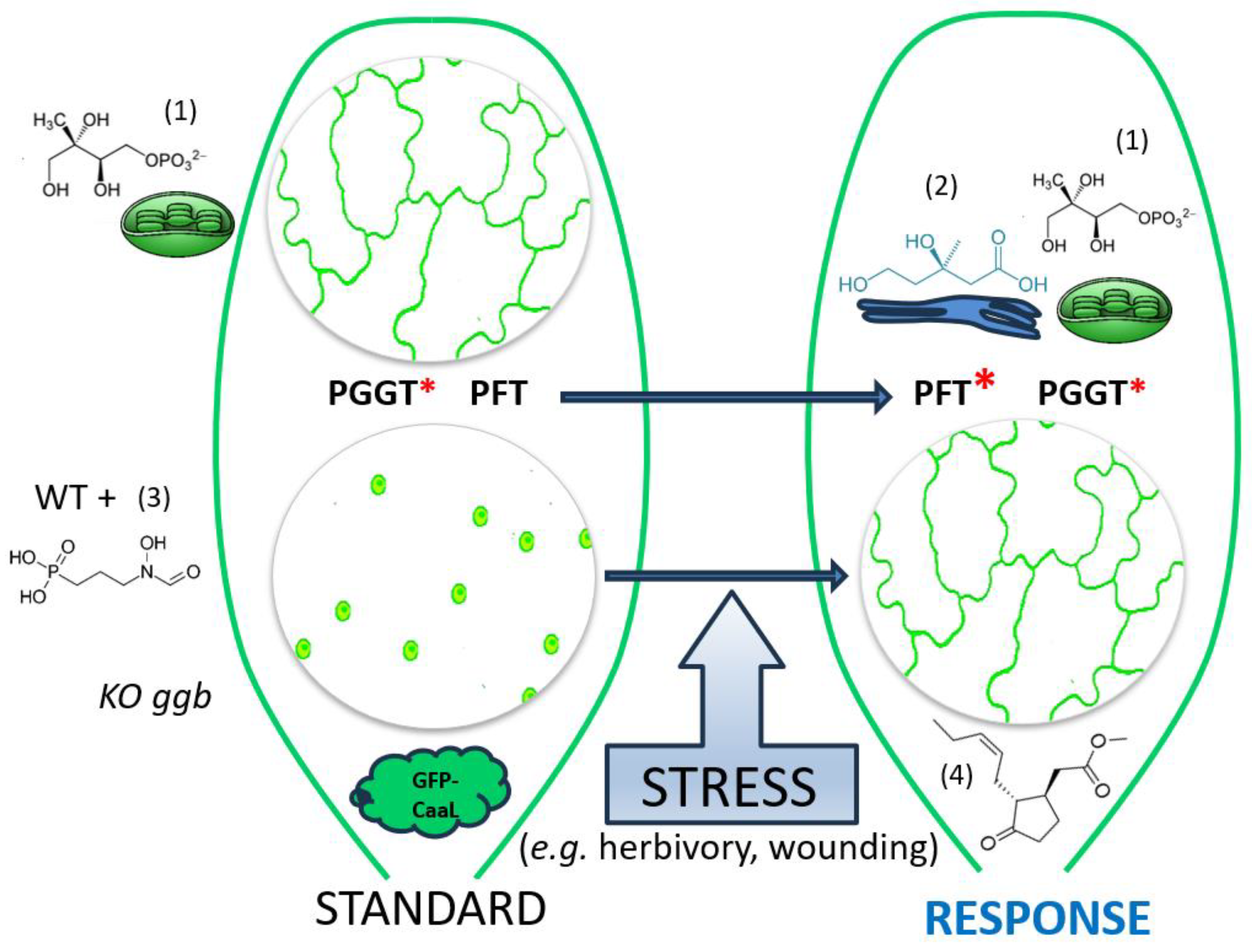
3.3. A Working Model of How Plants Benefit from Modifying PFT Substrate Selectivity
4. Materials and Methods
4.1. Chemical Materials
4.2. DNA Constructs, Biological Materials and Culture Conditions
4.3. Microscopy Analyses
4.4. Phytosterol Analyses
4.5. Protein Quantification, Enzyme Assays and Fluorography
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Fu, H.W.; Casey, P.J. Enzymology and biology of CaaX protein prenylation. Recent Prog. Horm. Res. 1999, 54, 315–342. [Google Scholar]
- Hála, M.; Žárský, V. Protein prenylation in plant stress responses. Molecules 2019, 24, 3906. [Google Scholar] [CrossRef] [PubMed]
- Hemsley, P.A. The importance of lipid modified proteins in plants. New Phytol. 2015, 205, 476–489. [Google Scholar] [CrossRef] [PubMed]
- Cutler, S.; Ghassemian, M.; Bonetta, D.; Cooney, S.; McCourt, P. A protein farnesyl transferase involved in abscisic acid signal transduction in Arabidopsis. Science 1996, 273, 1239–1241. [Google Scholar] [CrossRef] [PubMed]
- Pei, Z.M.; Ghassemian, M.; Kwak, C.M.; McCourt, P.; Schroeder, J.I. Role of farnesyltransferase in ABA regulation of guard cell anion channels and plant water loss. Science 1998, 282, 287–290. [Google Scholar] [CrossRef]
- Running, M.P.; Lavy, M.; Sternberg, H.; Galichet, A.; Gruissem, W.; Hake, S.; Ori, N.; Yalovsky, S. Enlarged meristems and delayed growth in plp mutants result from lack of CaaX prenyltransferases. Proc. Natl. Acad. Sci. USA 2004, 101, 7815–7820. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.D.; Chary, S.N.; Chernoff, E.A.; Zeng, Q.; Running, M.P.; Crowell, D.N. Protein geranylgeranyltransferase I is involved in specific aspects of abscisic acid and auxin signaling in Arabidopsis. Plant Physiol. 2005, 139, 722–733. [Google Scholar] [CrossRef] [PubMed]
- Running, M.P.; Fletcher, J.C.; Meyerowitz, E.M. The WIGGUM gene is required for proper regulation of floral meristem size in Arabidopsis. Development 1998, 125, 2545–2553. [Google Scholar] [CrossRef]
- Bonetta, D.; Bayliss, P.; Sun, S.; Sage, T.; McCourt, P. Farnesylation is involved in meristem organization in Arabidopsis. Planta 2000, 211, 182–190. [Google Scholar] [CrossRef]
- Yalovsky, S.; Rodríguez-Concepción, M.; Bracha, K.; Toledo-Ortiz, G.; Gruissem, W. Prenylation of the floral transcription factor APETALA1 modulates its function. Plant Cell 2000, 12, 1257–1266. [Google Scholar] [CrossRef]
- Ziegelhoffer, E.C.; Medrano, L.J.; Meyerowitz, E.M. Cloning of the Arabidopsis WIGGUM gene identifies a role for farnesylation in meristem development. Proc. Natl. Acad. Sci. USA 2000, 97, 7633–7638. [Google Scholar] [CrossRef] [PubMed]
- Vergès, V.; Dutilleul, C.; Godin, B.; Collet, B.; Lecureuil, A.; Rajjou, L.; Guimaraes, C.; Pinault, M.; Chevalier, S.; Giglioli-Guivarc’h, N.; Ducos, E. Protein farnesylation takes part in Arabidopsis seed development. Front. Plant Sci. 2021, 12, 620325. [Google Scholar] [CrossRef]
- Andrews, M.; Huizinga, D.H.; Crowell, D.N. The CaaX specificities of Arabidopsis protein prenyltransferases explain era1 and ggb phenotypes. BMC Plant Biol. 2010, 10, 118. [Google Scholar] [CrossRef]
- Barghetti, A.; Sjögren, L.; Floris, M.; Paredes, E.B.; Wenkel, S.; Brodersen, P. Heat-shock protein 40 is the key farnesylation target in meristem size control, abscisic acid signaling, and drought resistance. Genes Dev. 2017, 31, 2282–2295. [Google Scholar] [CrossRef] [PubMed]
- Brady, S.M.; Sarkar, S.F.; Bonetta, D.; McCourt, P. The ABSCISIC ACID INSENSITIVE 3 (ABI3) gene is modulated by farnesylation and is involved in auxin signaling and lateral root development in Arabidopsis. Plant J. 2003, 34, 67–75. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.R.; Wang, L.C.; Lin, Y.R.; Weng, C.P.; Yeh, C.H.; Wu, S.J. The Arabidopsis heat-intolerant 5 (hit5)/enhanced response to aba 1 (era1) mutant reveals the crucial role of protein farnesylation in plant responses to heat stress. New Phytol. 2017, 213, 1181–1193. [Google Scholar] [CrossRef]
- Yalovsky, S.; Kulukian, A.; Rodríguez-Concepción, M.; Young, C.A.; Gruissem, W. Functional requirement of plant farnesyltransferase during development in Arabidopsis. Plant Cell 2000, 12, 1267–1278. [Google Scholar] [CrossRef]
- Bracha-Drori, K.; Shichrur, K.; Lubetzky, T.C.; Yalovsky, S. Functional analysis of Arabidopsis postprenylation CaaX processing enzymes and their function in subcellular protein targeting. Plant Physiol. 2008; 148, 119–143. [Google Scholar] [CrossRef]
- Tugizimana, F.; Steenkamp, P.A.; Piater, L.A.; Dubery, I.A. Ergosterol-induced sesquiterpenoid synthesis in tobacco cells. Molecules 2012, 17, 1698–1715. [Google Scholar] [CrossRef]
- Palsuledesai, C.C.; Distefano, M.D. Protein prenylation: enzymes, therapeutics, and biotechnology applications. ACS Chem. Biol. 2015, 10, 51–62. [Google Scholar] [CrossRef]
- Jung, D.; Bachmann, H.S. Regulation of protein prenylation. Biomed. Pharmacother. 2023, 164, 114915. [Google Scholar] [CrossRef]
- Hemmerlin, A.; Harwood, J.L.; Bach, T.J. A raison d'être for two distinct pathways in the early steps of plant isoprenoid biosynthesis? Prog. Lipid Res. 2012, 51, 95–148. [Google Scholar] [CrossRef] [PubMed]
- Randall, S.K.; Marshall, M.S.; Crowell, D.N. Protein isoprenylation in suspension-cultured tobacco cells. Plant Cell 1993, 5, 433–442. [Google Scholar] [CrossRef]
- Morehead, T.A.; Biermann, B.J.; Crowell, D.N.; Randall, S.K. Changes in protein isoprenylation during the growth of suspension-cultured tobacco cells. Plant Physiol. 1995, 109, 277–284. [Google Scholar] [CrossRef]
- Swiezewska, E.; Thelin, A.; Dallner, G.; Andersson, B.; Ernster, L. Occurrence of prenylated proteins in plant cells. Biochem. Biophys. Res. Commun. 1993, 192, 161–166. [Google Scholar] [CrossRef] [PubMed]
- Galichet, A.; Gruissem, W. Developmentally controlled farnesylation modulates AtNAP1; 1 function in cell proliferation and cell expansion during Arabidopsis leaf development. Plant Physiol. 2006, 142, 1412–1426. [Google Scholar] [CrossRef]
- Hemmerlin, A.; Hoeffler, J.-F.; Meyer, O.; Tritsch, D.; Kagan, I.A.; Grosdemange-Billiard, C.; Rohmer, M.; Bach, T.J. Crosstalk between the cytosolic mevalonate and the plastidial methylerythritol phosphate pathways in tobacco Bright Yellow-2 cells. J. Biol. Chem. 2003, 278, 26666–26676. [Google Scholar] [CrossRef]
- Gerber, E.; Hemmerlin, A.; Hartmann, M.; Heintz, D.; Hartmann, M.A.; Mutterer, J.; Rodríguez-Concepción, M.; Boronat, A.; Van Dorsselaer, A.; Rohmer, M.; Crowell, D.N.; Bach, T.J. The plastidial 2-C-methyl-D-erythritol 4-phosphate pathway provides the isoprenyl moiety for protein geranylgeranylation in tobacco BY-2 cells. Plant Cell 2009, 21, 285–300. [Google Scholar] [CrossRef]
- Hartmann, M.; Hemmerlin, A.; Gas-Pascual, E.; Gerber, E.; Tritsch, D.; Rohmer, M.; Bach, T.J. The effect of MEP pathway and other inhibitors on the intracellular localization of a plasma membrane-targeted, isoprenylable GFP reporter protein in tobacco BY-2 cells. F1000Res. 2013, 2, 170. [Google Scholar] [CrossRef] [PubMed]
- Hemmerlin, A. Phosphorylation of metabolites involved in salvage pathways for isoprenoid biosynthesis in plants. Kinases Phosphatases 2023, 1, 151–166. [Google Scholar] [CrossRef]
- Nagel, R.; Schmidt, A.; Peters, R.J. Isoprenyl diphosphate synthases: the chain length determining step in terpene biosynthesis. Planta 2019, 249, 9–20. [Google Scholar] [CrossRef]
- Swiezewska, E.; Danikiewicz, W. Polyisoprenoids: structure, biosynthesis and function. Prog. Lipid Res. 2005, 44, 235–258. [Google Scholar] [CrossRef]
- Lipko, A.; Swiezewska, E. Isoprenoid generating systems in plants—a handy toolbox how to assess contribution of the mevalonate and methylerythritol phosphate pathways to the biosynthetic process. Prog. Lipid Res. 2016, 63, 70–92. [Google Scholar] [CrossRef] [PubMed]
- Vranová, E.; Coman, D.; Gruissem, W. Network analysis of the MVA and MEP pathways for isoprenoid synthesis. Annu. Rev. Plant Biol. 2013, 64, 665–700. [Google Scholar] [CrossRef]
- Wille, A.; Zimmermann, P.; Vranová, E.; Fürholz, A.; Laule, O.; Bleuler, S.; Hennig, L.; Prelić, A.; von Rohr, P.; Thiele, L.; Zitzler, E.; Gruissem, W.; Bühlmann, P. Sparse graphical Gaussian modeling of the isoprenoid gene network in Arabidopsis thaliana. Genome Biol. 2004, 5, R92. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, M.; Gas-Pascual, E.; Hemmerlin, A.; Rohmer, M.; Bach, T.J. Development of an image-based screening system for inhibitors of the plastidial MEP pathway and of protein geranylgeranylation. F1000Res. 2015, 4, 14. [Google Scholar] [CrossRef] [PubMed]
- Hemmerlin, A.; Bach, T.J. Farnesol induced cell death and stimulation of 3-hydroxy-3-methylglutaryl coenzyme A reductase activity in tobacco cv. Bright Yellow-2 cells. Plant Physiol. 2000, 123, 1257–1268. [Google Scholar] [CrossRef]
- Huchelmann, A.; Brahim, M.S.; Gerber, E.; Tritsch, D.; Bach, T.J.; Hemmerlin, A. Farnesol-mediated shift in the metabolic origin of prenyl groups used for protein prenylation in plants. Biochimie 2016, 127, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Huchelmann, A.; Gastaldo, C.; Veinante, M.; Zeng, Y.; Heintz, D.; Tritsch, D.; Schaller, H.; Rohmer, M.; Bach, T.J.; Hemmerlin, A. S-Carvone suppresses cellulase-induced capsidiol production in Nicotiana tabacum by interfering with protein isoprenylation. Plant Physiol. 2014, 164, 935–950. [Google Scholar] [CrossRef] [PubMed]
- Yalovsky, S. Protein lipid modifications and the regulation of ROP GTPase function. J. Exp. Bot. 2015, 66, 1617–1624. [Google Scholar] [CrossRef]
- Zhu, Z. Molecular basis for jasmonate and ethylene signal interactions in Arabidopsis. J. Exp. Bot. 2014, 65, 5743–5748. [Google Scholar] [CrossRef]
- Hampel, D.; Mosandl, A.; Wüst, M. Induction of de novo volatile terpene biosynthesis via cytosolic and plastidial pathways by methyl jasmonate in foliage of Vitis vinifera L. J. Agric. Food Chem. 2005, 53, 2652–2657. [Google Scholar] [CrossRef]
- Wei, Q.; Lan, K.; Liu, Y.; Chen, R.; Hu, T.; Zhao, S.; Yin, X.; Xie, T. Transcriptome analysis reveals regulation mechanism of methyl jasmonate-induced terpenes biosynthesis in Curcuma wenyujin. PLoS One 2022, 17, e0270309. [Google Scholar] [CrossRef]
- Wentzinger, L.F.; Bach, T.J.; Hartmann, M.A. Inhibition of squalene synthase and squalene epoxidase in tobacco cells triggers an up-regulation of 3-hydroxy-3-methylglutaryl coenzyme A reductase. Plant Physiol. 2002, 130, 334–346. [Google Scholar] [CrossRef]
- Hemmerlin, A.; Tritsch, D.; Hartmann, M.; Pacaud, K.; Hoeffler, J.-F.; Van Dorsselaer, A.; Rohmer, M.; Bach, T.J. A cytosolic Arabidopsis D-xylulose kinase catalyzes the phosphorylation of 1-deoxy-D-xylulose into a precursor of the plastidial isoprenoid pathway. Plant Physiol. 2006, 142, 441–457. [Google Scholar] [CrossRef]
- Kuchay, S.; Wang, H.; Marzio, A.; Jain, K.; Homer, H.; Fehrenbacher, N.; Philips, M.R.; Zheng, N.; Pagano, M. GGTase3 is a newly identified geranylgeranyltransferase targeting a ubiquitin ligase. Nat. Struct. Mol. Biol. 2019, 26, 628–636. [Google Scholar] [CrossRef]
- Shirakawa, R.; Goto-Ito, S.; Goto, K.; Wakayama, S.; Kubo, H.; Sakata, N.; Trinh, D.A.; Yamagata, A.; Sato, Y.; Masumoto, H.; Cheng, J.; Fujimoto, T.; Fukai, S.; Horiuchi, H. A SNARE geranylgeranyltransferase essential for the organization of the Golgi apparatus. EMBO J. 2020, 39, e104120. [Google Scholar] [CrossRef]
- Wasternack, C.; Hause, B. Jasmonates and octadecanoids: signals in plant stress responses and development. Prog. Nucleic Acid Res. Mol. Biol. 2002, 72, 165–221. [Google Scholar] [CrossRef]
- van der Fits, L.; Memelink, J. ORCA3, a jasmonate-responsive transcriptional regulator of plant primary and secondary metabolism. Science 2000, 289, 295–297. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Zhang, W.; Zhang, Y.; Zhang, X.; Lang, D.; Zhang, X. The roles of methyl jasmonate to stress in plants. Funct. Plant Biol. 2018, 46, 197–212. [Google Scholar] [CrossRef] [PubMed]
- Mertens, J.; Pollier, J.; Vanden Bossche, R.; Lopez-Vidriero, I.; Franco-Zorrilla, J.M.; Goossens, A. The bHLH transcription factors TSAR1 and TSAR2 regulate triterpene saponin biosynthesis in Medicago truncatula. Plant Physiol. 2016, 170, 194–210. [Google Scholar] [CrossRef]
- Yu, Z.X.; Li, J.X.; Yang, C.Q.; Hu, W.L.; Wang, L.J.; Chen, X.Y. The jasmonate-responsive AP2/ERF transcription factors AaERF1 and AaERF2 positively regulate artemisinin biosynthesis in Artemisia annua L. Mol. Plant 2012, 5, 353–365. [Google Scholar] [CrossRef] [PubMed]
- Perreca, E.; Eberl, F.; Santoro, M.V.; Wright, L.P.; Schmidt, A.; Gershenzon, J. Effect of drought and methyl jasmonate treatment on primary and secondary isoprenoid metabolites derived from the MEP pathway in the white spruce Picea glauca. Int. J. Mol. Sci. 2022, 23, 3838. [Google Scholar] [CrossRef] [PubMed]
- Ho, T.T.; Murthy, H.N.; Park, S.Y. Methyl jasmonate induced oxidative stress and accumulation of secondary metabolites in plant cell and organ cultures. Int. J. Mol. Sci. 2020, 21, 716. [Google Scholar] [CrossRef] [PubMed]
- Sasaki-Sekimoto, Y.; Taki, N.; Obayashi, T.; Aono, M.; Matsumoto, F.; Sakurai, N.; Suzuki, H.; Yokota Hirai, M.; Noji, M.; Saito, K.; Masuda, T.; Takamiya, K.I.; Shibata, D.; Ohta, H. Coordinated activation of metabolic pathways for antioxidants and defence compounds by jasmonates and their roles in stress tolerance in Arabidopsis. Plant J. 2005, 44, 653–668. [Google Scholar] [CrossRef] [PubMed]
- Sohn, S.I.; Pandian, S.; Rakkammal, K.; Largia, M.J.V.; Thamilarasan, S.K.; Balaji, S.; Zoclanclounon, Y.A.B.; Shilpha, J.; Ramesh, M. Jasmonates in plant growth and development and elicitation of secondary metabolites: An updated overview. Front. Plant Sci. 2022, 13, 942789. [Google Scholar] [CrossRef] [PubMed]
- Courdavault, V.; Burlat, V.; St-Pierre, B.; Giglioli-Guivarc’h, N. Proteins prenylated by type I protein geranylgeranyltransferase act positively on the jasmonate signalling pathway triggering the biosynthesis of monoterpene indole alkaloids in Catharanthus roseus. Plant Cell Rep. 2009, 28, 83–93. [Google Scholar] [CrossRef] [PubMed]
- Courdavault, V.; Thiersault, M.; Courtois, M.; Gantet, P.; Oudin, A.; Doireau, P.; St-Pierre, B.; Giglioli-Guivarc’h, N. CaaX-prenyltransferases are essential for expression of genes involved in the early stages of monoterpenoid biosynthetic pathway in Catharanthus roseus cells. Plant Mol. Biol. 2005, 57, 855–870. [Google Scholar] [CrossRef]
- Kumar, S.R.; Rai, A.; Bomzan, D.P.; Kumar, K.; Hemmerlin, A.; Dwivedi, V.; Godbole, R.C.; Barvkar, V.; Shanker, K.; Shilpashree, H.B.; Bhattacharya, A.; Smitha, A.R.; Hedge, N.; Nagegowda, D.A. A plastid-localized bona fide geranylgeranyl diphosphate synthase plays a necessary role in monoterpene indole alkaloid biosynthesis in Catharanthus roseus. Plant J. 2020, 103, 248–265. [Google Scholar] [CrossRef] [PubMed]
- Lacchini, E.; Goossens, A. Combinatorial control of plant specialized metabolism: mechanisms, functions, and consequences. Annu. Rev. Cell Dev. Biol. 2020, 36, 291–313. [Google Scholar] [CrossRef] [PubMed]
- Liao, P.; Hemmerlin, A.; Bach, T.J.; Chye, M.L. The potential of the mevalonate pathway for enhanced isoprenoid production. Biotechnol. Adv. 2016, 34, 697–713. [Google Scholar] [CrossRef]
- Opitz, S.; Nes, W.D.; Gershenzon, J. Both methylerythritol phosphate and mevalonate pathways contribute to biosynthesis of each of the major isoprenoid classes in young cotton seedlings. Phytochemistry 2014, 98, 110–119. [Google Scholar] [CrossRef]
- Schneider, M.M.; Hampp, R.; Ziegler, H. Envelope permeability to possible precursors of carotenoid biosynthesis during chloroplast-chromoplast transformation. Plant Physiol. 1977, 1977. 60, 518–520. [Google Scholar] [CrossRef]
- Li, W.; Liu, W.; Wei, H.; He, Q.; Chen, J.; Zhang, B.; Zhu, S. Species-specific expansion and molecular evolution of the 3-hydroxy-3-methylglutaryl coenzyme A reductase (HMGR) gene family in plants. PLoS One 2014, 9, e94172. [Google Scholar] [CrossRef]
- Chappell, J. Biochemistry and molecular biology of the isoprenoid biosynthetic pathway in plants. Annu. Rev. Plant Biol. 1995, 46, 521–547. [Google Scholar] [CrossRef]
- Gutensohn, M.; Hartzell, E.; Dudareva, N. Another level of complexity: The role of metabolic channeling and metabolons in plant terpenoid metabolism. Front. Plant Sci. 2022, 13, 954083. [Google Scholar] [CrossRef]
- Gong, L.; Xiao, Y.; Xia, F.; Wu, P.; Zhao, T.; Xie, S.; Wang, R.; Wen, Q.; Zhou, W.; Xu, H.; Zhu, L.; Zheng, Z.; Yang, T.; Chen, Z.; Duan, Q. The mevalonate coordinates energy input and cell proliferation. Cell Death Dis. 2019, 10, 327. [Google Scholar] [CrossRef]
- Cho, S.H.; Tóth, K.; Kim, D.; Vo, P.H.; Lin, C.H.; Handakumbura, P.P.; Ubach, A.R.; Evans, S.; Paša-Tolić, L.; Stacey, G. Activation of the plant mevalonate pathway by extracellular ATP. Nat. Commun. 2022, 13, 450. [Google Scholar] [CrossRef]
- Robertlee, J.; Kobayashi, K.; Suzuki, M.; Muranaka, T. AKIN 10, a representative Arabidopsis SNF 1-related protein kinase 1 (SnRK1), phosphorylates and downregulates plant HMG-CoA reductase. FEBS lett. 2017, 591, 1159–1166. [Google Scholar] [CrossRef]
- Hemmerlin, A.; Brown, S.C.; Bach, T.J. Function of mevalonate in tobacco cell proliferation. Acta Bot. Gall. 1999, 146, 85–100. [Google Scholar] [CrossRef]
- Sano, H.; Ohashi, Y. Involvement of small GTP-binding proteins in defense signal-transduction pathways of higher plants. Proc. Natl. Acad. Sci. USA 1995, 92, 4138–4144. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Huang, W. Calcium-dependent protein kinases in phytohormone signaling pathways. Int. J. Mol. Sci. 2017, 18, 2436. [Google Scholar] [CrossRef]
- Bomzan, D.P.; Sharma, A.; Cruz, P.L.; Carqueijeiro, I.; Bellenger, L.; Rai, A.; Kumar Thippesh, A.; Venkatesha, S.C.; Parihar, D.; Ducos, E.; Courdavault, V.; Nagegowda, D.A. Rho of plant GTPases with geranylgeranylation motif modulate monoterpene indole alkaloid biosynthesis in Catharanthus roseus. bioRxiv 2023, 2023–09. [Google Scholar] [CrossRef]
- Aoyama, T.; Chua, N.H. A glucocorticoid-mediated transcriptional induction system in transgenic plants. Plant J. 1997, 11, 605–612. [Google Scholar] [CrossRef]
- Rodríguez-Concepción, M.; Yalovsky, S.; Zik, M.; Fromm, H.; Gruissem, W. The prenylation status of a novel plant calmodulin directs plasma membrane or nuclear localization of the protein. EMBO J. 1999, 1999. 18, 1996–2007. [Google Scholar] [CrossRef]
- Hemmerlin, A.; Bach, T.J. Effects of mevinolin on cell cycle progression and viability of tobacco BY-2 cells. Plant J. 1998, 14, 65–74. [Google Scholar] [CrossRef]
- van der Fits, L.; Deakin, E.A.; Hoge, J.H.C.; Memelink, J. The ternary transformation system: constitutive virG on a compatible plasmid dramatically increases Agrobacterium-mediated plant transformation. Plant Mol. Biol. 2000, 43, 495–502. [Google Scholar] [CrossRef]
- Gallois, P.; Marinho, P. Leaf disk transformation using Agrobacterium tumefaciens - expression of heterologous genes in tobacco. Methods Mol. Biol. 1995, 49, 39–48. [Google Scholar]
- Darnet, S.; Martin, L.; Mercier, P.; Bracher, F.; Geoffroy, P.; Schaller, H. Inhibition of phytosterol biosynthesis by azasterols. Molecules 2020, 25, 1111. [Google Scholar] [CrossRef]
- Namdeo, A.G. Plant cell elicitation for production of secondary metabolites: A review. Pharmacogn. Rev. 2007, 1, 69–79. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



