Submitted:
20 December 2023
Posted:
20 December 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Experimental design
2.2. Experimental measurements
2.2.1. Phenotypic index
2.2.2. Transcriptome and proteome sequencing
2.3. Data analysis
3. Results
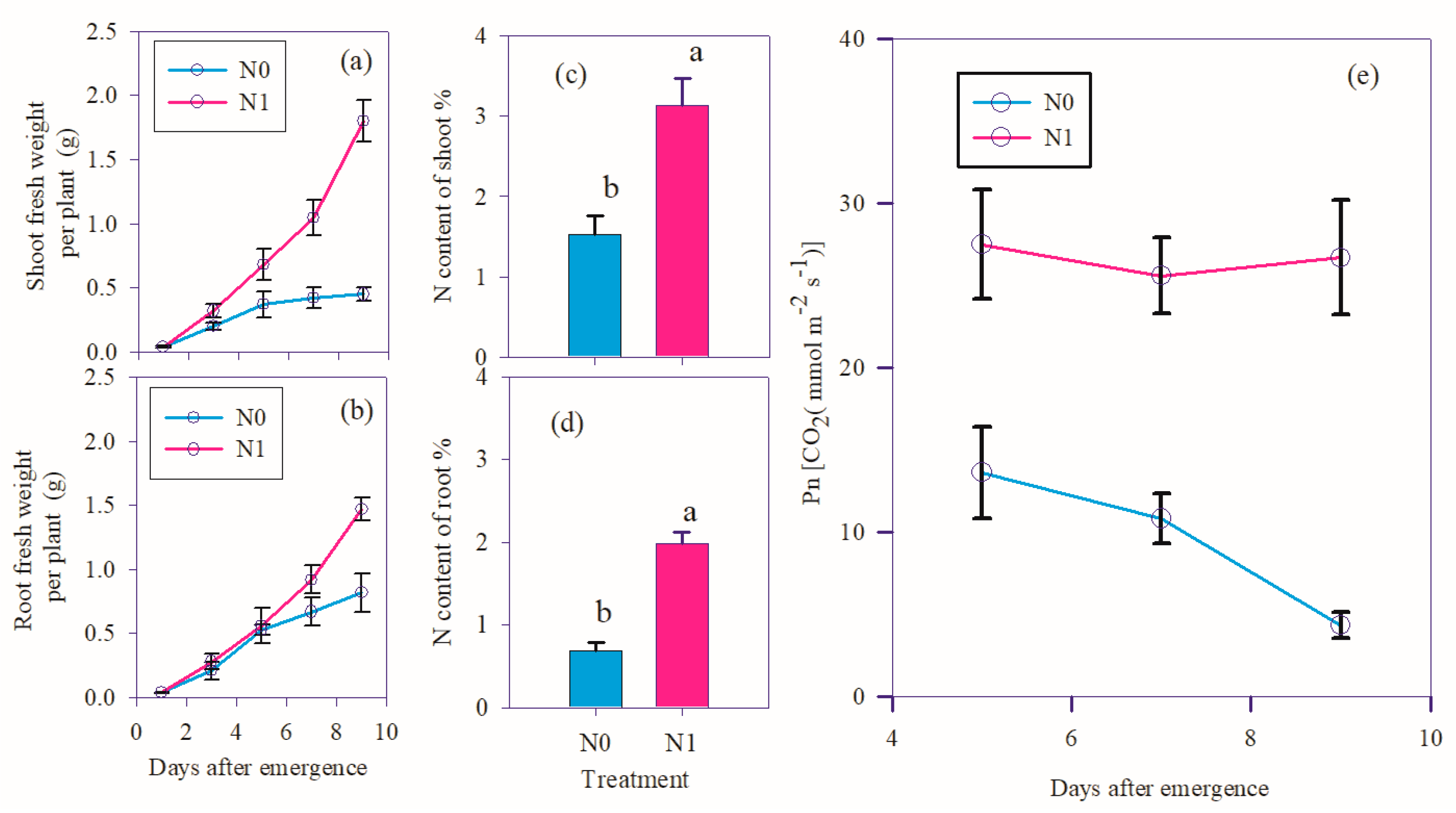
3.1. Phenotype of wheat under nitrogen deficit
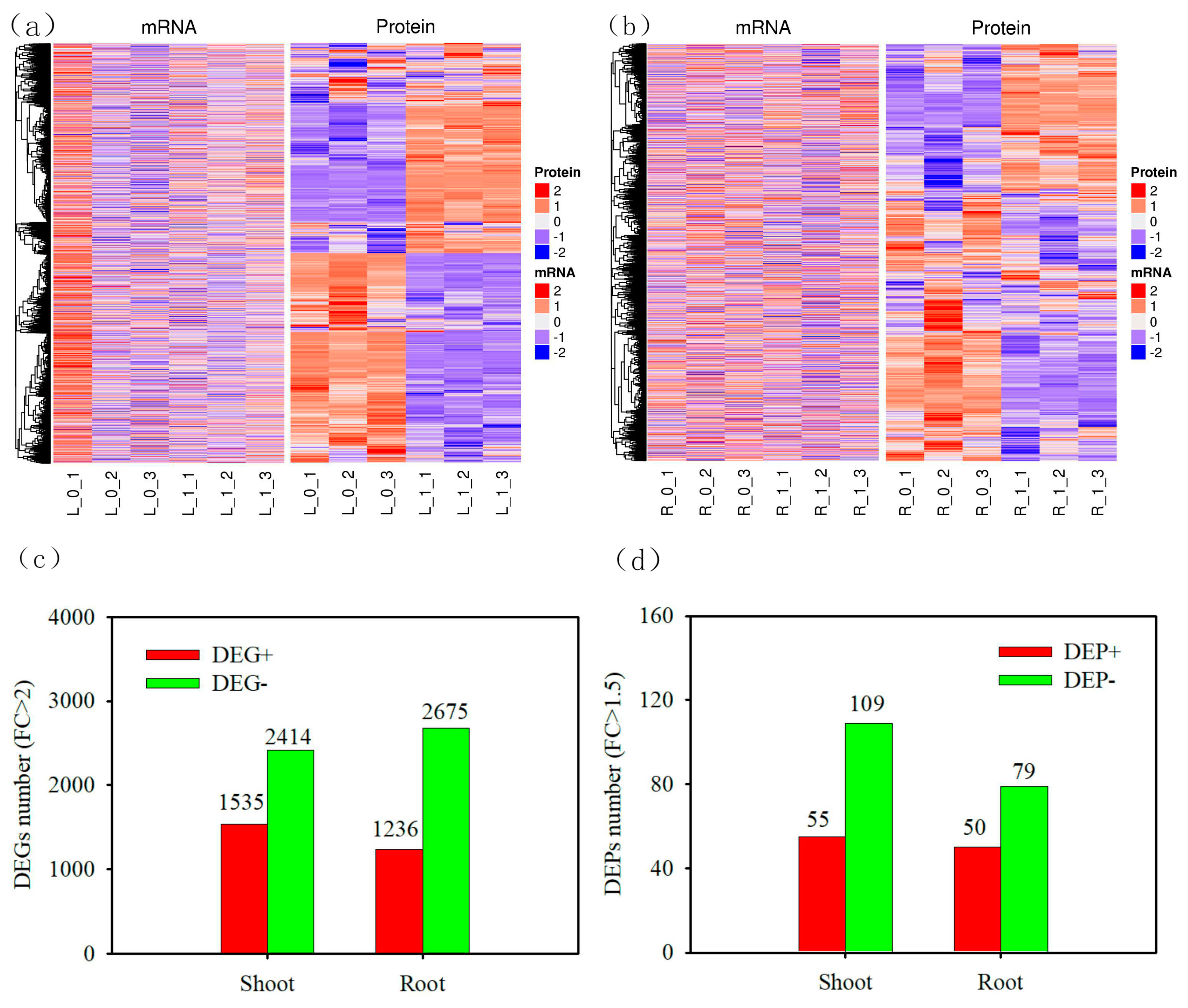
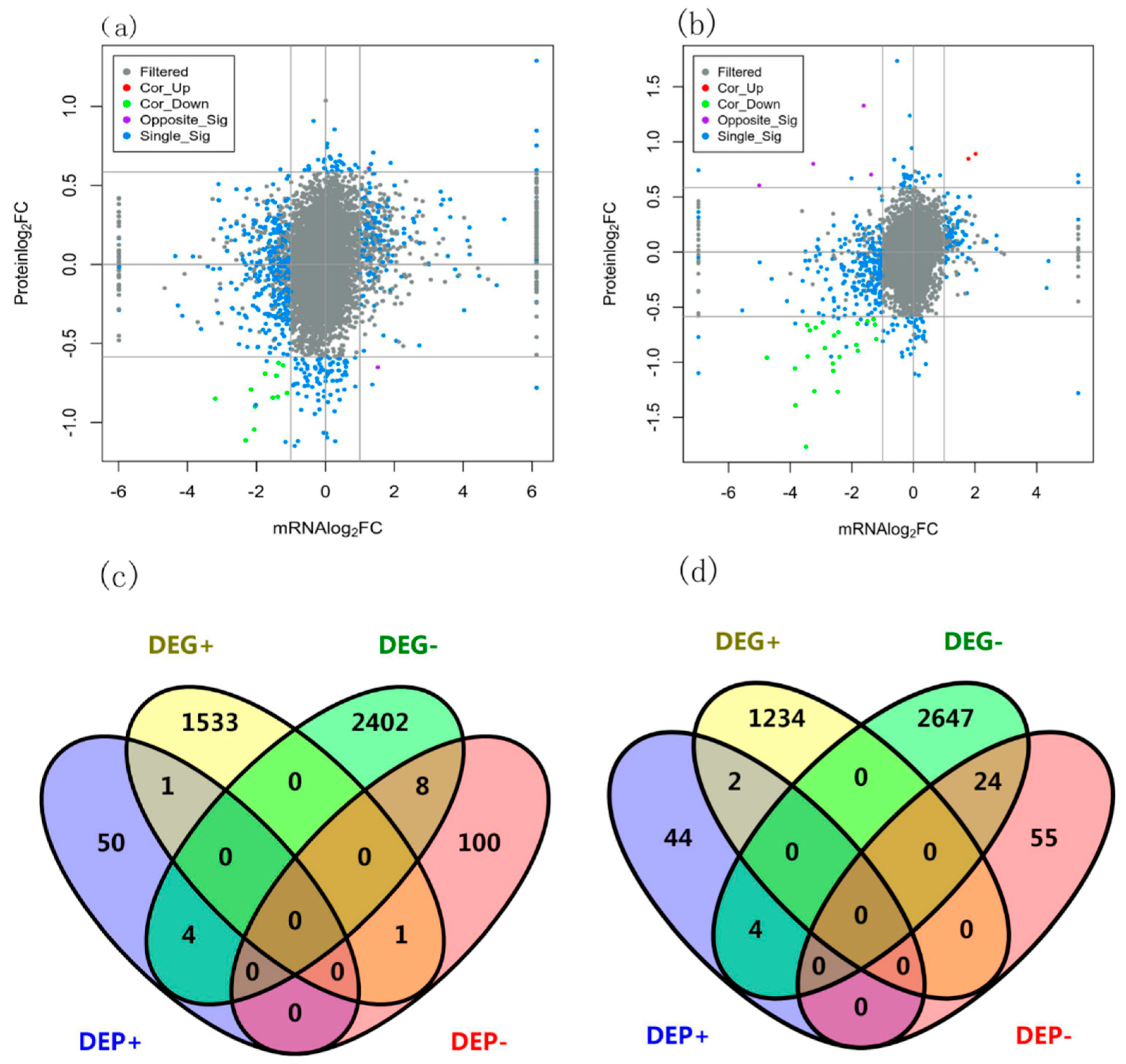
3.2. DEGs and DEPs of wheat under nitrogen deficit
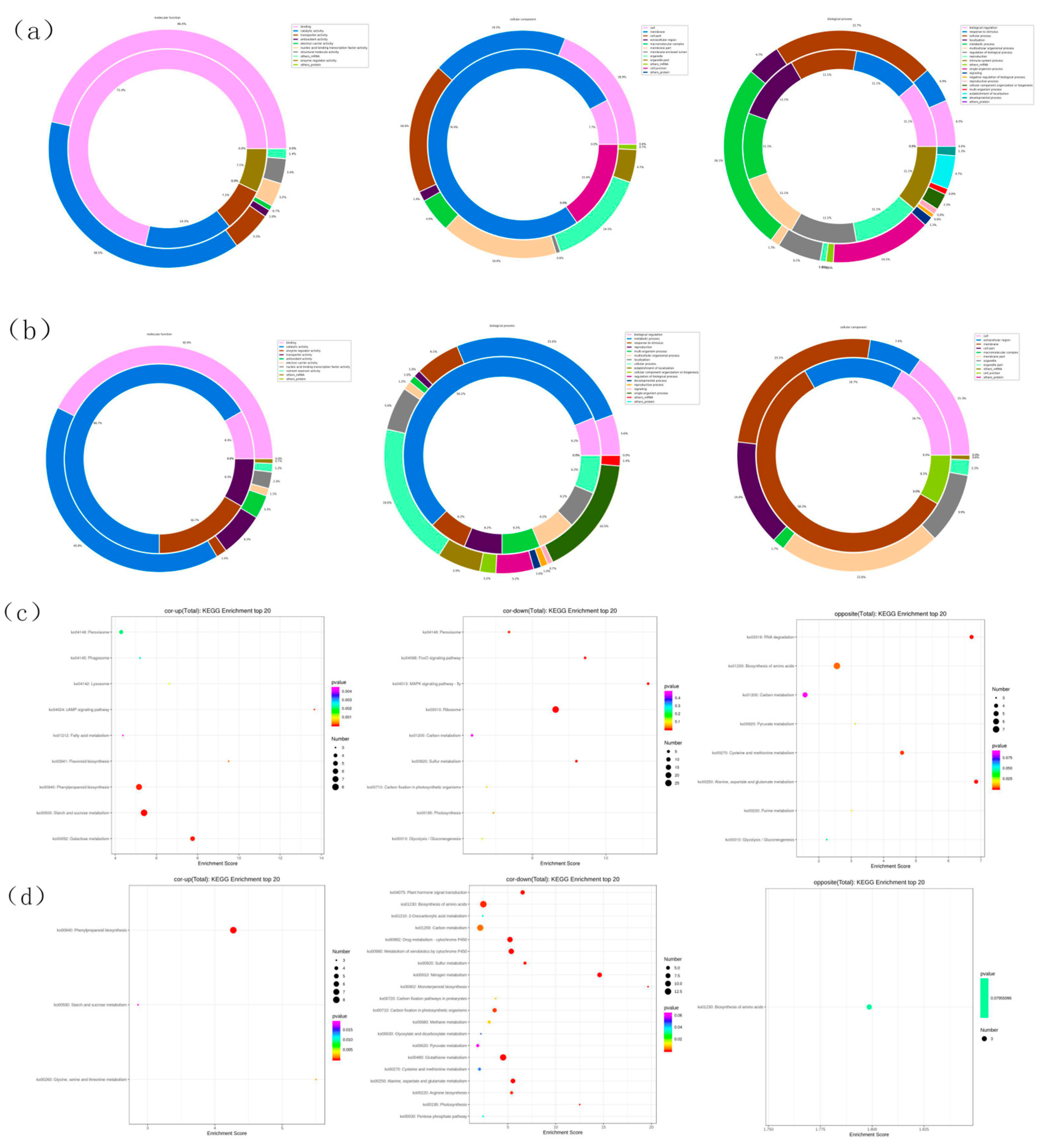
3.3. Pathway enrichment of wheat shoot and root under nitrogen deficit
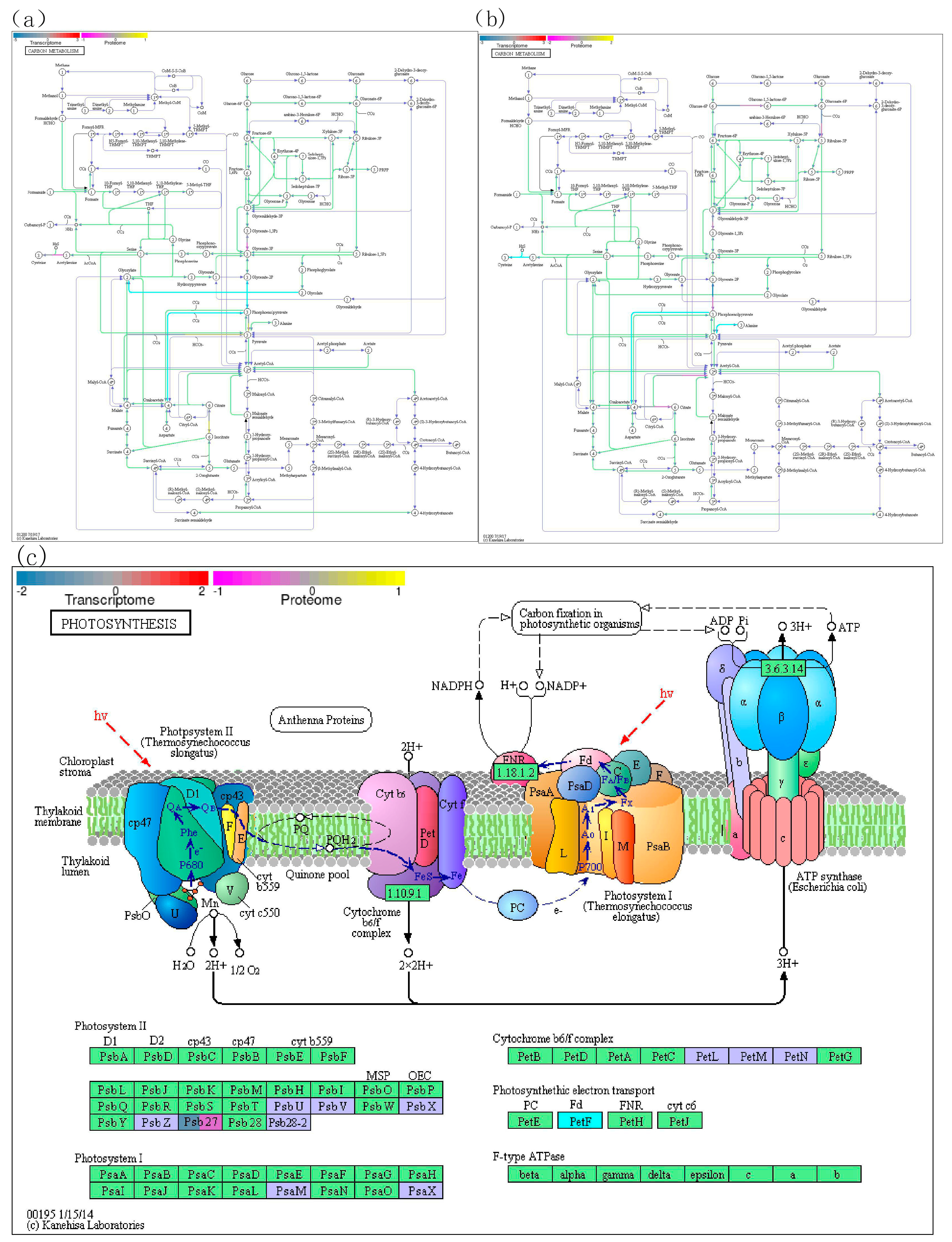
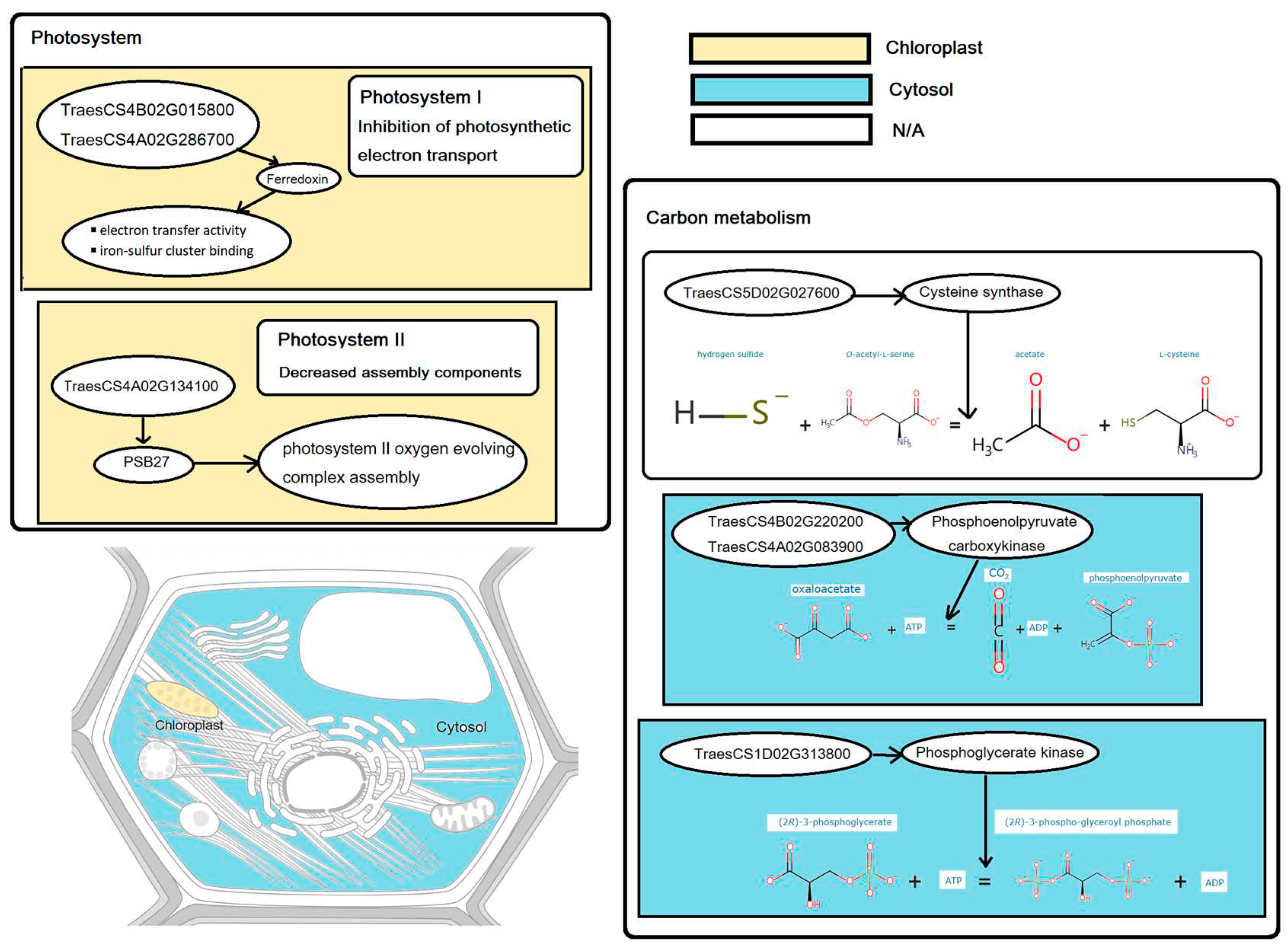
3.4. Differential genes in photosynthesis and carbon metabolism
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Catusse, J.; Job, C.; Job, D. Transcriptome- and proteome-wide analyses of seed germination. C R Biol. 2008, 331, 815–822. [Google Scholar] [CrossRef]
- Christian, H.; Lutterbey, M.C.; Lansing, H.; Meyer, T.; Schaewen, A.V. Defects in peroxisomal 6-phosphogluconate dehydrogenase isoform PGD2 prevent gametophytic interaction in Arabidopsis thaliana. Plant Physiol. 2016, 171. [Google Scholar]
- Dobakova, M.; Sobotka, R.; Tichy, M.; Komenda, J. Psb28 Protein Is Involved in the Biogenesis of the Photosystem II Inner Antenna CP47 (PsbB) in the Cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol. 2009, 149, 1076–1086. [Google Scholar]
- Du, J.L. Effects of Nitrogen Fertilizer Application on Groundwater Pollution and Analysis of Control Strategies. J Environ Sci Manag. 2019, 44, 22–27. [Google Scholar]
- Dustin, N.L.; Steven, G.R.; Michael, P.; Rick, W.; Derek, S.; Darryl, H.; Jonathan, G.; Kermit, R.; Christoph, H.B.; Jörg, B. Quantitative iTRAQ proteome and comparative transcriptome analysis of elicitor-induced Norway spruce (Picea abies) cells reveals elements of calcium signaling in the early conifer defense response. Proteomics. 2009, 9, 350–367. [Google Scholar]
- Efrati, A.; Tel-Vered, R.; Michaeli, D.; Nechushtai, R.; Willner, I. Cytochrome c-coupled photosystem I and photosystem II (PSI/PSII) photo-bioelectrochemical cells. Energ Environ Sci. 2013, 6, 2950–2956. [Google Scholar] [CrossRef]
- Fristedt, R.; Trotta, A.; Suorsa, M.; Nilsson, A.K.; Croce, R.; Aro, E.M.; Lundin, B.R. PSB33 sustains photosystem II D1 protein under fluctuating light conditions. J Exp Bot. 2017, 4281–4293. [Google Scholar] [CrossRef]
- Gao, J.; Wang, F.; Hu, H.; Jiang, S.; Muhammad, A.; Shao, Y.; Sun, C.; Tian, Z.; Jiang, D.; Dai, T. Improved leaf nitrogen reutilisation and Rubisco activation under short-term nitrogen-deficient conditions promotes photosynthesis in winter wheat (Triticum aestivum L.) at the seedling stage. Funct Plant Biol. 2018, 45. [Google Scholar] [CrossRef]
- Gerken, S.; Brettel, K.; Schlodder, E.; Witt, H.T. Optical characterization of the immediate electron donor to chlorophyll a+II in O 2-evolving photosystem II complexes Tyrosine as possible electron carrier between chlorophyll aII and the water-oxidizing manganese complex. Febs Lett. 1988, 237, 69–75. [Google Scholar] [CrossRef]
- Greenway, H.; Sims, A.P. Effects of High Concentrations of KC1 and NaCl on Responses of Malate Dehydrogenase (Decarboxylating) to Malate and Various Inhibitors. Funct Plant Biol. 1974, 1, 15–29. [Google Scholar] [CrossRef]
- Honoki, R.; Ono, S.; Oikawa, A.; Saito, K.; Masuda, S. Significance of accumulation of the alarmone (p)ppGpp in chloroplasts for controlling photosynthesis and metabolite balance during nitrogen starvation in Arabidopsis. Photosynth Res. 2018, 135, 299–308. [Google Scholar] [CrossRef] [PubMed]
- Ji, W.; Xiao, J.; Toor, G.S.; Li, Z. Nitrate-nitrogen transport in streamwater and groundwater in a loess covered region: Sources, drivers, and spatiotemporal variation. Sci Total Environ. 2020. [Google Scholar] [CrossRef] [PubMed]
- Jiao, C.; Gu, Z. iTRAQ-based proteomic analysis reveals changes in response to UV-B treatment in soybean sprouts. Food Chem. 2019, 275, 467–473. [Google Scholar] [CrossRef]
- Kanehisa, M.; Furumichi, M.; Tanabe, M.; Sato, y.; Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017, 45, 353–361. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Sato, Y.; Kawashima, M.; Furumichi, M.; Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016, 44, D457–D462. [Google Scholar] [CrossRef] [PubMed]
- Krzysztof, G.V.M.; Ramesh, A.N.; Melkozernov; Su, L. Excitation Dynamics in the Core Antenna of PS I from Chlamydomonas reinhardtii CC 2696 at Room Temperature. J Phys Chem B. 2001, 105, 11498–11506.
- Lan, G.; Jiao, C.; Wang, G.; Sun, Y.; Sun, Y. Effects of dopamine on growth, carbon metabolism, and nitrogen metabolism in cucumber under nitrate stress. Sci Hortic-amsterdam. 2020, 260, 108790. [Google Scholar] [CrossRef]
- Li, C.; Wu, G.; Fang, R.; Huang, X.; Ren, W. L-Cysteine metabolism and its nutritional implications. Mol Nutr Food Res. 2016, 60, 134–146. [Google Scholar]
- Li, N.; Zhao, J.; Warren, P.V.; Warden, J.T.; Bryant, D.A.; Golbeck, J.H. PsaD is required for the stable binding of PsaC to the photosystem I core protein of Synechococcus sp. PCC 6301. Biochemistry 1991, 30, 7863–7872. [Google Scholar]
- Liu, F.; Qian, H.; Shi, Z.; Wang, H. Long-term monitoring of hydrochemical characteristics and nitrogen pollution in the groundwater of Yinchuan area, Yinchuan basin of northwest China. Enviro Earth Sci. 2019, 78, 1–15. [Google Scholar] [CrossRef]
- Liu, S.; Xie, Z.; Zeng, Y.; Liu, B.; Li, R.; Wang, Y.; Wang, L.; Qin, P.; Jia, B.; Xie, J. Effects of anthropogenic nitrogen discharge on dissolved inorganic nitrogen transport in global rivers. Global Change Biol. 2019, 25, 1493–1513. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yin, C.; Xiang, L.; Jiang, W.; Xu, S.; Mao, Z. Transcription strategies related to photosynthesis and nitrogen metabolism of wheat in response to nitrogen deficiency. BMC Plant Biol. 2020, 20, 448. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yuan, Y.; Martinez, C.; Babu, R.; Antonio, E. Identification of QTL for early vigor and leaf senescence across two tropical maize doubled haploid populations under nitrogen deficiency conditions. Euphytica. 2020, 216, 1–14. [Google Scholar] [CrossRef]
- Lukáš, N.; Dmitry, S.; Egbert, J.B.; Petr, I.; Roman, K. Structural variability of plant photosystem II megacomplexes in thylakoid membranes. Global Change Biol. 2016, 89, 104–111. [Google Scholar]
- Lv, X.; Zhang, Y.; Hu, L.; Zhang, Y.; Zhang, B.; Xia, H.; Du, W.; Fan, S.; Kong, L. Low-Nitrogen Stress Stimulates Lateral Root Initiation and Nitrogen Assimilation in Wheat: Roles of Phytohormone Signaling. J Plant Growth Regul 2020. [Google Scholar] [CrossRef]
- Lyu, L.; Bi, Y.; Li, S.; Xue, H.; Li, Y.; Pruskyab, D.B. Sodium silicate prime defense responses in harvested muskmelon by regulating mitochondrial energy metabolism and reactive oxygen species production. Food Chem. 2019, 289, 369–376. [Google Scholar] [CrossRef] [PubMed]
- Milo, R. What governs the reaction center excitation wavelength of photosystems I and II Photosynth Res. 2009, 101, 59–67. 101.
- Ouyang, L.; Pei, H.; Xu, Z. Low nitrogen stress stimulating the indole-3-acetic acid biosynthesis of Serratia sp. ZM is vital for the survival of the bacterium and its plant growth-promoting characteristic. Arch Microbrol. 2016, 199, 425–432. [Google Scholar] [CrossRef] [PubMed]
- Post, A.F.; Arnold, V.; Mur, L.R. Regulation of cyanobacterial photosynthesis determined from variable fluorescence yields of photosystem II. Fems Microbiol Lett. 2010, 35, 129–133. [Google Scholar] [CrossRef]
- Qa, X.U.; La, Y.; Chitnis, V.P.; Chitnis, P.R. Function and organization of photosystem I in a cyanobacterial mutant strain that lacks PsaF and PsaJ subunits. J Biol Chem. 1994, 269, 3205–3211. [Google Scholar]
- Rosa-Tellez, S.; Djoro Anoman, A.; Flores-Tornero, M.; Toujani, W.; Alseek, S.; Fernie, A.R.; Nebauer, S.G.; Munoz-Bertomeu, J.; Segura, J.; Ros, R. Phosphoglycerate kinases are co-regulated to adjust metabolism and to optimize growth. Photosynth Res. 2018, 176, 1182. [Google Scholar] [CrossRef]
- Sharkey, T.D.; Weise, S.E. The glucose 6-phosphate shunt around the Calvin-Benson cycle. J Exp Bot. 2017, 68, 4731. [Google Scholar] [CrossRef] [PubMed]
- Sihvonen, M.; Pihlainen, S.; Lai, T.Y.; Salo, T.; Hyytiinen, K. Crop production, water pollution, or climate change mitigation—Which drives socially optimal fertilization management most. Agr Syst. 2021, 186. [Google Scholar] [CrossRef]
- Sonoike, K. Degradation of psaB gene product, the reaction center subunit of photosystem I, is caused during photoinhibition of photosystem I: Possible involvement of active oxygen species. Plant Sci. 1996, 115, 157–164. [Google Scholar] [CrossRef]
- Sonoike, K.; Kamo, M.; Hihara, Y.; Hiyama, T.; Enami, I. The mechanism of the degradation of psaB gene product, one of the photosynthetic reaction center subunits of Photosystem I, upon photoinhibition. Photosynth Res. 1997, 53, 55–63. [Google Scholar] [CrossRef]
- Sun, B.; Zhi, W.; Fei, Z. Molecular cloning of kuruma shrimp Marsupenaeus japonicus phosphopyruvate hydratase and its role in infection by white spot syndrome virus and Vibrio alginolyticus. Aquaculture. 2016, 455, 87–96. [Google Scholar] [CrossRef]
- Thomas, P.D. The Gene Ontology and the Meaning of Biological Function. Methods Mol Biol. 2017, 1446, 15. [Google Scholar]
- Thornton, L.E.; Ohkawa, H.; Roose, J.L.; Kashino, Y.; Keren, N.; Pakrasia, H.B. Homologs of Plant PsbP and PsbQ Proteins Are Necessary for Regulation of Photosystem II Activity in the Cyanobacterium Synechocystis 6803. Plant Cell. 2004, 16, 2164–2175. [Google Scholar] [CrossRef]
- Torabi, S.; Umate, P.; Manavski, N.; Plöchinger, M.; Kleinknecht, L.; Bogireddi, H.; Herrmann, R.G.; Wanner, G.; Schröder, W.P.; Meurer, J. PsbN Is Required for Assembly of the Photosystem II Reaction Center inNicotiana tabacum. Plant Cell. 2014, 26, 1183–1199. [Google Scholar] [CrossRef]
- Tsuchiya, K.; Tajima, H.; Kuwae, T.; Takeshima, T.; Ryoichi, I. Pro-apoptotic protein glyceraldehyde-3-phosphate dehydrogenase promotes the formation of Lewy body-like inclusions. Eur J Neurosci. 2015, 21, 317–326. [Google Scholar] [CrossRef]
- Walker, R.P.; Battistelli, A.; Moscatello, S.; Chen, Z.H.; Leegood, R.C.; Famiani, F. Phosphoenolpyruvate carboxykinase in cherry (L.) fruit during development. Ecol Eng. 2015, 52, 96–103. [Google Scholar]
- Wang, C.; Zheng, L.; Tang, Z.; Sun, S.; Ma, J.F.; Huang, X.Y.; Zhao, F.J. OASTL-A1 functions as a cytosolic cysteine synthase and affects arsenic tolerance in rice. J Exp Bot. 2020, 71, 3678–3689. [Google Scholar] [CrossRef] [PubMed]
- William, C.U. Modification of the Kjeldahl nitrogen determination method. Anal Chem. 1971, 43, 800–801. [Google Scholar]
- Wu, J.; Xu, Z.; Zhang, Y.; Chai, L.; Yi, H.; Deng, X. An integrative analysis of the transcriptome and proteome of the pulp of a spontaneous late-ripening sweet orange mutant and its wild type improves our understanding of fruit ripening in citrus. J Exp Bot. 2014, 65, 1651–1671. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Ying, H.; Xue, Y.; Zheng, H.; Zhang, Q.; Cui, Z. Calculating socially optimal nitrogen (N) fertilization rates for sustainable N management in China. Sci Total Environ. 2019, 688, 1162–1171. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Wang, J.; Xie, S.; Zhao, M.; Nie, L.; Zheng, Y.; Zhu, S.; Hou, J.; Chen, G.; Wang, C. Comparative Proteomics Indicates That Redox Homeostasis Is Involved in High- and Low-Temperature Stress Tolerance in a Novel Wucai (Brassica campestris L.) Genotype. Int J Mol Sci. 2019, 20, 3760. [Google Scholar] [CrossRef]
- Zhang, F.; Chen, X.; Vitousek, P. Chinese agriculture: An experiment for the world. Nature 2013, 497, 33–35. [Google Scholar] [CrossRef]
| Position | Pathway | Gene id | Gene name | Protein name | Chromosomal location | Subcellular localization |
|---|---|---|---|---|---|---|
| Shoot | Photosynthesis | TraesCS4A02G134100 | PSB27-1 | PSB27 | 4A:182128608-182129419 | Chloroplast |
| TraesCS4B02G015800 | PetF | Ferredoxin | 4B:11734874-11735616 | Chloroplast | ||
| TraesCS4A02G286700 | PetF | Ferredoxin | 4A:591919576-591921832 | Chloroplast | ||
| Carbon metabolism | TraesCS5D02G027600 | CYS1 | Cysteine synthase | 5D:25277866-25282033 | N/A | |
| TraesCS1D02G313800 | N/A | Phosphoglycerate kinase | 1D:409048850-409052150 | Cytosol | ||
| TraesCS4B02G220200 | N/A | Phosphoenolpyruvate carboxykinase | 4B:463208678-463213325 | Cytosol | ||
| TraesCS4A02G083900 | N/A | Phosphoenolpyruvate carboxykinase | 4A:88427962-88432958 | Cytosol | ||
| Root | Carbon metabolism | TraesCS2B02G080000 | G6PD2 | Glucose-6-phosphate 1-dehydrogenase | 2B:44424162-44429300 | N/A |
| TraesCS1A02G357200 | G6PGH2 | 6-phosphogluconate dehydrogenase, decarboxylating | 1A:539899281-539901954 | Cytosol | ||
| TraesCS7D02G309500 | GAPC | Glyceraldehyde-3-phosphate dehydrogenase | 7D:392821248-392824662 | N/A | ||
| TraesCS5D02G027600 | CYS1 | Cysteine synthase | 5D:25277866-25282033 | N/A | ||
| TraesCS4A02G401700 | CYSK | N/A | 4A:675600394-675603117 | Other locations | ||
| TraesCS7A02G075600 | ENO1 | Phosphopyruvate hydratase | 7A:41439093-41442254 | Cytosol | ||
| TraesCS5B02G179800 | N/A | Phosphoenolpyruvate carboxylase | 5B:327603933-327609868 | Other locations | ||
| TraesCS7D02G333900 | PEPC | Phosphoenolpyruvate carboxylase | 7D:425502517-425509832 | Other locations | ||
| TraesCS7B02G237900 | PEPC | Phosphoenolpyruvate carboxylase | 7B:443292417-443299286 | Other locations | ||
| TraesCS1B02G102700 | N/A | Aminotran_1_2 domain-containing protein | 1B:112748629-112753960 | N/A | ||
| TraesCS1A02G085600 | N/A | Aminotran_1_2 domain-containing protein | 1A:71689760-71695155 | N/A | ||
| TraesCS5D02G422000 | CSY3 | Citrate synthase | 5D:482102459-482108460 | Mitochondrion |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




