Submitted:
25 June 2024
Posted:
26 June 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
- The current study focuses on protein sequential data rather than image data.
- The most frequently mutated genes that were responsible for chronic myeloid leukemia were discovered through a literature review.
- Datasets were formulated from the most frequently muted gene data.
- Features were extracted through physicochemical properties of Amino Acid composition, Pseudo Amino Acid Composition, and di-peptide composition.
- The study focuses on enhancing early-stage prediction to improve patient recovery prospects significantly.
- Our proposed solution encompasses a user-friendly web application dashboard that presents an invaluable tool for early CML diagnosis, offering a deploy-able asset within healthcare institutions and hospitals.
2. Literature Review
3. Materials and Methods
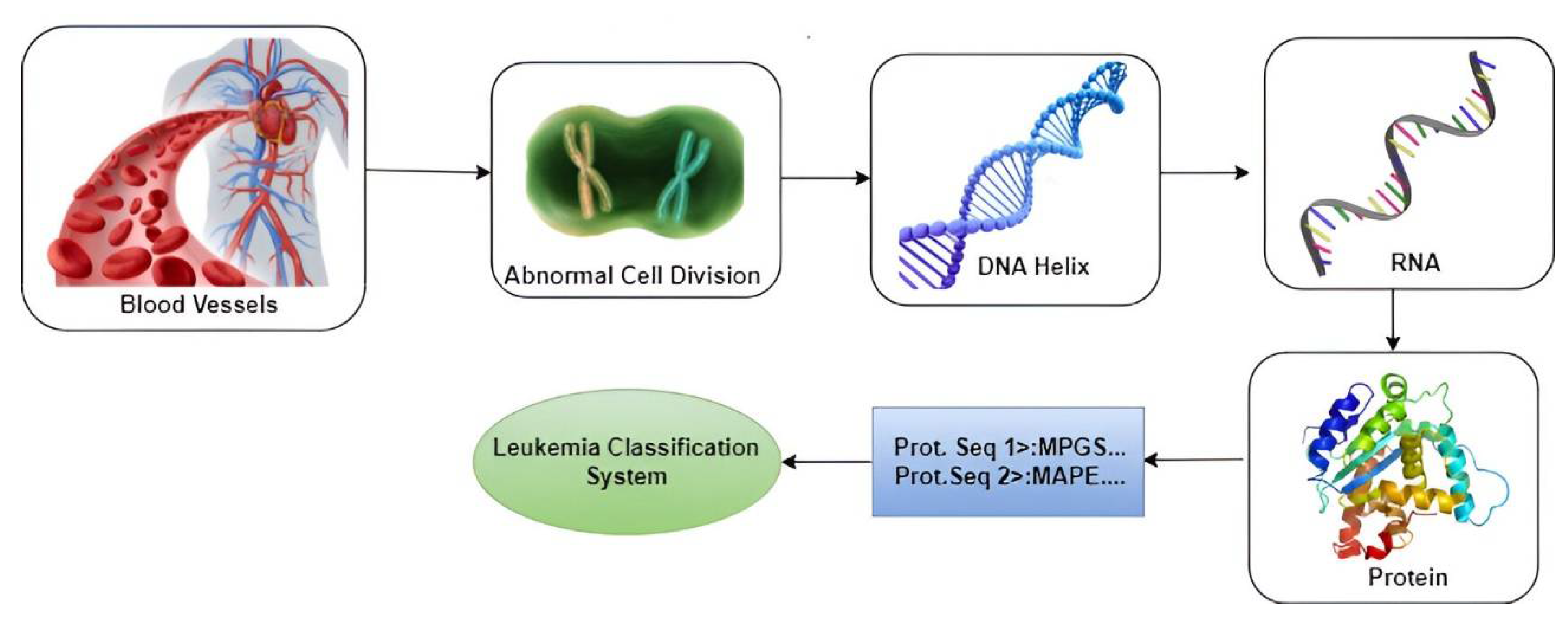
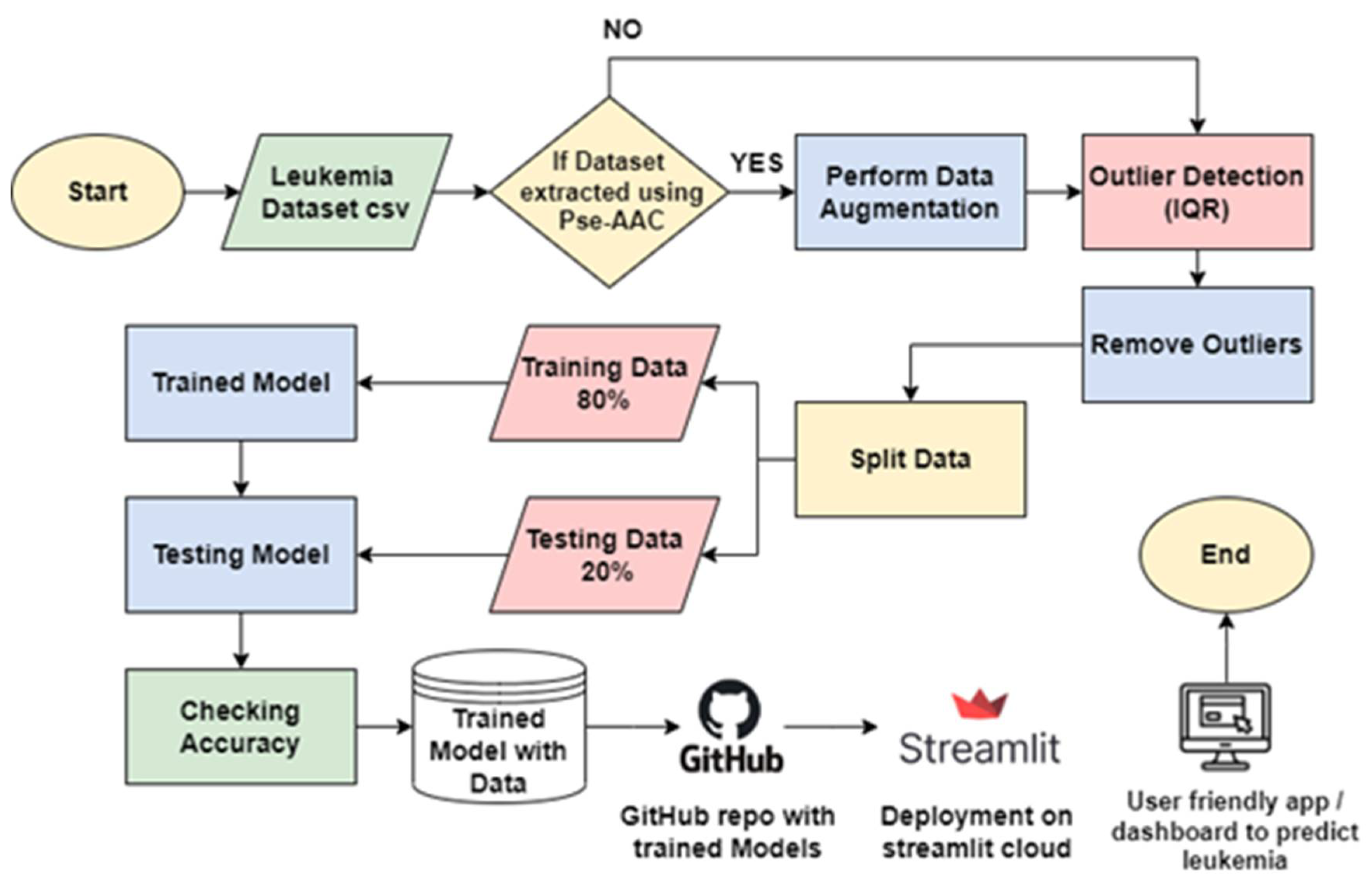
3.1. Block Diagram
3.2. Dataset Collection
3.2.1. Fasta Format
3.2.2. Sample of Protein Sequence (HSP90)
3.3. Feature Extraction
3.3.1. Amino Acid Composition
3.3.2. Pseudo Amino Acid Composition
3.3.3. Di-peptide Composition
3.3.4. Data Augmentation
4. Development of Individual Classifiers
4.1. Support Vector Machine
4.2. Random Forest
4.3. K-Nearest Neighbor (KNN)
4.4. Naïve Bayes
4.5. XGBoost
4.6. Logistic Regression
5. Results and Discussion
5.1. Results on Pseudo Amino Acid Composition (Pse-AAC) Data
5.2. Accuracy Result on Amino Acid Composition (AAC) Data
5.3. Accuracy Results on Di-Peptide Composition (DPC)
5.5. Machine Learning Based Dashboard
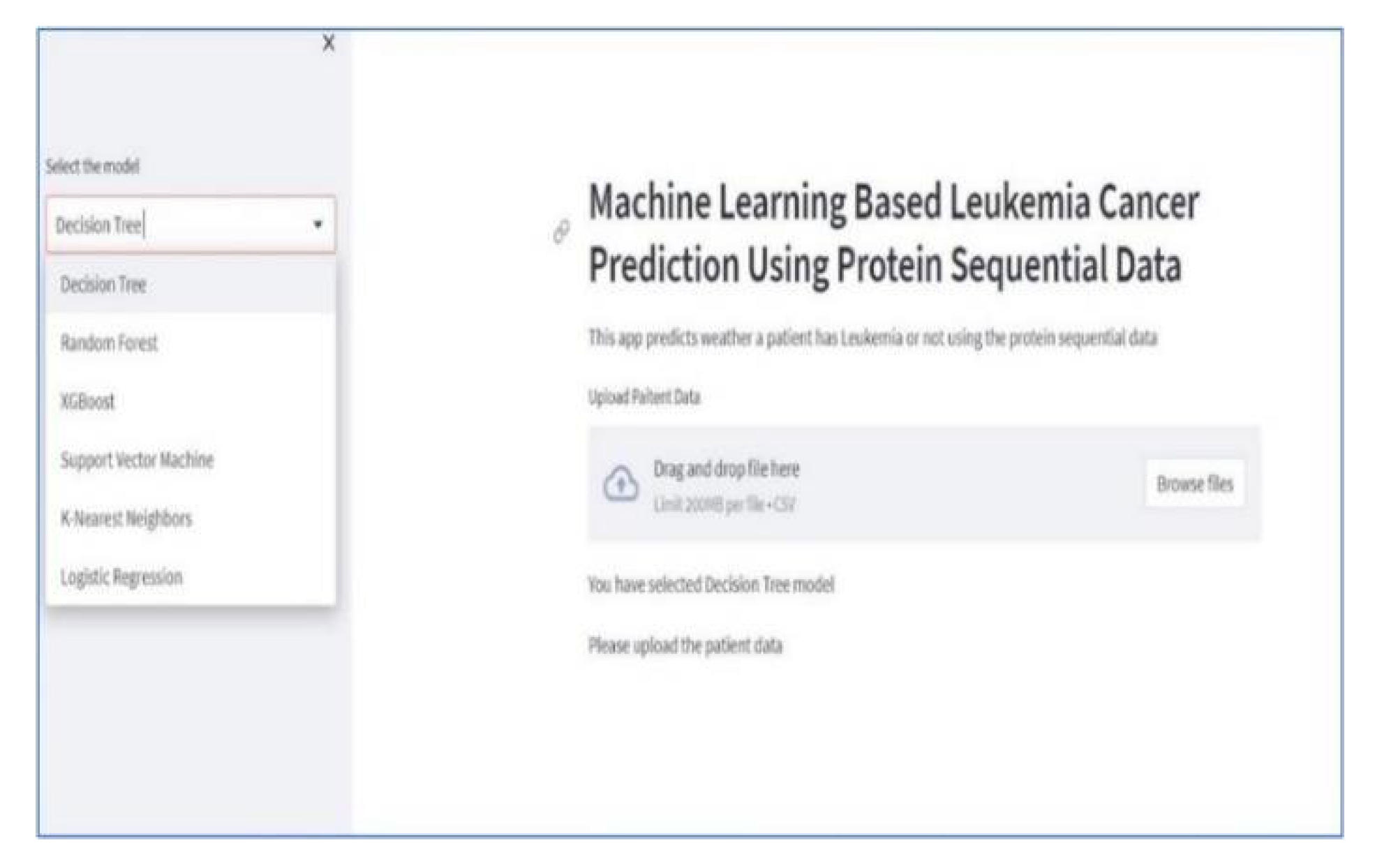
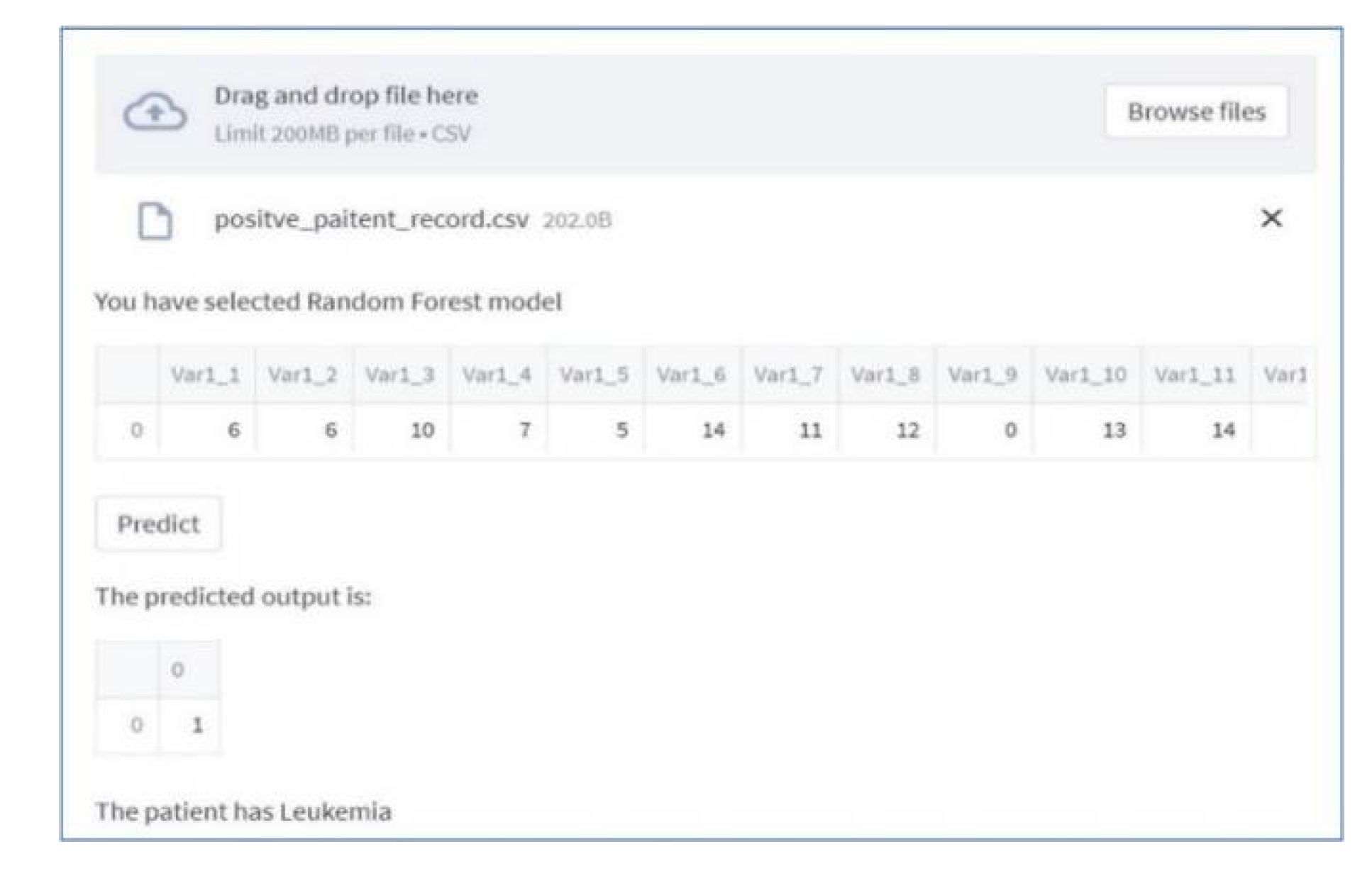
5. Conclusion
Funding
Acknowledgments
Conflicts of Interest
References
- R. L. Siegel, K. D. Miller, H. E. Fuchs, and A. Jemal, "Cancer statistics, 2021," Ca Cancer J Clin, vol. 71, no. 1, pp. 7-33 %@ 1542-4863, 2021.
- N. Bibi, M. Sikandar, I. Ud Din, A. Almogren, and S. Ali, "IoMT-based automated detection and classification of leukemia using deep learning," Journal of healthcare engineering, vol. 2020, pp. 1-12 %@ 2040-2309, 2020.
- I. IafRoC, "Leukaemia Source: Globocan 2020 2020 [Available from: https://gco. iarc. fr/today/data/factsheets/cancers/36-Leukaemia-fact-sheet. pdf," ed: Accessed, 2022.
- C. R. Munteanu, A. L. Magalhães, E. Uriarte, and H. González-Díaz, "Multi-target QPDR classification model for human breast and colon cancer-related proteins using star graph topological indices," Journal of theoretical biology, vol. 257, no. 2, pp. 303-311 %@ 0022-5193, 2009. [CrossRef]
- R. G. Ramani and S. G. Jacob, "Improved classification of lung cancer tumors based on structural and physicochemical properties of proteins using data mining models," PloS one, vol. 8, no. 3, pp. e58772 %@ 1932-6203, 2013. [CrossRef]
- J.-Y. Yang et al., "Predicting time to ovarian carcinoma recurrence using protein markers," The Journal of clinical investigation, vol. 123, no. 9, pp. 3740-3750 %@ 0021-9738, 2013.
- H. Mohamed et al., "Automated detection of white blood cells cancer diseases," 2018: IEEE, pp. 48-54 %@ 1538650835.
- S. Kumar, S. Mishra, P. Asthana, and Pragya, "Automated detection of acute leukemia using k-mean clustering algorithm," 2018: Springer, pp. 655-670 %@ 9811037728.
- R. Sharma and R. Kumar, "A novel approach for the classification of leukemia using artificial bee colony optimization technique and back-propagation neural networks," 2019: Springer, pp. 685-694 %@ 9811312168.
- G. Jothi, H. H. Inbarani, A. T. Azar, and K. R. Devi, "Rough set theory with Jaya optimization for acute lymphoblastic leukemia classification," Neural Computing and Applications, vol. 31, pp. 5175-5194 %@ 0941-0643, 2019. [CrossRef]
- Z. Moshavash, H. Z. Moshavash, H. Danyali, and M. S. Helfroush, "An automatic and robust decision support system for accurate acute leukemia diagnosis from blood microscopic images," Journal of digital imaging, vol. 31, pp. 702-717 %@ 0897-1889, 2018. [CrossRef]
- D. Umamaheswari and S. Geetha, "A framework for efficient recognition and classification of acute lymphoblastic leukemia with a novel customized-KNN classifier," Journal of computing and information technology, vol. 26, no. 2, pp. 131-140 %@ 1330-1136, 2018. [CrossRef]
- O. Gal, N. Auslander, Y. Fan, and D. Meerzaman, "Predicting complete remission of acute myeloid leukemia: machine learning applied to gene expression," Cancer informatics, vol. 18, pp. 1176935119835544 %@ 1176-9351, 2019. [CrossRef]
- E. Bostanci, E. Kocak, M. Unal, M. S. Guzel, K. Acici, and T. Asuroglu, "Machine learning analysis of RNA-seq data for diagnostic and prognostic prediction of colon cancer," Sensors, vol. 23, no. 6, pp. 3080 %@ 1424-8220, 2023. [CrossRef]
- F. Hosseinzadeh, A. H. KayvanJoo, M. Ebrahimi, and B. Goliaei, "Prediction of lung tumor types based on protein attributes by machine learning algorithms," SpringerPlus, vol. 2, pp. 1-14, 2013. [CrossRef]
- P. Dhakal, H. Tayara, and K. T. Chong, "An ensemble of stacking classifiers for improved prediction of miRNA–mRNA interactions," Computers in Biology and Medicine, vol. 164, pp. 107242 %@ 0010-4825, 2023. [CrossRef]
- M. Albitar et al., "Bone Marrow-Based Biomarkers for Predicting aGVHD Using Targeted RNA Next Generation Sequencing and Machine Learning," Blood, vol. 138, pp. 2892 %@ 0006-4971, 2021. [CrossRef]
- W. Ahmad, M. Hameed, M. Bilal, and A. Majid, "ML-Pred-CLL: Machine Learning based prediction of Chronic Lymphocytic Leukemia using protein sequential data," 2022: IEEE, pp. 1-7 %@ 1665491035.
- J. He, X. Pu, M. Li, C. Li, and Y. Guo, "Deep convolutional neural networks for predicting leukemia-related transcription factor binding sites from DNA sequence data," Chemometrics and Intelligent Laboratory Systems, vol. 199, pp. 103976 %@ 0169-7439, 2020. [CrossRef]
- D. Rodríguez et al., "Mutations in CHD2 cause defective association with active chromatin in chronic lymphocytic leukemia," Blood, The Journal of the American Society of Hematology, vol. 126, no. 2, pp. 195-202 %@ 0006-4971, 2015. [CrossRef]
- A. Ashraf, Q. Zhao, W. H. Bangyal, and M. Iqbal, "Analysis of Brain Imaging Data for the Detection of Early Age Autism Spectrum Disorder Using Transfer Learning Approaches for Internet of Things," IEEE Transactions on Consumer Electronics, 2023. [CrossRef]
- R. Apweiler et al., "UniProt: the universal protein knowledgebase," Nucleic acids research, vol. 32, no. suppl_1, pp. D115-D119 %@ 0305-1048, 2004. [CrossRef]
- W. Bangyal, J. Ahmad, and Q. Abbas, "Recognition of off-line isolated handwritten character using counter propagation network," International Journal of Engineering and Technology, vol. 5, no. 2, p. 227, 2013. [CrossRef]
- L. Fu, B. Niu, Z. Zhu, S. Wu, and W. Li, "CD-HIT: accelerated for clustering the next-generation sequencing data," Bioinformatics, vol. 28, no. 23, pp. 3150-3152 %@ 1367-4803, 2012. [CrossRef]
- P.-M. Feng, H. Lin, and W. Chen, "Identification of antioxidants from sequence information using naive Bayes," Computational and mathematical methods in medicine, vol. 2013 %@ 1748-670X, 2013. [CrossRef]
- P.-M. Feng, H. Ding, W. Chen, and H. Lin, "Naive Bayes classifier with feature selection to identify phage virion proteins," Computational and mathematical methods in medicine, vol. 2013 %@ 1748-670X, 2013. [CrossRef]
- J. Jia, Z. Liu, X. Xiao, B. Liu, and K.-C. Chou, "pSuc-Lys: predict lysine succinylation sites in proteins with PseAAC and ensemble random forest approach," Journal of theoretical biology, vol. 394, pp. 223-230 %@ 0022-5193, 2016.
- W.-Z. Lin, J.-A. Fang, X. Xiao, and K.-C. Chou, "iDNA-Prot: identification of DNA binding proteins using random forest with grey model," PloS one, vol. 6, no. 9, pp. e24756 %@ 1932-6203, 2011.
- A.M. Ali and M. A. Mohammed, "A Comprehensive Review of Artificial Intelligence Approaches in Omics Data Processing: Evaluating Progress and Challenges," International Journal of Mathematics, Statistics, and Computer Science, vol. 2, pp. 114-167, 2024. [CrossRef]
- Z. H. Arif and K. Cengiz, "Severity Classification for COVID-19 Infections based on Lasso-Logistic Regression Model," International Journal of Mathematics, Statistics, and Computer Science, vol. 1, pp. 25-32, 2022. [CrossRef]
- K. Qu, K. Han, S. Wu, G. Wang, and L. Wei, "Identification of DNA-binding proteins using mixed feature representation methods," Molecules, vol. 22, no. 10, pp. 1602 %@ 1420-3049, 2017. [CrossRef]
- K. V. Khajapeer and R. Baskaran, "Hsp90 inhibitors for the treatment of chronic myeloid leukemia," Leukemia research and treatment, vol. 2015 %@ 2090-3219, 2015.
- R. Alves et al., "Alvespimycin Inhibits Heat Shock Protein 90 and Overcomes Imatinib Resistance in Chronic Myeloid Leukemia Cell Lines," Molecules, vol. 28, no. 3, pp. 1210 %@ 1420-3049, 2023. [CrossRef]
- L. W. Ellisen, "PARP inhibitors in cancer therapy: promise, progress, and puzzles," Cancer cell, vol. 19, no. 2, pp. 165-167 %@ 1535-6108, 2011.
- Y. Liu, H. Song, H. Song, X. Feng, C. Zhou, and Z. Huo, "Targeting autophagy potentiates the anti-tumor effect of PARP inhibitor in pediatric chronic myeloid leukemia," AMB Express, vol. 9, pp. 1-9, 2019. [CrossRef]
- D. Kaloni, S. T. Diepstraten, A. Strasser, and G. L. Kelly, "BCL-2 protein family: Attractive targets for cancer therapy," Apoptosis, vol. 28, no. 1-2, pp. 20-38 %@ 1360-8185, 2023. [CrossRef]
- T. K. Ko, C. T. H. Chuah, J. W. J. Huang, K.-P. Ng, and S. T. Ong, "The BCL2 inhibitor ABT-199 significantly enhances imatinib-induced cell death in chronic myeloid leukemia progenitors," Oncotarget, vol. 5, no. 19, p. 9033, 2014. [CrossRef]
- L. Zhou et al., "Post-translational modifications on the retinoblastoma protein," Journal of Biomedical Science, vol. 29, no. 1, pp. 1-16 %@ 1423-0127, 2022. [CrossRef]
- D.-D. Yin et al., "Notch signaling inhibits the growth of the human chronic myeloid leukemia cell line K562," Leukemia research, vol. 33, no. 1, pp. 109-114 %@ 0145-2126, 2009. [CrossRef]
- Y.-D. Cai and K.-C. Chou, "Predicting subcellular localization of proteins in a hybridization space," Bioinformatics, vol. 20, no. 7, pp. 1151-1156 %@ 1367-4811, 2004. [CrossRef]
- K.-C. Chou, "Impacts of bioinformatics to medicinal chemistry," Medicinal chemistry, vol. 11, no. 3, pp. 218-234 %@ 1573-4064, 2015. [CrossRef]
- K. C. Chou, "Prediction of protein cellular attributes using pseudo-amino acid composition," Proteins: Structure, Function, and Bioinformatics, vol. 43, no. 3, pp. 246-255 %@ 0887-3585, 2001. [CrossRef]
- Y. D. Khan, F. Ahmad, and M. W. Anwar, "A neuro-cognitive approach for iris recognition using back propagation," World Applied Sciences Journal, vol. 16, no. 5, pp. 678-685 %@ 1818-4952, 2012.
- A.S. o. C. O. (ASCO). "Genes and Cancer." Cancer.net. https://www.cancer.net/navigating-cancer-care/cancer-basics/genetics/genes-and-cancer (accessed 11, 2023).
- A.H. Butt, S. A. Khan, H. Jamil, N. Rasool, and Y. D. Khan, "A prediction model for membrane proteins using moments based features," BioMed research international, vol. 2016 %@ 2314-6133, 2016. [CrossRef]
- Y. D. Khan, F. Ahmed, and S. A. Khan, "Situation recognition using image moments and recurrent neural networks," Neural Computing and Applications, vol. 24, pp. 1519-1529 %@ 0941-0643, 2014. [CrossRef]
- G. Hu, Y. Zheng, L. Abualigah, and A. G. Hussien, "DETDO: An adaptive hybrid dandelion optimizer for engineering optimization," Advanced Engineering Informatics, vol. 57, p. 102004, 2023. [CrossRef]
- L. Abualigah, S. Ekinci, D. Izci, and R. A. Zitar, "Modified elite opposition-based artificial hummingbird algorithm for designing FOPID controlled cruise control system," Intelligent Automation & Soft Computing, 2023. [CrossRef]
- A.H. Butt, N. Rasool, and Y. D. Khan, "A treatise to computational approaches towards prediction of membrane protein and its subtypes," The Journal of membrane biology, vol. 250, pp. 55-76 %@ 0022-2631, 2017. [CrossRef]
- W. Bangyal, J. Ahmad, and Q. Abbas, "Analysis of learning rate using CPN algorithm for hand written character recognition application," International Journal of Engineering and Technology, vol. 5, no. 2, p. 187, 2013. [CrossRef]
- Y. D. Khan, S. A. Khan, F. Ahmad, and S. Islam, "Iris recognition using image moments and k-means algorithm," The Scientific World Journal, vol. 2014 %@ 2356-6140, 2014. [CrossRef]
- M. Sugiyama, Introduction to statistical machine learning. Morgan Kaufmann, 2015.
- S. Theodoridis, Machine learning: a Bayesian and optimization perspective. Academic press, 2015.
- V. Vapnik, The nature of statistical learning theory. Springer science & business media, 1999.
- P. E. Hart, D. G. Stork, and R. O. Duda, Pattern classification. Wiley Hoboken, 2000.
- W. H. Bangyal et al., "Detection of fake news text classification on COVID-19 using deep learning approaches," Computational and mathematical methods in medicine, vol. 2021, pp. 1-14, 2021. [CrossRef]
- O.A. Montesinos López, A. Montesinos López, and J. Crossa, Multivariate statistical machine learning methods for genomic prediction. Springer Nature, 2022.
- Y. Jiao and P. Du, "Performance measures in evaluating machine learning based bioinformatics predictors for classifications," Quantitative Biology, vol. 4, pp. 320-330 %@ 2095-4689, 2016. [CrossRef]
- T. Fawcett, "ROC graphs: Notes and practical considerations for researchers," Machine learning, vol. 31, no. 1, pp. 1-38, 2004.

| Name of Algorithm | Accuracy | F1-Score | Recall | Specificity |
|---|---|---|---|---|
| Support Vector Classifier | 92~94% | 91~92% | 91~93% | 92~94% |
| Extreme Gradient Boost | 79~85% | 63~70% | 51~55% | 92~94% |
| Logistic Regression | 66~69% | 10~20% | 6~10% | 97~98% |
| Decision Tree | 81~84% | 73~76% | 74~76% | 84~86% |
| Random Forest | 87~91% | 85~87% | 80~83% | 96~97% |
| K Nearest Neighbor | 82~86% | 72~74% | 61~64% | 93~95% |
| Name of Algorithm | Confusion Matrix | |
|---|---|---|
| Support Vector Classifier | True Negative = 424 False Negative = 14 |
False Positive = 28 True Positive = 211 |
| Extreme Gradient Boost | True Negative = 26159 False Negative = 3435 |
False Positive = 2271 True Positive = 10890 |
| Logistic Regression | True Negative = 25817 False Negative = 11010 |
False Positive = 2849 True Positive = 3445 |
| Decision Tree | True Negative = 24388 False Negative = 3803 |
False Positive = 4278 True Positive = 10652 |
| Random Forest | True Negative = 28014 False Negative = 2753 |
False Positive = 808 True Positive = 11546 |
| K Nearest Neighbor | True Negative = 419 False Negative = 95 |
False Positive = 23 True Positive = 140 |
| Name of Algorithm | Accuracy | F1-Score | Recall | Specificity |
|---|---|---|---|---|
| Support Vector Classifier | 54.95% | 14.3% | 0.7% | 100% |
| Extreme Gradient Boost | 56.8% | 52.9% | 45.9% | 69% |
| Logistic Regression | 51.1% | 27.6% | 19.1% | 81.7% |
| Decision Tree | 54.4% | 52.25% | 52.9% | 55.8% |
| Random Forest | 50.6% | 41.1% | 35.4% | 64.9% |
| K Nearest Neighbor | 54.2% | 54.8% | 57% | 51% |
| Name of Algorithm | Confusion Matrix | |
|---|---|---|
| Support Vector Classifier | True Negative = 271 False Negative = 121 |
False Positive = 0 True Positive = 62 |
| Extreme Gradient Boost | True Negative = 409 False Negative = 119 |
False Positive = 23 True Positive = 103 |
| Logistic Regression | True Negative = 9028 False Negative = 8519 |
False Positive = 2022 True Positive = 2025 |
| Decision Tree | True Negative = 124 False Negative = 95 |
False Positive = 98 True Positive = 107 |
| Random Forest | True Negative = 12612 False Negative = 11832 |
False Positive = 6817 True Positive = 6510 |
| K Nearest Neighbor | True Negative = 112 False Negative = 89 |
False Positive = 105 True Positive = 118 |
| Name of Algorithm | Accuracy | F1-Score | Recall | Specificity |
|---|---|---|---|---|
| Support Vector Classifier | 92~94% | 87~88% | 91~93% | 90~93% |
| Extreme Gradient Boost | 79~84% | 66~68% | 55~57% | 92~94% |
| Logistic Regression | 66~69% | 0~0% | 6~10% | 100% |
| Decision Tree | 81~84% | 70~73% | 56~59% | 96~97% |
| Random Forest | 82~84% | 67~68% | 57~58% | 94~95% |
| K Nearest Neighbor | 72~73% | 31~32% | 20~21% | 95~97% |
| Name of Algorithm | Confusion Matrix | |
|---|---|---|
| Support Vector Classifier | True Negative = 416 False Negative = 17 |
False Positive = 37 True Positive = 207 |
| Extreme Gradient Boost | True Negative = 413 False Negative = 105 |
False Positive = 25 True Positive = 134 |
| Logistic Regression | True Negative = 453 False Negative = 224 |
False Positive = 0 True Positive = 0 |
| Decision Tree | True Negative = 433 False Negative = 54 |
False Positive = 16 True Positive = 134 |
| Random Forest | True Negative = 437 False Negative = 93 |
False Positive = 23 True Positive = 124 |
| K Nearest Neighbor | True Negative = 438 False Negative = 179 |
False Positive = 15 True Positive = 45 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




