Submitted:
10 October 2023
Posted:
12 October 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Patients
2.2. Generation of PDTOs
2.3. Flow cytometry
2.4. Mathematical approaches used in this study
3. Results
3.1. General work flow
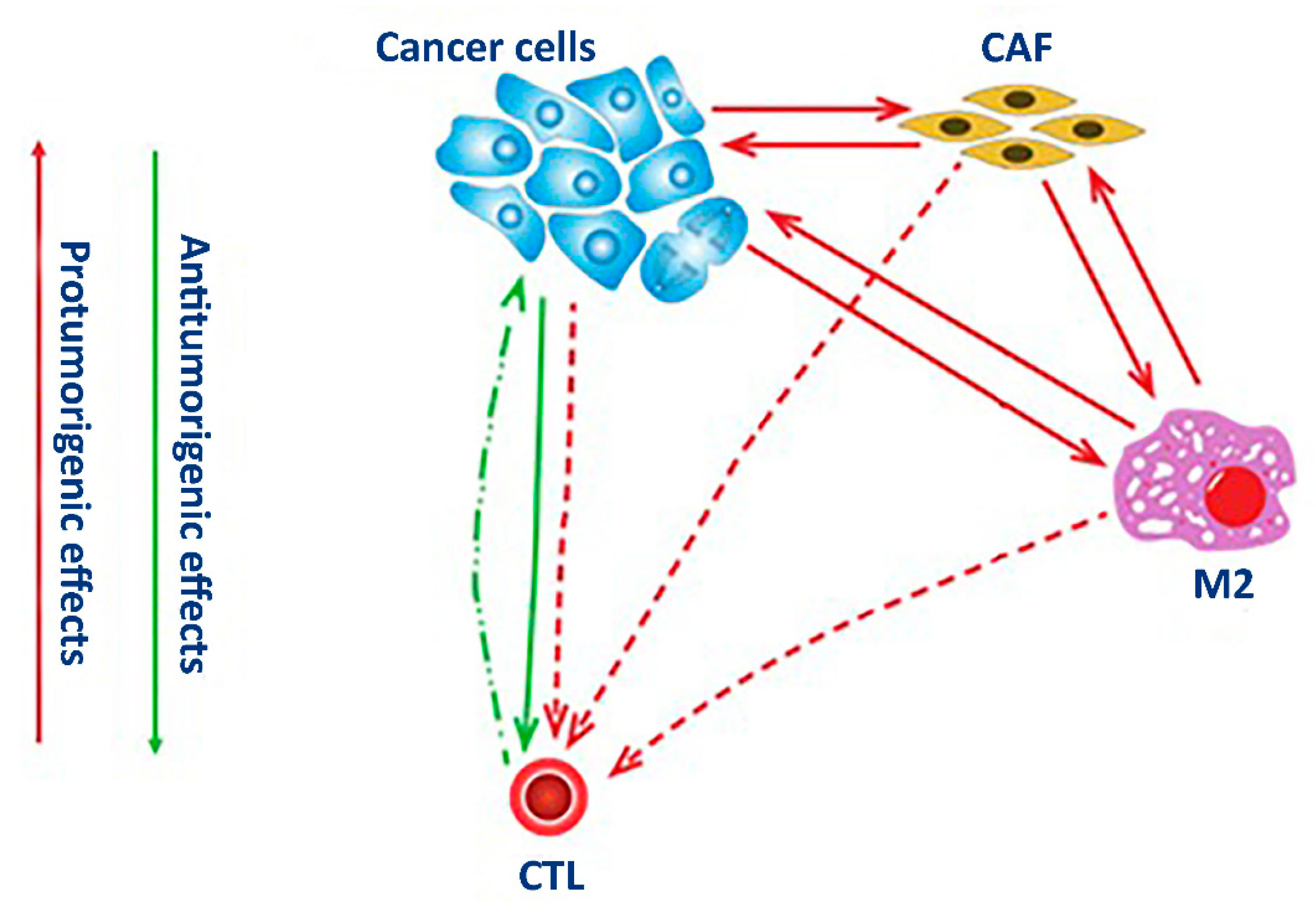
3.2. Mathematical model
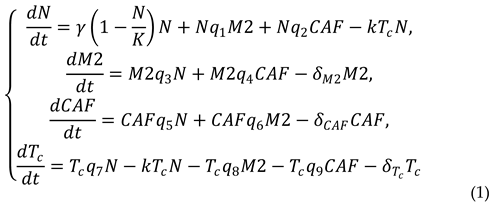
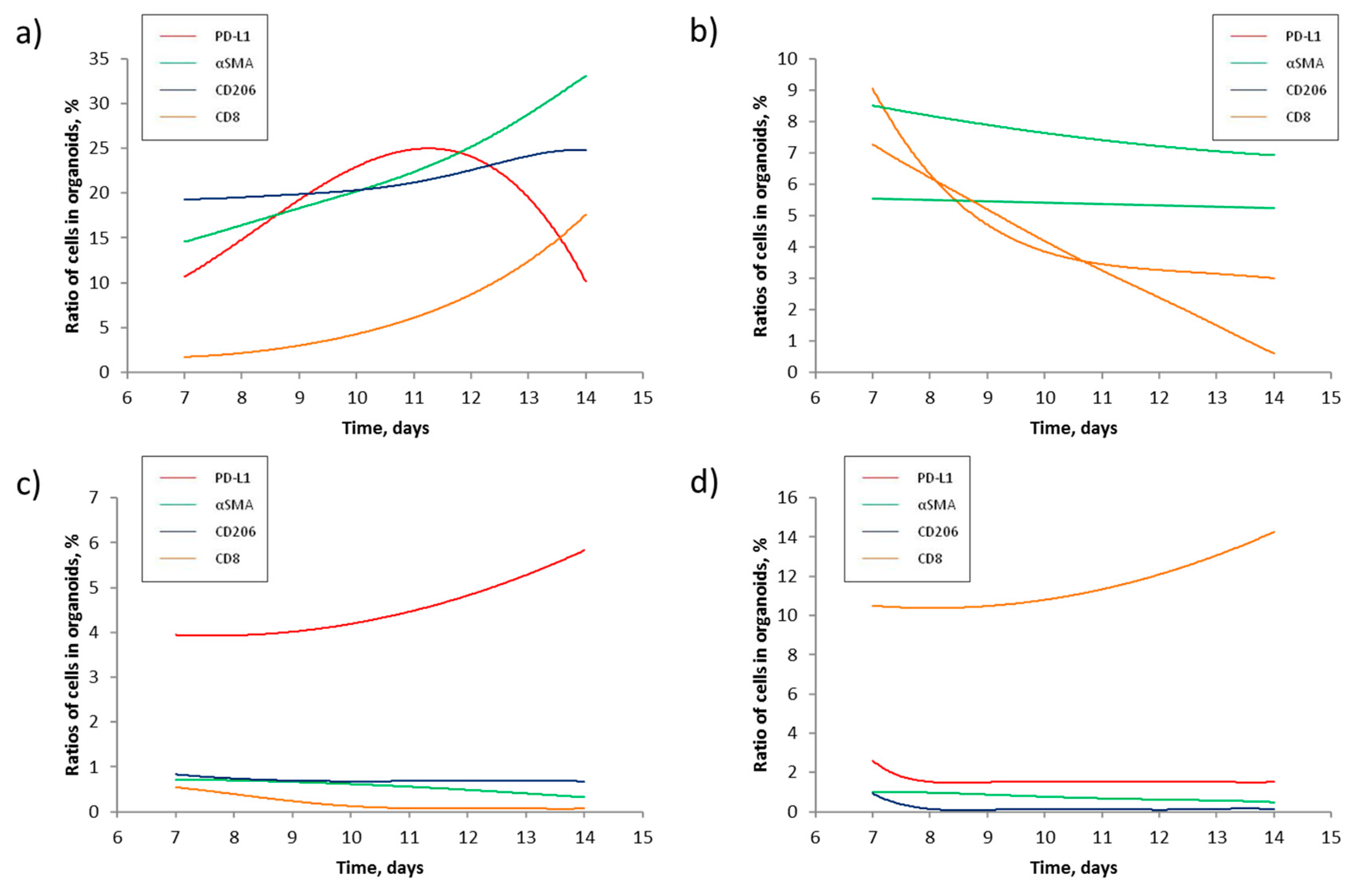
3.3. Solving the system of differential equations
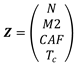
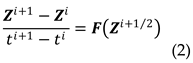
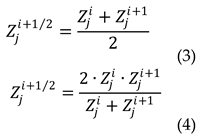
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Rossi, R.; De Angelis, M.L.; Xhelili, E.; Sette, G.; Eramo, A.; De Maria, R.; Cesta Incani, U.; Francescangeli, F.; Zeuner, A. Lung Cancer Organoids: The Rough Path to Personalized Medicine. Cancers (Basel) 2022, 14. [Google Scholar] [CrossRef]
- Li, Y.; Gao, X.; Ni, C.; Zhao, B.; Cheng, X. The application of patient-derived organoid in the research of lung cancer. Cell Oncol (Dordr) 2023, 46, 503–519. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Shen, J.; Zou, X. Mathematical modeling and dynamical analysis of anti-tumor drug dose-response. Math Biosci Eng 2022, 19, 4120–4144. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez Messan, M.; Yogurtcu, O.N.; McGill, J.R.; Nukala, U.; Sauna, Z.E.; Yang, H. Mathematical model of a personalized neoantigen cancer vaccine and the human immune system. PLoS Comput Biol 2021, 17, e1009318. [Google Scholar] [CrossRef] [PubMed]
- Voit, E.O. The best models of metabolism. Wiley Interdiscip Rev Syst Biol Med 2017, 9. [Google Scholar] [CrossRef] [PubMed]
- Bag, A.K.; Mandloi, S.; Jarmalavicius, S.; Mondal, S.; Kumar, K.; Mandal, C.; Walden, P.; Chakrabarti, S.; Mandal, C. Connecting signaling and metabolic pathways in EGF receptor-mediated oncogenesis of glioblastoma. PLoS Comput Biol 2019, 15, e1007090. [Google Scholar] [CrossRef] [PubMed]
- Santarpia, M.; Aguilar, A.; Chaib, I.; Cardona, A.F.; Fancelli, S.; Laguia, F.; Bracht, J.W.P.; Cao, P.; Molina-Vila, M.A.; Karachaliou, N.; et al. Non-Small-Cell Lung Cancer Signaling Pathways, Metabolism, and PD-1/PD-L1 Antibodies. Cancers (Basel) 2020, 12. [Google Scholar] [CrossRef] [PubMed]
- Tabassum, S.; Rosli, N.B.; Binti Mazalan, M.S.A. Mathematical Modeling of Cancer Growth Process: A Review. Journal of Physics: Conference Series 2019, 1366, 012018. [Google Scholar] [CrossRef]
- Feng, Y.; Li, X.; Wang, Y. Establishment of a mathematical model for predicting malignancy of lung cancer complicated with Talaromyces Marneffei infection and its chest imaging characteristics. Results in Physics 2021, 25, 104312. [Google Scholar] [CrossRef]
- Geng, C.; Paganetti, H.; Grassberger, C. Prediction of Treatment Response for Combined Chemo- and Radiation Therapy for Non-Small Cell Lung Cancer Patients Using a Bio-Mathematical Model. Sci Rep 2017, 7, 13542. [Google Scholar] [CrossRef]
- Popper, H.H. Manipulation of the immune system by non-small cell lung cancer and possible therapeutic interference. Cancer Drug Resist 2020, 3, 710–725. [Google Scholar] [CrossRef]
- Unni, P.; Seshaiyer, P. Mathematical Modeling, Analysis, and Simulation of Tumor Dynamics with Drug Interventions. Comput Math Methods Med 2019, 2019, 4079298. [Google Scholar] [CrossRef] [PubMed]
- Dritschel, H.; Waters, S.L.; Roller, A.; Byrne, H.M. A mathematical model of cytotoxic and helper T cellinteractions in a tumour microenvironment. Letters in Biomathematics 2018, 5, S36–S68. [Google Scholar] [CrossRef]
- Eftimie, R.; Eftimie, G. Investigating Macrophages Plasticity Following Tumour-Immune Interactions During Oncolytic Therapies. Acta Biotheor 2019, 67, 321–359. [Google Scholar] [CrossRef] [PubMed]
- Qin, W.; Hu, L.; Zhang, X.; Jiang, S.; Li, J.; Zhang, Z.; Wang, X. The Diverse Function of PD-1/PD-L Pathway Beyond Cancer. Front Immunol 2019, 10, 2298. [Google Scholar] [CrossRef] [PubMed]
- Hinz, B.; Celetta, G.; Tomasek, J.J.; Gabbiani, G.; Chaponnier, C. Alpha-smooth muscle actin expression upregulates fibroblast contractile activity. Mol Biol Cell 2001, 12, 2730–2741. [Google Scholar] [CrossRef]
- Zhang, W.; Huang, P. Cancer-stromal interactions: Role in cell survival, metabolism and drug sensitivity. Cancer Biol Ther 2011, 11, 150–156. [Google Scholar] [CrossRef]
- Sobolev, V.V.; Tchepourina, E.; Korsunskaya, I.M.; Geppe, N.A.; Chebysheva, S.N.; Soboleva, A.G.; Mezentsev, A. The Role of Transcription Factor PPAR-γ in the Pathogenesis of Psoriasis, Skin Cells, and Immune Cells. Int J Mol Sci 2022, 23. [Google Scholar] [CrossRef]
- Xu, Y.; Wang, X.; Liu, L.; Wang, J.; Wu, J.; Sun, C. Role of macrophages in tumor progression and therapy (Review). Int J Oncol 2022, 60. [Google Scholar] [CrossRef]
- Raskov, H.; Orhan, A.; Christensen, J.P.; Gögenur, I. Cytotoxic CD8(+) T cells in cancer and cancer immunotherapy. Br J Cancer 2021, 124, 359–367. [Google Scholar] [CrossRef]
- Guan, Y.; Kraus, S.G.; Quaney, M.J.; Daniels, M.A.; Mitchem, J.B.; Teixeiro, E. FOLFOX Chemotherapy Ameliorates CD8 T Lymphocyte Exhaustion and Enhances Checkpoint Blockade Efficacy in Colorectal Cancer. Front Oncol 2020, 10, 586. [Google Scholar] [CrossRef]
- Xie, Q.; Ding, J.; Chen, Y. Role of CD8(+) T lymphocyte cells: Interplay with stromal cells in tumor microenvironment. Acta Pharm Sin B 2021, 11, 1365–1378. [Google Scholar] [CrossRef] [PubMed]
- Dean, E.A.; Kimmel, G.J.; Frank, M.J.; Bukhari, A.; Hossain, N.M.; Jain, M.D.; Dahiya, S.; Miklos, D.B.; Altrock, P.M.; Locke, F.L. Circulating tumor DNA adds specificity to PET after axicabtagene ciloleucel in large B-cell lymphoma. Blood Adv 2023, 7, 4608–4618. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, R.K.; Andersen, M.; Skov, V.; Kjær, L.; Hasselbalch, H.C.; Ottesen, J.T.; Stiehl, T. HSC Niche Dynamics in Regeneration, Pre-malignancy, and Cancer: Insights From Mathematical Modeling. Stem Cells 2023, 41, 260–270. [Google Scholar] [CrossRef] [PubMed]
- Meade, W.; Weber, A.; Phan, T.; Hampston, E.; Resa, L.F.; Nagy, J.; Kuang, Y. High Accuracy Indicators of Androgen Suppression Therapy Failure for Prostate Cancer-A Modeling Study. Cancers (Basel) 2022, 14. [Google Scholar] [CrossRef] [PubMed]
- Ait Oumghar, I.; Barkaoui, A.; Ghazi, A.E.; Chabrand, P. Modeling and simulation of bone cells dynamic behavior under the late effect of breast cancer treatments. Med Eng Phys 2023, 115, 103982. [Google Scholar] [CrossRef] [PubMed]
- Larsson, I.; Dalmo, E.; Elgendy, R.; Niklasson, M.; Doroszko, M.; Segerman, A.; Jörnsten, R.; Westermark, B.; Nelander, S. Modeling glioblastoma heterogeneity as a dynamic network of cell states. Mol Syst Biol 2021, 17, e10105. [Google Scholar] [CrossRef]
- Schildhaus, H.U. [Predictive value of PD-L1 diagnostics]. Pathologe 2018, 39, 498–519. [Google Scholar] [CrossRef]
- Liu, J.; Li, C.; Seery, S.; Yu, J.; Meng, X. Identifying optimal first-line interventions for advanced non-small cell lung carcinoma according to PD-L1 expression: A systematic review and network meta-analysis. Oncoimmunology 2020, 9, 1746112. [Google Scholar] [CrossRef]
- Gordon, S.R.; Maute, R.L.; Dulken, B.W.; Hutter, G.; George, B.M.; McCracken, M.N.; Gupta, R.; Tsai, J.M.; Sinha, R.; Corey, D.; et al. PD-1 expression by tumour-associated macrophages inhibits phagocytosis and tumour immunity. Nature 2017, 545, 495–499. [Google Scholar] [CrossRef]
- Li, Y.; Chan, J.W.Y.; Lau, R.W.H.; Cheung, W.W.Y.; Wong, A.M.; Wong, A.M.; Wong, N.; Ng, C.S.H. Organoids in Lung Cancer Management. Front Surg 2021, 8, 753801. [Google Scholar] [CrossRef] [PubMed]
- Qi, X.; Prokhorova, A.V.; Mezentsev, A.V.; Shen, N.; Trofimenko, A.V.; Filkov, G.I.; Sulimanov, R.A.; Makarov, V.A.; Durymanov, M.O. Comparison of EMT-Related and Multi-Drug Resistant Gene Expression, Extracellular Matrix Production, and Drug Sensitivity in NSCLC Spheroids Generated by Scaffold-Free and Scaffold-Based Methods. Int J Mol Sci 2022, 23. [Google Scholar] [CrossRef] [PubMed]
| Type of cells | Cell-specific biomarker |
|---|---|
| Cancer cells | PD-L1 |
| Cancer-associated fibroblasts | αSMA |
| M2-polarized macrophages | CD206 |
| Cytotoxic lymphocytes | CD8 |
| Parameter | Definition | Published value | References | Adjusted value |
|---|---|---|---|---|
| γ | Growth rate of cancer cells | 0.05 – 0.44 day−1 | [12,13] | 0.05 day−1 |
| K | Final number of cancer cells | 109 – 3.3 × 109 day | [14] | 106 day−1 |
| q1 | Stimulation of cancer cells by M2-polarized macrophages | 0.4 day−1 | [14] | 4 x 10−5 day−1 |
| q3 | Stimulation of M2 macrophages by cancer cells | 4 x 10−8 day−1 | [14] | 4×10−8 day−1 |
| δM2 | Death rate of M2-polarized macrophages from natural causes | 0.2 day−1 | [13] | 0.2 day−1 |
| k | Number of cancer cells eliminated by cytotoxic cells | 3.4 x 10−10 - 1 x 10−3 cell −1 day −1 | [13] | 0.001 cell−1 day−1 |
| δTc | Death rate of cytotoxic cells | 2 x 10−3 – 1 day −1 | [13] | 0.1 day−1 |
| Parameter | Definition | Dimension |
|---|---|---|
| q2 | Stimulation of cancer cells by cancer-associated fibroblasts | day−1 |
| q4 | Stimulation of M2-polarized macrophages by cancer-associated fibroblasts | day−1 |
| q5 | Stimulation of cancer-associated fibroblasts by cancer cells | day−1 |
| q6 | Stimulation of cancer-associated fibroblasts by M2-polarized macrophages | day−1 |
| q7 | Stimulation of cytotoxic T cells by cancer cells | day−1 |
| q8 | Suppression of cytotoxic T cells by M2-polarized macrophages | day−1 |
| q9 | Suppression of cytotoxic T cells by tumor-associated macrophages | day−1 |
| δCAF | Death rate of cancer-associated fibroblasts | day−1 |
| Parameter | Calculated values, day−1 |
|---|---|
| q2 | 0.0001–0.005 |
| q4 | 0.0001–0.001 |
| q5 | 0–0.00001 |
| q6 | 0.00001–0.001 |
| q7 | 0.0009–0.0015 |
| q8 | 0–0.00001 |
| q9 | 0–0.00001 |
| δCAF | 0.1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




