Submitted:
01 June 2023
Posted:
08 June 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Structure of ORF6 in Water Solvent
2.2. Structure of ORF6 in ER Membrane
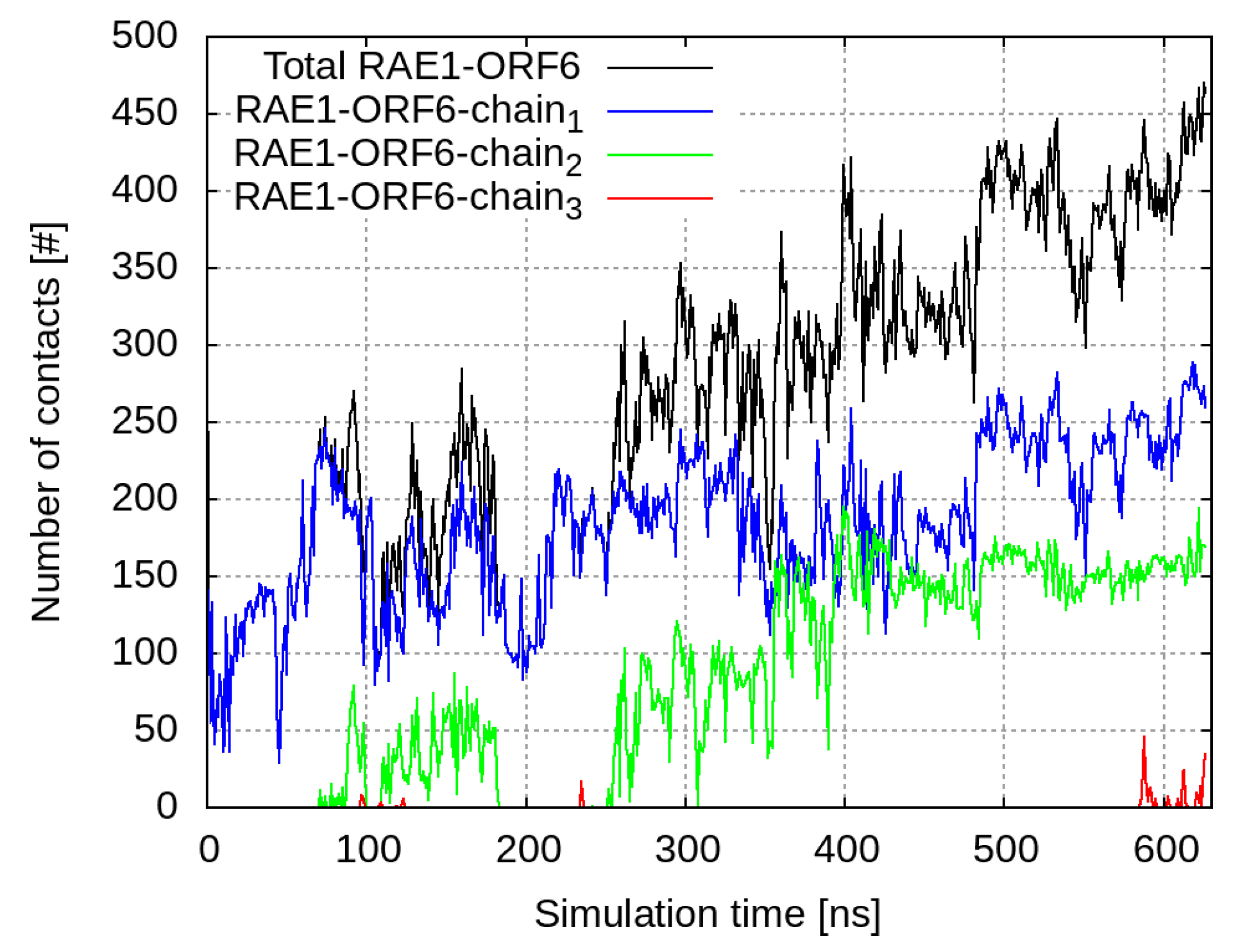
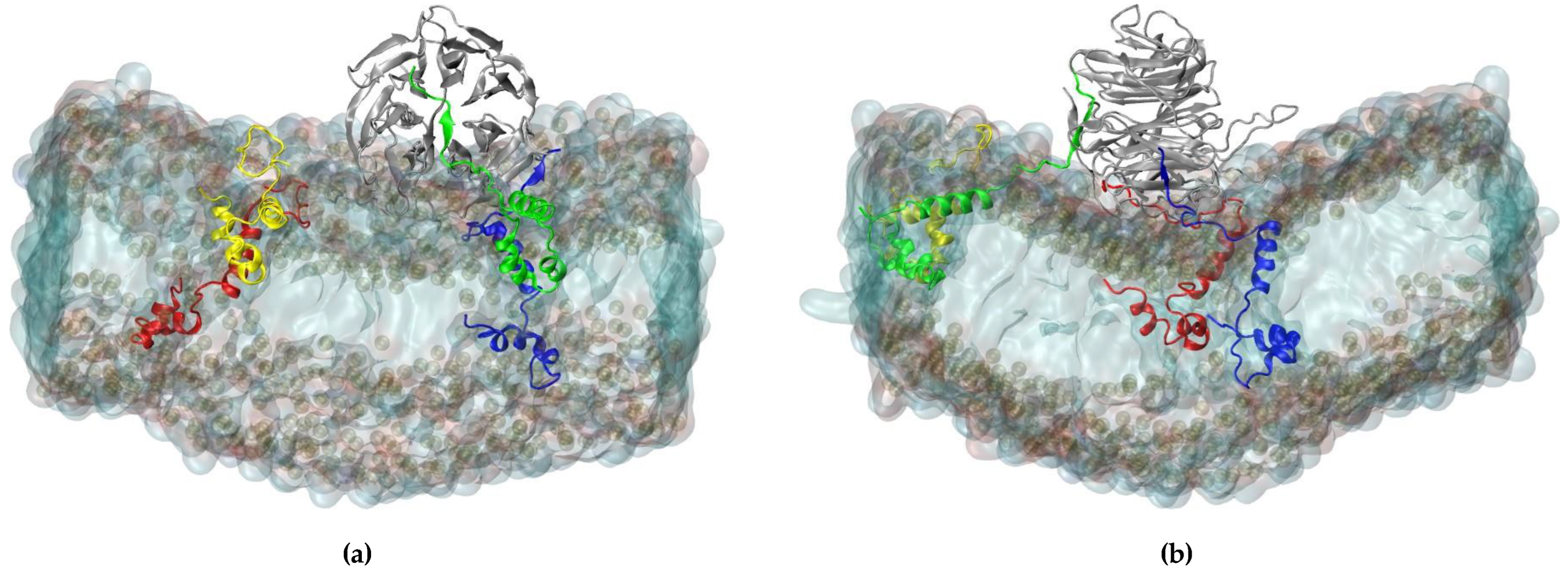
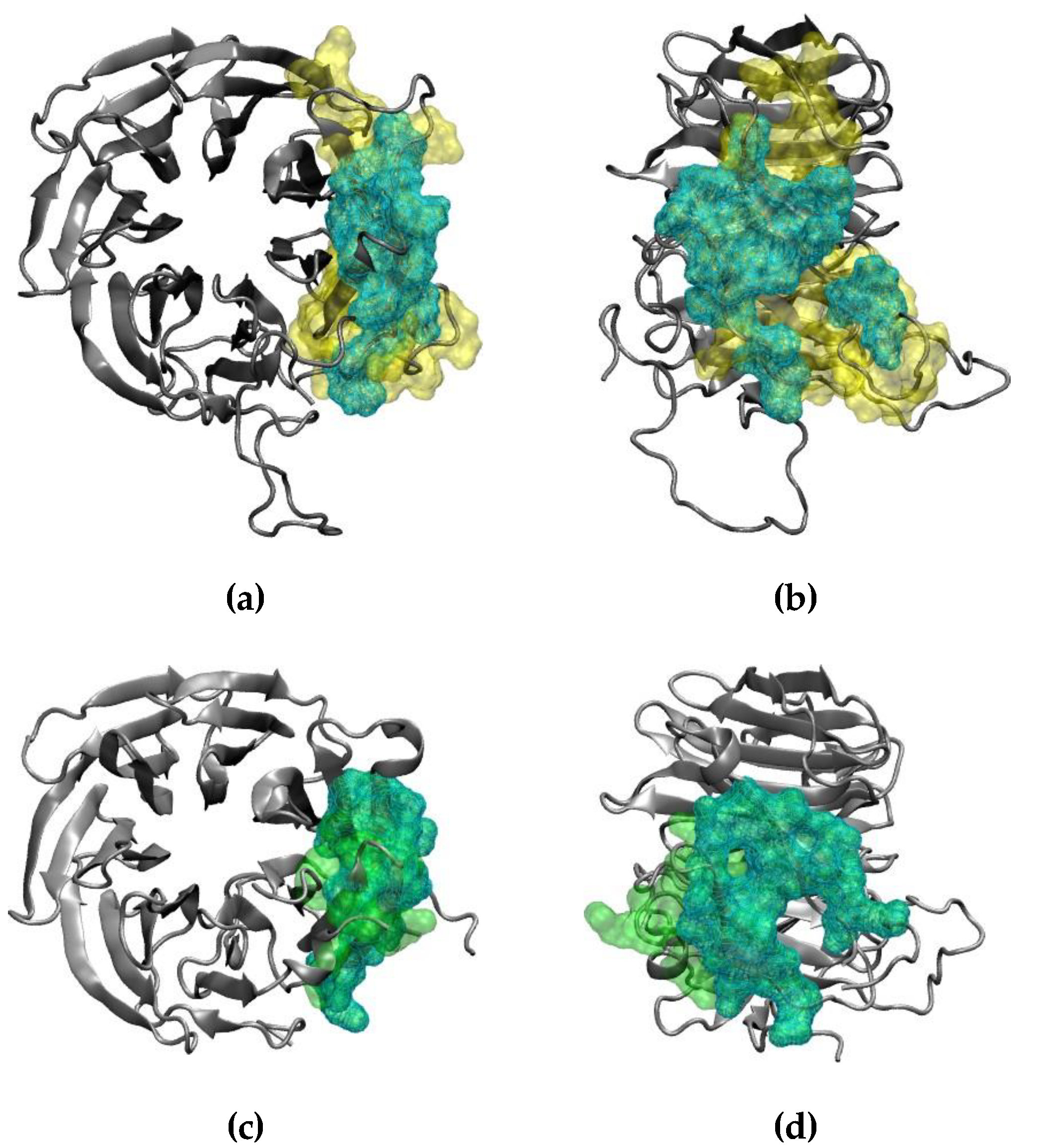
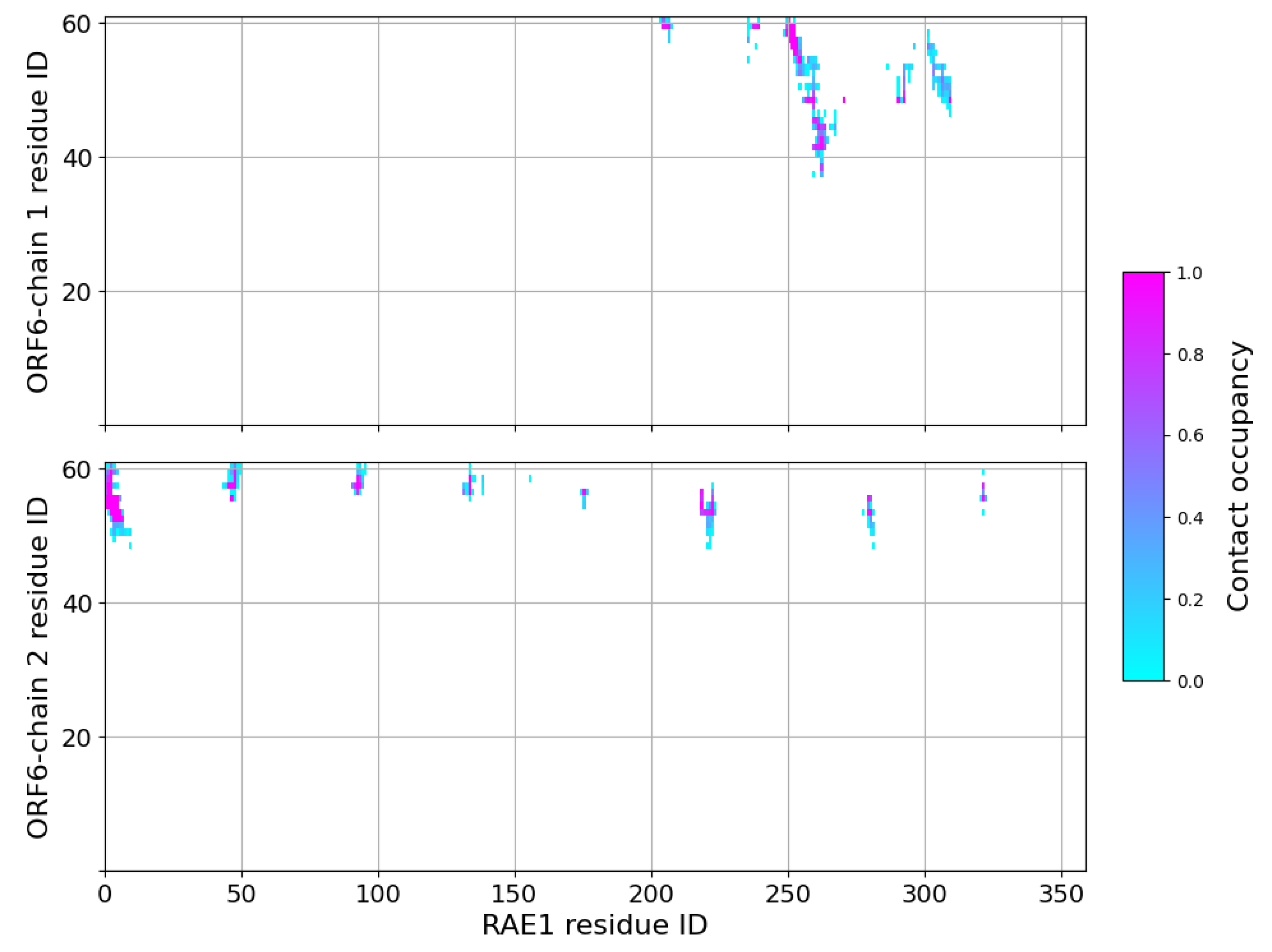
2.3. Interaction of RAE1 and ORF6
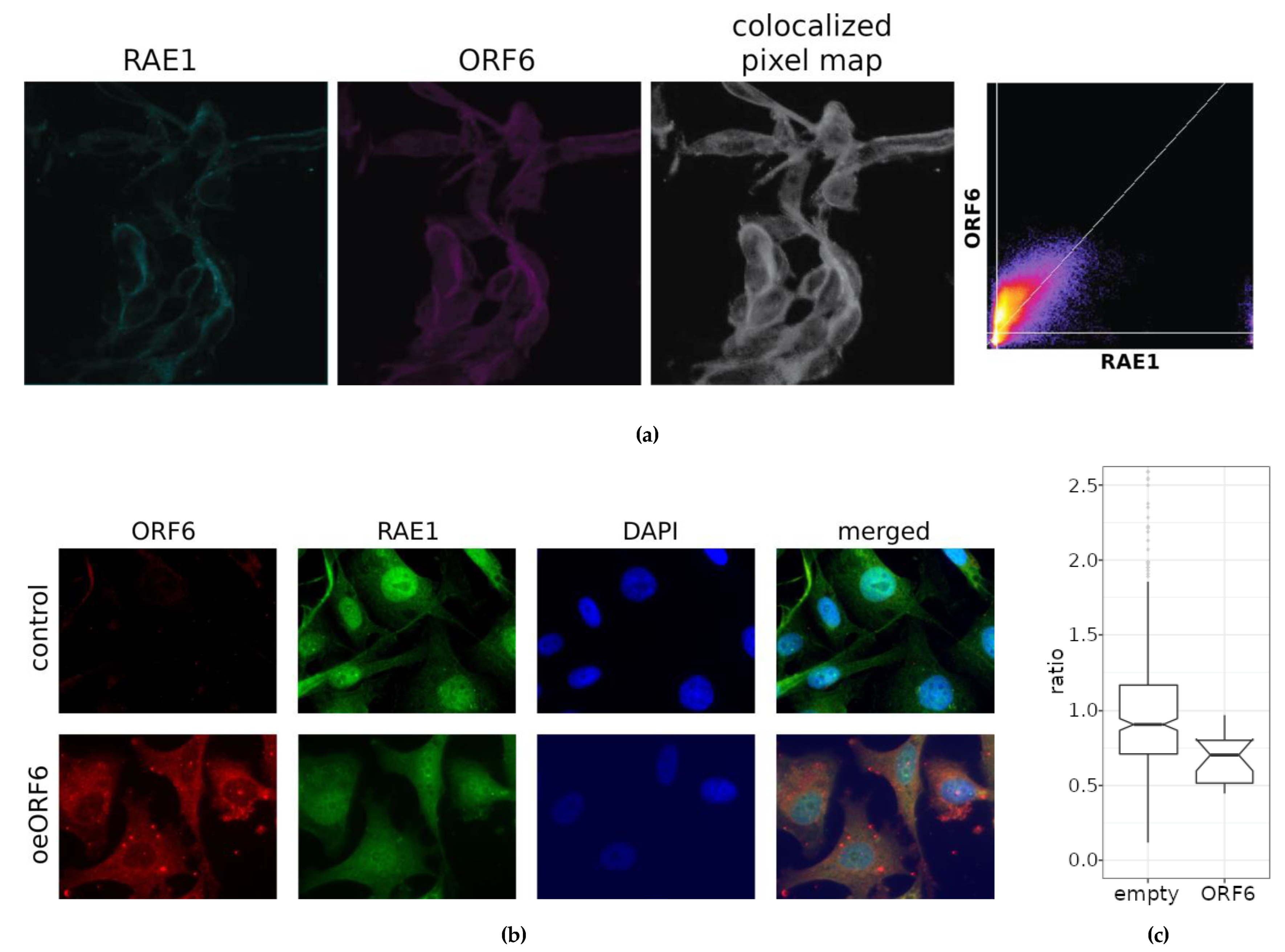
2.4. Co-Localization Between RAE1 and ORF6
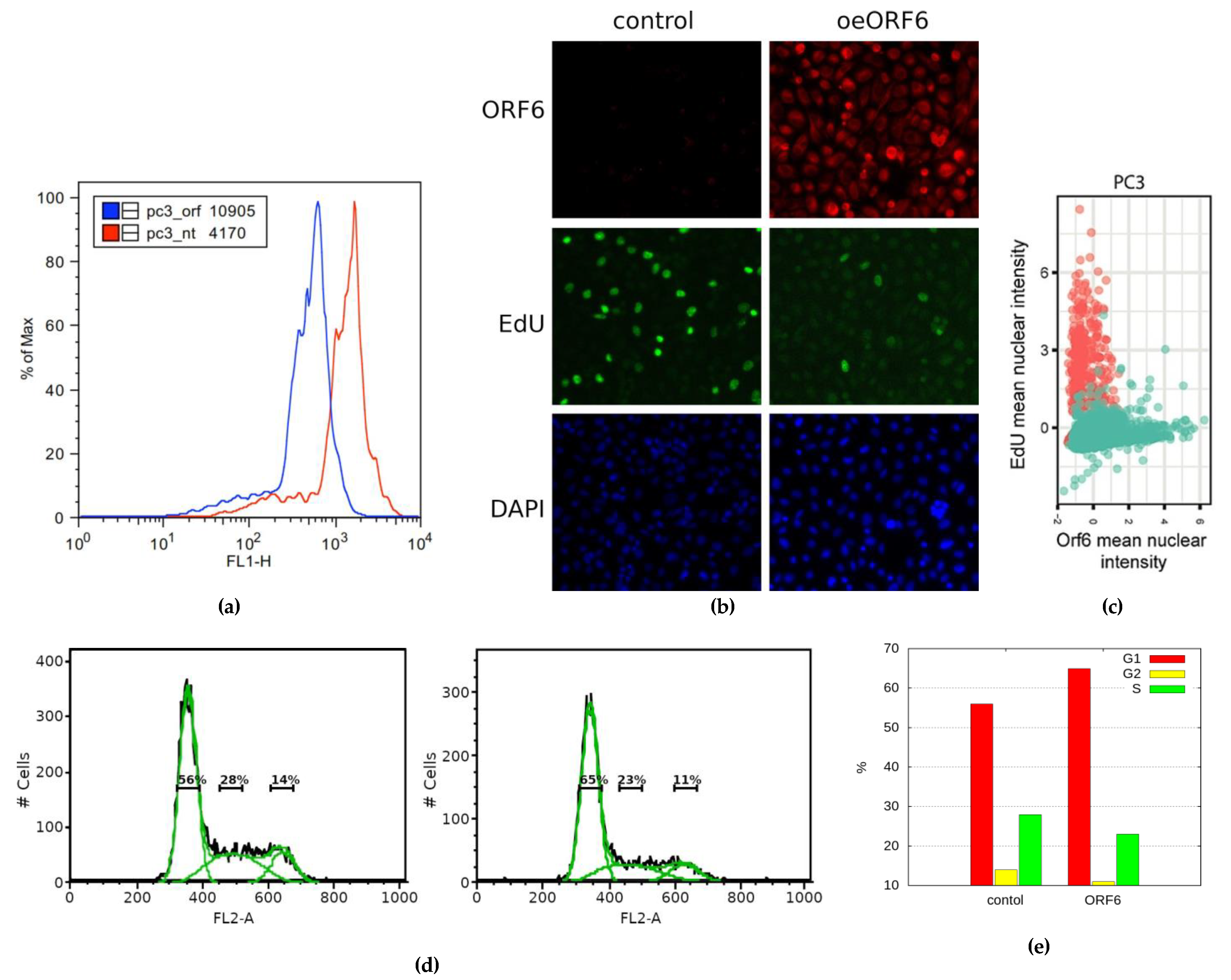
2.5. Overexpression of SARS-CoV-2 ORF6 Inhibits Cell Cycle Progression
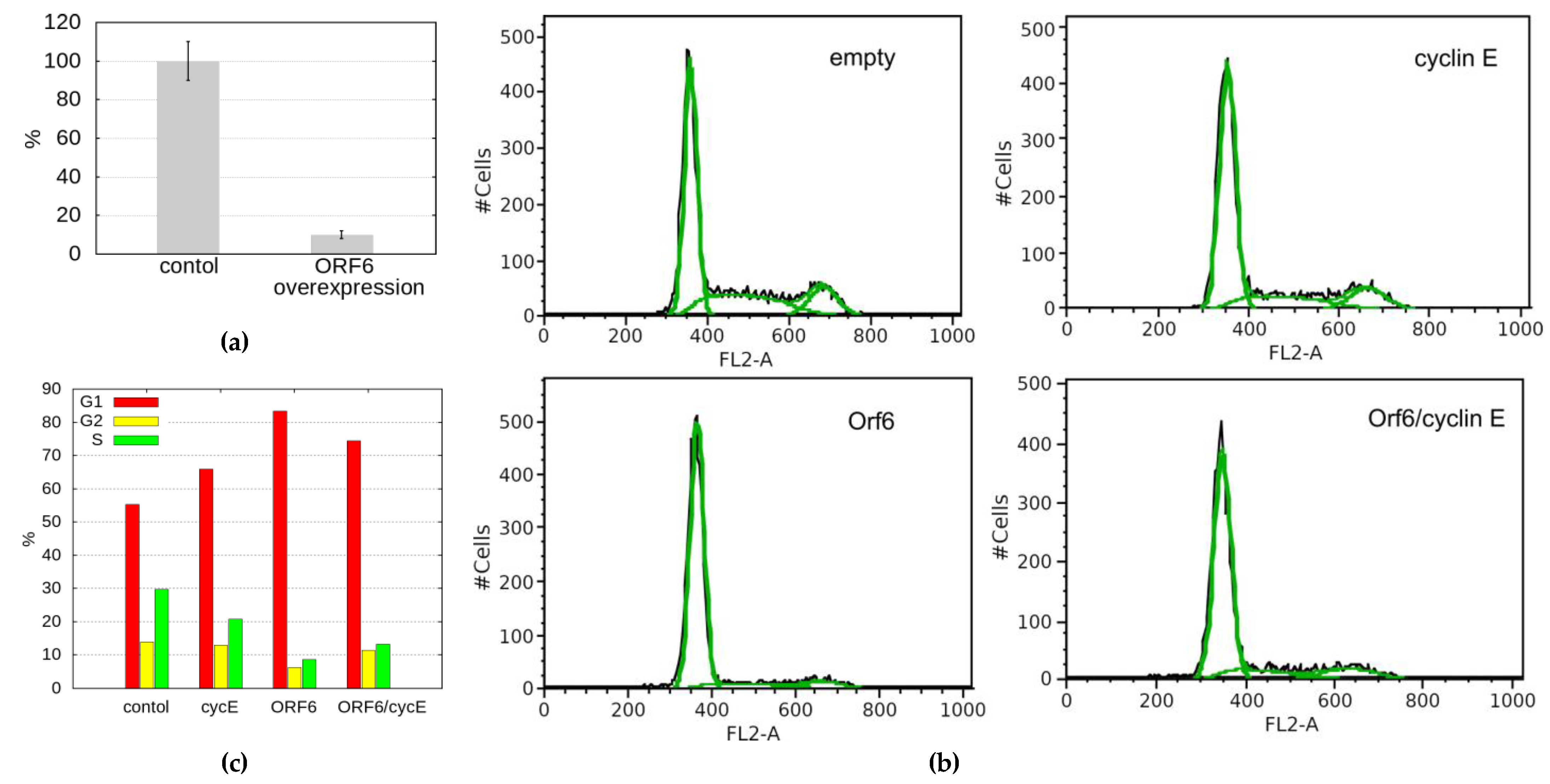
2.6. ORF6 Overexpression Inhibits Proliferation by Decreasing Cyclin E Level
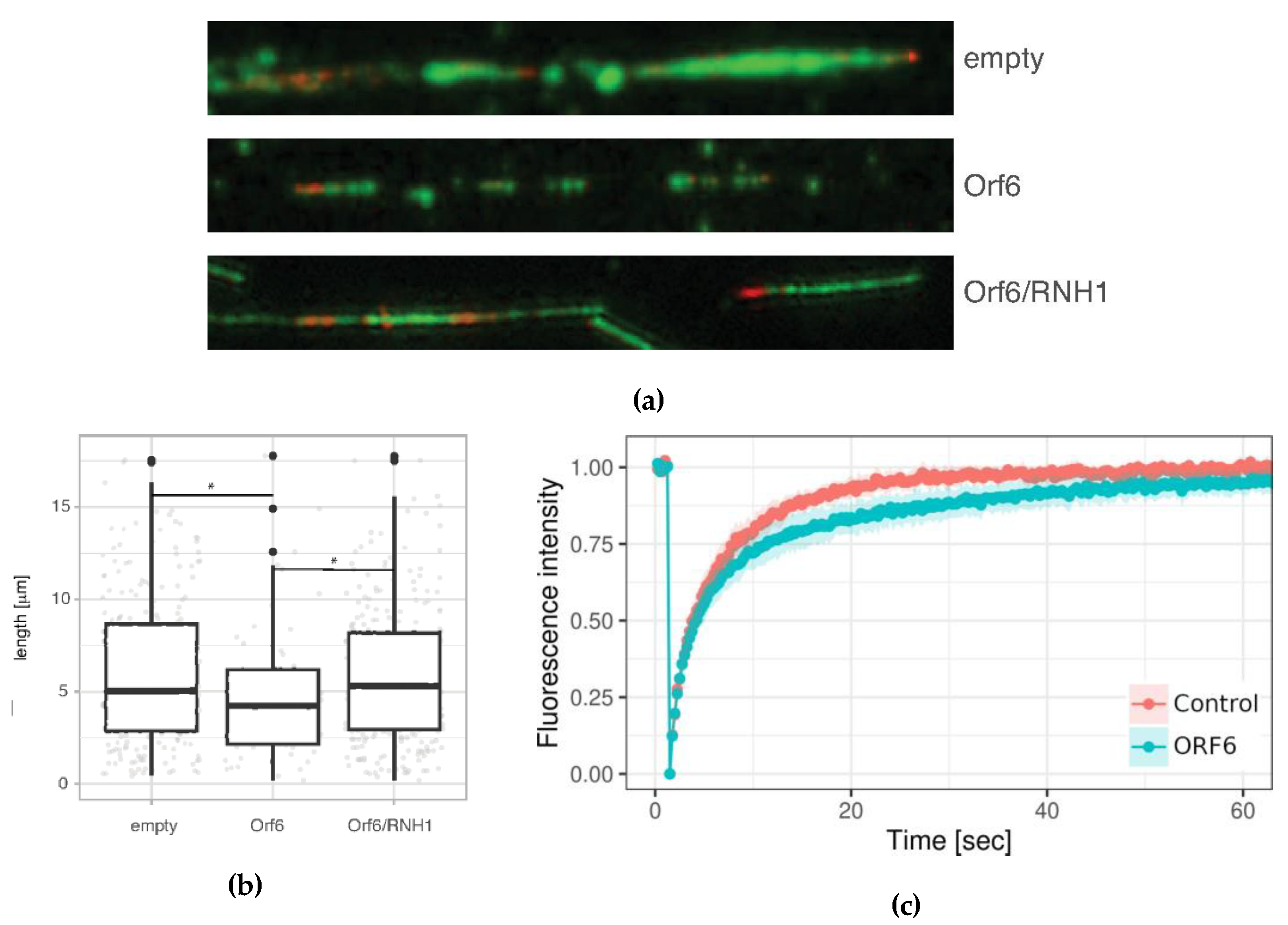
2.7. ORF6 Overexpression Causes Accumulation of R-loops, Which Impedes Progression of Active Replication Forks
3. Discussion
4. Materials and Methods
4.1. MD Simulation Protocol
4.2. Input Structural Models
4.2.1. SARS-CoV-2 ORF6 Protein
4.2.2. SARS-CoV-2 ORF6 Embedded in a Model ER Membrane
4.3. Modelling the Interaction of SARS-CoV-2 ORF6 and RAE1
4.4. MD Data Analysis
4.5. Cell culture, Plasmids, and Antibodies
4.6. Quantitative Real-Time PCR Analysis
4.7. 5-Ethynyl-2’-deoxyuridine (EdU) Labelling
4.8. Flow Cytometry
4.9. Immunofluorescent Microscopy
4.10. Fluorescence Recovery After Photobleaching (FRAP) Analysis
4.11. DNA Fibre Labeling
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| aa | amino acids |
| CHOL | cholesterol |
| ER | endoplasmic reticulum |
| GA | Golgi apparatus |
| MD | molecular dynamics |
| NUP98 | nucleoporin 98 |
| oeORF6 | ORF6-overexpressing cells |
| ORF6 | Open reading frame 6 |
| POPC | 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine |
| POPE | 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine |
| POPI | 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoinositol |
| RAE1 | ribonucleic acid export 1 |
| Rg | radius of gyration |
| RMSD | root mean square deviation |
| RMSF | root mean square fluctuations |
| SASA | solvent accessible surface area |
References
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y.M.; Wang, W.; Song, Z.G.; Hu, Y.; Tao, Z.W.; Tian, J.H.; Pei, Y.Y.; Yuan, M.L.; Zhang, Y.L.; Dai, F.H.; Liu, Y.; Wang, Q.M.; Zheng, J.J.; Xu, L.; Holmes, E.C.; Zhang, Y.Z. A new coronavirus associated with human respiratory disease in China. Nature 2020, Nature, 265–269. [Google Scholar] [CrossRef]
- Bai, C.; Zhong, Q.; Gao, G. Overview of SARS-CoV-2 genome-encoded proteins. Science China Life Sciences 2022, 65, 280–294. [Google Scholar] [CrossRef]
- Zandi, M.; Shafaati, M.; Kalantar-Neyestanaki, D.; Pourghadamyari, H.; Fani, M.; Soltani, S.; Kaleji, H.; Abbasi, S. The role of SARS-CoV-2 accessory proteins in immune evasion. Biomedicine & Pharmacotherapy 2022, 156, 113889. [Google Scholar] [CrossRef]
- Redondo, N.; Zaldívar-López, S.; Garrido, J.J.; Montoya, M. SARS-CoV-2 Accessory Proteins in Viral Pathogenesis: Knowns and Unknowns. Frontiers in Immunology 2021, 12. [Google Scholar] [CrossRef]
- Kimura, I.; Konno, Y.; Uriu, K.; Hopfensperger, K.; Sauter, D.; Nakagawa, S.; Sato, K. Sarbecovirus ORF6 proteins hamper induction of interferon signaling. Cell Reports 2021, 34, 108916. [Google Scholar] [CrossRef]
- Frieman, M.; Yount, B.; Heise, M.; Kopecky-Bromberg, S.A.; Palese, P.; Baric, R.S. Severe Acute Respiratory Syndrome Coronavirus ORF6 Antagonizes STAT1 Function by Sequestering Nuclear Import Factors on the Rough Endoplasmic Reticulum/Golgi Membrane. Journal of Virology 2007, 81, 9812–9824. [Google Scholar] [CrossRef]
- Kopecky-Bromberg, S.A.; Martínez-Sobrido, L.; Frieman, M.; Baric, R.A.; Palese, P. Severe Acute Respiratory Syndrome Coronavirus Open Reading Frame (ORF) 3b, ORF 6, and Nucleocapsid Proteins Function as Interferon Antagonists. Journal of Virology 2007, 81, 548–557. [Google Scholar] [CrossRef]
- Miorin, L.; Kehrer, T.; Sanchez-Aparicio, M.T.; Zhang, K.; Cohen, P.; Patel, R.S.; Cupic, A.; Makio, T.; Mei, M.; Moreno, E.; Danziger, O.; White, K.M.; Rathnasinghe, R.; Uccellini, M.; Gao, S.; Aydillo, T.; Mena, I.; Yin, X.; Martin-Sancho, L.; Krogan, N.J.; Chanda, S.K.; Schotsaert, M.; Wozniak, R.W.; Ren, Y.; Rosenberg, B.R.; Fontoura, B.M.A.; García-Sastre, A. SARS-CoV-2 Orf6 hijacks Nup98 to block STAT nuclear import and antagonize interferon signaling. Proceedings of the National Academy of Sciences 2020, 117, 28344–28354. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Wen, Y.; Guo, H.; Yang, T.; Yang, H.; Ji, X. Molecular Mechanism of SARS-CoVs Orf6 Targeting the Rae1–Nup98 Complex to Compete With mRNA Nuclear Export. Frontiers in Molecular Biosciences 2022, 8. [Google Scholar] [CrossRef] [PubMed]
- Yuen, C.K.; Lam, J.Y.; Wong, W.M.; Mak, L.F.; Wang, X.; Chu, H.; Cai, J.P.; Jin, D.Y.; To, K.K.W.; Chan, J.F.W.; Yuen, K.Y.; Kok, K.H. SARS-CoV-2 nsp13, nsp14, nsp15 and orf6 function as potent interferon antagonists. Emerging Microbes & Infections 2020, 9, 1418–1428. [Google Scholar] [CrossRef]
- Miyamoto, Y.; Itoh, Y.; Suzuki, T.; Tanaka, T.; Sakai, Y.; Koido, M.; Hata, C.; Wang, C.X.; Otani, M.; Moriishi, K.; Tachibana, T.; Kamatani, Y.; Yoneda, Y.; Okamoto, T.; Oka, M. SARS-CoV-2 ORF6 disrupts nucleocytoplasmic trafficking to advance viral replication. Communications Biology 2022, 5, 483. [Google Scholar] [CrossRef]
- Lee, J.G.; Huang, W.; Lee, H.; van de Leemput, J.; Kane, M.A.; Han, Z. Characterization of SARS-CoV-2 proteins reveals Orf6 pathogenicity, subcellular localization, host interactions and attenuation by Selinexor. Cell & Bioscience 2021, 11, 58. [Google Scholar] [CrossRef]
- Netland, J.; Ferraro, D.; Pewe, L.; Olivares, H.; Gallagher, T.; Perlman, S. Enhancement of Murine Coronavirus Replication by Severe Acute Respiratory Syndrome Coronavirus Protein 6 Requires the N-Terminal Hydrophobic Region but Not C-Terminal Sorting Motifs. Journal of Virology 2007, 81, 11520–11525. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Falcón, A.; Zhou, H.; Netland, J.; Enjuanes, L.; Breña, P.P.; Perlman, S. Severe Acute Respiratory Syndrome Coronavirus Protein 6 Is Required for Optimal Replication. Journal of Virology 2009, 83, 2368–2373. [Google Scholar] [CrossRef] [PubMed]
- Gordon, D.E.; Jang, G.M.; Bouhaddou, M.; Xu, J.; Obernier, K.; White, K.M.; O’Meara, M.J.; Rezelj, V.V.; Guo, J.Z.; Swaney, D.L.; Tummino, T.A.; Hüttenhain, R.; Kaake, R.M.; Richards, A.L.; Tutuncuoglu, B.; Foussard, H.; Batra, J.; Haas, K.; Modak, M.; Kim, M.; Haas, P.; Polacco, B.J.; Braberg, H.; Fabius, J.M.; Eckhardt, M.; Soucheray, M.; Bennett, M.J.; Cakir, M.; McGregor, M.J.; Li, Q.; Meyer, B.; Roesch, F.; Vallet, T.; Mac Kain, A.; Miorin, L.; Moreno, E.; Naing, Z.Z.C.; Zhou, Y.; Peng, S.; Shi, Y.; Zhang, Z.; Shen, W.; Kirby, I.T.; Melnyk, J.E.; Chorba, J.S.; Lou, K.; Dai, S.A.; Barrio-Hernandez, I.; Memon, D.; Hernandez-Armenta, C.; Lyu, J.; Mathy, C.J.P.; Perica, T.; Pilla, K.B.; Ganesan, S.J.; Saltzberg, D.J.; Rakesh, R.; Liu, X.; Rosenthal, S.B.; Calviello, L.; Venkataramanan, S.; Liboy-Lugo, J.; Lin, Y.; Huang, X.P.; Liu, Y.; Wankowicz, S.A.; Bohn, M.; Safari, M.; Ugur, F.S.; Koh, C.; Savar, N.S.; Tran, Q.D.; Shengjuler, D.; Fletcher, S.J.; O’Neal, M.C.; Cai, Y.; Chang, J.C.J.; Broadhurst, D.J.; Klippsten, S.; Sharp, P.P.; Wenzell, N.A.; Kuzuoglu-Ozturk, D.; Wang, H.Y.; Trenker, R.; Young, J.M.; Cavero, D.A.; Hiatt, J.; Roth, T.L.; Rathore, U.; Subramanian, A.; Noack, J.; Hubert, M.; Stroud, R.M.; Frankel, A.D.; Rosenberg, O.S.; Verba, K.A.; Agard, D.A.; Ott, M.; Emerman, M.; Jura, N.; von Zastrow, M.; Verdin, E.; Ashworth, A.; Schwartz, O.; d’Enfert, C.; Mukherjee, S.; Jacobson, M.; Malik, H.S.; Fujimori, D.G.; Ideker, T.; Craik, C.S.; Floor, S.N.; Fraser, J.S.; Gross, J.D.; Sali, A.; Roth, B.L.; Ruggero, D.; Taunton, J.; Kortemme, T.; Beltrao, P.; Vignuzzi, M.; García-Sastre, A.; Shokat, K.M.; Shoichet, B.K.; Krogan, N.J. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 2020, 583, 459–468. [Google Scholar] [CrossRef]
- Overexpression of SARS-CoV-2 protein ORF6 dislocates RAE1 and NUP98 from the nuclear pore complex. Biochemical and Biophysical Research Communications 2021, 536, 59–66. [CrossRef] [PubMed]
- Addetia, A.; Lieberman, N.A.P.; Phung, Q.; Hsiang, T.Y.; Xie, H.; Roychoudhury, P.; Shrestha, L.; Loprieno, M.A.; Huang, M.L.; Gale, M.; Jerome, K.R.; Greninger, A.L. SARS-CoV-2 ORF6 Disrupts Bidirectional Nucleocytoplasmic Transport through Interactions with Rae1 and Nup98. mBio 2021, 12, e00065–21. [Google Scholar] [CrossRef]
- Wong, H.T.; Cheung, V.; Salamango, D.J. Decoupling SARS-CoV-2 ORF6 localization and interferon antagonism. Journal of Cell Science 2022, 135. [Google Scholar] [CrossRef]
- Gao, X.; Tian, H.; Zhu, K.; Li, Q.; Hao, W.; Wang, L.; Qin, B.; Deng, H.; Cui, S. Structural basis for Sarbecovirus ORF6 mediated blockage of nucleocytoplasmic transport. Nature Communications 2022, 13, 4782. [Google Scholar] [CrossRef]
- Ren, Y.; Seo, H.S.; Blobel, G.; Hoelz, A. Structural and functional analysis of the interaction between the nucleoporin Nup98 and the mRNA export factor Rae1. Proceedings of the National Academy of Sciences 2010, 107, 10406–10411. [Google Scholar] [CrossRef] [PubMed]
- Quan, B.; Seo, H.S.; Blobel, G.; Ren, Y. Vesiculoviral matrix (M) protein occupies nucleic acid binding site at nucleoporin pair (Rae1•Nup98). Proceedings of the National Academy of Sciences 2014, 111, 9127–9132. [Google Scholar] [CrossRef] [PubMed]
- Salic, A.; Mitchison, T.J. A chemical method for fast and sensitive detection of DNA synthesis <i>in vivo</i>. Proceedings of the National Academy of Sciences 2008, 105, 2415–2420. [Google Scholar] [CrossRef]
- Sunwoo, H.H.; Suresh, M.R. Chapter 9.13 - Cancer Markers. In The Immunoassay Handbook (Fourth Edition), Fourth Edition ed.; Wild, D., Ed.; Elsevier: Oxford, 2013; pp. 833–856. [Google Scholar] [CrossRef]
- Characterization of the Drosophila Rae1 protein as a G1 phase regulator of the cell cycle. Gene 2004, 326, 107–116. [CrossRef]
- Jimeno, S.; Rondón, A.G.; Luna, R.; Aguilera, A. The yeast THO complex and mRNA export factors link RNA metabolism with transcription and genome instability. The EMBO Journal 2002, 21, 3526–3535. [Google Scholar] [CrossRef] [PubMed]
- Zenklusen, D.; Vinciguerra, P.; Wyss, J.C.; Stutz, F. Stable mRNP Formation and Export Require Cotranscriptional Recruitment of the mRNA Export Factors Yra1p and Sub2p by Hpr1p. Molecular and Cellular Biology 2002, 22, 8241–8253. [Google Scholar] [CrossRef] [PubMed]
- Domínguez-Sánchez, M.S.; Barroso, S.; Gímez-González, B.; Luna, R.; Aguilera, A. Genome Instability and Transcription Elongation Impairment in Human Cells Depleted of THO/TREX. PLOS Genetics 2011, 7, 1–17. [Google Scholar] [CrossRef]
- Gómez-González, B.; García-Rubio, M.; Bermejo, R.; Gaillard, H.; Shirahige, K.; Marín, A.; Foiani, M.; Aguilera, A. Genome-wide function of THO/TREX in active genes prevents R-loop-dependent replication obstacles. The EMBO Journal 2011, 30, 3106–3119. [Google Scholar] [CrossRef]
- Sollier, J.; Cimprich, K.A. Breaking bad: R-loops and genome integrity. Trends in Cell Biology 2015, 25, 514–522. [Google Scholar] [CrossRef]
- Gaillard, H.; Aguilera, A. Transcription as a Threat to Genome Integrity. Annual Review of Biochemistry 2016, 85, 291–317. [Google Scholar] [CrossRef]
- Tuduri, S.; Crabbé, L.; Conti, C.; Tourrière, H.; Holtgreve-Grez, H.; Jauch, A.; Pantesco, V.; De Vos, J.; Thomas, A.; Theillet, C.; Pommier, Y.; Tazi, J.; Coquelle, A.; Pasero, P. Topoisomerase I suppresses genomic instability by preventing interference between replication and transcription. Nature Cell Biology 2009, 11, 1315–1324. [Google Scholar] [CrossRef]
- Pritchard, C.E.; Fornerod, M.; Kasper, L.H.; van Deursen, J.M. RAE1 Is a Shuttling mRNA Export Factor That Binds to a GLEBS-like NUP98 Motif at the Nuclear Pore Complex through Multiple Domains. Journal of Cell Biology 1999, 145, 237–254. [Google Scholar] [CrossRef] [PubMed]
- Hall, R.; Guedán, A.; Yap, M.W.; Young, G.R.; Harvey, R.; Stoye, J.P.; Bishop, K.N. SARS-CoV-2 ORF6 disrupts innate immune signalling by inhibiting cellular mRNA export. PLOS Pathogens 2022, 18, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Feng, H.; Tian, H.; Wang, Y.; Zhang, Q.; Lin, N.; Liu, S.; Yu, Y.; Deng, H.; Gao, P. Molecular mechanism underlying selective inhibition of mRNA nuclear export by herpesvirus protein ORF10. Proceedings of the National Academy of Sciences 2020, 117, 26719–26727. [Google Scholar] [CrossRef] [PubMed]
- Skourti-Stathaki, K.; Proudfoot, N.; Gromak, N. Human Senataxin Resolves RNA/DNA Hybrids Formed at Transcriptional Pause Sites to Promote Xrn2-Dependent Termination. Molecular Cell 2011, 42, 794–805. [Google Scholar] [CrossRef] [PubMed]
- Crossley, M.P.; Bocek, M.; Cimprich, K.A. R-Loops as Cellular Regulators and Genomic Threats. Molecular Cell 2019, 73, 398–411. [Google Scholar] [CrossRef] [PubMed]
- Santos-Pereira, J.M.; Aguilera, A. R loops: New modulators of genome dynamics and function. Nature Reviews Genetics 2015, 16, 583–597. [Google Scholar] [CrossRef]
- Gan, W.; Guan, Z.; Liu, J.; Gui, T.; Shen, K.; Manley, J.L.; Li, X. R-loop-mediated genomic instability is caused by impairment of replication fork progression. Genes & Development 2011, 25, 2041–56. [Google Scholar] [CrossRef]
- Helmrich, A.; Ballarino, M.; Tora, L. Collisions between Replication and Transcription Complexes Cause Common Fragile Site Instability at the Longest Human Genes. Molecular Cell 2011, 44, 966–977. [Google Scholar] [CrossRef]
- Hamperl, S.; Cimprich, K.A. The contribution of co-transcriptional RNA:DNA hybrid structures to DNA damage and genome instability. DNA Repair 2014, 19, 84–94. [Google Scholar] [CrossRef]
- Jackson, B.R.; Noerenberg, M.; Whitehouse, A. A Novel Mechanism Inducing Genome Instability in Kaposi’s Sarcoma-Associated Herpesvirus Infected Cells. PLOS Pathogens 2014, 10, 1–15. [Google Scholar] [CrossRef]
- Gioia, U.; Tavella, S.; Martínez-Orellana, P.; Cicio, G.; Colliva, A.; Ceccon, M.; Cabrini, M.; Henriques, A.C.; Fumagalli, V.; Paldino, A.; Presot, E.; Rajasekharan, S.; Iacomino, N.; Pisati, F.; Matti, V.; Sepe, S.; Conte, M.I.; Barozzi, S.; Lavagnino, Z.; Carletti, T.; Volpe, M.C.; Cavalcante, P.; Iannacone, M.; Rampazzo, C.; Bussani, R.; Tripodo, C.; Zacchigna, S.; Marcello, A.; d’Adda di Fagagna, F. SARS-CoV-2 infection induces DNA damage, through CHK1 degradation and impaired 53BP1 recruitment, and cellular senescence. Nature Cell Biology 2023, 25, 550–564. [Google Scholar] [CrossRef]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 1-2, 19–25. [Google Scholar] [CrossRef]
- Huang, J.; Rauscher, S.; Nawrocki, G.; Ran, T.; Feig, M.; de Groot, B.L.; Grubmüller, H.; MacKerell Jr, A.D. CHARMM36m: An improved force field for folded and intrinsically disordered proteins. Nature Methods 2016, 14, 71–73. [Google Scholar] [CrossRef] [PubMed]
- Klauda, J.B.; Venable, R.M.; Freites, J.A.; O’Connor, J.W.; Tobias, D.J.; Mondragon-Ramirez, C.; Vorobyov, I.; MacKerell, A.D.J.; Pastor, R.W. Update of the CHARMM All-Atom Additive Force Field for Lipids: Validation on Six Lipid Types. The Journal of Physical Chemistry B 2010, 114, 7830–7843. [Google Scholar] [CrossRef] [PubMed]
- Berendsen, H.J.C.; Postma, J.P.M.; van Gunsteren, W.F.; DiNola, A.; Haak, J.R. Molecular dynamics with coupling to an external bath. The Journal of Chemical Physics 1984, 81, 3684. [Google Scholar] [CrossRef]
- Bussi, G.; Donadio, D.; Parrinello, M. Canonical sampling through velocity rescaling. The Journal of chemical physics 2007, 126, 014101. [Google Scholar] [CrossRef] [PubMed]
- Parrinello, M.; Rahman, A. Crystal Structure and Pair Potentials: A Molecular-Dynamics Study. Physical Review Letters 1980, 45, 1196. [Google Scholar] [CrossRef]
- Hess, B. P-LINCS: A Parallel Linear Constraint Solver for Molecular Simulation. Journal of Chemical Theory and Computation 2008, 4, 116–122. [Google Scholar] [CrossRef]
- https://ds-814.cr.cnaf.infn.it:8443/ex4cov-public/Molecular_Dynamics/ORF6/, 2020. Accessed:.
- Casares, D.; Escribá, P.V.; Rosselló, C.A. Membrane Lipid Composition: Effect on Membrane and Organelle Structure, Function and Compartmentalization and Therapeutic Avenues. International Journal of Molecular Sciences 2019, 20. [Google Scholar] [CrossRef]
- Wu, E.L.; Cheng, X.; Jo, S.; Rui, H.; Song, K.C.; Dávila-Contreras, E.M.; Qi, Y.; Lee, J.; Monje-Galvan, V.; Venable, R.M.; Klauda, J.B.; Im, W. CHARMM-GUI Membrane Builder toward realistic biological membrane simulations. Journal of Computational Chemistry 2014, 35, 1997–2004. [Google Scholar] [CrossRef]
- Jo, S.; Kim, T.; Iyer, V.G.; Im, W. CHARMM-GUI: A web-based graphical user interface for CHARMM. Journal of Computational Chemistry 2008, 29, 1859–1865. [Google Scholar] [CrossRef]
- Emsley, P.; Cowtan, K. Coot: Model-building tools for molecular graphics. Acta Crystallographica Section D 2004, 60, 2126–2132. [Google Scholar] [CrossRef] [PubMed]
- Daura, X.; Gademann, K.; Jaun, B.; Seebach, D.; van Gunsteren, W.F.; Mark, A.E. Peptide Folding: When Simulation Meets Experiment. Angewandte Chemie International Edition 1999, 38, 236–240. [Google Scholar] [CrossRef]
- Frishman, D.; Argos, P. Knowledge-based secondary structure assignment. Proteins: Structure, function and genetics 1995, 23, 566–579. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD – Visual Molecular Dynamics. Journal of Molecular Graphics 1996, 14, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Prendergast, L.; McClurg, U.L.; Hristova, R.; Berlinguer-Palmini, R.; Greener, S.; Veitch, K.; Hernandez, I.; Pasero, P.; Rico, D.; Higgins, J.M.G.; Gospodinov, A.; Papamichos-Chronakis, M. Resolution of R-loops by INO80 promotes DNA replication and maintains cancer cell proliferation and viability. Nature Communications 2020, 11, 4534. [Google Scholar] [CrossRef] [PubMed]
- Stirling, D.R.; Swain-Bowden, M.J.; Lucas, A.M.; Carpenter, A.E.; Cimini, B.A.; Goodman, A. CellProfiler 4: Improvements in speed, utility and usability. BMC Bioinformatics 2021, 22, 433. [Google Scholar] [CrossRef] [PubMed]
- Danovski, G.; Dyankova, T.; Stoynov, S. CellTool: An open source software combining bio-image analysis and mathematical modeling. bioRxiv 2018. [Google Scholar] [CrossRef]
- Schwab, R.A.; Niedzwiedz, W. Visualization of DNA Replication in the Vertebrate Model System DT40 using the DNA Fiber Technique. Journal of Visualized Experiments 2011, 56, e3255. [Google Scholar] [CrossRef]
- Kirilov, T.; Gospodinov, A.; Kirilov, K. An algorithm and application to efficiently analyse DNA fibre data. Biotechnology & Biotechnological Equipment 2023, 37, 2206488. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



