Submitted:
26 May 2023
Posted:
30 May 2023
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Results
2.1. Specific methylated cores mark the promoters of genes involved in murine postnatal brain differentiation.
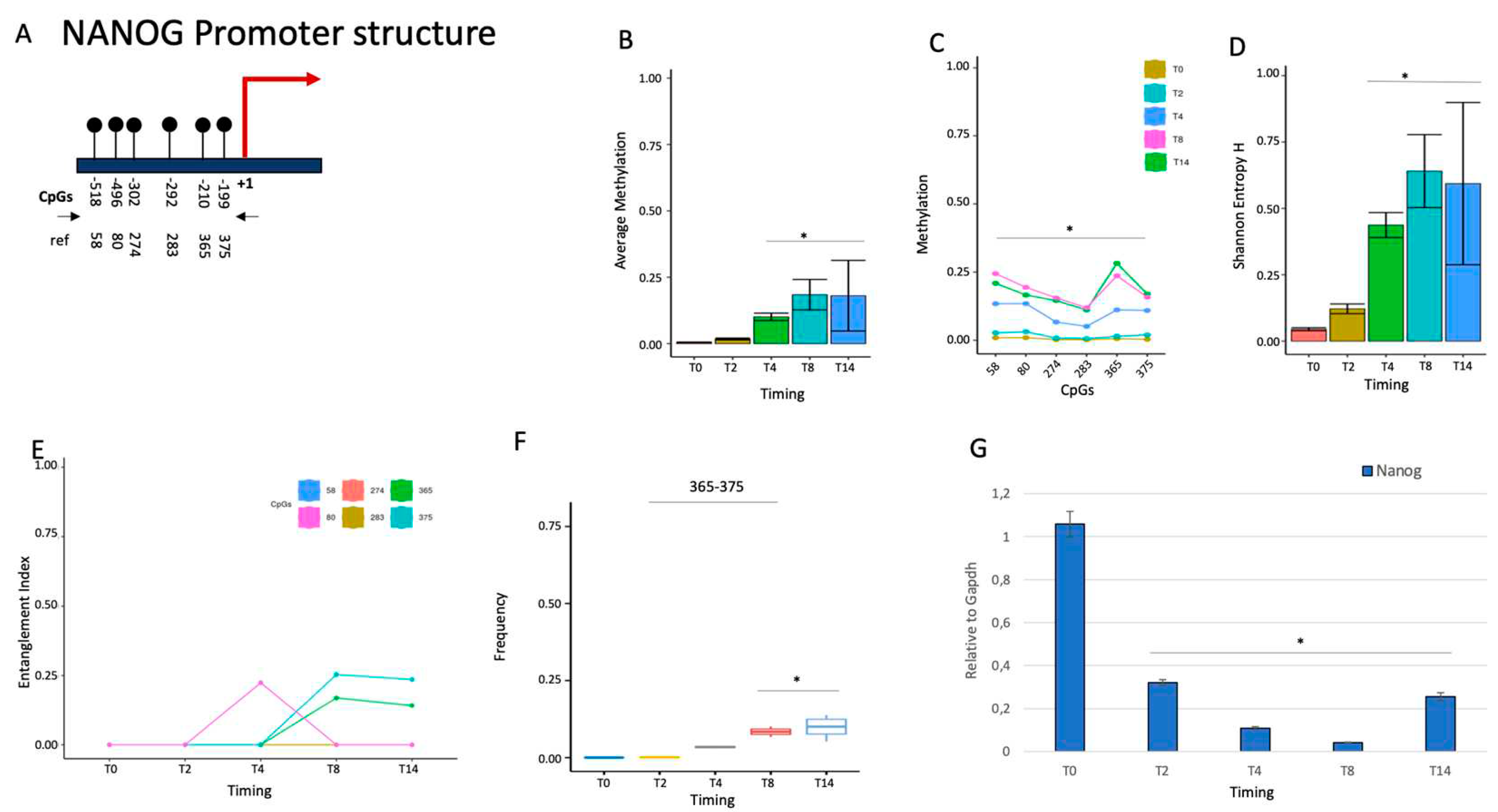
2.1.1. NANOG repression is associated with de novo formation of a methylated core in the promoter.
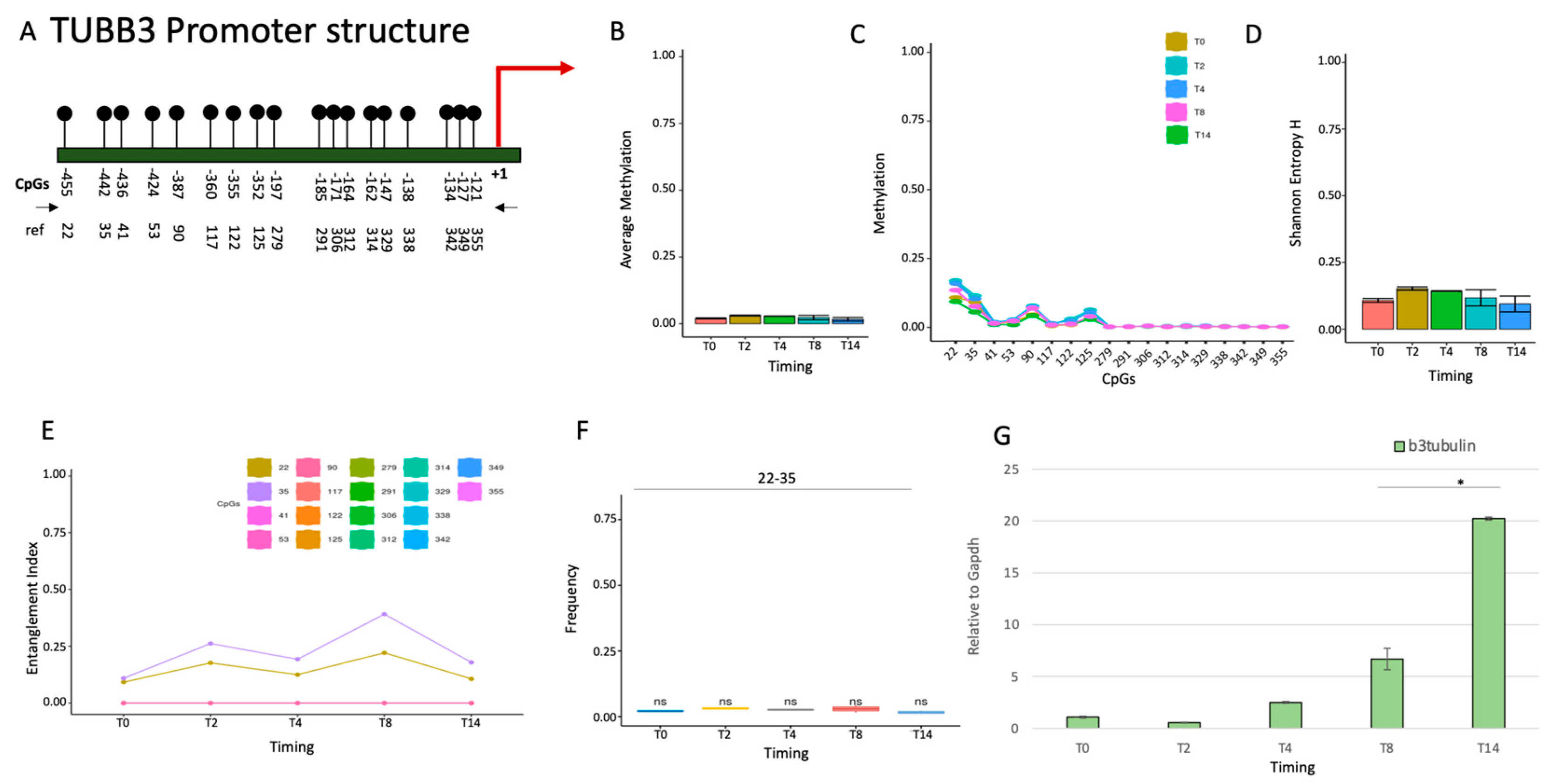
2.1.2. TUBB3 promoter methylation does not control the expression during neuronal differentiation.
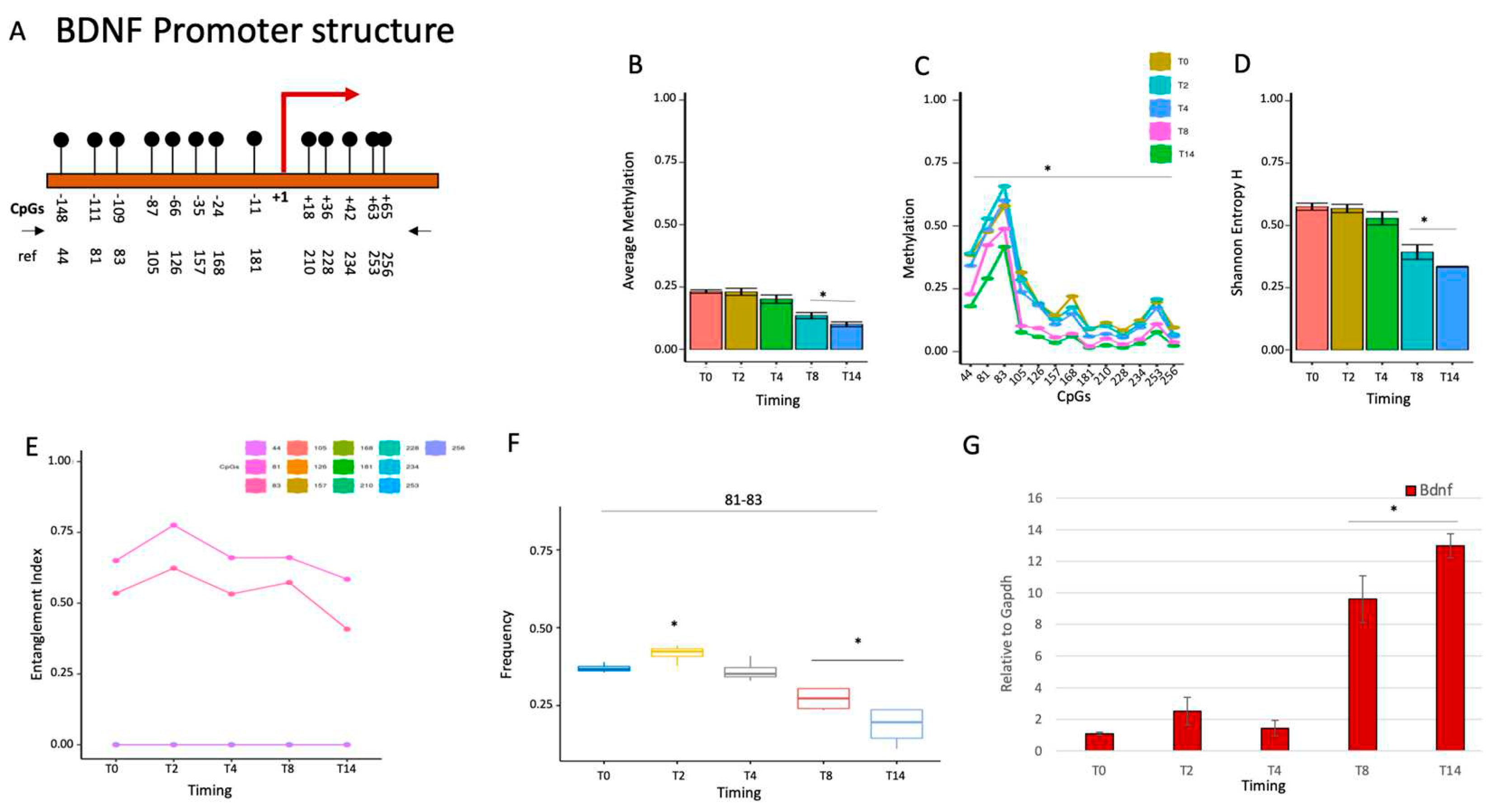
2.1.3. BDNF gene activation parallels the loss of the methylated core at the promoter during brain differentiation.
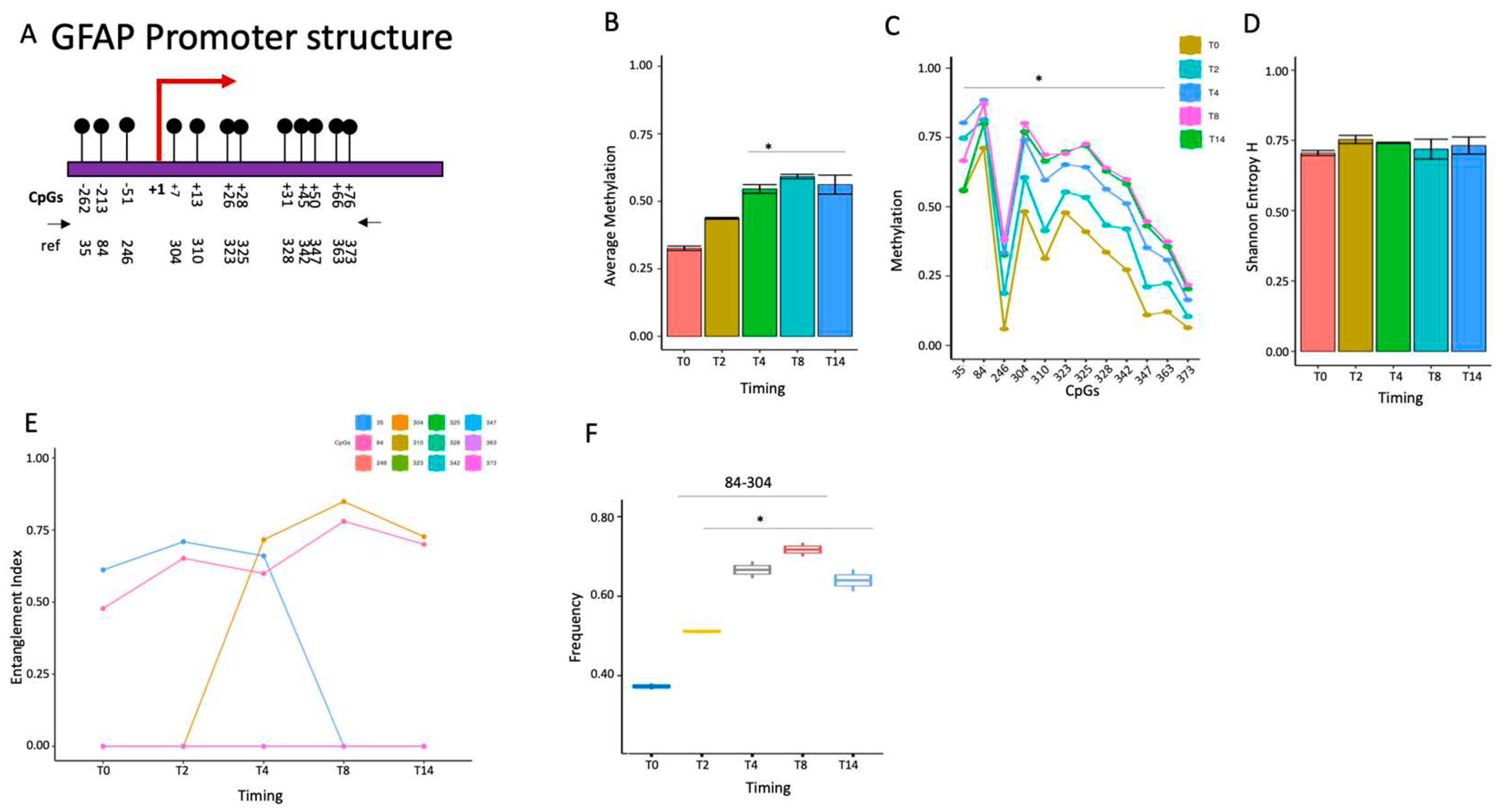
2.1.4. GFAP gene activation changes the configuration of the promoter methylated core.
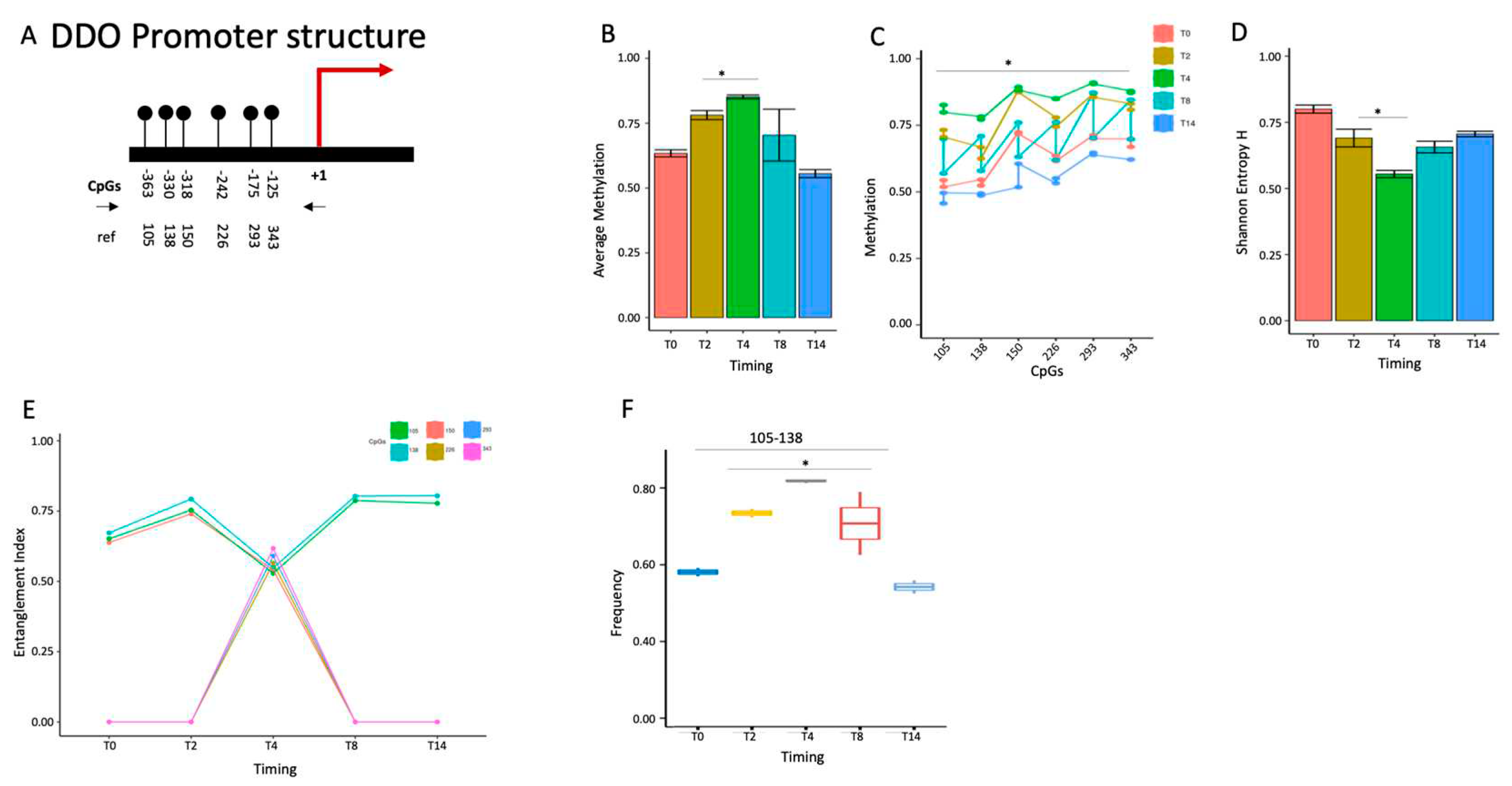
2.1.5. DDO gene promoter methylation profile and expression during postnatal brain differentiation.
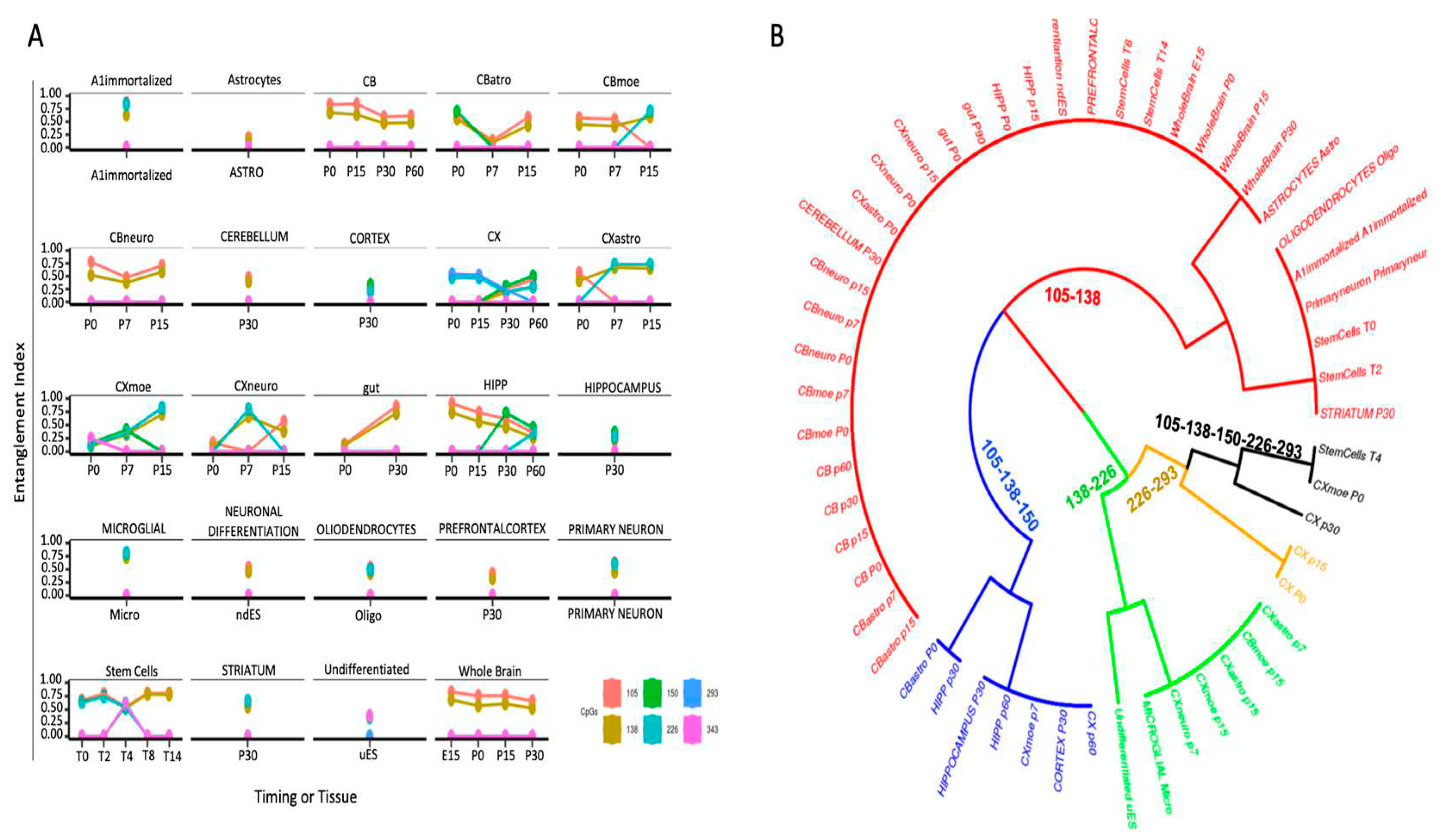
2.2. CpG methylation cores define cell identity.
3. Discussion
4. Materials and Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Moore, L., Le, T. & Fan, G. DNA Methylation and Its Basic Function. Neuropsychopharmacol 38, 23–38 (2013). [CrossRef]
- Tramontano A., Boffo F.L., Russo G., De Rosa M., Iodice I., Pezone A. Methylation of the suppressor gene: mechanism and consequences. Biomolecules. 2020; 10:446.
- Jin B, Ernst J, Tiedemann RL, et al. Linking DNA methyltransferases to epigenetic marks and nucleosome structure genome-wide in human tumor cells. Cell Rep. 2012;2(5):1411-1424. [CrossRef]
- Huang Y., Pastor W.A., Shen Y., Tahiliani M., Liu D.R., Rao A. The behavior of 5-hydroxymethylcytosine in bisulfite sequencing. PLoS One. 2010; 5:e8888.
- Rasmussen KD, Helin K. Role of TET enzymes in DNA methylation, development, and cancer. Genes Dev. 2016 Apr 1;30(7):733-50. PMID: 27036965; PMCID: PMC4826392. [CrossRef]
- Huang Y, Pastor WA, Shen Y, Tahiliani M, Liu DR, Rao A. The behavior of 5-hydroxymethylcytosine in bisulfite sequencing. PLoS One. 2010;5(1):e8888. Published 2010 Jan 26. [CrossRef]
- Song CX, He C. Potential functional roles of DNA demethylation intermediates. Trends Biochem Sci. 2013;38(10):480-484. [CrossRef]
- Xie H., Wang M., de Andrade A., Bonaldo M. de F., Galat V., Arndt K., Soares M.B. Genome-wide quantitative assessment of variation in DNA methylation patterns. Nucleic Acids Res. 2011; 39:4099–4108. [CrossRef]
- Barros-Silva D, Marques CJ, Henrique R, Jerónimo C. Profiling DNA Methylation Based on Next-Generation Sequencing Approaches: New Insights and Clinical Applications. Genes (Basel). 2018 Aug 23;9(9):429. PMID: 30142958; PMCID: PMC6162482. [CrossRef]
- Pezone A, Tramontano A, Scala G, et al. Tracing and tracking epiallele families in complex DNA populations. NAR Genom Bioinform. 2020;2(4):lqaa096. Published 2020 Nov 16. [CrossRef]
- Podobinska M, Szablowska-Gadomska I, Augustyniak J, Sandvig I, Sandvig A, Buzanska L. Epigenetic Modulation of Stem Cells in Neurodevelopment: The Role of Methylation and Acetylation. Front Cell Neurosci. 2017;11:23. Published 2017 Feb 7. [CrossRef]
- Kim M., Costello J. DNA methylation: an epigenetic mark of cellular memory. Exp. Mol. Med. 2017; 49:e322.
- Basu, A., Tiwari, V.K. Epigenetic reprogramming of cell identity: lessons from development for regenerative medicine. Clin Epigenet 13, 144 (2021). [CrossRef]
- Yang, Y.; Fan, X.; Yan, J.; Chen, M.; Zhu, M.; Tang, Y.; Liu, S.; Tang, Z. A comprehensive epigenome atlas reveals DNA methylation regulating skeletal muscle development. Nucleic Acids Res. 2021, 49, 1313–1329. [CrossRef]
- Li S, Garrett-Bakelman F, Perl AE, et al. Dynamic evolution of clonal epialleles revealed by methclone. Genome Biol. 2014;15(9):472. Published 2014 Sep 27. [CrossRef]
- Li S, Garrett-Bakelman FE, Chung SS, et al. Distinct evolution and dynamics of epigenetic and genetic heterogeneity in acute myeloid leukemia. Nat Med. 2016;22(7):792-799. [CrossRef]
- Shannon C.E. A mathematical theory of communication. Bell Syst. Tech. J. 1948; 27:379–423.
- Verma R, Liu J, Holland MK, Temple-Smith P, Williamson M, Verma PJ. Nanog is an essential factor for induction of pluripotency in somatic cells from endangered felids. Biores Open Access. 2013;2(1):72-76. [CrossRef]
- Radwitz J, Hausrat TJ, Heisler FF, Janiesch PC, Pechmann Y, Rübhausen M, Kneussel M. Tubb3 expression levels are sensitive to neuronal activity changes and determine microtubule growth and kinesin-mediated transport. Cell Mol Life Sci. 2022 Oct 29;79(11):575. PMID: 36309617; PMCID: PMC9617967. [CrossRef]
- Errico F, Cuomo M, Canu N, Caputo V, Usiello A. New insights on the influence of free d-aspartate metabolism in the mammalian brain during prenatal and postnatal life. Biochim Biophys Acta Proteins Proteom. 2020;1868(10):140471. [CrossRef]
- Jurga AM, Paleczna M, Kadluczka J, Kuter KZ. Beyond the GFAP-Astrocyte Protein Markers in the Brain. Biomolecules. 2021 Sep 14;11(9):1361. PMID: 34572572; PMCID: PMC8468264. [CrossRef]
- Bathina S, Das UN. Brain-derived neurotrophic factor and its clinical implications. Arch Med Sci. 2015 Dec 10;11(6):1164-78. Epub 2015 Dec 11. PMID: 26788077; PMCID: PMC4697050. [CrossRef]
- Busskamp V, Lewis NE, Guye P, et al. Rapid neurogenesis through transcriptional activation in human stem cells. Mol Syst Biol. 2014;10(11):760. Published 2014 Nov 17. [CrossRef]
- Pang ZP, Yang N, Vierbuchen T, et al. Induction of human neuronal cells by defined transcription factors. Nature. 2011;476(7359):220-223. Published 2011 May 26. [CrossRef]
- Raspaglio G, Petrillo M, Martinelli E, et al. Sox9 and Hif-2α regulate TUBB3 gene expression and affect ovarian cancer aggressiveness. Gene. 2014;542(2):173-181. [CrossRef]
- He M, Wang J. Decreased serum brain-derived neurotrophic factor in Chinese patients with Type 2 diabetes mellitus. Acta Biochim Biophys Sin (Shanghai). 2014;46(5):426-427. [CrossRef]
- Mamber C, Kamphuis W, Haring NL, Peprah N, Middeldorp J, Hol EM. GFAPδ expression in glia of the developmental and adolescent mouse brain. PLoS One. 2012;7(12):e52659. [CrossRef]
- Ahmed AI, Shtaya AB, Zaben MJ, Owens EV, Kiecker C, Gray WP. Endogenous GFAP-positive neural stem/progenitor cells in the postnatal mouse cortex are activated following traumatic brain injury. J Neurotrauma. 2012;29(5):828-842. [CrossRef]
- Koga R., Miyoshi Y., Sakaue H., Hamase K., Konno R. Mouse D-amino-acid oxidase: distribution and physiological substrates. Front.Mol. Biosci. 2017; 4:82. [CrossRef]
- Florio E, Keller S, Coretti L, et al. Tracking the evolution of epialleles during neural differentiation and brain development: D-Aspartate oxidase as a model gene. Epigenetics. 2017;12(1):41-54. [CrossRef]
- Punzo D, Errico F, Cristino L, et al. Age-Related Changes in D-Aspartate Oxidase Promoter Methylation Control Extracellular D-Aspartate Levels and Prevent Precocious Cell Death during Brain Aging. J Neurosci. 2016;36(10):3064-3078. [CrossRef]
- Breiling, A., Lyko, F. Epigenetic regulatory functions of DNA modifications: 5-methylcytosine and beyond. Epigenetics & Chromatin 8, 24 (2015). [CrossRef]
- Luo C, Hajkova P, Ecker JR. Dynamic DNA methylation: In the right place at the right time. Science. 2018;361(6409):1336-1340. [CrossRef]
- Fuentealba L.C., Rompani S.B., Parraguez J.I., Obernier K., Romero R., Cepko C.L., Alvarez-Buylla A. Embryonic origin of postnatal neural stem cells. Cell. 2015; 161:1644–1655. [CrossRef]
- Anderson RB, Newgreen DF, Young HM. Neural Crest and the Development of the Enteric Nervous System. In: Madame Curie Bioscience Database [Internet]. Austin (TX): Landes Bioscience; 2000-2013. Available from: https://www.ncbi.nlm.nih.gov/books/NBK6273/.
- Niwa, H., Miyazaki, Ji. & Smith, A. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat Genet 24, 372–376 (2000). [CrossRef]
- Maltsev VA, Rohwedel J, Hescheler J, Wobus AM. Embryonic stem cells differentiate in vitro into cardiomyocytes representing sinusnodal, atrial and ventricular cell types. Mech Dev. 1993;44(1):41-50. [CrossRef]
- Mònica Suelves, Elvira Carrió, Yaiza Núñez-Álvarez, Miguel A. Peinado, DNA methylation dynamics in cellular commitment and differentiation, Briefings in Functional Genomics, Volume 15, Issue 6, November 2016, Pages 443–453. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



