Submitted:
04 May 2023
Posted:
08 May 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. RNAi-mediated downregulation of the evolutionary conserved dSAMTOR protein causes either lethal or viable phenotypes: a gene-targeting efficiency dependence
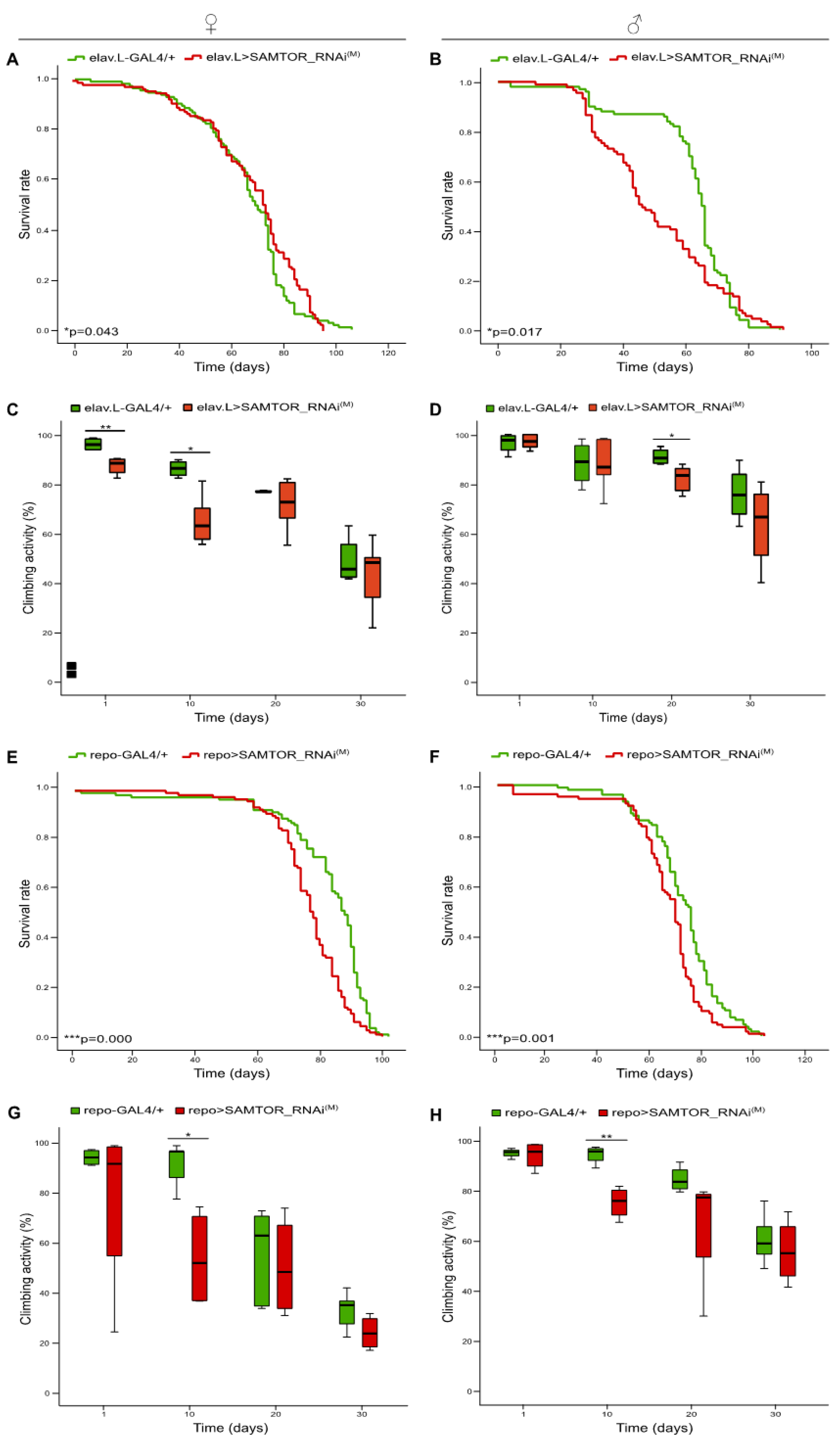
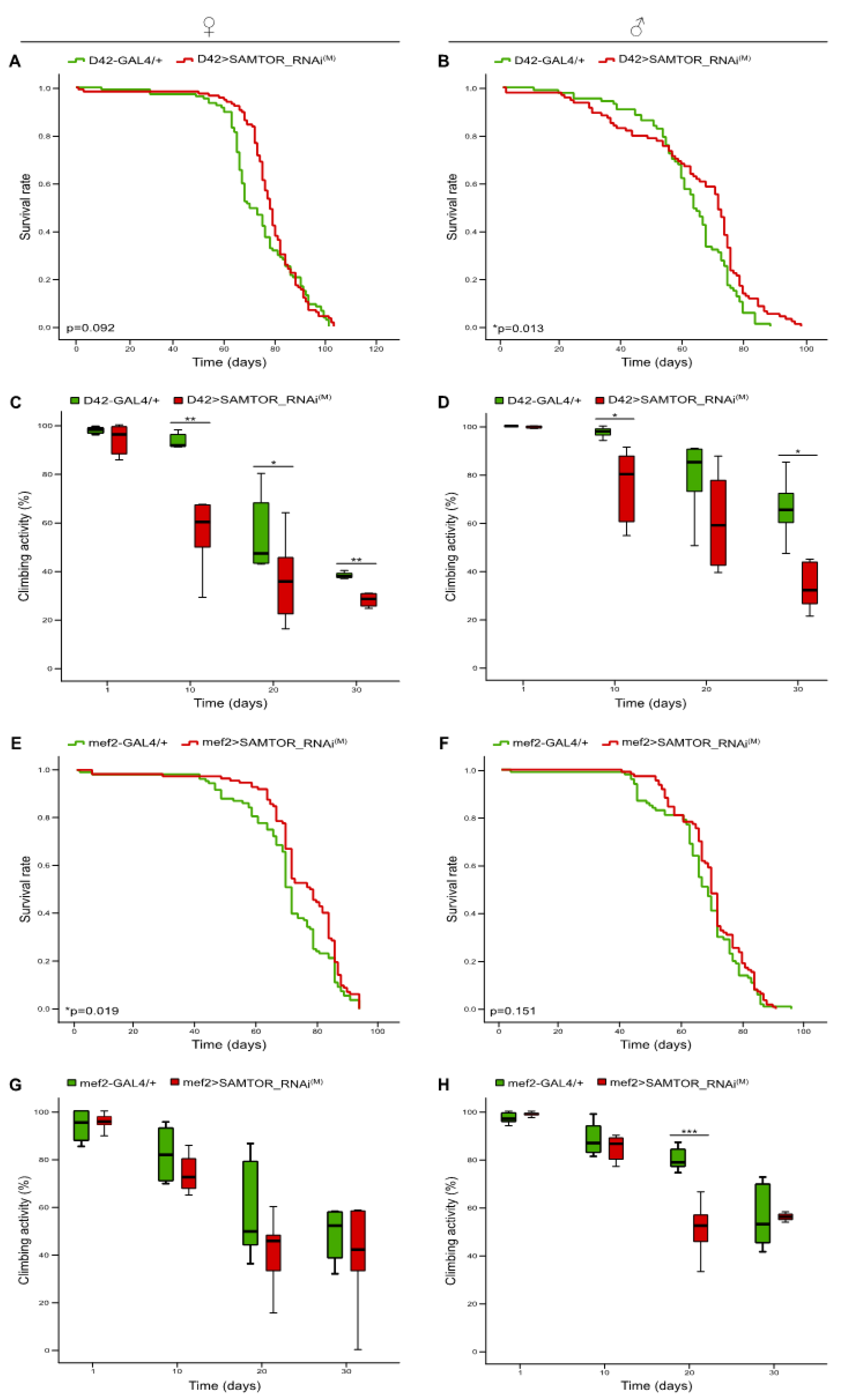
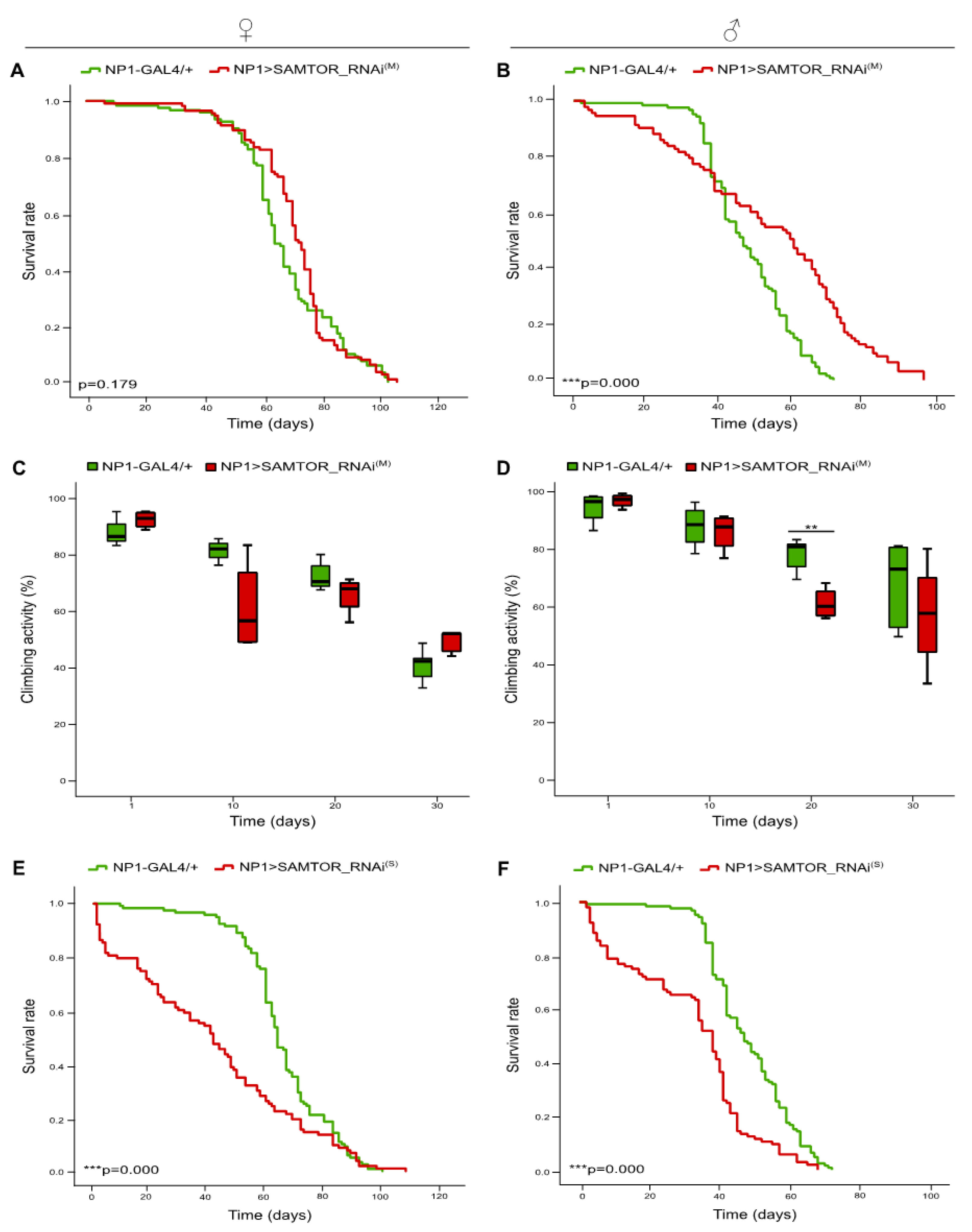
2.2. Differential contribution of neuronal tissue-, glia cell-, motor neuron- and muscle tissue-specific moderate silencing of dSAMTOR gene to fly survival and kinetic capacity, during aging
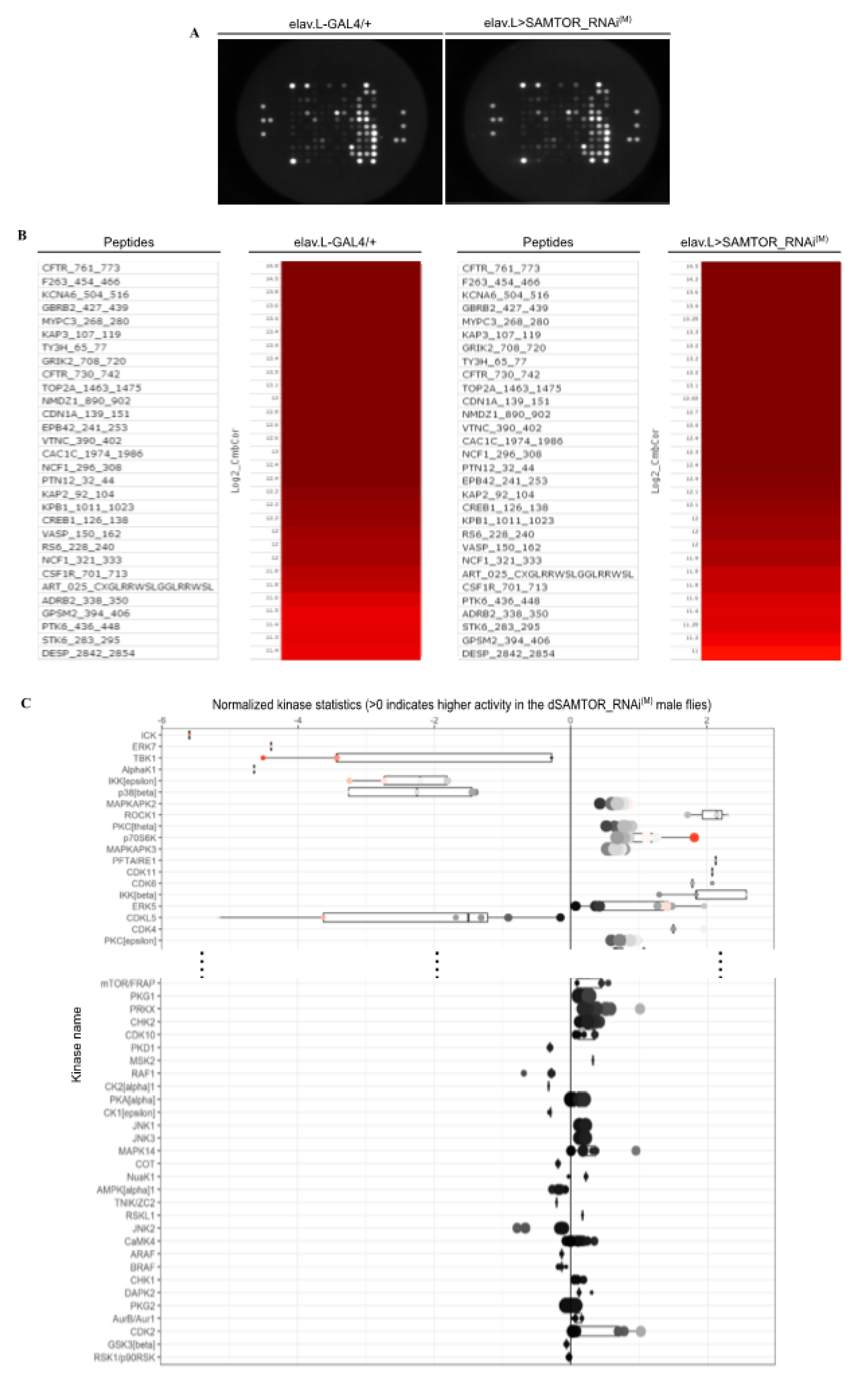
2.4. Functional kinome profiling of mildly dSAMTOR-downregulated transgenic flies, specifically in neuronal tissues
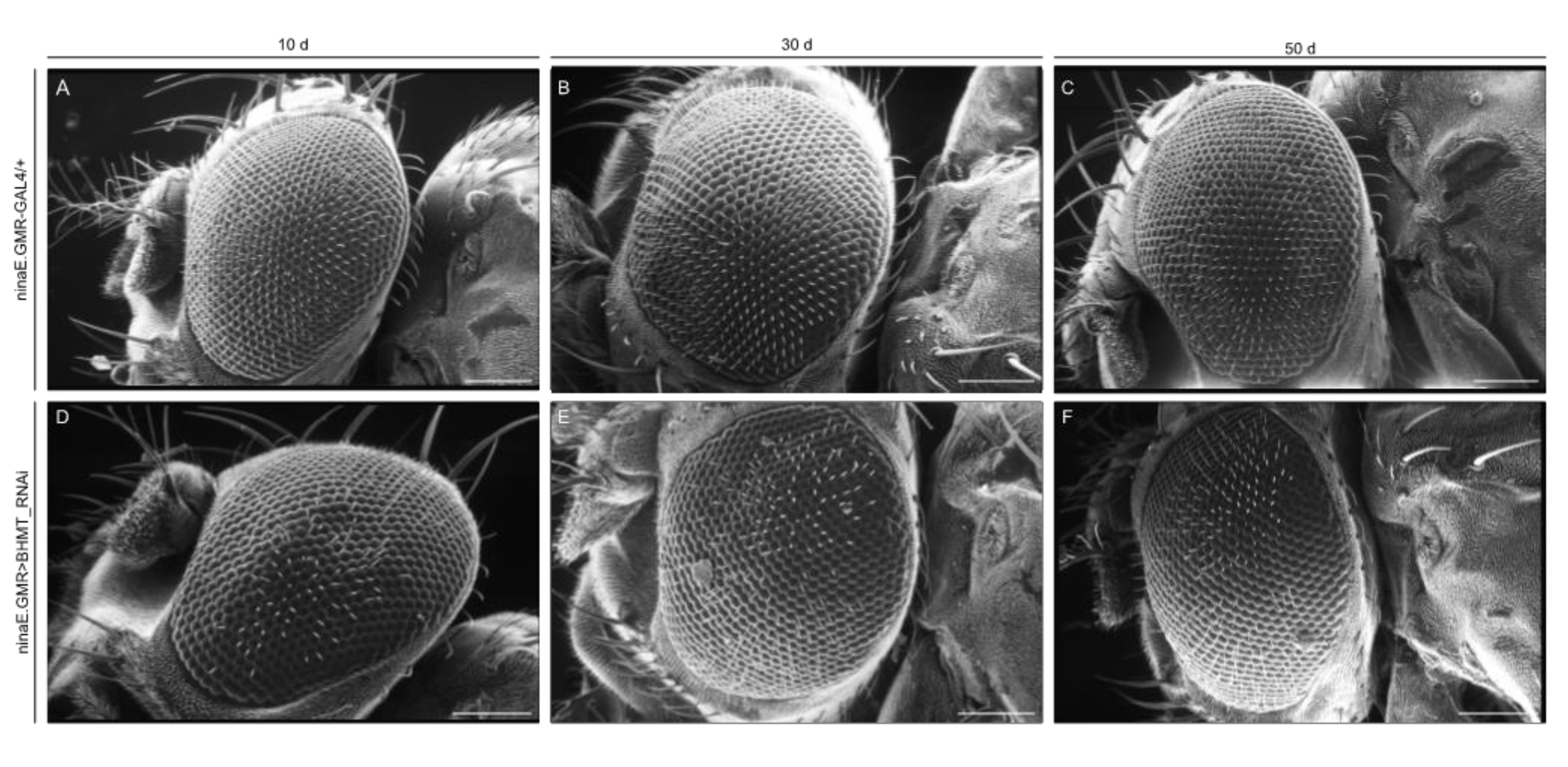
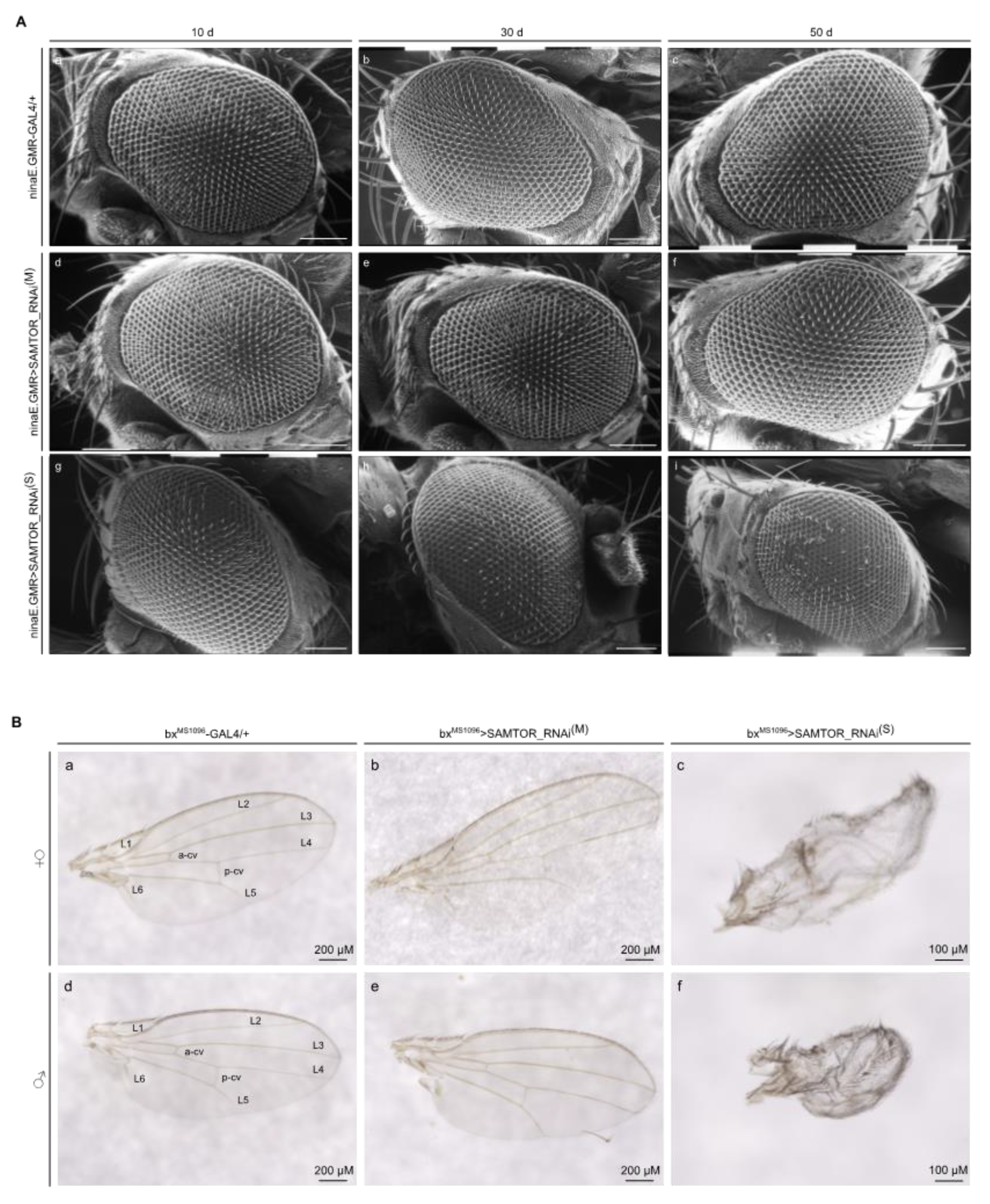
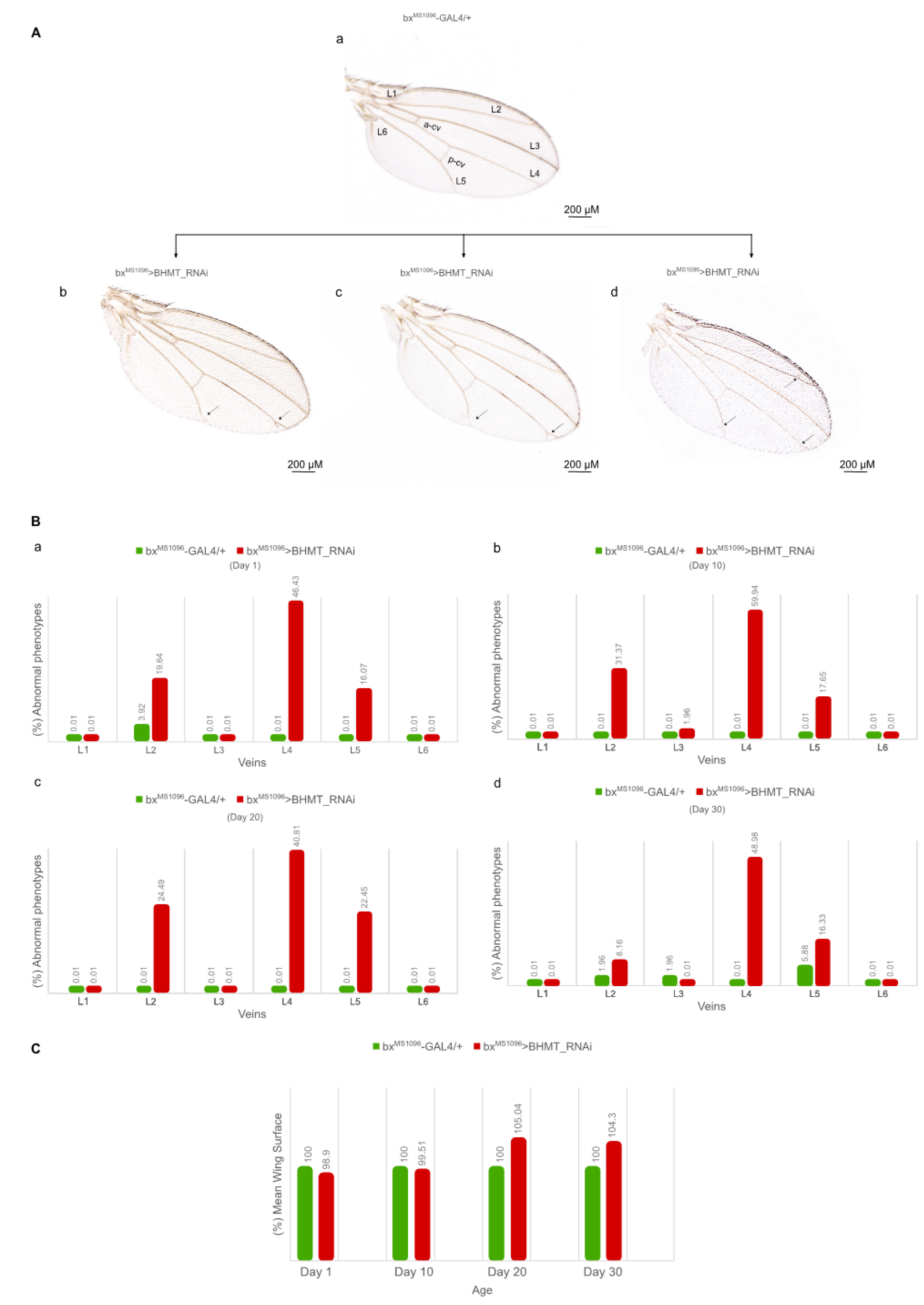
2.5. Strong downregulation of dSAMTOR gene expression causes eye and wing dysmorphic phenotypes in Drosophila
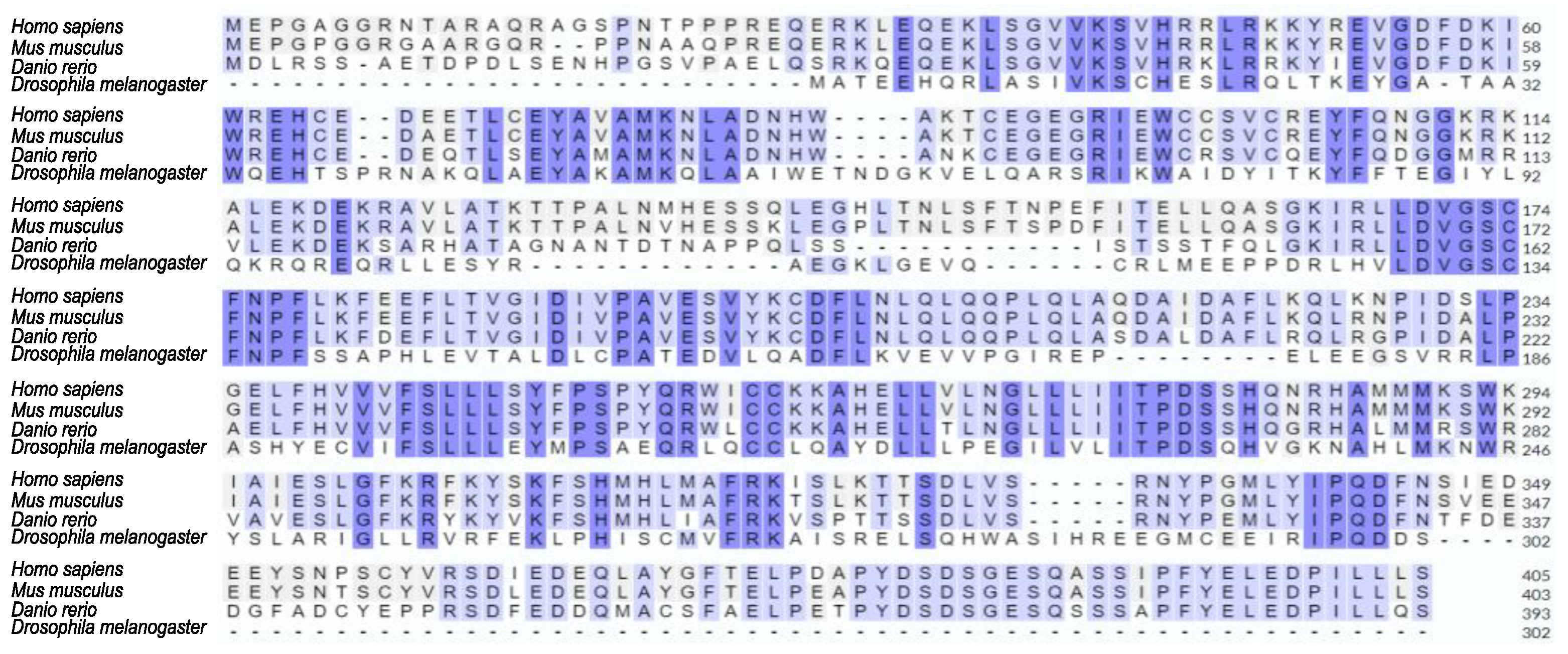
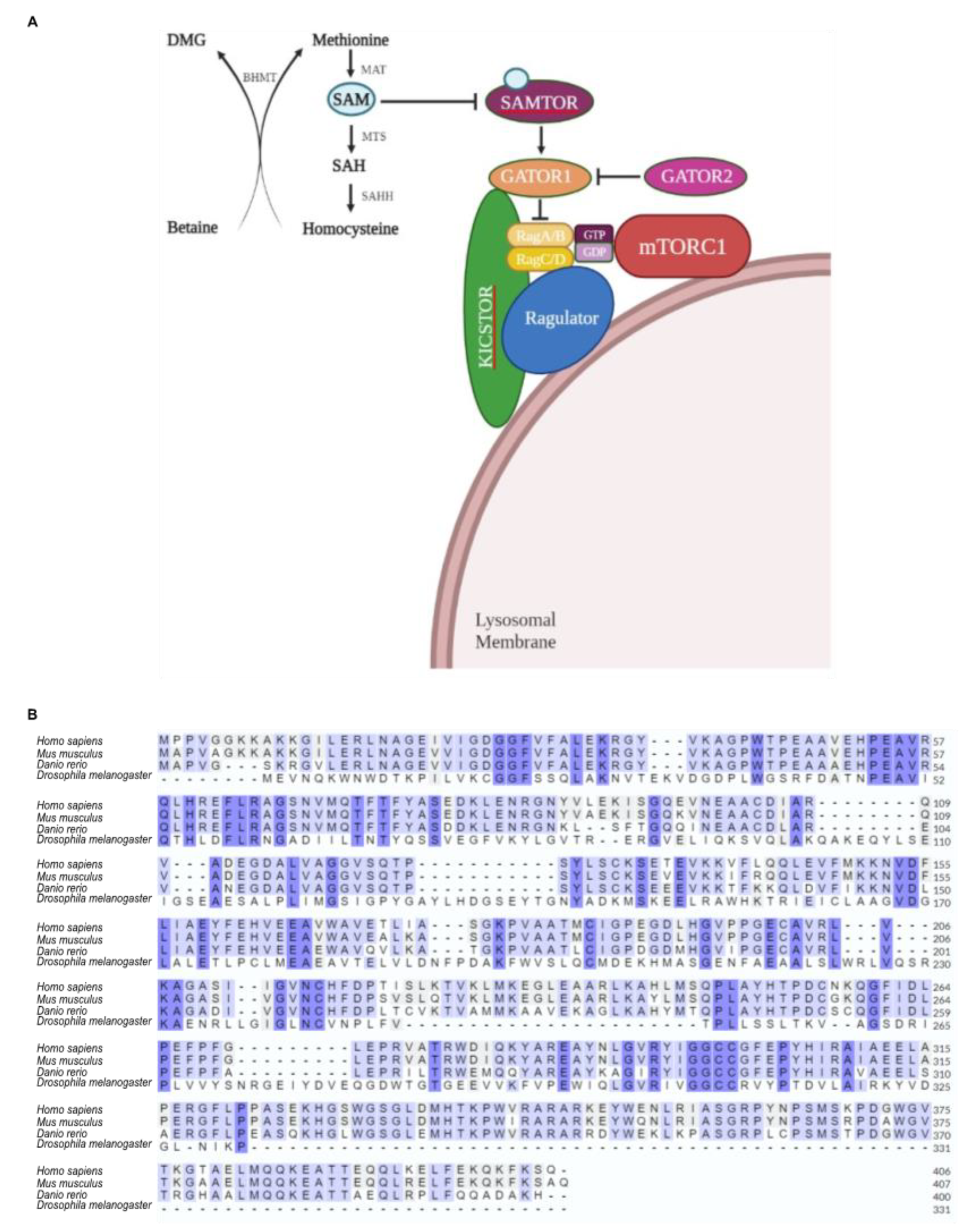
2.6. Sensing and metabolism of Methionine are evolutionary conserved pathways
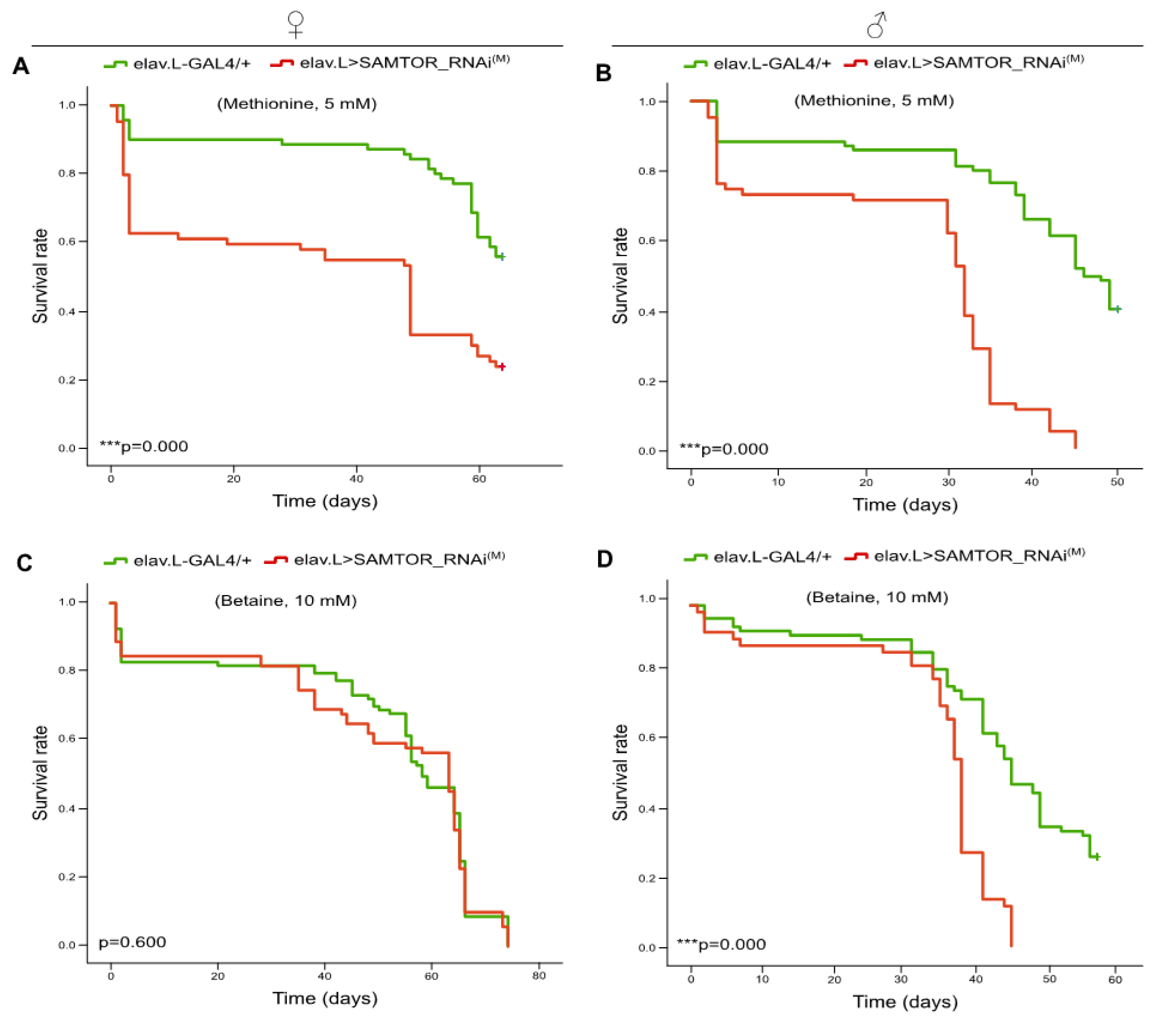
2.7. Methionine and Betaine administration induce reduced viabilities in transgenic flies, bearing moderately suppressed dSAMTOR gene expression, specifically in neuronal tissues
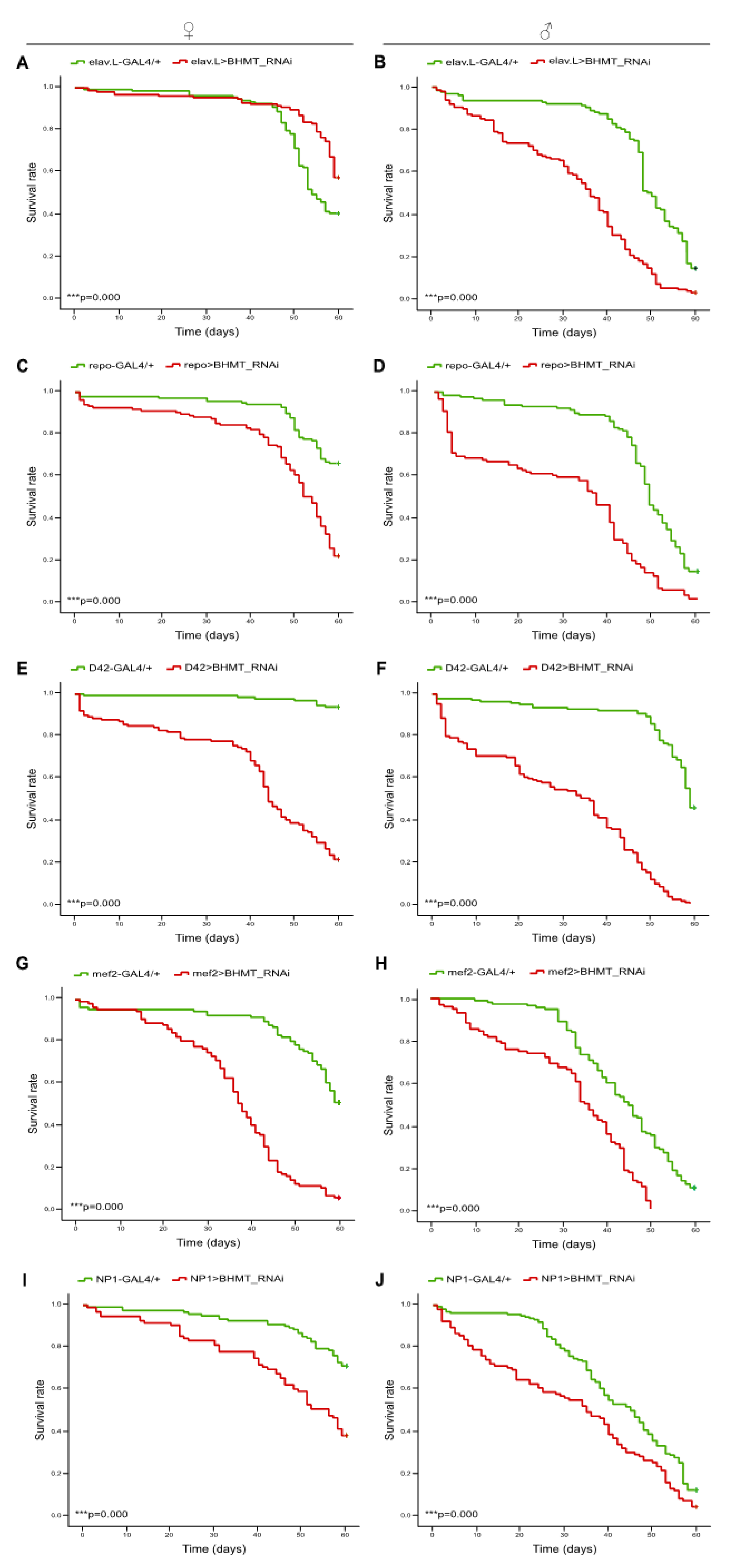
2.8. Lack of dBHMT gene expression, specifically in neuronal-, glial-, motor neuron- muscle- and midgut-tissues shortens Drosophila’s longevity
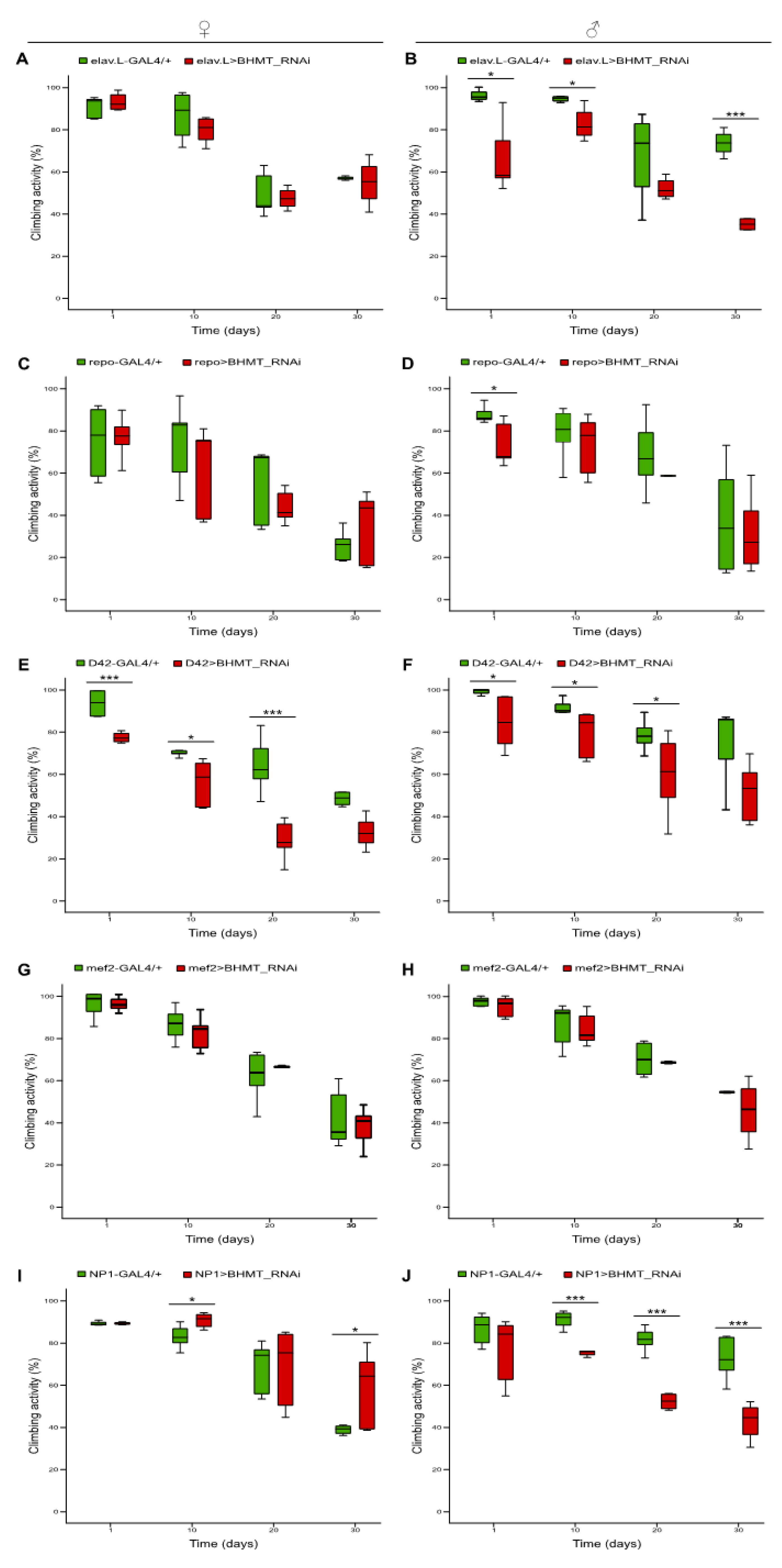
2.9. Differential contribution of neuronal, glial, motor neuron, muscle and midgut tissue-specific silencing of dBHMT gene expression to Drosophila locomotion, during aging
3. Discussion
4. Materials and Methods
4.1. Fly stocks and culturing conditions (Drosophila husbandry)
4.2. Chemicals and reagents
4.3. Longevity measurement
4.4. Negative geotaxis assay
4.5. Statistical analysis
4.6. Light microscopy (LM)
4.7. Scanning electron microscopy (SEM)
4.8. Kinase activity profiling
4.9. RNA extraction and RT-qPCR
Supplementary Materials
Author Contributions
Data Availability Statement
Conflicts of Interest
References
- Sabatini, D.M. , Twenty-five years of mTOR: Uncovering the link from nutrients to growth. Proc Natl Acad Sci U S A 2017, 114, 11818–11825. [Google Scholar] [CrossRef] [PubMed]
- Saxton, R.A.; Sabatini, D.M. , mTOR Signaling in Growth, Metabolism, and Disease. Cell 2017, 168, 960–976. [Google Scholar] [CrossRef] [PubMed]
- Laplante, M.; Sabatini, D.M. , mTOR signaling in growth control and disease. Cell 2012, 149, 274–293. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.C.; Guan, K.L. , mTOR: a pharmacologic target for autophagy regulation. J Clin Invest 2015, 125, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.Y.; Sabatini, D.M. , mTOR at the nexus of nutrition, growth, ageing and disease. Nat Rev Mol Cell Biol 2020, 21, 183–203. [Google Scholar] [CrossRef] [PubMed]
- Deleyto-Seldas, N.; Efeyan, A. , The mTOR-Autophagy Axis and the Control of Metabolism. Front Cell Dev Biol 2021, 9, 655731. [Google Scholar] [CrossRef]
- Gu, X.; Orozco, J.M.; Saxton, R.A.; Condon, K.J.; Liu, G.Y.; Krawczyk, P.A.; Scaria, S.M.; Harper, J.W.; Gygi, S.P.; Sabatini, D.M. , SAMTOR is an S-adenosylmethionine sensor for the mTORC1 pathway. Science 2017, 358, 813–818. [Google Scholar] [CrossRef]
- Hesketh, G.G.; Papazotos, F.; Pawling, J.; Rajendran, D.; Knight, J.D.R.; Martinez, S.; Taipale, M.; Schramek, D.; Dennis, J.W.; Gingras, A.C. , The GATOR-Rag GTPase pathway inhibits mTORC1 activation by lysosome-derived amino acids. Science 2020, 370, 351–356. [Google Scholar] [CrossRef]
- Bar-Peled, L.; Sabatini, D.M. , Regulation of mTORC1 by amino acids. Trends Cell Biol 2014, 24, 400–406. [Google Scholar] [CrossRef]
- Anandapadamanaban, M.; Masson, G.R.; Perisic, O.; Berndt, A.; Kaufman, J.; Johnson, C.M.; Santhanam, B.; Rogala, K.B.; Sabatini, D.M.; Williams, R.L. , Architecture of human Rag GTPase heterodimers and their complex with mTORC1. Science 2019, 366, 203–210. [Google Scholar] [CrossRef]
- Wolfson, R.L.; Chantranupong, L.; Saxton, R.A.; Shen, K.; Scaria, S.M.; Cantor, J.R.; Sabatini, D.M. , Sestrin2 is a leucine sensor for the mTORC1 pathway. Science 2016, 351, 43–48. [Google Scholar] [CrossRef] [PubMed]
- Wolfson, R.L.; Sabatini, D.M. , The Dawn of the Age of Amino Acid Sensors for the mTORC1 Pathway. Cell Metab 2017, 26, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Kitada, M.; Xu, J.; Ogura, Y.; Monno, I.; Koya, D. , Mechanism of Activation of Mechanistic Target of Rapamycin Complex 1 by Methionine. Front Cell Dev Biol 2020, 8, 715. [Google Scholar] [CrossRef] [PubMed]
- Coolon, J.D.; Stevenson, K.R.; McManus, C.J.; Graveley, B.R.; Wittkopp, P.J. , Genomic imprinting absent in Drosophila melanogaster adult females. Cell Rep 2012, 2, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Guan, K.L. , mTOR as a central hub of nutrient signalling and cell growth. Nat Cell Biol 2019, 21, 63–71. [Google Scholar] [CrossRef] [PubMed]
- Lauinger, L.; Kaiser, P. , Sensing and Signaling of Methionine Metabolism. Metabolites.
- Law, J.A.; Jacobsen, S.E. , Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet 2010, 11, 204–220. [Google Scholar] [CrossRef] [PubMed]
- Mentch, S.J.; Mehrmohamadi, M.; Huang, L.; Liu, X.; Gupta, D.; Mattocks, D.; Gomez Padilla, P.; Ables, G.; Bamman, M.M.; Thalacker-Mercer, A.E.; Nichenametla, S.N.; Locasale, J.W. , Histone Methylation Dynamics and Gene Regulation Occur through the Sensing of One-Carbon Metabolism. Cell Metab 2015, 22, 861–873. [Google Scholar] [CrossRef]
- Locasale, J.W. , Serine, glycine and one-carbon units: cancer metabolism in full circle. Nat Rev Cancer 2013, 13, 572–583. [Google Scholar] [CrossRef]
- Mossmann, D.; Park, S.; Hall, M.N. , mTOR signalling and cellular metabolism are mutual determinants in cancer. Nat Rev Cancer 2018, 18, 744–757. [Google Scholar] [CrossRef]
- Zeng, J.D.; Wu, W.K.K.; Wang, H.Y.; Li, X.X. , Serine and one-carbon metabolism, a bridge that links mTOR signaling and DNA methylation in cancer. Pharmacol Res 2019, 149, 104352. [Google Scholar] [CrossRef]
- Menon, S.; Manning, B.D. , Common corruption of the mTOR signaling network in human tumors. Oncogene 2008, (0 2), S43–51. [Google Scholar] [CrossRef]
- Lipton, J.O.; Sahin, M. , The neurology of mTOR. Neuron 2014, 84, 275–291. [Google Scholar] [CrossRef] [PubMed]
- Efeyan, A.; Zoncu, R.; Chang, S.; Gumper, I.; Snitkin, H.; Wolfson, R.L.; Kirak, O.; Sabatini, D.D.; Sabatini, D.M. , Regulation of mTORC1 by the Rag GTPases is necessary for neonatal autophagy and survival. Nature 2013, 493, 679–683. [Google Scholar] [CrossRef] [PubMed]
- Peng, M.; Yin, N.; Li, M.O. , Sestrins function as guanine nucleotide dissociation inhibitors for Rag GTPases to control mTORC1 signaling. Cell 2014, 159, 122–133. [Google Scholar] [CrossRef] [PubMed]
- Nixon, R.A. , The role of autophagy in neurodegenerative disease. Nat Med 2013, 19, 983–997. [Google Scholar] [CrossRef] [PubMed]
- Hara, T.; Nakamura, K.; Matsui, M.; Yamamoto, A.; Nakahara, Y.; Suzuki-Migishima, R.; Yokoyama, M.; Mishima, K.; Saito, I.; Okano, H.; Mizushima, N. , Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 2006, 441, 885–889. [Google Scholar] [CrossRef] [PubMed]
- Finkelstein, J.D. , Methionine metabolism in mammals. J Nutr Biochem 1990, 1, 228–237. [Google Scholar] [CrossRef] [PubMed]
- Ye, C.; Sutter, B.M.; Wang, Y.; Kuang, Z.; Tu, B.P. , A Metabolic Function for Phospholipid and Histone Methylation. Mol Cell 2017, 66, 180–193. [Google Scholar] [CrossRef]
- Barth, T.K.; Imhof, A. , Fast signals and slow marks: the dynamics of histone modifications. Trends Biochem. Sci. 2010, 35, 618–626. [Google Scholar] [CrossRef]
- Obata, F.; Miura, M. , Enhancing S-adenosyl-methionine catabolism extends Drosophila lifespan. Nature communications 2015, 6, 8332. [Google Scholar] [CrossRef]
- Bjedov, I.; Toivonen, J.M.; Kerr, F.; Slack, C.; Jacobson, J.; Foley, A.; Partridge, L. , Mechanisms of life span extension by rapamycin in the fruit fly Drosophila melanogaster. Cell Metab 2010, 11, 35–46. [Google Scholar] [CrossRef] [PubMed]
- Kitada, M.; Ogura, Y.; Monno, I.; Koya, D. , The impact of dietary protein intake on longevity and metabolic health. EBioMedicine 2019, 43, 632–640. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.C.; Kaya, A.; Ma, S.; Kim, G.; Gerashchenko, M.V.; Yim, S.H.; Hu, Z.; Harshman, L.G.; Gladyshev, V.N. , Methionine restriction extends lifespan of Drosophila melanogaster under conditions of low amino-acid status. Nature communications 2014, 5, 3592. [Google Scholar] [CrossRef] [PubMed]
- Miller, R.A.; Buehner, G.; Chang, Y.; Harper, J.M.; Sigler, R.; Smith-Wheelock, M. , Methionine-deficient diet extends mouse lifespan, slows immune and lens aging, alters glucose, T4, IGF-I and insulin levels, and increases hepatocyte MIF levels and stress resistance. Aging Cell 2005, 4, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Parkhitko, A.A.; Binari, R.; Zhang, N.; Asara, J.M.; Demontis, F.; Perrimon, N. , Tissue-specific down-regulation of S-adenosyl-homocysteine via suppression of dAhcyL1/dAhcyL2 extends health span and life span in Drosophila. Genes Dev 2016, 30, 1409–1422. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.P.; Simpson, S.J.; Clissold, F.J.; Brooks, R.; Ballard, J.W.; Taylor, P.W.; Soran, N.; Raubenheimer, D. , Lifespan and reproduction in Drosophila: New insights from nutritional geometry. Proc Natl Acad Sci U S A 2008, 105, 2498–2503. [Google Scholar] [CrossRef] [PubMed]
- Harper, A.E.; Benevenga, N.J.; Wohlhueter, R.M. , Effects of ingestion of disproportionate amounts of amino acids. Physiol Rev 1970, 50, 428–558. [Google Scholar] [CrossRef]
- Pajares, M.A.; Perez-Sala, D. , Betaine homocysteine S-methyltransferase: just a regulator of homocysteine metabolism? Cell Mol Life Sci 2006, 63, 2792–2803. [Google Scholar] [CrossRef]
- Jensen, K.; McClure, C.; Priest, N.K.; Hunt, J. , Sex-specific effects of protein and carbohydrate intake on reproduction but not lifespan in Drosophila melanogaster. Aging Cell 2015, 14, 605–615. [Google Scholar] [CrossRef]
- Zajitschek, F.; Zajitschek, S.R.; Friberg, U.; Maklakov, A.A. , Interactive effects of sex, social environment, dietary restriction, and methionine on survival and reproduction in fruit flies. Age (Dordr) 2013, 35, 1193–1204. [Google Scholar] [CrossRef]
- Shiraki, N.; Shiraki, Y.; Tsuyama, T.; Obata, F.; Miura, M.; Nagae, G.; Aburatani, H.; Kume, K.; Endo, F.; Kume, S. , Methionine metabolism regulates maintenance and differentiation of human pluripotent stem cells. Cell Metab 2014, 19, 780–794. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. , Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]

| DRIVER | dSAMTOR_RNAi(S) | dSAMTOR_RNAi(M) | |
| STRAIN | |||
| Act5C-GAL4 (whole body) | Lethal | Viable | |
| elav.L-GAL4 (nervous system) | Lethal | Viable | |
| repo-GAL4 (glial cells) | Lethal | Viable | |
| D42-GAL4 (motor neurons) | Lethal | Viable | |
| Mef2.R-GAL4 (muscles) | Lethal | Viable | |
| NP1-GAL4 (midgut) | Viable | Viable | |
| ninaE.GMR-GAL4 (eyes) | Viable | Viable | |
| bxMS1096-GAL4 (wings) | Viable | Viable | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions, or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




