Submitted:
10 March 2023
Posted:
10 March 2023
You are already at the latest version
Abstract

Keywords:
1. Introduction
2. Materials and Methods
2.1. Half-Space Proximal Networks Building
2.2. Metadata Networks
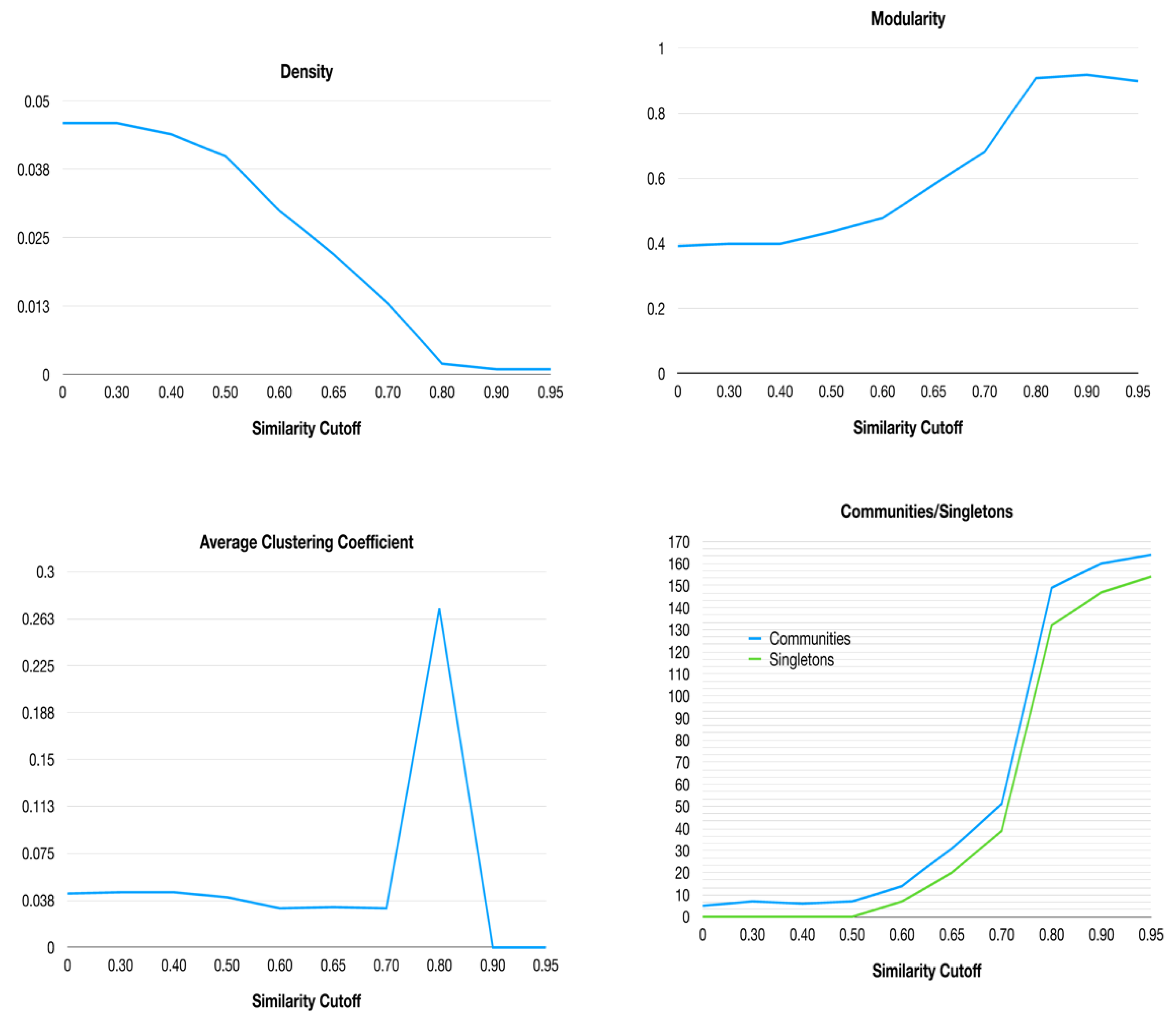
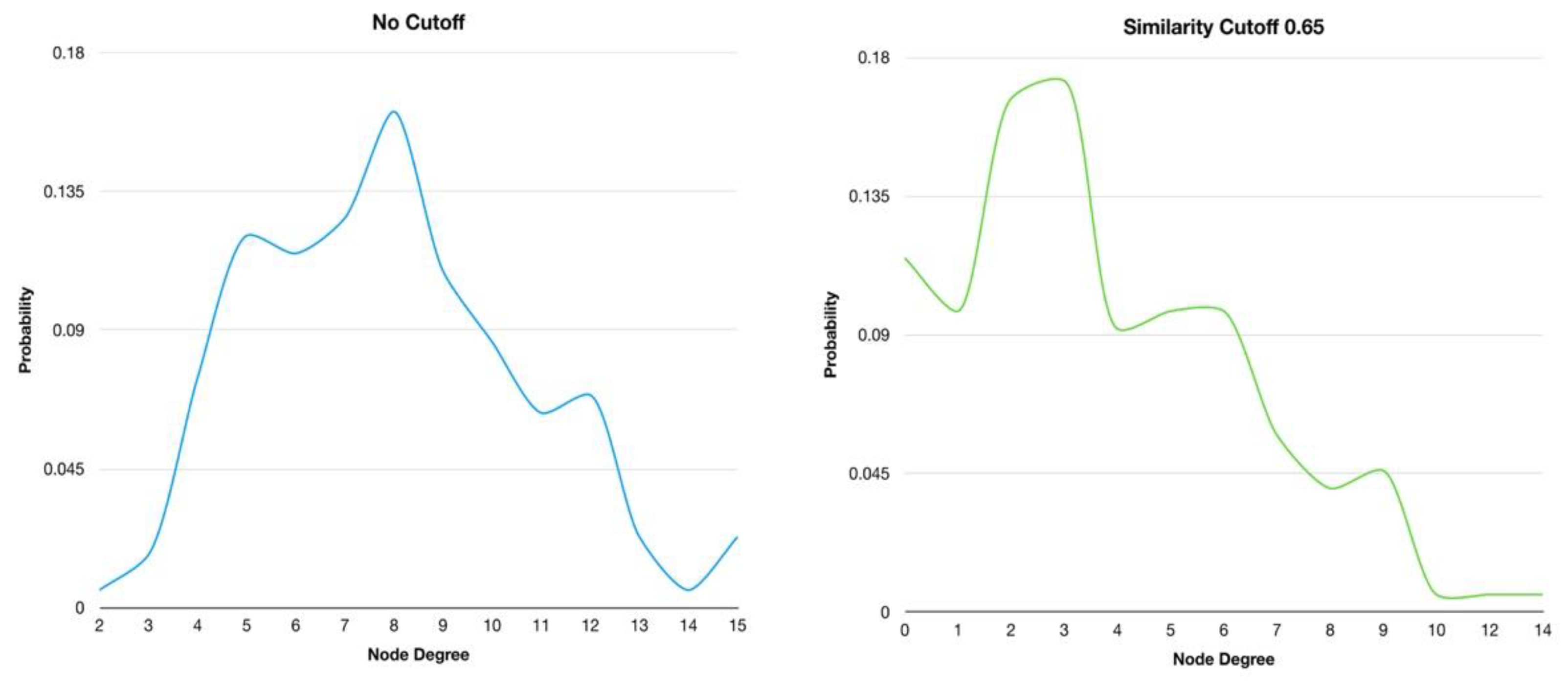
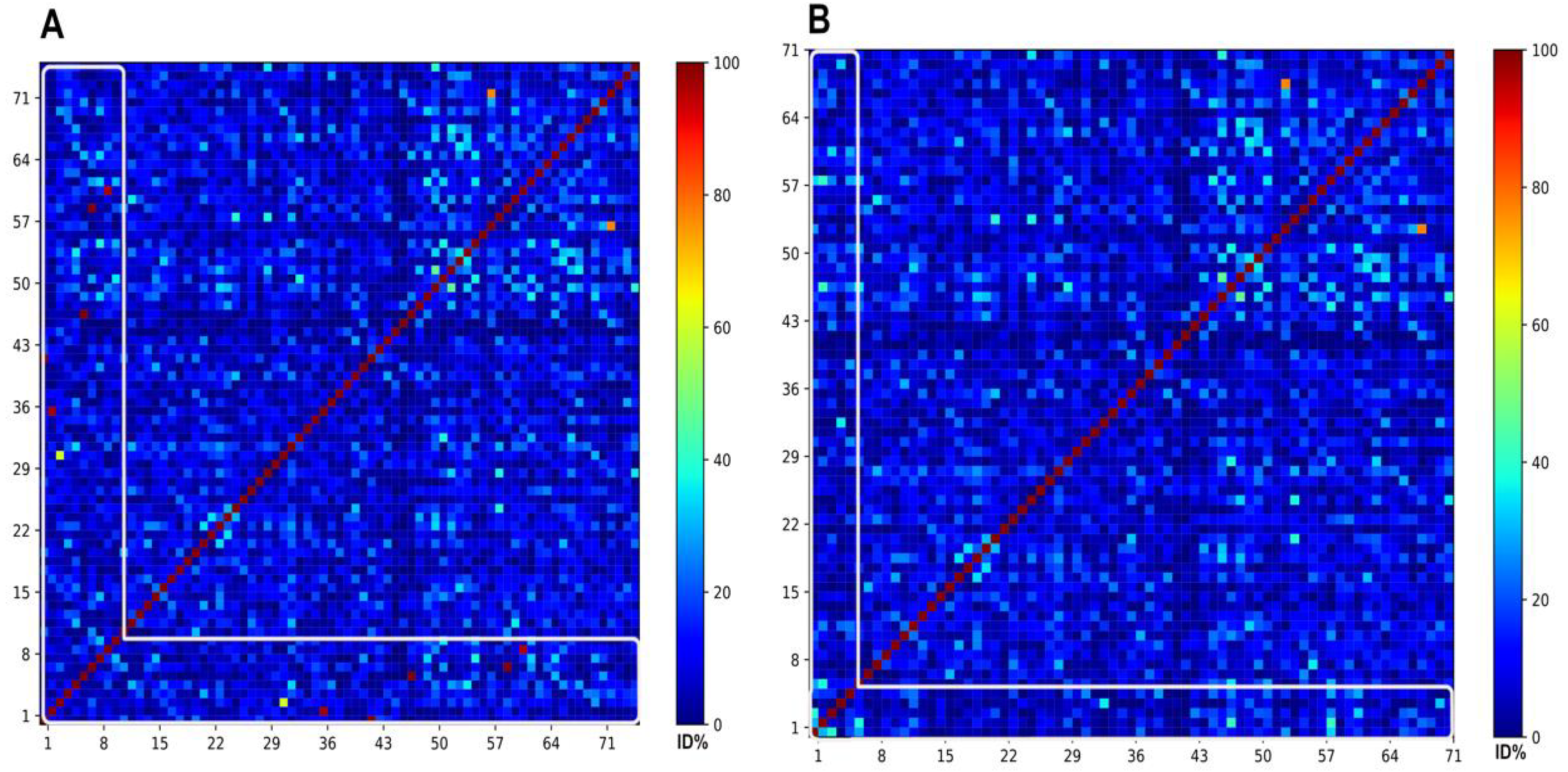
2.3. Networks similarity cutoff analysis
2.4. Network Visualization
2.4. Scaffold Extraction by Centrality Measures
2.5. Selection of the Most Representative Extracted Subset
2.6. Motif Discovery
2.6.1. Multiple Sequence Alignments
- -
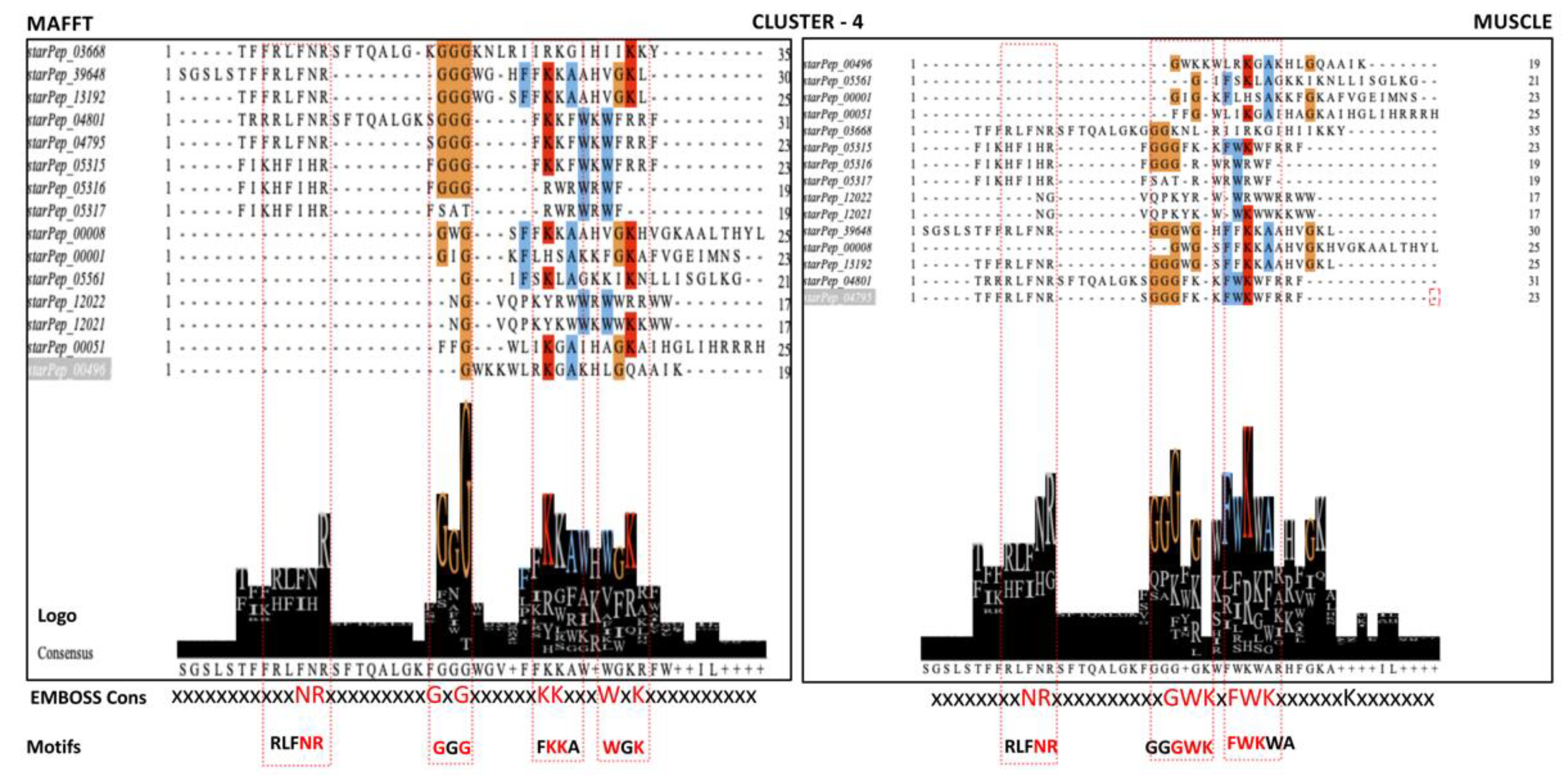
- Communities with more than 2 ABFPs including the one containing the 20 singletons were aligned independently using multiple sequence alignment (MSA) algorithms. The algorithms of choice were MAFFT (Multiple Alignment using Fast Fourier Transform) v7.487 with the iterative refinement FFT-NS-i option [39] and MUSCLE (Multiple Sequence Comparison by Log- Expectation) v3.8 [40], publicly available at https://www.ebi.ac.uk/Tools/msa/. The conserved motifs were detected by jointly analyzing the consensus sequences and Seq2Logo, implemented in the Jalview v2.11.2.5 program [41] and EMBOSS Cons v6.6.0 available at https://www.ebi.ac.uk/Tools/msa/emboss_cons.
- -
2.6.2. Alignment-Free (AF) Detection
2.6.3. Motif Enrichment Analysis
3. Results and Discussion
3.1. Half-Space Proximal Network Model
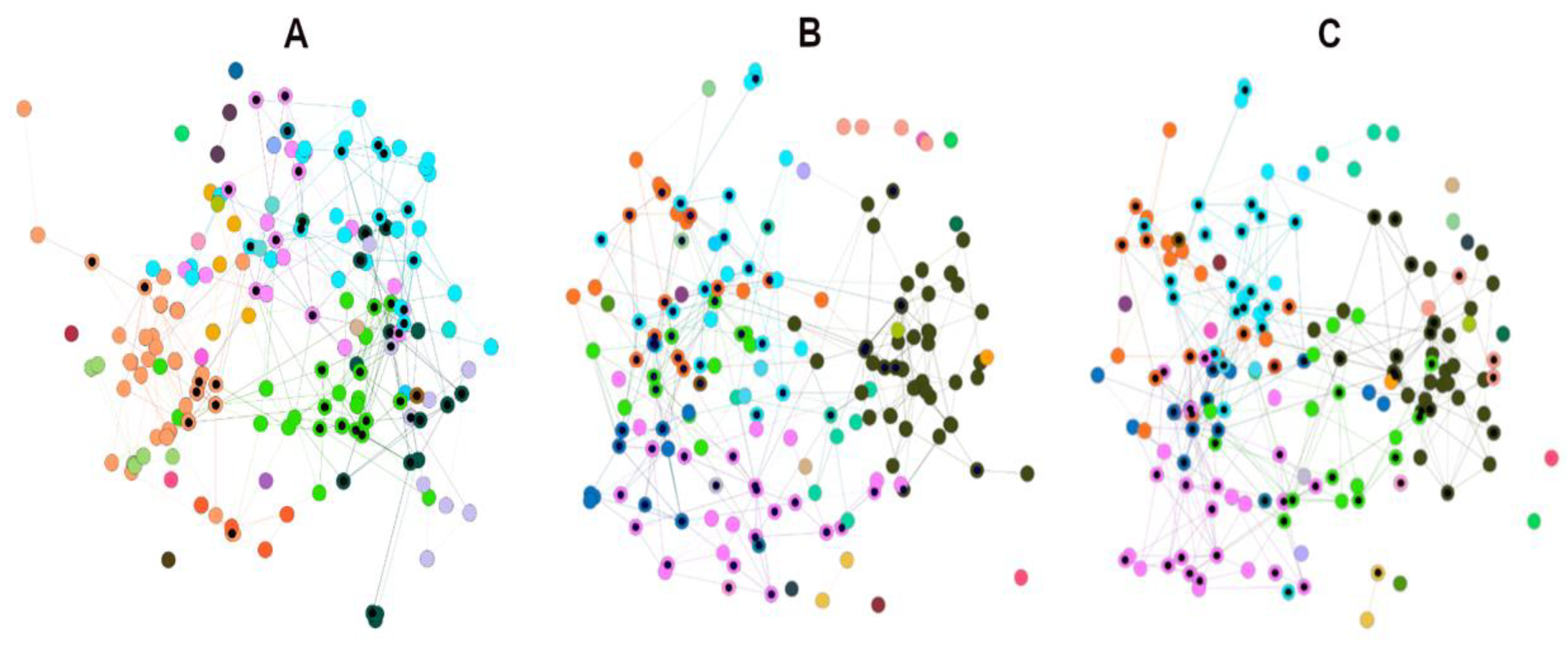
3.2. Network Visual Mining
3.2.1. Visual Mining of HSPNs, the Most Central and Atypical ABFPs
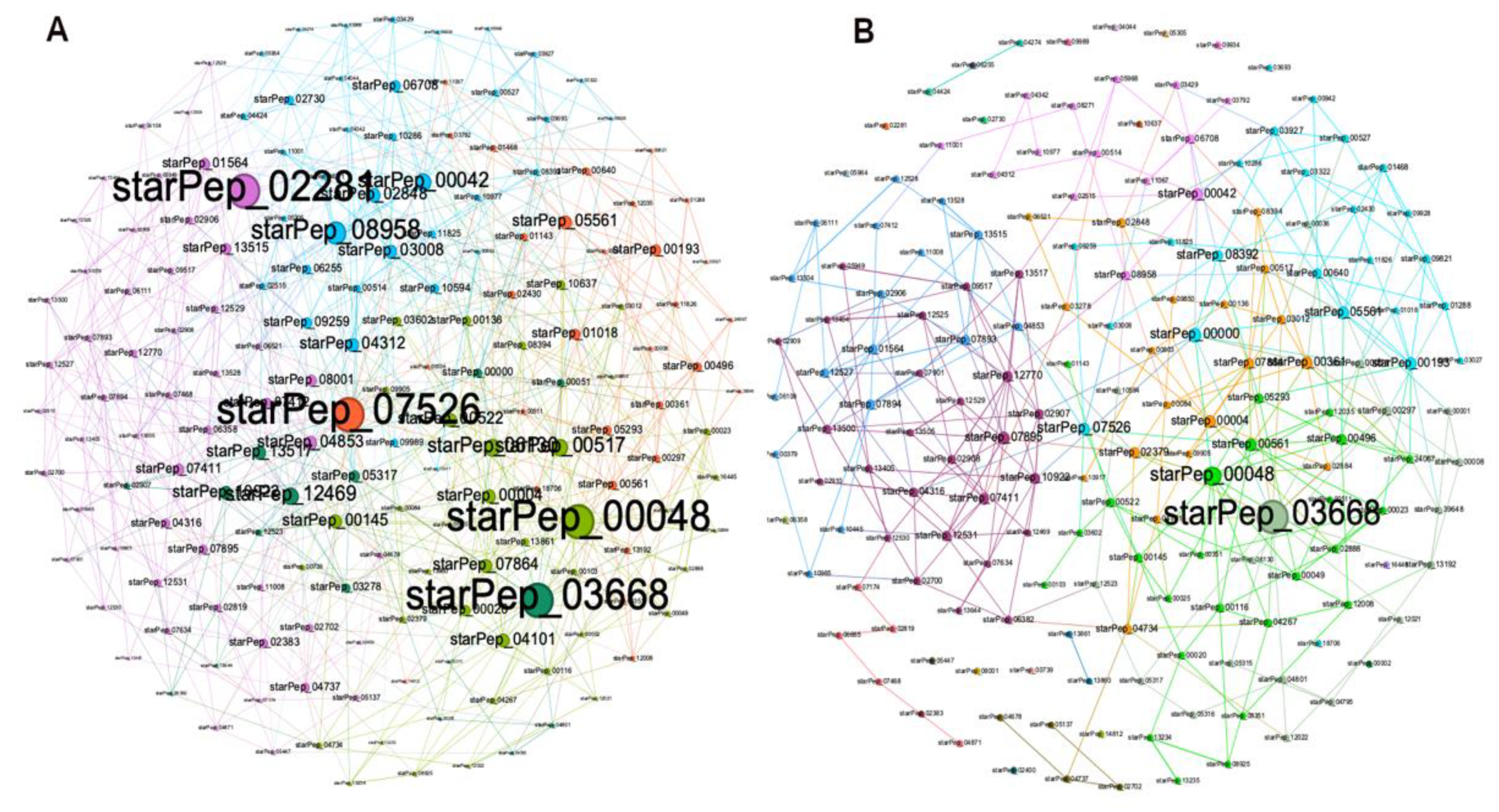

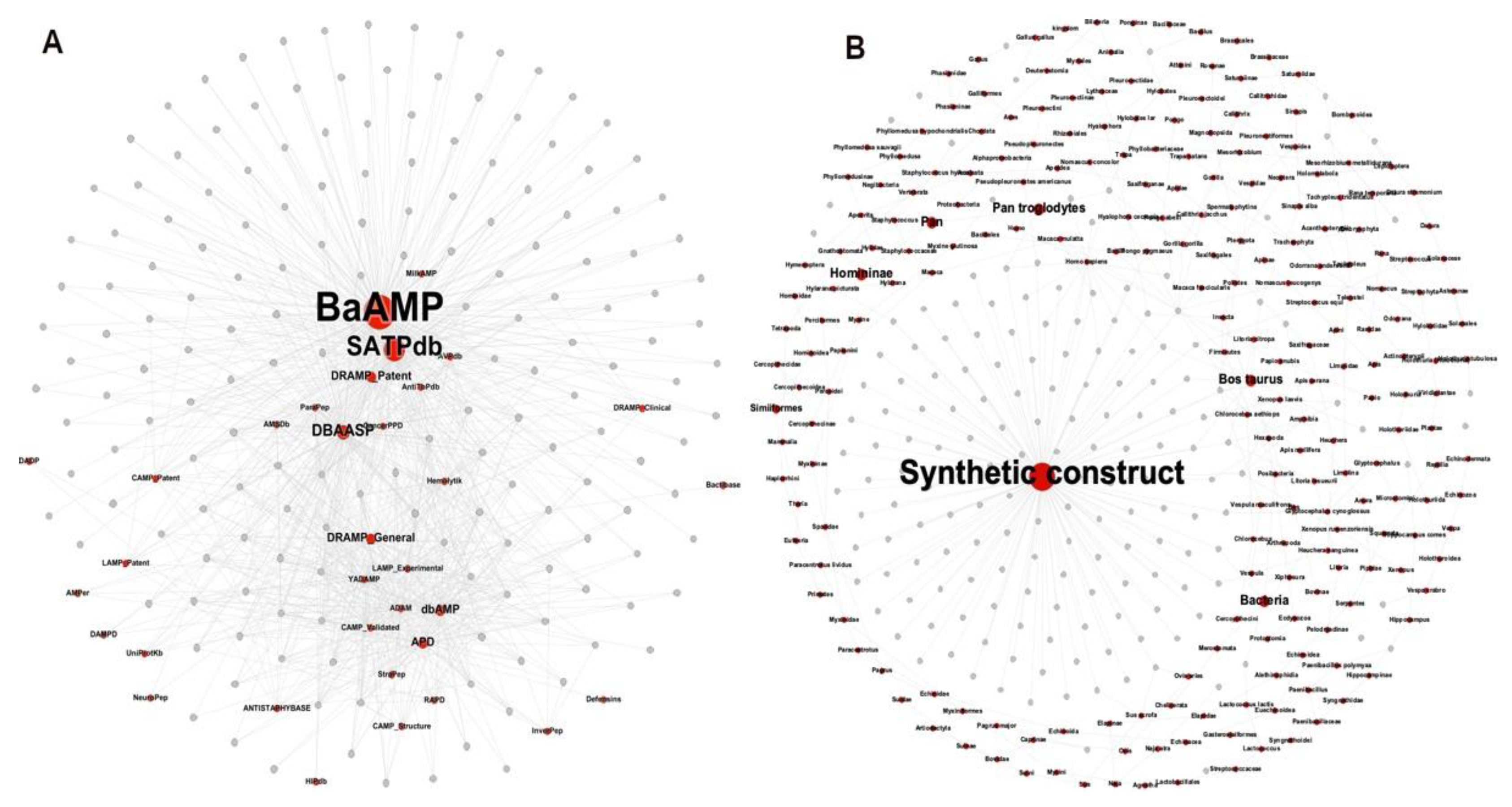
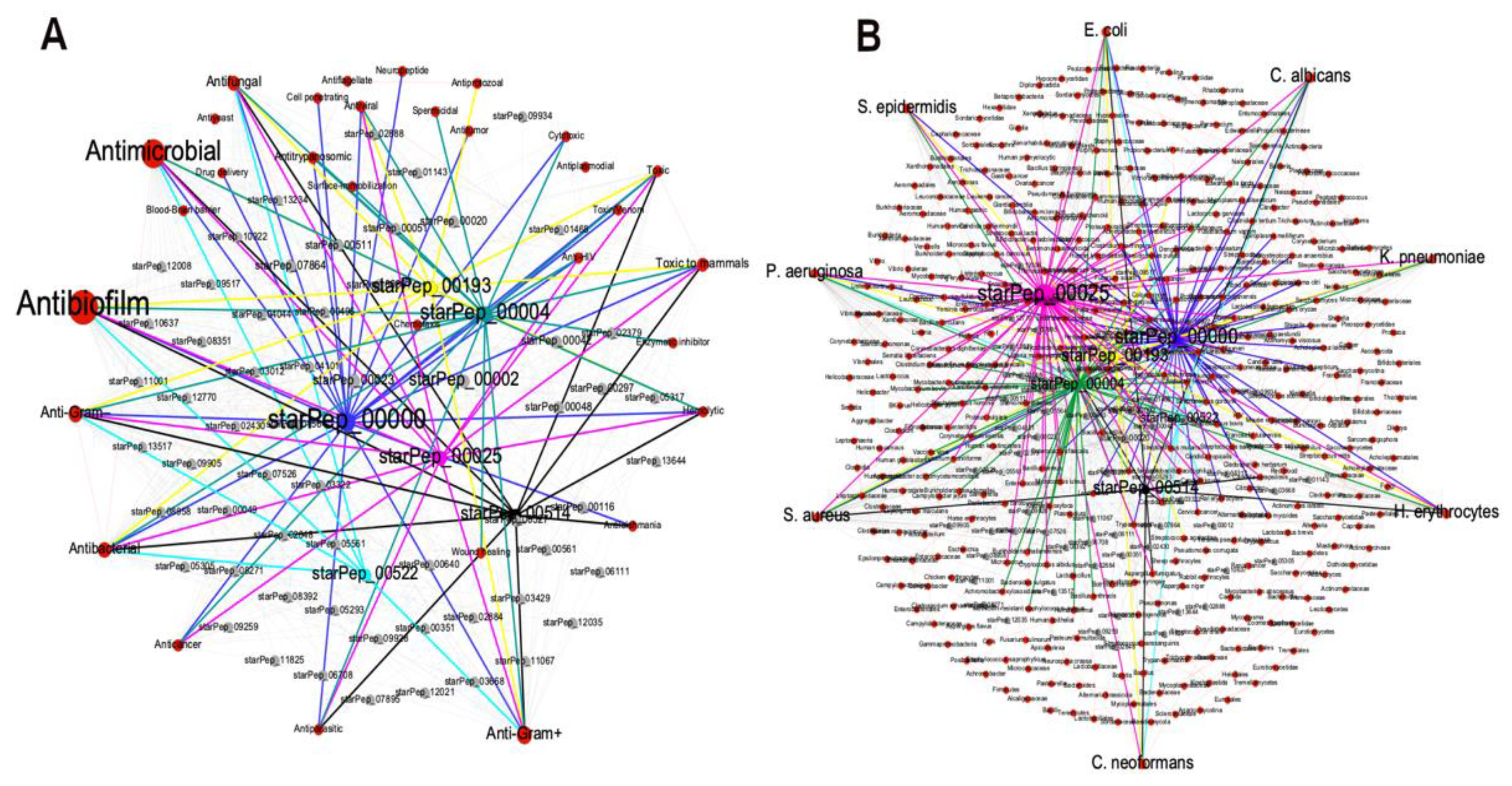
3.2.2. Metadata Analysis by Visual Mining
3.3. Representing the ABFPs with a reduced subset
3.3.1. The selection of the best representative subset
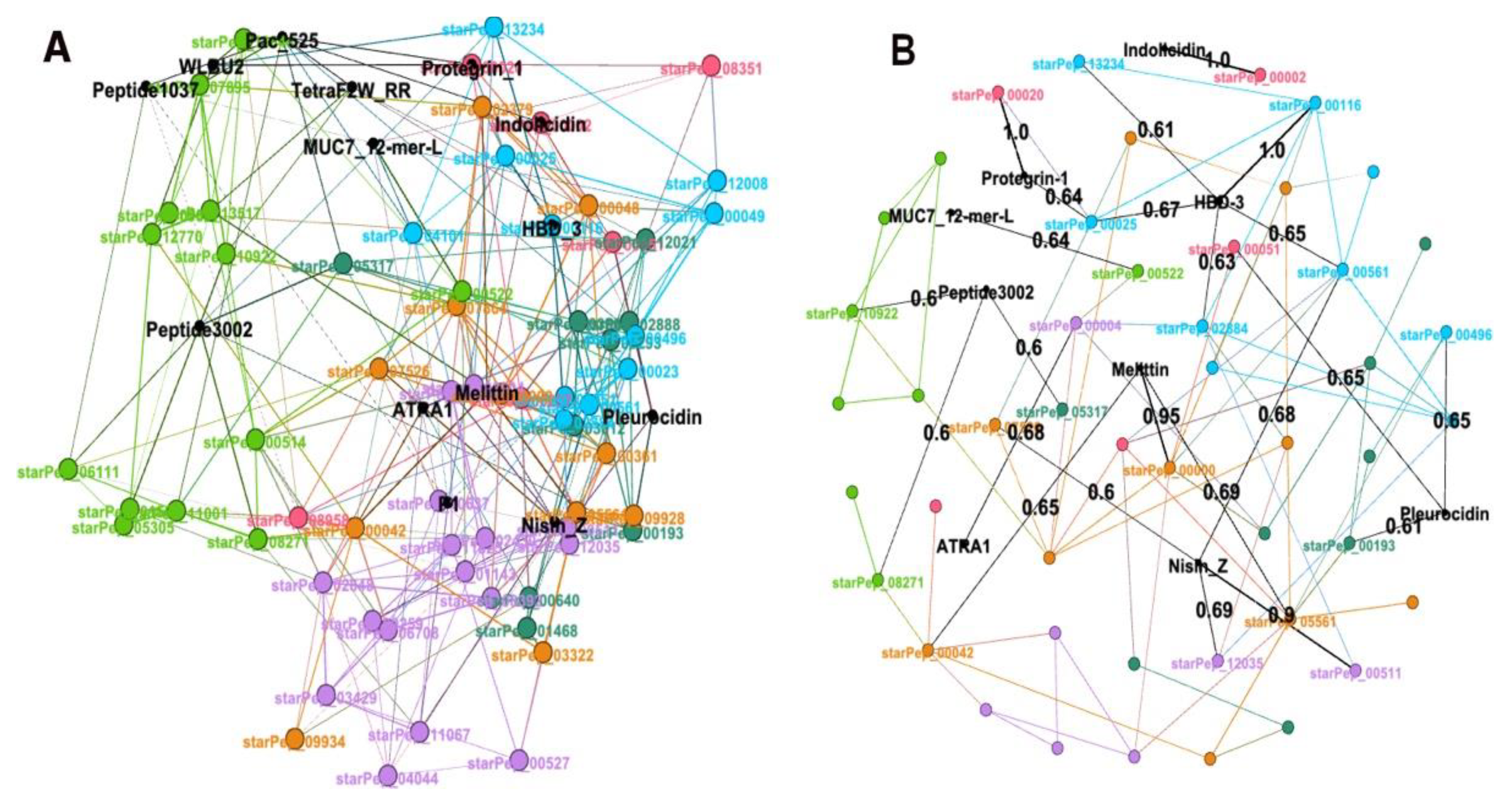
3.3.2. Visualizing/Analysing the best representative subset with HSPNs
3.3.3. Visualizing Mining of the METNs
3.4. External Representative ABFPs on the representative Antibiofilm HSPN
3.5. Motif Discovery Assisted by Complex Networks
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Vestby, L.K.; Gronseth, T.; Simm, R.; Nesse, L.L. Bacterial Biofilm and its Role in the Pathogenesis of Disease. Antibiotics (Basel) 2020, 9. [Google Scholar] [CrossRef]
- Gloag, E.S.; Fabbri, S.; Wozniak, D.J.; Stoodley, P. Biofilm mechanics: Implications in infection and survival. Biofilm 2020, 2, 100017. [Google Scholar] [CrossRef] [PubMed]
- de la Fuente-Nunez, C.; Reffuveille, F.; Fernandez, L.; Hancock, R.E. Bacterial biofilm development as a multicellular adaptation: antibiotic resistance and new therapeutic strategies. Curr Opin Microbiol 2013, 16, 580–589. [Google Scholar] [CrossRef] [PubMed]
- Sauer, K.; Stoodley, P.; Goeres, D.M.; Hall-Stoodley, L.; Burmolle, M.; Stewart, P.S.; Bjarnsholt, T. The biofilm life cycle: expanding the conceptual model of biofilm formation. Nat Rev Microbiol 2022, 20, 608–620. [Google Scholar] [CrossRef] [PubMed]
- An, A.Y.; Choi, K.G.; Baghela, A.S.; Hancock, R.E.W. An Overview of Biological and Computational Methods for Designing Mechanism-Informed Anti-biofilm Agents. Front Microbiol 2021, 12, 640787. [Google Scholar] [CrossRef]
- Jamal, M.; Ahmad, W.; Andleeb, S.; Jalil, F.; Imran, M.; Nawaz, M.A.; Hussain, T.; Ali, M.; Rafiq, M.; Kamil, M.A. Bacterial biofilm and associated infections. J Chin Med Assoc 2018, 81, 7–11. [Google Scholar] [CrossRef] [PubMed]
- Bryers, J.D. Medical biofilms. Biotechnol Bioeng 2008, 100, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Veerachamy, S.; Yarlagadda, T.; Manivasagam, G.; Yarlagadda, P.K. Bacterial adherence and biofilm formation on medical implants: a review. Proc Inst Mech Eng H 2014, 228, 1083–1099. [Google Scholar] [CrossRef] [PubMed]
- Fleming, D.; Rumbaugh, K. The Consequences of Biofilm Dispersal on the Host. Sci Rep 2018, 8, 10738. [Google Scholar] [CrossRef]
- Rumbaugh, K.P.; Sauer, K. Biofilm dispersion. Nat Rev Microbiol 2020, 18, 571–586. [Google Scholar] [CrossRef]
- Breidenstein, E.B.; de la Fuente-Nunez, C.; Hancock, R.E. Pseudomonas aeruginosa: all roads lead to resistance. Trends Microbiol 2011, 19, 419–426. [Google Scholar] [CrossRef] [PubMed]
- Romling, U.; Balsalobre, C. Biofilm infections, their resilience to therapy and innovative treatment strategies. J Intern Med 2012, 272, 541–561. [Google Scholar] [CrossRef]
- Barraud, N.; Hassett, D.J.; Hwang, S.H.; Rice, S.A.; Kjelleberg, S.; Webb, J.S. Involvement of nitric oxide in biofilm dispersal of Pseudomonas aeruginosa. J Bacteriol 2006, 188, 7344–7353. [Google Scholar] [CrossRef]
- Xiong, Y.Q.; Estelles, A.; Li, L.; Abdelhady, W.; Gonzales, R.; Bayer, A.S.; Tenorio, E.; Leighton, A.; Ryser, S.; Kauvar, L.M. A Human Biofilm-Disrupting Monoclonal Antibody Potentiates Antibiotic Efficacy in Rodent Models of both Staphylococcus aureus and Acinetobacter baumannii Infections. Antimicrob Agents Chemother 2017, 61. [Google Scholar] [CrossRef] [PubMed]
- Verderosa, A.D.; Totsika, M.; Fairfull-Smith, K.E. Bacterial Biofilm Eradication Agents: A Current Review. Front Chem 2019, 7, 824. [Google Scholar] [CrossRef] [PubMed]
- Overhage, J.; Campisano, A.; Bains, M.; Torfs, E.C.; Rehm, B.H.; Hancock, R.E. Human host defense peptide LL-37 prevents bacterial biofilm formation. Infect Immun 2008, 76, 4176–4182. [Google Scholar] [CrossRef] [PubMed]
- de la Fuente-Nunez, C.; Reffuveille, F.; Haney, E.F.; Straus, S.K.; Hancock, R.E. Broad-spectrum anti-biofilm peptide that targets a cellular stress response. PLoS Pathog 2014, 10, e1004152. [Google Scholar] [CrossRef] [PubMed]
- Chavez de Paz, L.E.; Lemos, J.A.; Wickstrom, C.; Sedgley, C.M. Role of (p)ppGpp in biofilm formation by Enterococcus faecalis. Appl Environ Microbiol 2012, 78, 1627–1630. [Google Scholar] [CrossRef] [PubMed]
- Di Somma, A.; Moretta, A.; Cane, C.; Cirillo, A.; Duilio, A. Antimicrobial and Antibiofilm Peptides. Biomolecules 2020, 10. [Google Scholar] [CrossRef]
- Ma, L.; Ye, X.; Sun, P.; Xu, P.; Wang, L.; Liu, Z.; Huang, X.; Bai, Z.; Zhou, C. Antimicrobial and antibiofilm activity of the EeCentrocin 1 derived peptide EC1-17KV via membrane disruption. EBioMedicine 2020, 55, 102775. [Google Scholar] [CrossRef]
- UniProt, C. UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res 2019, 47, D506–D515. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Li, X.; Wang, Z. APD3: the antimicrobial peptide database as a tool for research and education. Nucleic Acids Res 2016, 44, D1087–1093. [Google Scholar] [CrossRef] [PubMed]
- Shi, G.; Kang, X.; Dong, F.; Liu, Y.; Zhu, N.; Hu, Y.; Xu, H.; Lao, X.; Zheng, H. DRAMP 3.0: an enhanced comprehensive data repository of antimicrobial peptides. Nucleic Acids Res 2022, 50, D488–D496. [Google Scholar] [CrossRef]
- Di Luca, M.; Maccari, G.; Maisetta, G.; Batoni, G. BaAMPs: the database of biofilm-active antimicrobial peptides. Biofouling 2015, 31, 193–199. [Google Scholar] [CrossRef]
- Sharma, A.; Gupta, P.; Kumar, R.; Bhardwaj, A. dPABBs: A Novel in silico Approach for Predicting and Designing Anti-biofilm Peptides. Sci Rep 2016, 6, 21839. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Sharma, A.K.; Jaiswal, S.K.; Sharma, V.K. Prediction of Biofilm Inhibiting Peptides: An In silico Approach. Front Microbiol 2016, 7, 949. [Google Scholar] [CrossRef] [PubMed]
- Fallah Atanaki, F.; Behrouzi, S.; Ariaeenejad, S.; Boroomand, A.; Kavousi, K. BIPEP: Sequence-based Prediction of Biofilm Inhibitory Peptides Using a Combination of NMR and Physicochemical Descriptors. ACS Omega 2020, 5, 7290–7297. [Google Scholar] [CrossRef]
- Bose, B.; Downey, T.; Ramasubramanian, A.K.; Anastasiu, D.C. Identification of Distinct Characteristics of Antibiofilm Peptides and Prospection of Diverse Sources for Efficacious Sequences. Front Microbiol 2021, 12, 783284. [Google Scholar] [CrossRef]
- Aguero-Chapin, G.; Galpert-Canizares, D.; Dominguez-Perez, D.; Marrero-Ponce, Y.; Perez-Machado, G.; Teijeira, M.; Antunes, A. Emerging Computational Approaches for Antimicrobial Peptide Discovery. Antibiotics (Basel) 2022, 11. [Google Scholar] [CrossRef]
- Romero, M.; Marrero-Ponce, Y.; Rodriguez, H.; Aguero-Chapin, G.; Antunes, A.; Aguilera-Mendoza, L.; Martinez-Rios, F. A Novel Network Science and Similarity-Searching-Based Approach for Discovering Potential Tumor-Homing Peptides from Antimicrobials. Antibiotics (Basel) 2022, 11. [Google Scholar] [CrossRef]
- Ayala-Ruano, S.; Marrero-Ponce, Y.; Aguilera-Mendoza, L.; Perez, N.; Aguero-Chapin, G.; Antunes, A.; Aguilar, A.C. Network Science and Group Fusion Similarity-Based Searching to Explore the Chemical Space of Antiparasitic Peptides. ACS Omega 2022, 7, 46012–46036. [Google Scholar] [CrossRef] [PubMed]
- Aguilera-Mendoza, L.; Marrero-Ponce, Y.; Beltran, J.A.; Tellez Ibarra, R.; Guillen-Ramirez, H.A.; Brizuela, C.A. Graph-based data integration from bioactive peptide databases of pharmaceutical interest: toward an organized collection enabling visual network analysis. Bioinformatics 2019, 35, 4739–4747. [Google Scholar] [CrossRef] [PubMed]
- Aguilera-Mendoza, L.; Marrero-Ponce, Y.; Garcia-Jacas, C.R.; Chavez, E.; Beltran, J.A.; Guillen-Ramirez, H.A.; Brizuela, C.A. Automatic construction of molecular similarity networks for visual graph mining in chemical space of bioactive peptides: an unsupervised learning approach. Sci Rep 2020, 10, 18074. [Google Scholar] [CrossRef] [PubMed]
- Chavez, E.; Dobrev, S.; Kranakis, E.; Opatrny, J.; Stacho, L.; Tejeda, H.; Urrutia, J. Half-space proximal: A new local test for extracting a bounded dilation spanner of a unit disk graph. In Proceedings of the Principles of Distributed Systems: 9th International Conference, OPODIS 2005, Revised Selected Papers 9, 2006. Pisa, Italy, December 12-14, 2005; pp. 235–245. [Google Scholar]
- Cherven, K. Network Graph Analysis and Visualization with Gephi; Packt Publishing: 2013.
- Smith, T.F.; Waterman, M.S. Identification of common molecular subsequences. J Mol Biol 1981, 147, 195–197. [Google Scholar] [CrossRef] [PubMed]
- Blondel, V.D.; Guillaume, J.-L.; Lambiotte, R.; Lefebvre, E. Fast unfolding of communities in large networks. Journal of statistical mechanics: theory and experiment 2008, 2008, P10008. [Google Scholar] [CrossRef]
- Fruchterman, T.M.; Reingold, E.M. Graph drawing by force-directed placement. Software: Practice and experience 1991, 21, 1129–1164. [Google Scholar] [CrossRef]
- Katoh, K.; Misawa, K.; Kuma, K.; Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 2002, 30, 3059–3066. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.; Clamp, M.; Barton, G.J. Jalview Version 2--a multiple sequence alignment editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef]
- Needleman, S.B.; Wunsch, C.D. A general method applicable to the search for similarities in the amino acid sequence of two proteins. J Mol Biol 1970, 48, 443–453. [Google Scholar] [CrossRef]
- Bailey, T.L. STREME: Accurate and versatile sequence motif discovery. Bioinformatics 2021, 37, 2834–2840. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.L.; Grant, C.E. SEA: Simple enrichment analysis of motifs. BioRxiv, 2008. [Google Scholar]
- Newman, M. Networks; Oxford university press: 2018.
- Martinez, L.R.; Casadevall, A. Cryptococcus neoformans cells in biofilms are less susceptible than planktonic cells to antimicrobial molecules produced by the innate immune system. Infection and immunity 2006, 74, 6118–6123. [Google Scholar] [CrossRef] [PubMed]
- Moazzezy, N.; Asadi Karam, M.R.; Rafati, S.; Bouzari, S.; Oloomi, M. Inhibition and eradication activity of truncated α-defensin analogs against multidrug resistant uropathogenic Escherichia coli biofilm. PLoS One 2020, 15, e0235892. [Google Scholar] [CrossRef] [PubMed]
- Eckert, R.; He, J.; Yarbrough, D.K.; Qi, F.; Anderson, M.H.; Shi, W. Targeted killing of Streptococcus mutans by a pheromone-guided "smart" antimicrobial peptide. Antimicrob Agents Chemother 2006, 50, 3651–3657. [Google Scholar] [CrossRef] [PubMed]
- de la Fuente-Nunez, C.; Reffuveille, F.; Mansour, S.C.; Reckseidler-Zenteno, S.L.; Hernandez, D.; Brackman, G.; Coenye, T.; Hancock, R.E. D-enantiomeric peptides that eradicate wild-type and multidrug-resistant biofilms and protect against lethal Pseudomonas aeruginosa infections. Chem Biol 2015, 22, 196–205. [Google Scholar] [CrossRef] [PubMed]
- Guha, S.; Ferrie, R.P.; Ghimire, J.; Ventura, C.R.; Wu, E.; Sun, L.; Kim, S.Y.; Wiedman, G.R.; Hristova, K.; Wimley, W.C. Applications and evolution of melittin, the quintessential membrane active peptide. Biochem Pharmacol 2021, 193, 114769. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Hirt, H.; Li, Y.; Gorr, S.U.; Aparicio, C. Antimicrobial GL13K peptide coatings killed and ruptured the wall of Streptococcus gordonii and prevented formation and growth of biofilms. PLoS One 2014, 9, e111579. [Google Scholar] [CrossRef] [PubMed]
- Holmberg, K.V.; Abdolhosseini, M.; Li, Y.; Chen, X.; Gorr, S.U.; Aparicio, C. Bio-inspired stable antimicrobial peptide coatings for dental applications. Acta Biomater 2013, 9, 8224–8231. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Chaudhary, K.; Dhanda, S.K.; Bhalla, S.; Usmani, S.S.; Gautam, A.; Tuknait, A.; Agrawal, P.; Mathur, D.; Raghava, G.P. SATPdb: a database of structurally annotated therapeutic peptides. Nucleic Acids Res 2016, 44, D1119–1126. [Google Scholar] [CrossRef]
- Pirtskhalava, M.; Gabrielian, A.; Cruz, P.; Griggs, H.L.; Squires, R.B.; Hurt, D.E.; Grigolava, M.; Chubinidze, M.; Gogoladze, G.; Vishnepolsky, B.; et al. DBAASP v.2: an enhanced database of structure and antimicrobial/cytotoxic activity of natural and synthetic peptides. Nucleic Acids Res 2016, 44, 6503. [Google Scholar] [CrossRef]
- Qureshi, A.; Thakur, N.; Tandon, H.; Kumar, M. AVPdb: a database of experimentally validated antiviral peptides targeting medically important viruses. Nucleic Acids Res 2014, 42, D1147–1153. [Google Scholar] [CrossRef]
- Mehta, D.; Anand, P.; Kumar, V.; Joshi, A.; Mathur, D.; Singh, S.; Tuknait, A.; Chaudhary, K.; Gautam, S.K.; Gautam, A.; et al. ParaPep: a web resource for experimentally validated antiparasitic peptide sequences and their structures. Database (Oxford) 2014, 2014. [Google Scholar] [CrossRef]
- Gautam, A.; Chaudhary, K.; Singh, S.; Joshi, A.; Anand, P.; Tuknait, A.; Mathur, D.; Varshney, G.C.; Raghava, G.P. Hemolytik: a database of experimentally determined hemolytic and non-hemolytic peptides. Nucleic Acids Res 2014, 42, D444–449. [Google Scholar] [CrossRef] [PubMed]
- Tyagi, A.; Tuknait, A.; Anand, P.; Gupta, S.; Sharma, M.; Mathur, D.; Joshi, A.; Singh, S.; Gautam, A.; Raghava, G.P. CancerPPD: a database of anticancer peptides and proteins. Nucleic Acids Res 2015, 43, D837–843. [Google Scholar] [CrossRef]
- Seshadri Sundararajan, V.; Gabere, M.N.; Pretorius, A.; Adam, S.; Christoffels, A.; Lehvaslaiho, M.; Archer, J.A.; Bajic, V.B. DAMPD: a manually curated antimicrobial peptide database. Nucleic Acids Res 2012, 40, D1108–1112. [Google Scholar] [CrossRef] [PubMed]
- Waghu, F.H.; Barai, R.S.; Gurung, P.; Idicula-Thomas, S. CAMPR3: a database on sequences, structures and signatures of antimicrobial peptides. Nucleic Acids Res 2016, 44, D1094–1097. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Wu, H.; Lu, H.; Li, G.; Huang, Q. LAMP: A Database Linking Antimicrobial Peptides. PLoS One 2013, 8, e66557. [Google Scholar] [CrossRef] [PubMed]
- Fan, L.; Sun, J.; Zhou, M.; Zhou, J.; Lao, X.; Zheng, H.; Xu, H. DRAMP: a comprehensive data repository of antimicrobial peptides. Sci Rep 2016, 6, 24482. [Google Scholar] [CrossRef] [PubMed]
- Reza, A.; Sutton, J.M.; Rahman, K.M. Effectiveness of Efflux Pump Inhibitors as Biofilm Disruptors and Resistance Breakers in Gram-Negative (ESKAPEE) Bacteria. Antibiotics (Basel) 2019, 8. [Google Scholar] [CrossRef]
- Li, J.; Chen, D.; Lin, H. Antibiofilm peptides as a promising strategy: comparative research. Appl Microbiol Biotechnol 2021, 105, 1647–1656. [Google Scholar] [CrossRef]
- Juretic, D. Designed Multifunctional Peptides for Intracellular Targets. Antibiotics (Basel) 2022, 11. [Google Scholar] [CrossRef] [PubMed]
- Agüero-Chapin, G.; Galpert, D.; Molina-Ruiz, R.; Ancede-Gallardo, E.; Pérez-Machado, G.; De la Riva, G.A.; Antunes, A. Graph Theory-Based Sequence Descriptors as Remote Homology Predictors. Biomolecules 2020, 10, 26. [Google Scholar] [CrossRef] [PubMed]
- Vens, C.; Rosso, M.N.; Danchin, E.G. Identifying discriminative classification-based motifs in biological sequences. Bioinformatics 2011, 27, 1231–1238. [Google Scholar] [CrossRef]
- de Souza, C.M.; da Silva, Á.P.; Júnior, N.G.O.; Martínez, O.F.; Franco, O.L. Peptides as a therapeutic strategy against Klebsiella pneumoniae. Trends in Pharmacological Sciences 2022, 43, 335–348. [Google Scholar] [CrossRef] [PubMed]
- Hee, C.S.; Habazettl, J.; Schmutz, C.; Schirmer, T.; Jenal, U.; Grzesiek, S. Intercepting second-messenger signaling by rationally designed peptides sequestering c-di-GMP. Proc Natl Acad Sci U S A 2020, 117, 17211–17220. [Google Scholar] [CrossRef] [PubMed]
- Edwards-Gayle, C.J.C.; Barrett, G.; Roy, S.; Castelletto, V.; Seitsonen, J.; Ruokolainen, J.; Hamley, I.W. Selective Antibacterial Activity and Lipid Membrane Interactions of Arginine-Rich Amphiphilic Peptides. ACS Appl Bio Mater 2020, 3, 1165–1175. [Google Scholar] [CrossRef] [PubMed]
- Zarena, D.; Mishra, B.; Lushnikova, T.; Wang, F.; Wang, G. The pi Configuration of the WWW Motif of a Short Trp-Rich Peptide Is Critical for Targeting Bacterial Membranes, Disrupting Preformed Biofilms, and Killing Methicillin-Resistant Staphylococcus aureus. Biochemistry 2017, 56, 4039–4043. [Google Scholar] [CrossRef]
- Mishra, B.; Lushnikova, T.; Golla, R.M.; Wang, X.; Wang, G. Design and surface immobilization of short anti-biofilm peptides. Acta Biomater 2017, 49, 316–328. [Google Scholar] [CrossRef]

| HSPN – No Cutoff | |||
| Centrality Measure | Total | Peptide Name | Cluster |
| Node Degree Harmonic Betweenness Hub-Bridge | 1 | starPep_03668 | (1) |
| Node Degree Betweenness Hub-Bridge | 1 | starPep_00048 | (1) |
| Harmonic Hub-Bridge | 2 | starPep_00000 starPep_10922 | (1) (3) |
| Node Degree Betweenness | 7 | starPep_12469 starPep_07526 starPep_00145 starPep_06130 starPep_00042 starPep_08958 starPep_02281 | (3) (2)(2) (3) (0) (2) (2) |
| Node Degree | 1 | starPep_00517 | (0) |
| Harmonic | 7 | starPep_07864 starPep_07895 starPep_02907 starPep_12770 starPep_12531 starPep_13517 starPep_07893 | (0) (3)(3) (3)(3) (3) (2) |
| Betweenness | 1 | starPep_08001 | (3) |
| Hub-Bridge | 6 | starPep_00496 starPep_00561 starPep_00361 starPep_13515 starPep_00193 starPep_05561 | (1) (1) (1) (2) (1) (1) |
| HSPN – Cutoff 0.65 | |||
| Node Degree Harmonic Betweenness Hub-Bridge | 1 | starPep_00048 | (9) |
| Node Degree Betweenness Hub-Bridge | 1 | starPep_00042 | (11) |
| Harmonic Betweenness Hub-Bridge | 1 | starPep_10922 | (15) |
| Node Degree Harmonic Betweenness | 2 | starPep_00000 starPep_03668 | (15) (4) |
| Node Degree Harmonic | 1 | starPep_00361 | (9) |
| Node Degree Betweenness | 1 | starPep_07526 | (17) |
| Node Degree Hub-Bridge | 2 | starPep_00004 starPep_00193 | (11) (9) |
| Harmonic Betweenness | 1 | starPep_02379 | (17) |
| Harmonic Hub-Bridge | 3 | starPep_07895 starPep_02907 starPep_12531 | (15) (15) (15) |
| Node Degree | 2 | starPep_00561 starPep_05561 | (9) (4) |
| Harmonic | 1 | starPep_07893 | (14) |
| Betweenness | 3 | starPep_12530 starPep_04734 starPep_08958 | (15) (17) (15) |
| Hub-Bridge | 2 | starPep_12770 starPep_02908 | (15) (15) |
| HSPN – Cutoff 0.65 | |||
| Atypical peptides | Total | Peptide Name | Cluster |
| Singletons | 20 | starPep_00002 starPep_00739 starPep_02281 starPep_02383 starPep_02400 starPep_02730 starPep_03693 starPep_04044* starPep_05305* starPep_05447* starPep_05964 starPep_06255 starPep_06358* starPep_08001 starPep_09934* starPep_09989* starPep_10637* starPep_14812* starPep_16445* starPep_18706* | (10) (18) (20) (25) (0) (5) (19) (13) (8) (27) (26) (21) (23) (24) (16) (29) (1) (12) (2) (3) |
| Isolated Community | 2 | starPep_04274 – starPep_04424 ϦstarPep_13860* – starPep_13861* | (6)Ϧ(28) |
| Harmonic (HC) | Hub-Bridge (HB) | ||||||
| Subsets | Cutoff | Edges | Nodes | Coverage1% | Edges | Nodes | Coverage1% |
| 1 | 0.90 | 227 | 154 | 89 | 276 | 154 | 89 |
| 2 | 0.80 | 230 | 138 | 79 | 235 | 137 | 79 |
| 3 | 0.70 | 199 | 122 | 70 | 201 | 125 | 72 |
| 4 | 0.60 | 167 | 103 | 59 | 162 | 104 | 60 |
| 5 | 0.50 | 128 | 80 | 46 | 112 | 80 | 46 |
| 6 | 0.45 | 115 | 74 | 43 | 88 | 68 | 39 |
| 7 | 0.40 | 74 | 54 | 31 | 63 | 51 | 29 |
| 8 | 0.35 | 62 | 45 | 26 | 44 | 40 | 23 |
| 9 | 0.30 | 40 | 32 | 22 | 35 | 34 | 20 |
| Class-Peptides | Action Mode | Sequence |
| Class1- HBD-3 | Influence on icaAD and icaR genes transcription levels | GIINTLQKYYCRVRGGRCAVLSCLPKEEQIGKCSTRGRKCCRRKK |
| Class2- Nisin Z | Decrease adhesion, kill bacteria, reduce biofilm formation | ITSISLCTPGCKTGALM GCNMKTATCNCSIHVSK |
| Class3- MUC7 12-mer-L | Attracted to bacterial surfaces by the electrostatic bonding | RKSYKCLHKRCR |
| Class4- ATRA1 | Promote biofilm dispersal | KRFKKFFKKLKNSVK KRFKKFFKKLKVIGVT FPF |
| Class5- Pleurocidin | Induces disturbance/permeabilization of the membranes and bind to bacterial DNA, causing interference with cellular functions | GWGSFFKKAAHVGK HVGKAALTHYL |
| Class6- Pac-525 | Ability to enter membranes and to affect the lipopolysaccharides of Gram-negative bacteria | KWRRWVRWI |
| Class7- peptide 1037 | Decreases the attachment of bacterial cells, stimulates twitching motility, and influenced two major quorum sensing systems | KRFRIRVRV |
| Class8- Indolicidin | Induces lipid removal and that mixed indolicidin-lipid patches; membrane permeabilization | ILPWKWPWWPWRR |
| Class9- Protegrin-1* | Forms amyloid fibers to associate with the bacterial membrane and product transmembrane pores | RGGRLCYCRRRFCVCVGR |
| Class10- Peptide 3002 | Blocking (p)ppGpp | ILVRWIRWRIQW |
| Class11-TetraF2W-RR* | Disrupts membranes to kill bacteria rapidly | WWWLRRIW |
| Class12-P1 | Interferes with the proper secretion and/or intermolecular interaction of key extracellular polymers in the biofilm matrix | PARKARAATAATAATAATAATAAT |
| Class13-WLBU2* | LPS-binding property, interfers with bacterial attachment; destroying the bacterial membrane | RRWVRRVRRVWRRVVRVVRRWVRR |
| Class14-Melittin* | Inhibits the expression of biofilm-associated bap genes | GIGAVLKVLTTGLPALISWIKRKRQQ |
| No | Motif | EMBOSS Cons. | Cluster | Cluster size | MSA Method | Enrichment ratio* |
| 1 | RLFNR | xxxNR | 4 | 15 | MAFFT/MUSCLE | – |
| 2 | GGG | GxG | MAFFT | – | ||
| 3 | GGGWK | xxGWK | MUSCLE | ( –)/(2.25) | ||
| 4 | FKKA | xKKx | MAFFT | (–)/(4.0) | ||
| 5 | FWKWA | FWK | MAFFT | (3.0)/(2.83) | ||
| 6 | WGK | WxK | MAFFT | (–)/(1.41) | ||
| 7 | LLLLLKKK | LLLLLKKK | 6 | 2 | Pair-Aligned | – |
| 8 | LISWIK | lisxik | 7 | 8 | MAFFT | (–)/(2.71) |
| 9 | KNKRK | knkxk | MUSCLE | (3.0)/(2.22) | ||
| 10 | KRKQ | kxkQ | MAFFT | (3.0)/(–) | ||
| 11 | R[GP]RVS | rxRVS | MAFFT | (–)/(3.0) | ||
| 12 | RRPR | RRxR | MUSCLE | – | ||
| 13 | [GR]GG | xGG | MAFFT | (3.0)/(1.90) | ||
| 14 | GGRRRR | GGrrRR | MUSCLE | (–)/(2.0) | ||
| 15 | RRRRR | RRRRR | MAFFT/MUSCLE | – | ||
| 16 | ISGI | Ixxx | 9 | 23 | MAFFT | – |
| 17 | FKKLL | xKKLL | 11 | 27 | MAFFT/MUSCLE | (–)/(2.25) |
| 18 | KKLK | MAFFT | – | |||
| 19 | KKL | MUSCLE | – | |||
| 20 | LKK | LKK | MUSCLE | – | ||
| 21 | RIRVR | RIRVR | 14 | 23 | MAFFT | (–)/(1.58) |
| 22 | RVIR | xRVIR | MAFFT | (–)/(1.32) | ||
| 23 | VRVIR | MUSCLE | (–)/(2.83) | |||
| 24 | R[WL]R | RxR | MUSCLE | (1.57)/(–) | ||
| 25 | RIRRW | RIxRW | 15 | 26 | MAFFT/MUSCLE | (–)/(4.0) |
| 26 | RI[VR]W | (–)/(1.67) | ||||
| 27 | WVV | WVV | MAFFT | (–)/(1.44) | ||
| 28 | I[IR]R | IIxR | MUSCLE | – | ||
| 29 | WLRK | Wxxx | 17 | 23 | MAFFT | (–)/(2.50) |
| 30 | RWK | Rxx | MUSCLE | – | ||
| 31 | KKL | Kxx | MAFFT | – | ||
| 32 | KR[AKL]RK | KRxRK | MUSCLE | (6.0)/(3.0) | ||
| 33 | WR[IV]R | xRWR[IV]R | 22 | 5 | MAFFT/MUSCLE | – |
| 32 | FRWRI | MAFFT | (3.0)/ (–) | |||
| 33 | RWRVR | MUSCLE | (–)/(1.63) | |||
| 34 | YAPWYN | YAPWYN | 28 | 2 | Pair-Aligned | – |
| 35 | [FI][KW]RK | iKrK | Singletons | 20 | MAFFT/MUSCLE | (–)/(1.46) |
| No | Motif | Cluster | Cluster size | Matches in ABFPs | Matches in control | Sites (%) | Score | Enrichment ratio* |
| 1 | FKKA | 4 | 15 | 7 | 0 | 46.7 | 3.3e-003 | (–)/(3.33) |
| 2 | GGGR | 7 | 0 | 46.7 | 3.3e-003 | (–)/(2.11) | ||
| 3 | W[KR]WF | 7 | 0 | 46.7 | 3.3e-003 | (–)/(1.38) | ||
| 4 | FIH | 6 | 0 | 40.0 | 8.4e-002 | – | ||
| 5 | RLFNR | 5 | 0 | 33.3 | 2.1e-003 | – | ||
| 6 | KKK | 6 | 2 | 2 | 0 | 100 | 1.7e-001 | – |
| 7 | LLLLL | 2 | 0 | 100 | 1.7e-001 | – | ||
| 8 | RGG | 7 | 8 | 8 | 0 | 100 | 7.8e-005 | (3.0)/(1.56) |
| 9 | ISWIK | 4 | 0 | 50 | 3.8e-002 | (–)/(2.83) | ||
| 10 | NKRKQ | 4 | 0 | 50 | 3.8e-002 | – | ||
| 11 | RPRVS | 3 | 0 | 37.5 | 1.0e-001 | (–)/(3.71) | ||
| 12 | RRRRR | 3 | 0 | 37.5 | 1.0e-001 | – | ||
| 13 | SAC | 9 | 23 | 16 | 1 | 69.6 | 3.3e-006 | – |
| 14 | AKA | 5 | 0 | 21.7 | 2.85e-002 | – | ||
| 15 | CD[VI] | 5 | 0 | 21.7 | 2.85e-002 | – | ||
| 16 | IA[GVK] | 5 | 0 | 21.7 | 2.85e-002 | – | ||
| 17 | LFKKL | 11 | 27 | 9 | 0 | 33.3 | 8.8e-004 | (–)/(2.40) |
| 18 | KVLK | 8 | 0 | 29.6 | 2.1e-003 | (3.0)/(4.0) | ||
| 19 | KRFL | 6 | 0 | 22.2 | 1.1e-002 | (3.0)/(1.8) | ||
| 20 | VRLRI | 14 | 23 | 12 | 0 | 52.2 | 3.5e-005 | – |
| 21 | RVIR | 10 | 0 | 43.5 | 2.8e-004 | (–)/(1.32) | ||
| 22 | VWVI | 15 | 26 | 14 | 3 | 53.8 | 1.3e-003 | (3.0)/(3.0) |
| 23 | VIWRR | 8 | 0 | 30.8 | 2.1e-003 | (–)/(2.50) | ||
| 24 | LRK | 17 | 23 | 9 | 0 | 39.1 | 7.4e-004 | (3.0)/(1.27) |
| 25 | WRRK | 6 | 0 | 26.1 | 1.1e-002 | (–)/(1.67) | ||
| 26 | WRIR | 22 | 5 | 5 | 1 | 100 | 2.4e-002 | (–)/(3.25) |
| 27 | IRR | 2 | 3 | 40.0 | 9.0e-001 | (1.67)/(–) | ||
| 28 | APWTN | 28 | 2 | 2 | 0 | 100 | 1.7e-001 | (–)/(3.0) |
| 29 | KKRK | Singletons | 20 | 2 | 0 | 10.0 | 2.3e-001 | – |
| 30 | KKVVF | 2 | 0 | 10.0 | 2.4e-001 | – | ||
| 31 | LLKLL | 2 | 0 | 10.0 | 2.4e-001 | – | ||
| 32 | VKFK | 2 | 0 | 10.0 | 2.4e-001 | – | ||
| 33 | WRWR | 2 | 0 | 10.0 | 2.4e-001 | (–)/(1.64) |
| No | Motif | Cluster | Method | Enrichment ratio* |
| 1 | FWKWA | 4 | MAFFT | (3.0)/(2.83) |
| 2 | KNKRK | 7 | MUSCLE | (3.0)/(2.22) |
| 3 | [GR]GG | 7 | MAFFT/STREME | (3.0)/(1.90) |
| 4 | KVLK | 11 | STREME | (3.0)/(4.0) |
| 5 | KRFL | 11 | STREME | (3.0)/(1.8) |
| 6 | VWVI | 15 | STREME | (3.0)/(3.0) |
| 7 | KR[AKL]RK | 17 | MUSCLE | (6.0)/(3.0) |
| 8 | LRK | 17 | STREME | (3.0)/(1.27) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




