Submitted:
16 February 2023
Posted:
17 February 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
- Chromosomal diseases occur when the complete chromosome or large sections of a chromosome are lost, duplicated, or otherwise changed. For example, Down’s syndrome is an example of chromosomal irregularity.
- Single-gene disorders occur when gene alteration affects one gene, like sickle-cell anemia, where a defect in the Hemoglobin gene affects the red blood cells.
- Multifactorial disorders occur as a result of mutations in several genes, generally linked with conservational reasons. For example, diabetes is a multifactorial syndrome.
- Mitochondrial syndromes: are rare illnesses caused by mutations in non-chromosomal DNA found within the subcellular organelles, i.e., mitochondria. These disorders can be found to affect any part of the body like the brain and the muscles.
2. Background Study
3. Catalog of Human Variation Databases
3.1. Cenetral Databases
3.2. Locus-Specific Mutation Databases (LSDBs)
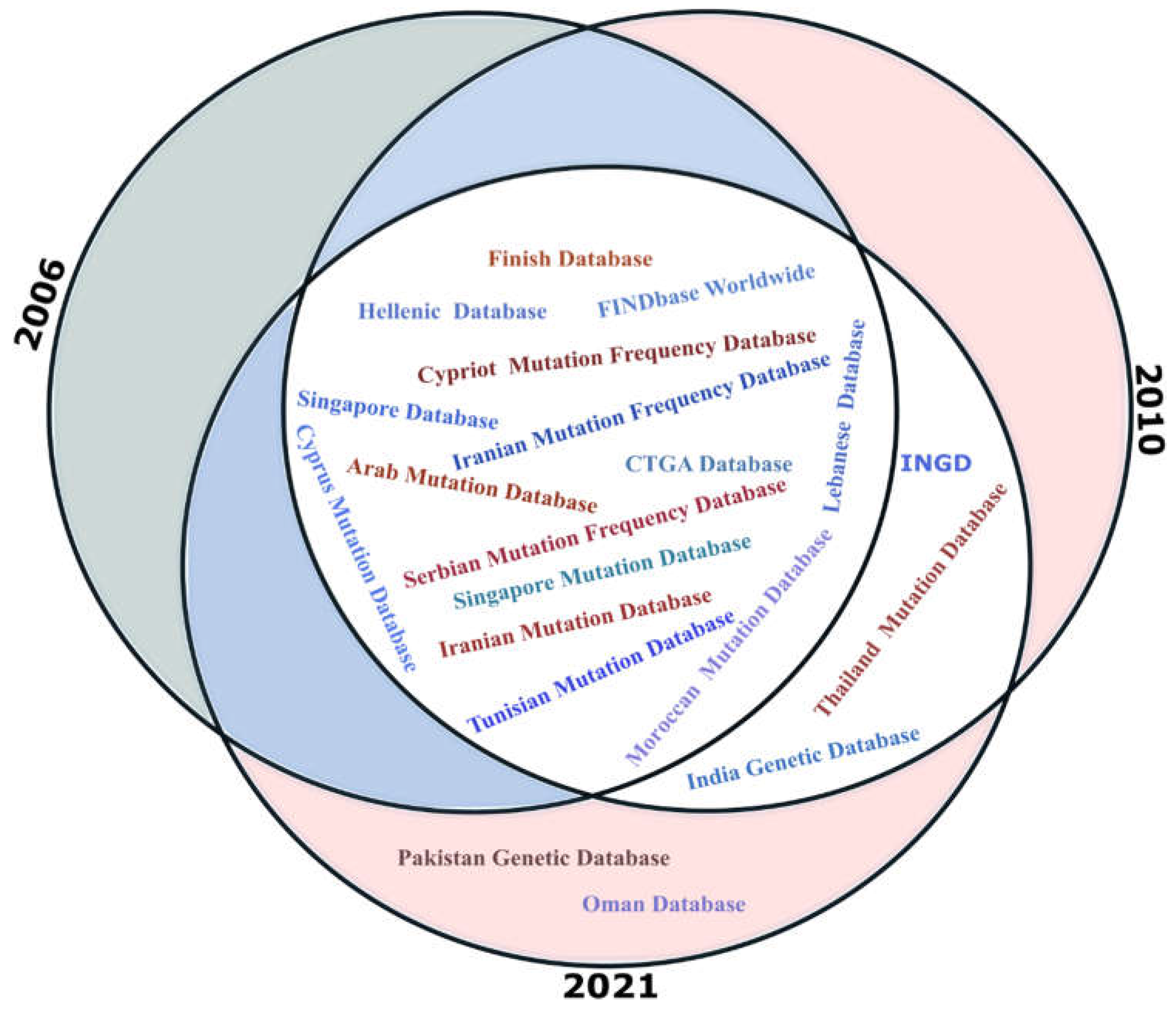
3.3. The National and Ethnic Mutation Databases (NEMDBs)
4. Materials and Methods
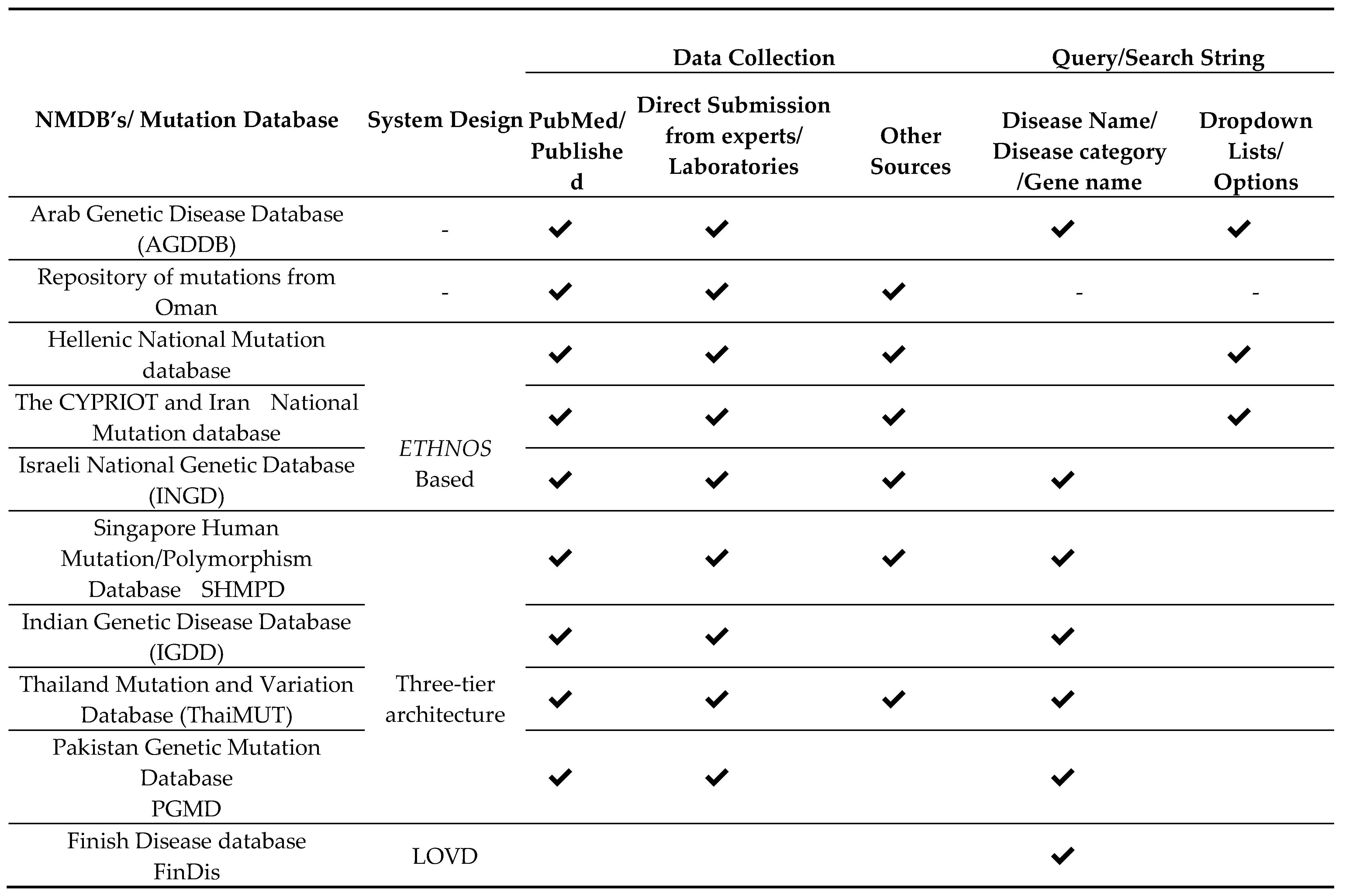
4.1. System Design
4.2. The Quality Data Collection
4.3. Querying the Database
4.4. Disease-Related Content
5. Discussion
- The mutation databases (NEMDBs) have used different platforms for the development and hence the data querying, data representation, adding new records, and all the other features of these databases are different from each other.
- Most of these databases are linked to the central databases but not with each other and due to this reason, these databases may have duplicate data. The data duplication may occur due to the same/similar ethnic boundaries of different nations.
- Some of the databases are not been updated over time, hence not covering the new mutation disorders if occurred.
- Some databases provide details about a very limited number of mutation disorders. These databases may inherit records from other databases as the same population can have similar mutation disorders as in other databases, though not included.
6. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
| 1 | https://coggle.it/diagram/YUgtHQ9uj-ii0cse/t/ncbi. |
| 2 | https://coggle.it/diagram/YTSDZgEJq_PwvBo1/t/the-national-and-mutation-frequency-databases-nemdbs. |
References
- Agris, P.F., The importance of being modified: roles of modified nucleosides and Mg2+ in RNA structure and function, in Progress in nucleic acid research and molecular biology. 1996, Elsevier. p. 79-129.
- Baltimore, D., Our genome unveiled. Nature, 2001. 409(6822): p. 814.
- Venter, J.C., et al., The sequence of the human genome. science, 2001. 291(5507): p. 1304-1351.
- Adams, P.C., et al., Hemochromatosis and iron-overload screening in a racially diverse population. New England Journal of Medicine, 2005. 352(17): p. 1769-1778. [CrossRef]
- Bhardwaj, U., Y.-H. Zhang, and E.R. McCabe, Neonatal hemoglobinopathy screening: molecular genetic technologies. Molecular genetics and metabolism, 2003. 1(80): p. 129-137. [CrossRef]
- Pradhan, S., et al., Indian genetic disease database. Nucleic acids research, 2010. 39(suppl_1): p. D933-D938. [CrossRef]
- Bhardwaj, U., et al., Molecular genetic confirmatory testing from newborn screening samples for the common African-American, Asian Indian, Southeast Asian, and Chinese β-thalassemia mutations. American journal of hematology, 2005. 78(4): p. 249-255.
- Hoodfar, E. and A.S. Teebi, Genetic referrals of Middle Eastern origin in a western city: inbreeding and disease profile. Journal of medical genetics, 1996. 33(3): p. 212-215. [CrossRef]
- Consortium, I.H.G.S., Correction: initial sequencing and analysis of the human genome. Nature, 2001. 412(6846): p. 565.
- Stenson, P.D., et al., Human gene mutation database (HGMD®): 2003 update. Human mutation, 2003. 21(6): p. 577-581. [CrossRef]
- Hamosh, A., et al., Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic acids research, 2005. 33(suppl_1): p. D514-D517. [CrossRef]
- Claustres, M., et al., Time for a unified system of mutation description and reporting: a review of locus-specific mutation databases. Genome research, 2002. 12(5): p. 680-688. [CrossRef]
- Bianco, A.M., et al., Database tools in genetic diseases research. Genomics, 2013. 101(2): p. 75-85. [CrossRef]
- Patrinos, G.P., et al., Hellenic National Mutation database: a prototype database for mutations leading to inherited disorders in the Hellenic population. Human mutation, 2005. 25(4): p. 327-333. [CrossRef]
- Kleanthous, M., et al., The cypriot and Iranian national mutation frequency databases. Human mutation, 2006. 27(6): p. 598-599. [CrossRef]
- Hunter, L. and K.B. Cohen, Biomedical language processing: what's beyond PubMed? Molecular cell, 2006. 21(5): p. 589-594.
- Lu, Z., PubMed and beyond: a survey of web tools for searching biomedical literature. Database, 2011. 2011. [CrossRef]
- Ding, J., et al., PubMed Assistant: a biologist-friendly interface for enhanced PubMed search. Bioinformatics, 2006. 22(3): p. 378-380. [CrossRef]
- Plake, C., et al., AliBaba: PubMed as a graph. Bioinformatics, 2006. 22(19): p. 2444-2445. [CrossRef]
- Tsai, R.T.-H., et al., PubMed-EX: a web browser extension to enhance PubMed search with text mining features. Bioinformatics, 2009. 25(22): p. 3031-3032. [CrossRef]
- Galperin, M.Y. and G.R. Cochrane, Nucleic acids research annual database issue and the NAR online molecular biology database collection in 2009. Nucleic Acids Research, 2009. 37(suppl_1): p. D1-D4. [CrossRef]
- Scriver, C.R., et al., PAHdb: a locus-specific knowledgebase. Human mutation, 2000. 15(1): p. 99-104. [CrossRef]
- Landrum, M.J., et al., ClinVar: improvements to accessing data. Nucleic acids research, 2020. 48(D1): p. D835-D844. [CrossRef]
- Sayers, E.W., et al., Database resources of the national center for biotechnology information. Nucleic acids research, 2012. 40(D1): p. D13-D25.
- Boguski, M.S., T.M. Lowe, and C.M. Tolstoshev, dbEST—database for “expressed sequence tags”. Nature genetics, 1993. 4(4): p. 332-333.
- Church, D.M., et al., Public data archives for genomic structural variation. Nature genetics, 2010. 42(10): p. 813-814. [CrossRef]
- Louhichi, A., A. Fourati, and A. Rebaï, IGD: a resource for intronless genes in the human genome. Gene, 2011. 488(1-2): p. 35-40. [CrossRef]
- Mailman, M., et al., Bagoutdinov r, hao l. Kiang a, Paschall J, Phan l, Popova n, Pretel s, Ziyabari l, lee M, shao Y, Wang ZY, sirotkin K, Ward M, Kholodov M, Zbicz K, Beck J, Kimelman M, shevelev s, Preuss D, Yaschenko e, graeff a, Ostell J, sherry sT. The ncBi dbgaP database of genotypes and phenotypes. Nat Genet, 2007. 39: p. 1181-6. [CrossRef]
- Manolio, T.A., et al., New models of collaboration in genome-wide association studies: the Genetic Association Information Network. Nature genetics, 2007. 39(9).
- Sherry, S.T., et al., dbSNP: the NCBI database of genetic variation. Nucleic acids research, 2001. 29(1): p. 308-311. [CrossRef]
- Horaitis, O. and R.G. Cotton, Human mutation databases. Current protocols in bioinformatics, 2005. 9(1): p. 1.10. 1-1.10. 13.
- Hamosh, A., et al., Online Mendelian inheritance in man (OMIM). Human mutation, 2000. 15(1): p. 57-61. [CrossRef]
- Cooper, D.N., et al., Genes, mutations, and human inherited disease at the dawn of the age of personalized genomics. Human mutation, 2010. 31(6): p. 631-655. [CrossRef]
- Stenson, P.D., et al., The Human Gene Mutation Database: towards a comprehensive repository of inherited mutation data for medical research, genetic diagnosis and next-generation sequencing studies. Human genetics, 2017. 136(6): p. 665-677. [CrossRef]
- Cooper, D.N., et al., On the sequence-directed nature of human gene mutation: the role of genomic architecture and the local DNA sequence environment in mediating gene mutations underlying human inherited disease. Human mutation, 2011. 32(10): p. 1075-1099. [CrossRef]
- Stenson, P.D., et al., The Human Gene Mutation Database: building a comprehensive mutation repository for clinical and molecular genetics, diagnostic testing and personalized genomic medicine. Human genetics, 2014. 133(1): p. 1-9. [CrossRef]
- Samuels, M.E. and G.A. Rouleau, The case for locus-specific databases. Nature Reviews Genetics, 2011. 12(6): p. 378-379. [CrossRef]
- Celli, J., et al., Curating gene variant databases (LSDBs): toward a universal standard. Human mutation, 2012. 33(2): p. 291-297. [CrossRef]
- Vihinen, M., et al., Guidelines for establishing locus specific databases. Human mutation, 2012. 33(2): p. 298-305. [CrossRef]
- Dalgleish, R., LSDBs and how they have evolved. Human Mutation, 2016. 37(6): p. 532-539. [CrossRef]
- Béroud, C., et al., UMD (Universal mutation database): a generic software to build and analyze locus-specific databases. Human mutation, 2000. 15(1): p. 86-94. [CrossRef]
- Riikonen, P. and M. Vihinen, MUTbase: maintenance and analysis of distributed mutation databases. Bioinformatics (Oxford, England), 1999. 15(10): p. 852-859. [CrossRef]
- Brown, A.F. and M.A. McKie, MuStaR™ and other software for locus-specific mutation databases. Human mutation, 2000. 15(1): p. 76-85. [CrossRef]
- Fokkema, I.F., J.T. den Dunnen, and P.E. Taschner, LOVD: easy creation of a locus-specific sequence variation database using an “LSDB-in-a-box” approach. Human mutation, 2005. 26(2): p. 63-68.
- Scriver, C.R., Human genetics: lessons from Quebec populations. Annual review of genomics and human genetics, 2001. 2(1): p. 69-101. [CrossRef]
- Qasim, I., et al., Pakistan genetic mutation database (PGMD); a centralized Pakistani mutome data source. European journal of medical genetics, 2018. 61(4): p. 204-208. [CrossRef]
- Clark, B. and S. Thein, Molecular diagnosis of haemoglobin disorders. Clinical & Laboratory Haematology, 2004. 26(3): p. 159-176. [CrossRef]
- Tan, E.c., et al., Singapore Human Mutation/Polymorphism Database: a country-specific database for mutations and polymorphisms in inherited disorders and candidate gene association studies. Human mutation, 2006. 27(3): p. 232-235. [CrossRef]
- Patrinos, G.P., National and ethnic mutation databases: recording populations' genography. Human mutation, 2006. 27(9): p. 879-887. [CrossRef]
- Teebi, A.S., et al., Arab genetic disease database (AGDDB): A population-specific clinical and mutation database. Human mutation, 2002. 19(6): p. 615-621. [CrossRef]
- Tadmouri, G.O., et al., CTGA: the database for genetic disorders in Arab populations. Nucleic acids research, 2006. 34(suppl_1): p. D602-D606. [CrossRef]
- Peltonen, L., A. Jalanko, and T. Varilo, Molecular genetics the Finnish disease heritage. Human molecular genetics, 1999. 8(10): p. 1913-1923.
- van Baal, S., et al., ETHNOS: a versatile electronic tool for the development and curation of National Genetic databases. Human genomics, 2010. 4(5): p. 1-8.
- Zlotogora, J. and G.P. Patrinos, The Israeli National Genetic database: a 10-year experience. Human genomics, 2017. 11(1): p. 1-5. [CrossRef]
- Nakouzi, G., K. Kreidieh, and S. Yazbek, A review of the diverse genetic disorders in the Lebanese population: highlighting the urgency for community genetic services. Journal of community genetics, 2015. 6(1): p. 83-105. [CrossRef]
- Sefiani, A., Genetic disorders in Morocco, in Genetic disorders Among Arab populations. 2010, Springer. p. 455-472.
- Ruangrit, U., et al., Thailand mutation and variation database (ThaiMUT). Human mutation, 2008. 29(8): p. E68-E75. [CrossRef]
- Romdhane, L., et al., Genetic diseases in the Tunisian population. American Journal of Medical Genetics Part A, 2011. 155(1): p. 238-267. [CrossRef]
- Rajab, A., et al., Repository of mutations from Oman: The entry point to a national mutation database. F1000Research, 2015. 4. [CrossRef]
- Megarbane, A., et al., The Lebanese National Mutation Frequency database. Eur. J. Hum. Genet, 2006. 14(Suppl 1): p. 365.
- van Baal, S., et al., FINDbase: a relational database recording frequencies of genetic defects leading to inherited disorders worldwide. Nucleic acids research, 2007. 35(suppl_1): p. D690-D695.
- Dalabira, E., et al., DruGeVar: an online resource triangulating drugs with genes and genomic biomarkers for clinical pharmacogenomics. Public Health Genomics, 2014. 17(5-6): p. 265-271. [CrossRef]
- Fokkema, I.F., et al., LOVD v. 2.0: the next generation in gene variant databases. Human mutation, 2011. 32(5): p. 557-563. [CrossRef]
- Coordinators, N.R., Database resources of the national center for biotechnology information. Nucleic acids research, 2016. 44(Database issue): p. D7. [CrossRef]
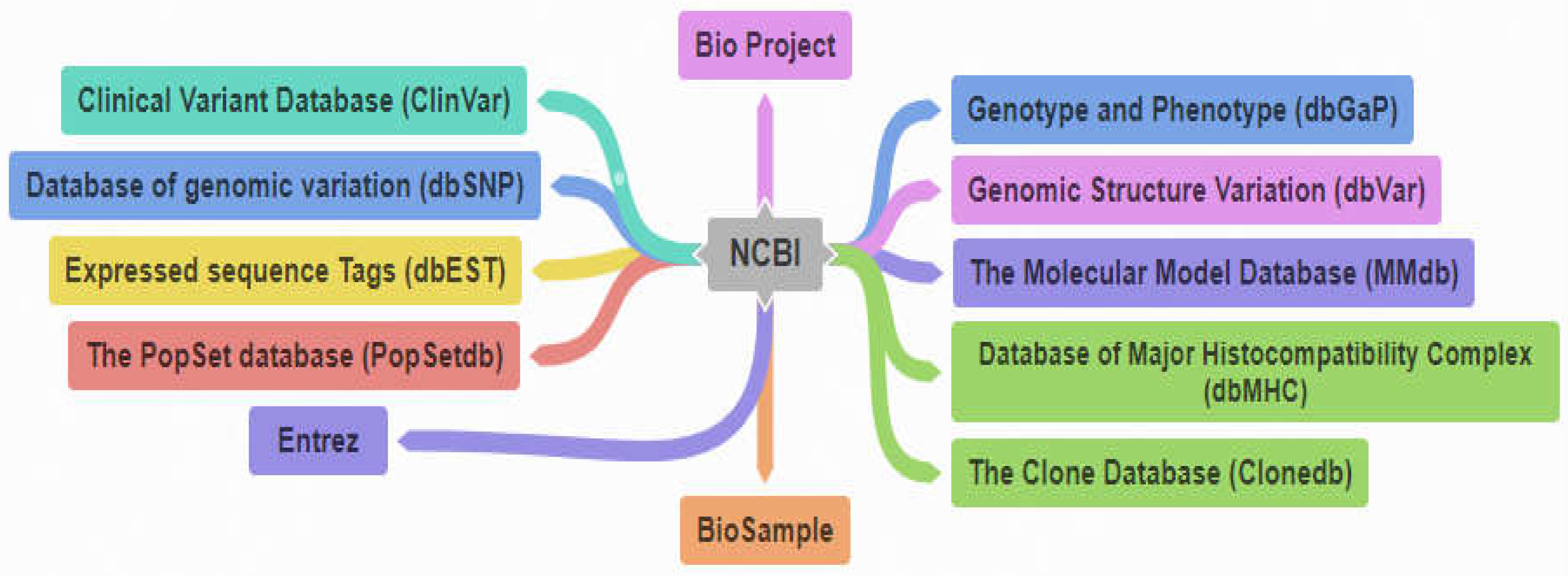
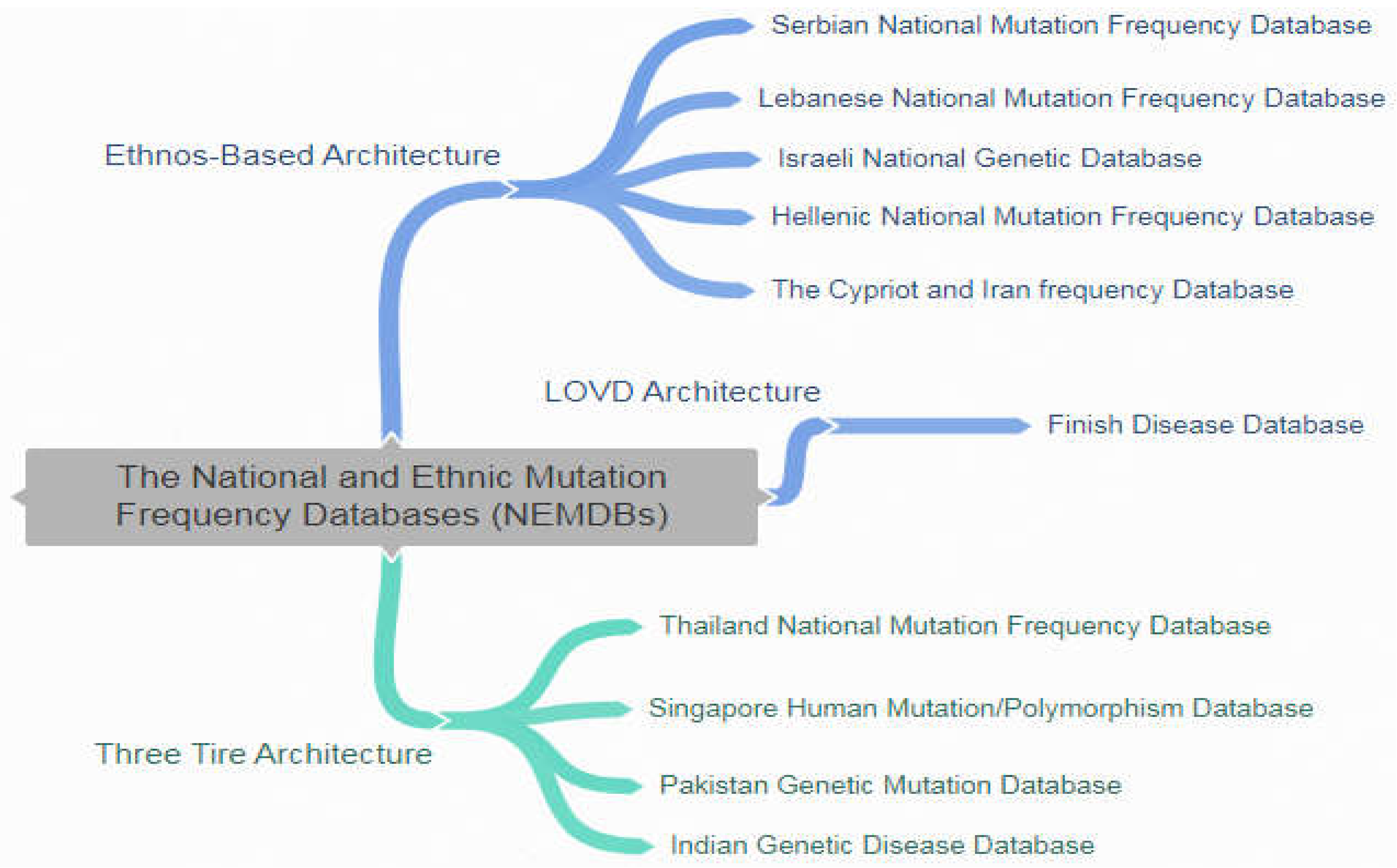
| References | Database Name | Brief Description |
|---|---|---|
| [13] | Bio Project Database | www.ncbi.nlm.nih.gov/bioproject/) The database allows users for submitting detailed research studies from intensive genome sequences projects to huge worldwide associations. |
| [13] | BioSample Database | The Biosample Database (www.ncbi.nlm.nih.gov/biosample/) is a new resource that provides annotation for biological samples used in a variety of NCBI-submitted studies, including genome-wide association study (GWAS), epigenetics, genomics sequencing, and microarrays. |
| [23] | Clinical variant database( ClinVar) | ClinVar (1,2) is a database that contains human genomic variants and its relevant disease. The database is publically available. |
| [24] | PopSet Database | (www.ncbi.nlm.nih.gov/popset/) This database contains different sets of data submitted to GenBank. The data is about the gene-related sequence data and their alignments of a certain population, phylogenetic, mutation, and study of the ecosystem. |
| [24] | Clone database (CloneDB) | (www.ncbi.nlm.nih.gov/clone/) The database is about Incorporating clones and libraries information, which includes sequence data, the position of maps, and information distribution. It also provides filtering through organism and vector types. |
| [24] | MMDB (Molecular Modeling Database) | (www.ncbi.nlm.nih.gov/Structure/CN3D/cn3d.shtml) It contain details about sequence alignments and profiles for the representation of protein spheres preserved in the evolution of molecule. |
| [25] | Database of expressed sequence tags (dbEST) Nucleotide EST Database | This database holds the collection of Sequence Tags and covers short details about cDNA (transcript) sequences. dbEST is accessible directly via Nucleotide EST Database. |
| [26] | Database of Genomic Structural Variation (dbVar) | It was designed for collecting details about large-scale genomic variation that includes large insertions, deletions, translocations, and inversions. It also contains the relations of different variants to their phenotype. |
| [27] | Entrez | Entrez is a rich database that integrates information from 35 databases containing records of 570 million. The database provides a graphical representation of sequences and chromosome maps and that's why it is considered to be favorable in genetic research. |
| [28,29] | Databases of Genotypes and Phenotypes (dbGaP) | The database contains information about genotype and phenotype and gathers it using studies such as GWAS, medical resequencing, and molecular diagnostic assays. |
| [30] | Database of Short Genetic Variations (dbSNP) | This database was developed for supporting large-scale polymorphism detection such as HapMap. It has been then updated as a collection for other classes such as insertions/deletions, microsatellites, and non-polymorphic variants. |
| [30] | Database of Major Histocompatibility Complex (dbMHC) | An interactive alignment viewer for HLA and related genes, as well as MHC microsatellite database are included. |
| Databases | References |
|---|---|
| Turkish Human Mutation Database | Unpublished |
| Cyprus Gene Mutation Database | Unpublished |
| Iranian Human Mutation Database | Unpublished |
| Singapore Human Mutation and Polymorphism Database | [48] |
| Arab Disease Mutation Database | [50] |
| Catalog of Transmission Genetics in Arabs (CTGA) | [51] |
| Finnish Disease Heritage | [52] |
| Ref | Database | Brief Description |
|---|---|---|
| [52] | Finnish Disease Heritage 2002 | This database contains the gene mutations and its related comprehensive information of the Finnish population. Mutant allele frequencies are typically reported for Finnish mutations together with multiple external links (OMIM; GeneTests; www.genetests.org) and references. The database was initially published in 2004 and has been updated by adding more genes and mutation disorders. This database has been designed using the LOVD platform. |
| [15] | The Iranian National Mutation Frequency Database (Iran) NEMDBs 2006 Cypriot National Mutation Frequency Database 2006 |
Here two similar databases are presented, one for the population of Cyprus and the other for the Iranian population. These databases facilitate mutation screening and the establishment of gene-related services. Both of the databases are developed using the ETHNOS platform. |
| [14] | Hellenic National Mutation database 2005 | Hellenic national mutation database aims to provide qualitative and updated reports of genetic disorders of Greece population. They have reported various diseases along with related information occurring in the Hellenic (Greece) population. |
| [54] | Israeli National Genetic Database | The database has documented all the genetic disorders happening in the Jewish and non-Jewish populations of Israel. The database has been developed based on the ETHNOS platform. Moreover, the Israeli NEMDB offers a detailed list of all the registered laboratories that provide genetic testing facilities of the Israeli population through a separate query interface. |
| [55] | The Lebanese National Mutation Frequency Database Lebanon 2006 | This database was designed to analyze the genetic diseases in the population of Lebanon. |
| [56] | The Moroccan Human Mutation Database (Morocco) 2010 | The Moroccan mutation database was developed to report the various mutation disorders found in the population of morocco. The Moroccan human mutation database is available online and a report in book chapter containing the details of various genetic disorders have also been published. |
| [57] | Thailand Human Mutation and Variation database (Thailand) 2008 | ThaiMUT is an online ethnical database reporting the mutation disorders of the population of Thailand. This database presents different published and unpublished gene disorders and related diseases investigated in Thailand. |
| [6] | Indian Genetic Disease Database (India) 2010 | A database containing the gene-related disease integrated from the Indian population. The diseases of this database have been curated by domain experts. The database was developed using three-tier architecture. |
| [46] | Pakistan Genetic Mutation Database (PGMD) | The database contains information about different disorders occurring in the Pakistani population. Pakistan Gene Mutation Database have currently two versions: one is the public version, which has used relational database and is available for the public. The second version is developed using ontology as a knowledge base. |
| [58] | Tunisian National Mutation Frequency Database | This database was developed to collect data about the different genetic disorders found in Tunisian population. |
| [51] | Catalog of Arab Disease Mutation Database (CTGA) 2006 | The CTGA database is an open-access repository of information and findings on human gene variations and inherited, heritable, genetic disorders in Arabs that is constantly updated. |
| [49] | Singapore human mutation database 2006 | The database contains mutations found in Singapore for Mendelian diseases. They present the mutation disorders and their frequency of polymorphisms examined based on phenotypes. |
| [59] | Oman 2015 | The database was developed for collecting and managing the mutations found in the population of Oman. In this database, the mutations were collected from the scientific literature and service provision. |
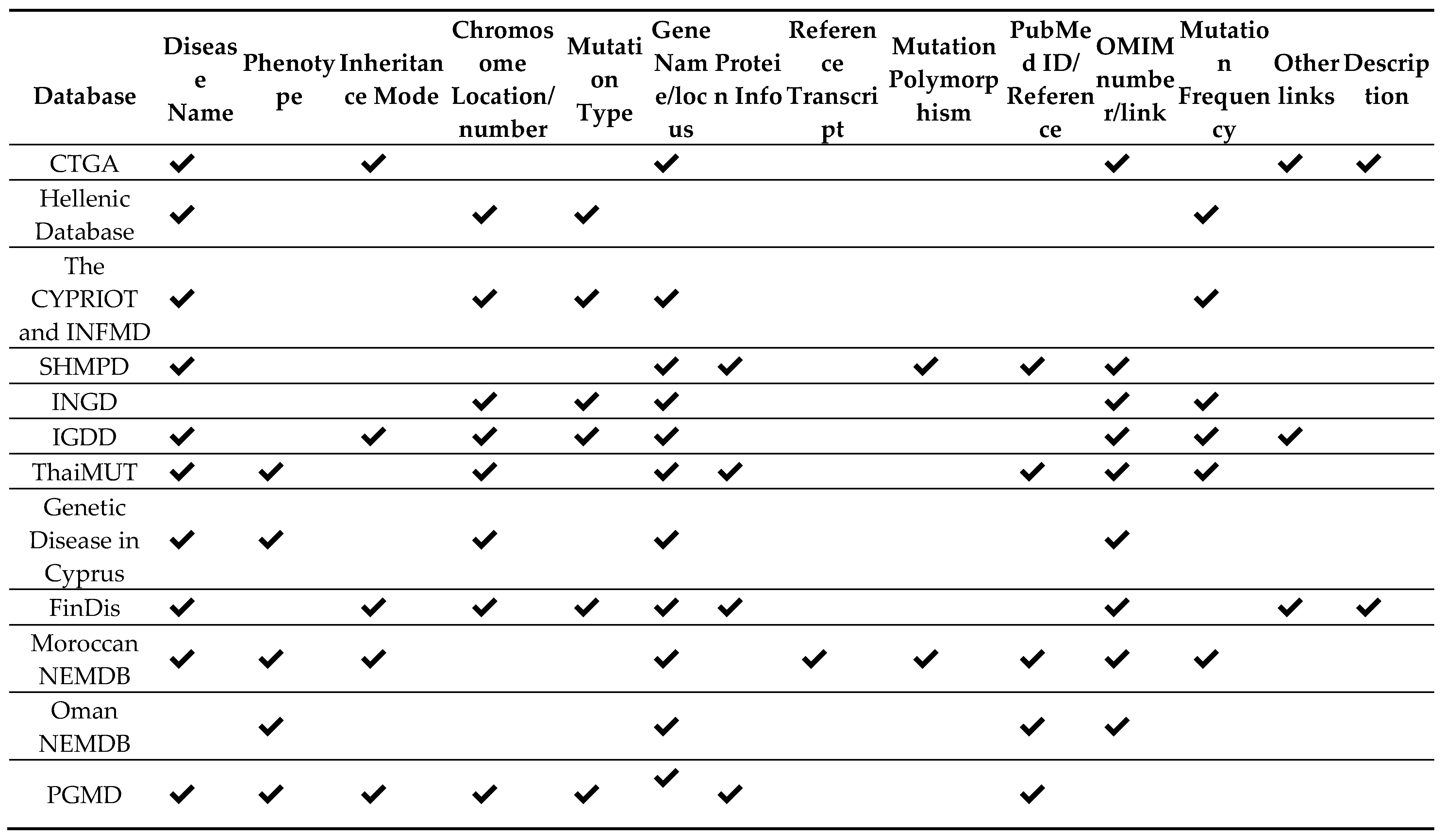

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




