Submitted:
13 January 2023
Posted:
17 January 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Data
3. Materials and Methods
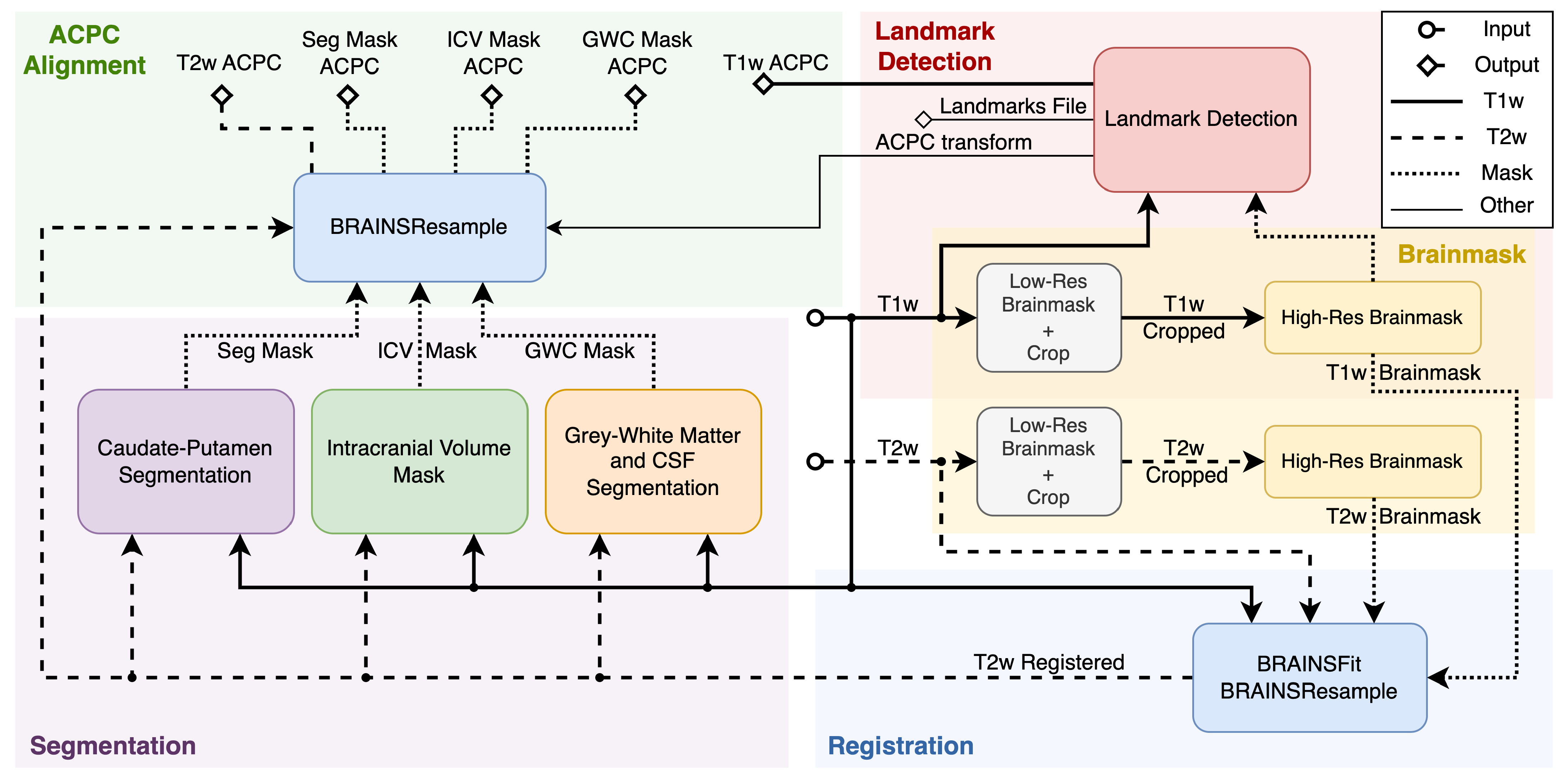
3.1. Pipeline Overview
3.2. Deep Learning Segmentation Model
3.3. Landmarks Detection
3.4. Image Registration and AC-PC Alignment
4. Results
4.1. Deep-Learning Model Accuracy
4.1.1. Brainmask
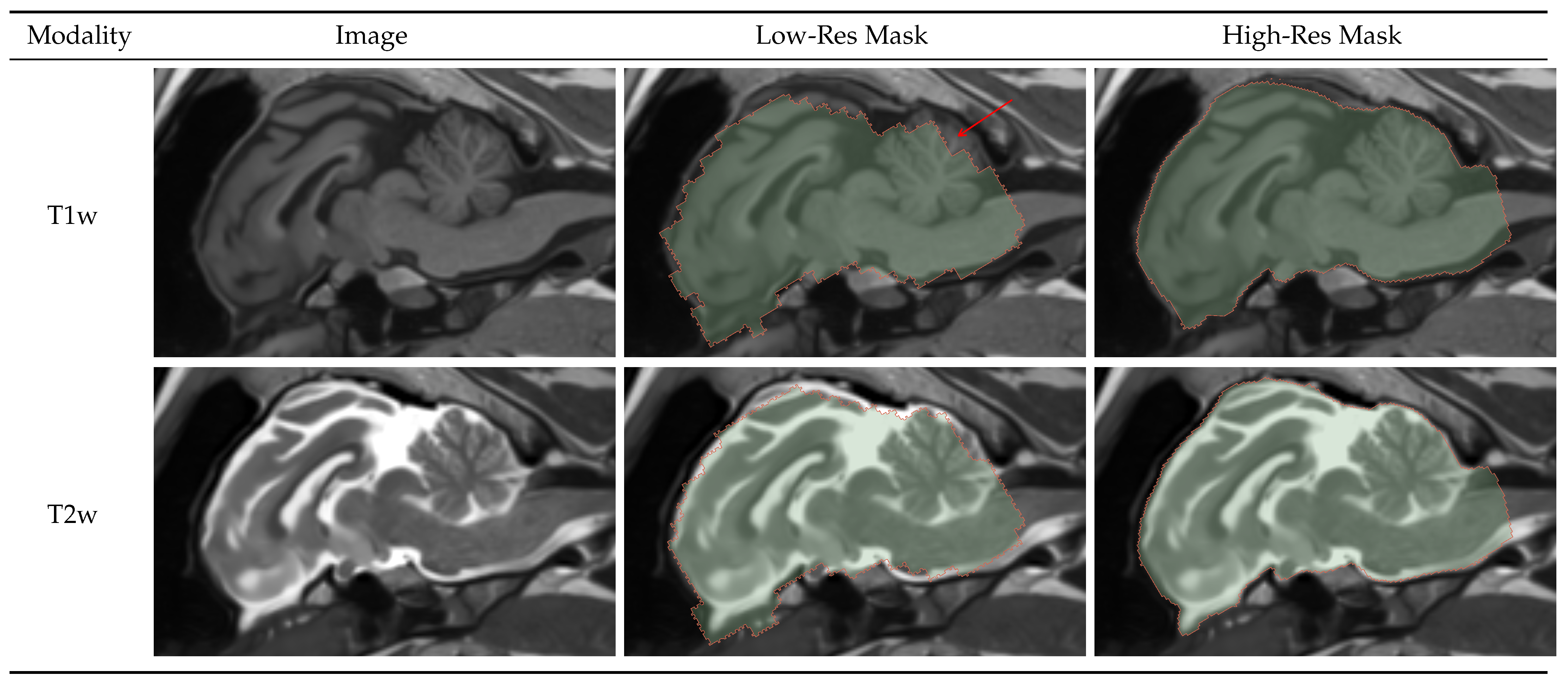
4.1.2. Intracranial Volume Mask
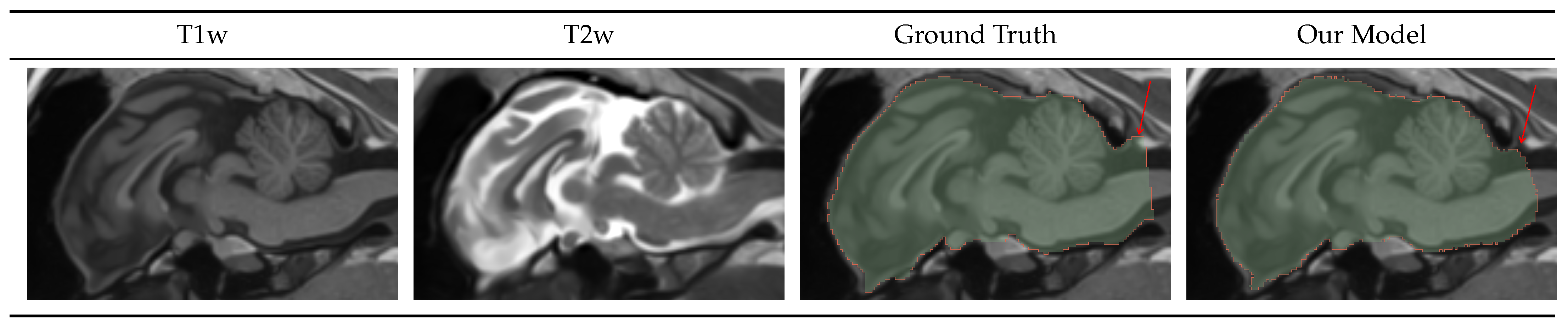
4.1.3. Caudate-Putamen Segmentation Mask
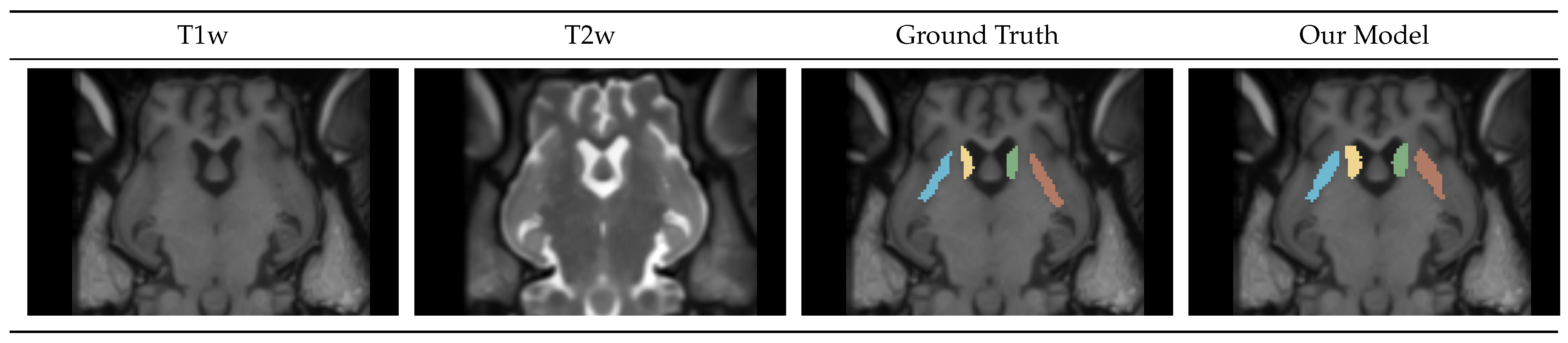
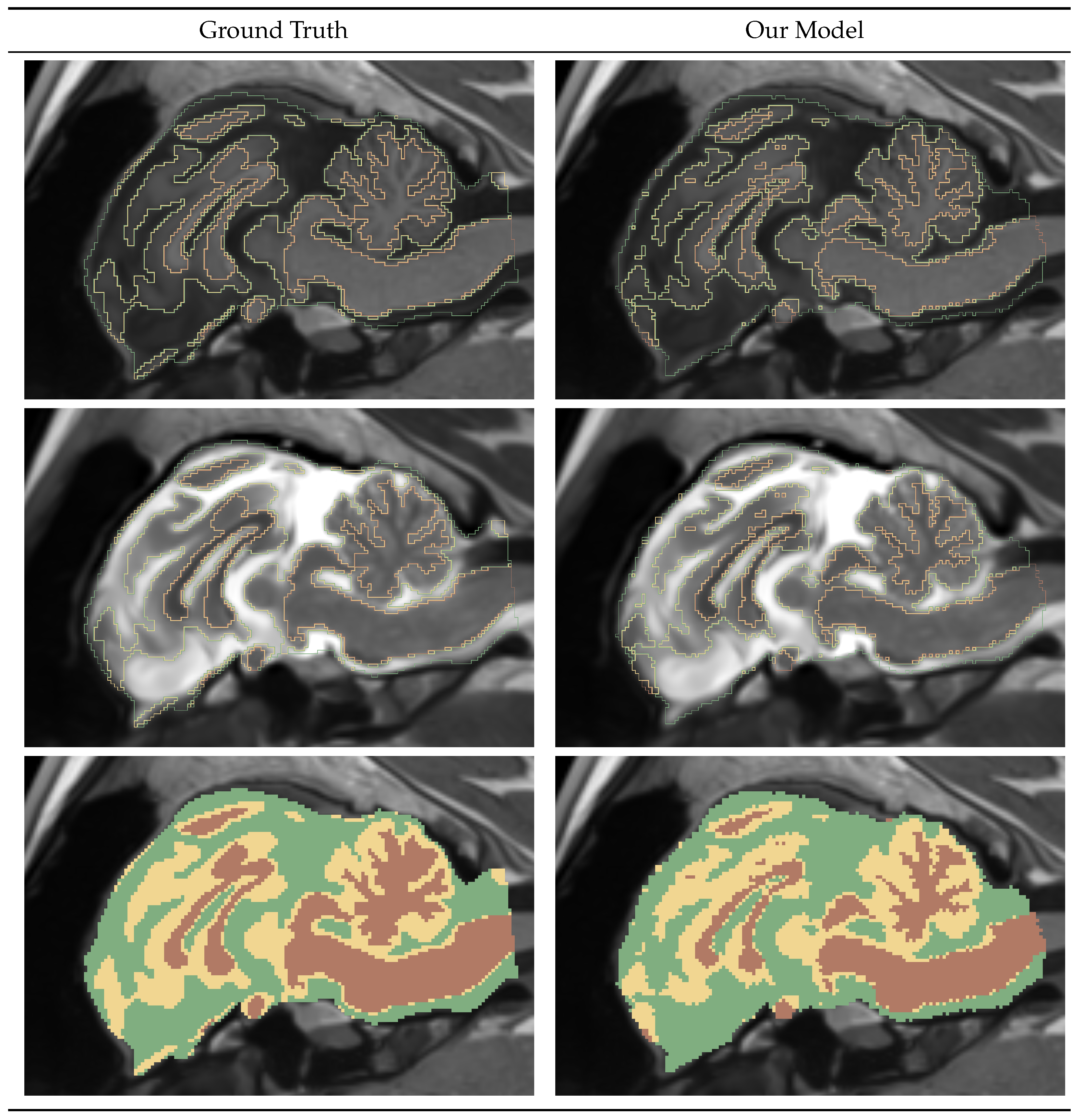
4.1.4. Gray-White-CSF Segmentation Mask
4.2. Pipeline Performance
4.2.1. Runtime
4.2.2. Memory and Hardware
5. Discussion
5.1. Deep Learning Models
5.2. Future Work
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| T1w | T1-weighted MRI sequence |
| T2w | T2-weighted MRI sequence |
| CSF | Cerebrospinal Fluid |
| GWC | Gray Matter-White Matter-CSF |
| Seg | Caudate-Putamen Segmentation |
| bAVD | balanced Average Hausdorff Distance |
References
- Neurological disorders : public health challenges; World Health Organization: Geneva, 2006.
- Moon, C. New Insights into and Emerging Roles of Animal Models for Neurological Disorders. International Journal of Molecular Sciences 2022, 23. [Google Scholar] [CrossRef]
- Leung, C.; Jia, Z. Mouse Genetic Models of Human Brain Disorders. Frontiers in Genetics 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- McGarry, A.; McDermott, M.; Kieburtz, K.; de Blieck, E.A.; Beal, F.; Marder, K.; Ross, C.; Shoulson, I.; Gilbert, P.; Mallonee, W.M.; et al. A randomized, double-blind, placebo-controlled trial of coenzyme Q10 in Huntington disease. Neurology 2016, 88, 152–159. [Google Scholar] [CrossRef]
- Hersch, S.M.; Schifitto, G.; Oakes, D.; Bredlau, A.L.; Meyers, C.M.; Nahin, R.; and, H.D.R. The CREST-E study of creatine for Huntington disease. Neurology 2017, 89, 594–601. [Google Scholar] [CrossRef]
- Eaton, S.L.; Wishart, T.M. Bridging the gap: large animal models in neurodegenerative research. Mammalian Genome 2017, 28, 324–337. [Google Scholar] [CrossRef] [PubMed]
- Ardan, T.; Baxa, M.; Levinská, B.; Sedláčková, M.; Nguyen, T.D.; Klíma, J.; Juhás, Š.; Juhásová, J.; Šmatlíková, P.; Vochozková, P.; Motlík, J.; Ellederová, Z. Transgenic minipig model of Huntington’s disease exhibiting gradually progressing neurodegeneration. Disease Models & Mechanisms, 2019. [Google Scholar] [CrossRef]
- VODIČKA, P.; SMETANA, K.; DVOŘÁNKOVÁ, B.; EMERICK, T.; XU, Y.Z.; OUREDNIK, J.; OUREDNIK, V.; MOTLÍK, J. The Miniature Pig as an Animal Model in Biomedical Research. Annals of the New York Academy of Sciences 2005, 1049, 161–171. [Google Scholar] [CrossRef] [PubMed]
- Hoopes, A.; Mora, J.S.; Dalca, A.V.; Fischl, B.; Hoffmann, M. SynthStrip: skull-stripping for any brain image. NeuroImage 2022, 260, 119474. [Google Scholar] [CrossRef]
- Pierson, R.; Johnson, H.; Harris, G.; Keefe, H.; Paulsen, J.; Andreasen, N.; Magnotta, V. Fully automated analysis using BRAINS: autoWorkup. NeuroImage 2011, 54, 328–36. [Google Scholar] [CrossRef]
- Brzus, M.; Powers, A.B.; Knoernschild, K.S.; Sieren, J.C.; Johnson, H.J. Multi-agent reinforcement learning pipeline for anatomical landmark detection in minipigs. In Proceedings of the Medical Imaging 2022: Image Processing; Colliot, O.; Išgum, I., Eds. International Society for Optics and Photonics, SPIE, 2022, Vol. 12032, p. 1203229. [CrossRef]
- Schubert, R.; Frank, F.; Nagelmann, N.; Liebsch, L.; Schuldenzucker, V.; Schramke, S.; Wirsig, M.; Johnson, H.; Kim, E.Y.; Ott, S.; Hölzner, E.; Demokritov, S.O.; Motlik, J.; Faber, C.; Reilmann, R. Neuroimaging of a minipig model of Huntington’s disease: Feasibility of volumetric, diffusion-weighted and spectroscopic assessments. Journal of Neuroscience Methods 2016, 265, 46–55, Current Methods in Huntington’s Disease Research. [Google Scholar] [CrossRef]
- Swier, V.J.; White, K.A.; Johnson, T.B.; Sieren, J.C.; Johnson, H.J.; Knoernschild, K.; Wang, X.; Rohret, F.A.; Rogers, C.S.; Pearce, D.A.; Brudvig, J.J.; Weimer, J.M. A Novel Porcine Model of CLN2 Batten Disease that Recapitulates Patient Phenotypes. Neurotherapeutics 2022, 19, 1905–1919. [Google Scholar] [CrossRef]
- Avants, B.B.; Tustison, N.J.; Wu, J.; Cook, P.A.; Gee, J.C. An Open Source Multivariate Framework for n-Tissue Segmentation with Evaluation on Public Data. Neuroinformatics 2011, 9, 381–400. [Google Scholar] [CrossRef] [PubMed]
- Kerfoot, E.; Clough, J.; Oksuz, I.; Lee, J.; King, A.P.; Schnabel, J.A. Left-Ventricle Quantification Using Residual U-Net. Statistical Atlases and Computational Models of the Heart. Atrial Segmentation and LV Quantification Challenges; Pop, M., Sermesant, M., Zhao, J., Li, S., McLeod, K., Young, A., Rhode, K., Mansi, T., Eds.; Springer International Publishing: Cham, 2019; pp. 371–380. [Google Scholar]
- Consortium, T.M. Project MONAI, 2020. [CrossRef]
- Paszke, A.; Gross, S.; Massa, F.; Lerer, A.; Bradbury, J.; Chanan, G.; Killeen, T.; Lin, Z.; Gimelshein, N.; Antiga, L.; Desmaison, A.; Kopf, A.; Yang, E.; DeVito, Z.; Raison, M.; Tejani, A.; Chilamkurthy, S.; Steiner, B.; Fang, L.; Bai, J.; Chintala, S. PyTorch: An Imperative Style, High-Performance Deep Learning Library. Advances in Neural Information Processing Systems; Wallach, H.; Larochelle, H.; Beygelzimer, A.; d’Alché-Buc, F.; Fox, E.; Garnett, R., Eds. Curran Associates, Inc., 2019, Vol. 32.
- Falcon, W.; Borovec, J.; Wälchli, A.; Eggert, N.; Schock, J.; Jordan, J.; Skafte, N.; Ir1dXD.; Bereznyuk, V.; Harris, E.; et al. PyTorchLightning/pytorch-lightning: 0.7.6 release, 2020. [CrossRef]
- Johnson, H.J.; McCormick, M.M.; Ibanez, L. Template:The ITK Software Guide Book 1: Introduction and Development Guidelines-Volume 1, 2015.
- Chollet, M.B.; Aldridge, K.; Pangborn, N.; Weinberg, S.M.; DeLeon, V.B. Landmarking the Brain for Geometric Morphometric Analysis: An Error Study. PLoS ONE 2014, 9, e86005. [Google Scholar] [CrossRef] [PubMed]
- da Silva, Jr, E. B.; Leal, A.G.; Milano, J.B.; da Silva, Jr, L.F.M.; Clemente, R.S.; Ramina, R. Image-guided surgical planning using anatomical landmarks in the retrosigmoid approach. Acta Neurochir. (Wien) 2010, 152, 905–910. [Google Scholar] [CrossRef]
- Visser, M.; Petr, J.; Müller, D.M.J.; Eijgelaar, R.S.; Hendriks, E.J.; Witte, M.; Barkhof, F.; van Herk, M.; Mutsaerts, H.J.M.M.; Vrenken, H.; de Munck, J.C.; De Witt Hamer, P.C. Accurate MR Image Registration to Anatomical Reference Space for Diffuse Glioma. Frontiers in Neuroscience 2020, 14. [Google Scholar] [CrossRef] [PubMed]
- François-Lavet, V.; Henderson, P.; Islam, R.; Bellemare, M.G.; Pineau, J. An Introduction to Deep Reinforcement Learning. CoRR 2018, abs/1811.12560, [1811.12560].
- Ruder, S. An Overview of Multi-Task Learning in Deep Neural Networks. ArXiv 2017, abs/1706.05098.
- Caruana, R. Multitask Learning. Machine Learning 1997, 28, 41–75. [Google Scholar] [CrossRef]
- Watkins, C.J.C.H.; Dayan, P. Q-learning. Machine Learning 1992, 8, 279–292. [Google Scholar] [CrossRef]
- Huber, P.J. Robust Estimation of a Location Parameter. The Annals of Mathematical Statistics 1964, 35, 73–101. [Google Scholar] [CrossRef]
- Kingma, D.P.; Ba, J. Adam: A Method for Stochastic Optimization. 3rd International Conference on Learning Representations, ICLR 2015, San Diego, CA, USA, May 7-9, 2015, Conference Track Proceedings; Bengio, Y.; LeCun, Y., Eds., 2015.
- Thrun, S. Efficient Exploration In Reinforcement Learning. Technical Report CMU-CS-92-102, Carnegie Mellon University, Pittsburgh, PA, 1992.
- BRAINSTools. https://github.com/BRAINSia/BRAINSTools.
- Dice, L.R. Measures of the Amount of Ecologic Association Between Species. Ecology 1945, 26, 297–302. [Google Scholar] [CrossRef]
- Aydin, O.U.; Taha, A.A.; Hilbert, A.; Khalil, A.A.; Galinovic, I.; Fiebach, J.B.; Frey, D.; Madai, V.I. On the usage of average Hausdorff distance for segmentation performance assessment: hidden error when used for ranking. European Radiology Experimental 2021, 5. [Google Scholar] [CrossRef] [PubMed]
- Taha, A.A.; Hanbury, A. Metrics for Evaluating 3D Medical Image Segmentation: analysis, selection, and tool. BMC Medical Imaging 2015, 15, 29. [Google Scholar] [CrossRef]
- Detlefsen, N.S.; Borovec, J.; Schock, J.; Jha, A.H.; Koker, T.; Liello, L.D.; Stancl, D.; Quan, C.; Grechkin, M.; Falcon, W. TorchMetrics - Measuring Reproducibility in PyTorch. Journal of Open Source Software 2022, 7, 4101. [Google Scholar] [CrossRef]
| Germany Dataset | Iowa Dataset | Total | ||||
|---|---|---|---|---|---|---|
| Number of | Images | Subjects | Images | Subjects | Images | Subjects |
| Training | 164 | 26 | 243 | 18 | 407 | 44 |
| Validation | 30 | 4 | 39 | 3 | 69 | 7 |
| Test | 18 | 3 | 30 | 2 | 48 | 5 |
| Germany Dataset | Iowa Dataset | Total | ||||
|---|---|---|---|---|---|---|
| Number of | Image Pairs | Subjects | Image Pairs | Subjects | Image Pairs | Subjects |
| Training | 78 | 26 | 113 | 18 | 191 | 44 |
| Validation | 15 | 4 | 19 | 3 | 34 | 7 |
| Test | 8 | 3 | 14 | 2 | 22 | 5 |
| Low-Resolution | High-Resolution | |||
|---|---|---|---|---|
| Dataset | DICE | bAVD | DICE | bAVD |
| Iowa | 0.89 | 0.21 | 0.97 | 0.03 |
| Germany | 0.88 | 0.32 | 0.97 | 0.06 |
| Total | 0.88 | 0.25 | 0.97 | 0.04 |
| Dataset | DICE | bAVD |
|---|---|---|
| Iowa | 0.97 | 0.03 |
| Germany | 0.98 | 0.03 |
| Total | 0.98 | 0.03 |
| Dataset | Left Caudate | Right Caudate | Left Putamen | Right Putamen | Global bAVD |
|---|---|---|---|---|---|
| Germany | 0.82 | 0.79 | 0.79 | 0.81 | 0.29 |
| Iowa | 0.80 | 0.83 | 0.78 | 0.80 | 0.27 |
| Total | 0.81 | 0.82 | 0.78 | 0.80 | 0.28 |
| Dataset | Gray Matter | White Matter | CSF | Global bAVD |
|---|---|---|---|---|
| Germany | 0.79 | 0.89 | 0.71 | 0.06 |
| Iowa | 0.81 | 0.89 | 0.78 | 0.05 |
| Total | 0.80 | 0.89 | 0.76 | 0.05 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




