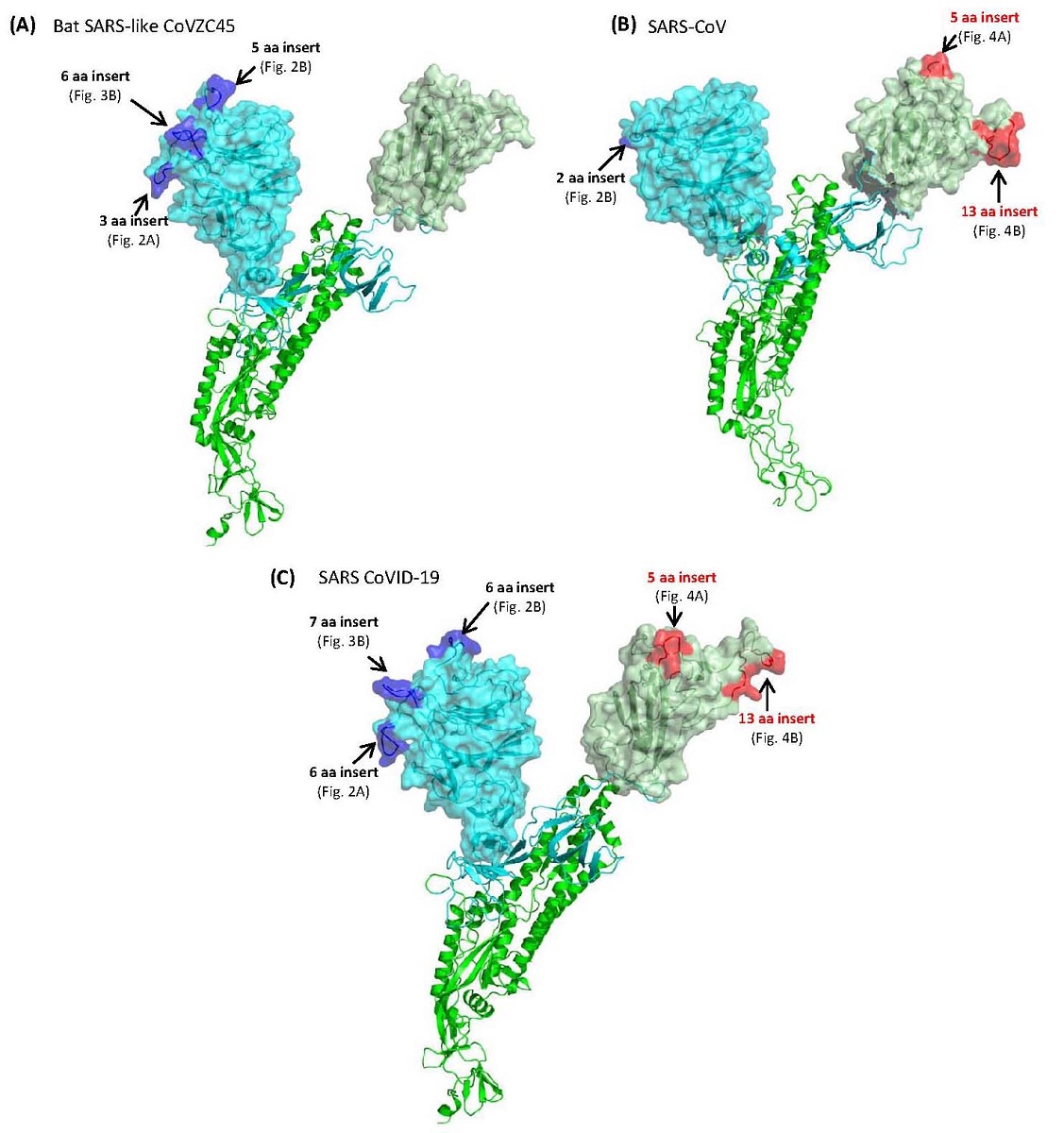
Both SARS-CoV-2 (COVID-19) and SARS coronaviruses (CoVs) are members of the subgenus Sarbecovirus. To understand the origin of SARS-CoV-2, protein sequences from sarbecoviruses were analyzed to identify highly-specific molecular markers consisting of conserved inserts or deletions (termed CSIs) in the spike (S) and nucleocapsid (N) proteins that are specific for either particular clusters/lineages of these viruses or are commonly shared by specific lineages. Three novel CSIs in the N-terminal domain of the spike protein S1-subunit (S1-NTD) are uniquely shared by the SARS-CoV-2, BatCoV-RaTG13 and most pangolin CoVs, distinguishing this cluster of viruses (SARS-CoV-2r) from all others. In the same positions, where these CSIs are found, related CSIs are also present in two other sarbecoviruses (viz. CoVZXC21 and CoVZC45 forming CoVZC cluster), which form an out group of the SARS-CoV-2r cluster. These three CSIs are not found in the SARS-CoVs. However, both SARS and SARS-CoV-2r CoVs contain two large CSIs in the C-terminal domain of S1 (S1-CTD), which binds the human ACE-2 receptor, that are absent in the CoVZC cluster of CoVs. These results indicate that while the S1-NTD of the SARS-CoV-2r viruses possesses the sequence characteristics of the CoVZC cluster of CoVs, their S1-CTD resembles the SARS viruses. Thus, the spike protein of SARS-CoV-2r viruses has likely originated from a recombination event between the S1-NTD of the CoVZC viruses and the S1-CTD of SARS viruses. This inference is also supported by the amino acid sequence similarity of the S1-NTD and S1-CTD from SARS-CoV-2 compared to the CoVZC and SARS CoVs. We also present evidence that one of the pangolin-CoV_MP789, whose receptor-binding domain is most similar to the SARS-CoV-2, is also derived by a recent recombination between the S1-NTD of the CoVZC CoVs and the S1-CTD of a SARS-CoV-2 related virus. Several other identified CSIs are specific for others clusters of sarbecoviruses including a clade consisting of bat SARS-CoVs (BM48-31/BGR/2008 and SARS_BtKY72). Structural mappings studies show that the identified CSIs are located within surface-exposed loops and form distinct patches on the surface of the spike protein. These surface loops/patches are predicted to interact with other host components and play important role in the biology/pathology of SARS-CoV-2 virus. Lastly, the CSIs specific for the SARS-CoV-2r clade provide novel means for development of new diagnostic and therapeutic targets for these viruses.