Submitted:
25 December 2025
Posted:
26 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
- A domain-aware taxonomy of contrastive learning methods is proposed, detailing their key components, loss functions, and adaptation strategies for medical data.
- A unified analysis of contrastive learning applications across medical imaging, electronic health records, genomics, and multimodal systems is presented, highlighting cross-domain similarities and differences.
- The challenges of evaluation, robustness, and generalization in medical contrastive learning are critically examined, with insights into reproducibility and transferability.
- Practical guidelines and future research directions are discussed, focusing on interpretability, fairness, and integration with federated and causal learning frameworks.
2. Methodology
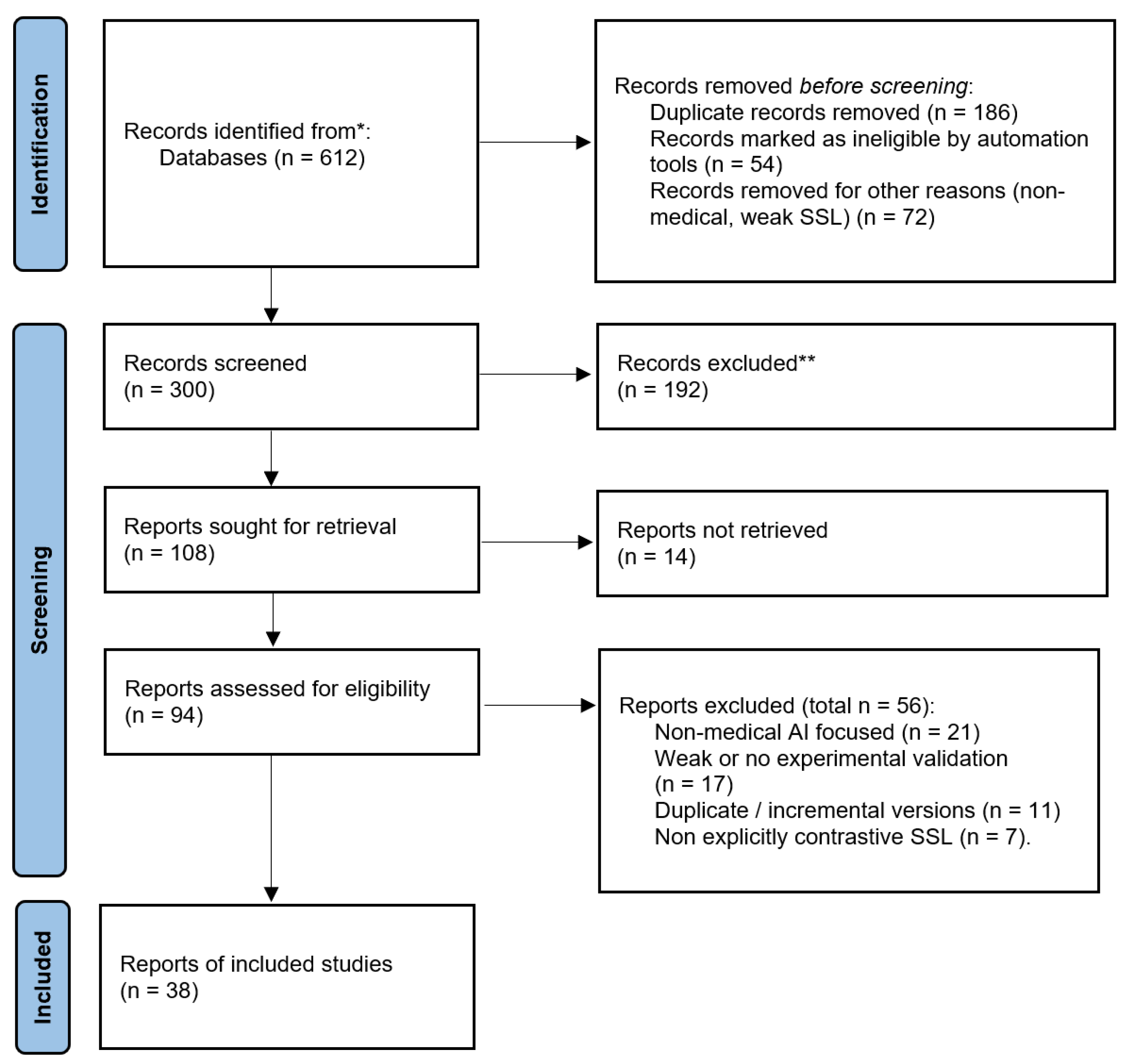
2.1. Databases and Search Strategy
- ("contrastive learning" OR "self-supervised") AND (medical OR healthcare OR clinical)
- ("contrastive learning" OR SimCLR OR MoCo OR BYOL) AND (radiology OR "chest x-ray" OR MRI OR CT)
- ("contrastive learning" OR "self-supervised") AND ("electronic health record" OR EHR OR "clinical notes")
- ("contrastive learning" OR "representation learning") AND (ECG OR EEG OR "physiological signals")
- ("contrastive learning" OR "self-supervised") AND (genomics OR proteomics OR "single-cell")
2.2. Inclusion and Exclusion Criteria
- Peer-reviewed journal articles or full-length conference papers, with a small number of influential preprints included when they introduced widely adopted methods or benchmarks.
- Work that explicitly employs contrastive, self-supervised, or closely related representation-learning objectives (e.g., InfoNCE, supervised contrastive loss, MoCo, SimCLR, BYOL- or CLIP-style frameworks).
- Applications involving medical, clinical, or biomedical data (e.g., medical imaging, EHRs, physiological time series, genomics, proteomics, or pathology).
- Preprints were included selectively when they introduced methods or benchmarks that have been widely adopted or cited in subsequent peer-reviewed literature.
- Studies that use contrastive objectives solely for non-medical domains (e.g., natural images, generic NLP) without any medical or biomedical application.
- Short abstracts, workshop posters without sufficient methodological detail, editorials, commentaries, and theses.
- Non–peer-reviewed technical reports and preprints that did not provide experimental validation or that were superseded by later peer-reviewed versions.
2.3. Screening and Synthesis
- Data modality (e.g., imaging, EHRs, physiological signals, genomics/pathology).
- Contrastive learning formulation (e.g., unsupervised, supervised, multimodal, temporal or patient-level objectives).
- Architectural choices (e.g., CNNs, Transformers, multimodal encoders).
- Downstream tasks and evaluation metrics (e.g., classification, segmentation, retrieval, risk prediction).
2.4. Protocol Registration
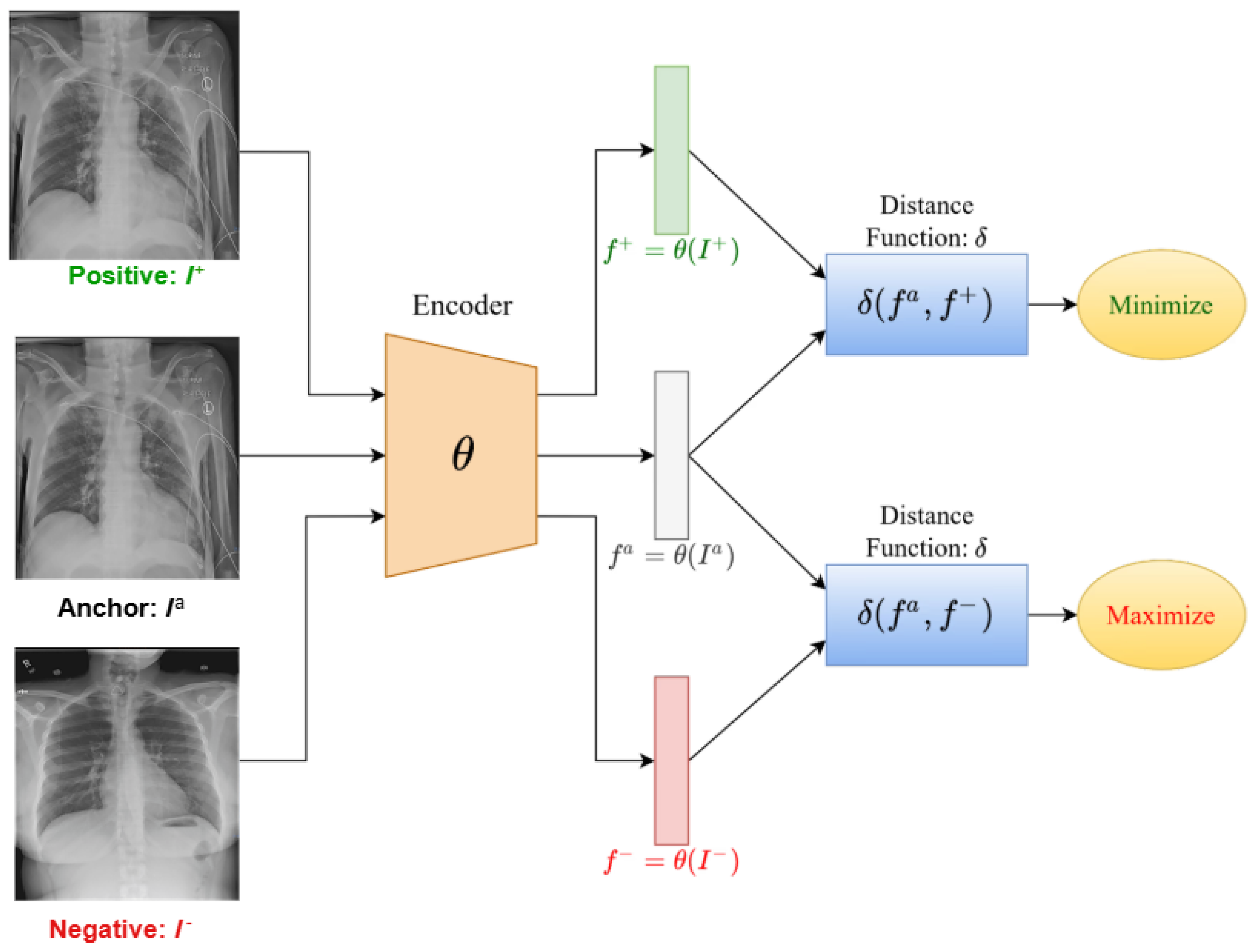
3. Overview of Contrastive Learning
3.1. Variants and Extensions of Contrastive Learning
3.1.1. Supervised Contrastive Learning
3.1.2. Self-Distillation with No Labels
3.1.3. Momentum Contrast
3.1.4. Simple Framework for Contrastive Learning of Visual Representations
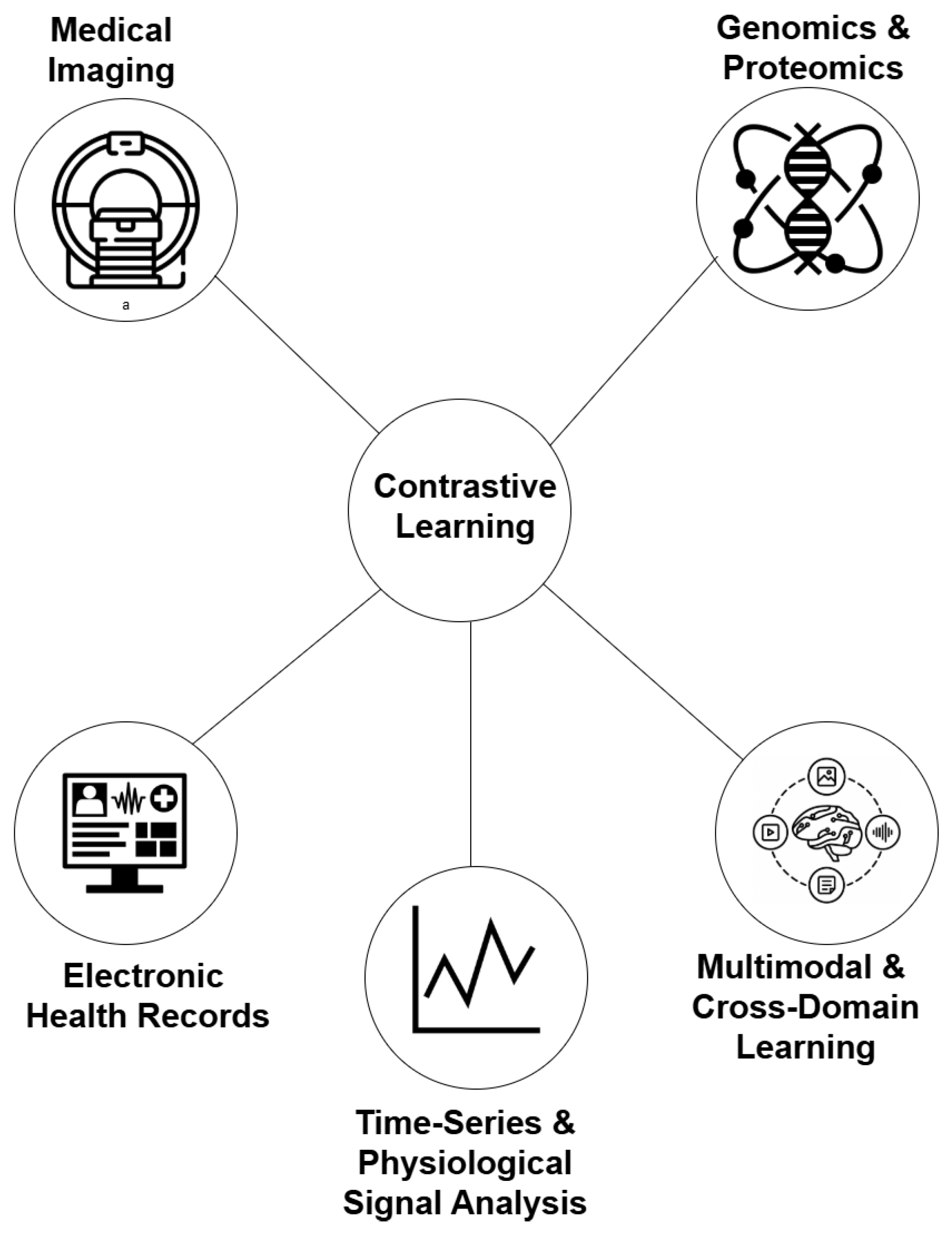
4. Applications of Contrastive Learning in Medical AI
4.1. Medical Imaging
4.2. Electronic Health Records
4.3. Genomics and Proteomics
4.4. Multimodal and Cross-Domain Learning
4.5. Time-Series and Physiological Signal Analysis
5. Challenges and Limitations
6. Discussion and Future Research Direction
7. Conclusion
Author Contributions
Conflicts of Interest
Abbreviations
| AI | Artificial Intelligence |
| AUROC | Area Under the Receiver Operating Characteristic |
| BioViL | Biomedical Vision–Language |
| BioViL-T | Biomedical Vision–Language with Temporal alignment |
| BiomedCLIP | Biomedical CLIP |
| BYOL | Bootstrap Your Own Latent |
| CheXpert | Chest X-ray benchmark dataset |
| CL | Contrastive Learning |
| CLOCS | Contrastive Learning of Cardiac Signals |
| COMET | Hierarchical contrastive framework |
| CONCH | Histopathology foundation model |
| ConVIRT | Contrastive Learning of Visual Representations from Text |
| CPC | Contrastive Predictive Coding |
| CXR-CLIP | Chest X-Ray CLIP |
| DINO | Self-Distillation with No Labels |
| ECG | Electrocardiogram |
| EEG | Electroencephalogram |
| EHR(s) | Electronic Health Record(s) |
| F1 | F1 score |
| GLoRIA | Global–Local image–text alignment in radiology |
| InfoNCE | Noise-Contrastive Estimation (loss) |
| MaCo | Masked Contrastive Learning |
| MBSL | Multi-Scale and Multi-Modal Contrastive Learning |
| MedCLIP | Medical CLIP |
| MIMIC-CXR | Medical Information Mart for Intensive Care—Chest X-Ray |
| MoCo | Momentum Contrast |
| MRI | Magnetic Resonance Imaging |
| MSCL | Multi-Scale Contrastive Learning |
| PCLR | Patient Contrastive Learning of Representations |
| PLIP | Pathology Language–Image Pretraining (model) |
| PMQ | Patient Memory Queue |
| PPI(s) | Protein–Protein Interaction(s) |
| PMC-CLIP | PubMed Central CLIP |
| RSNA | Radiological Society of North America |
| SAM | Segment Anything Model |
| scRNA-seq | single-cell RNA sequencing |
| SimCLR | Simple Framework for Contrastive Learning of Visual Representations |
| SwAV | Swapping Assignments between Multiple Views |
| VQA | Visual Question Answering |
| 1D CNN | One-dimensional Convolutional Neural Network |
References
- Parvin, N.; Joo, S.W.; Jung, J.H.; Mandal, T.K. Multimodal AI in Biomedicine: Pioneering the Future of Biomaterials, Diagnostics, and Personalized Healthcare. Nanomaterials 2025, 15, 895. [Google Scholar] [CrossRef]
- Nazir, A.; Hussain, A.; Singh, M.; Assad, A. Deep learning in medicine: advancing healthcare with intelligent solutions and the future of holography imaging in early diagnosis. Multimedia Tools and Applications 2025, 84, 17677–17740. [Google Scholar] [CrossRef]
- Mienye, I.D.; Swart, T.G.; Obaido, G.; Jordan, M.; Ilono, P. Deep convolutional neural networks in medical image analysis: A review. Information 2025, 16, 195. [Google Scholar] [CrossRef]
- Mienye, I.D.; Jere, N.; Obaido, G.; Ogunruku, O.O.; Esenogho, E.; Modisane, C. Large language models: an overview of foundational architectures, recent trends, and a new taxonomy. Discover Applied Sciences 2025, 7, 1027. [Google Scholar] [CrossRef]
- Nichyporuk, B.; Cardinell, J.; Szeto, J.; Mehta, R.; Falet, J.P.R.; Arnold, D.L.; Tsaftaris, S.A.; Arbel, T. Rethinking generalization: The impact of annotation style on medical image segmentation. arXiv 2022, arXiv:2210.17398. [Google Scholar] [CrossRef]
- Daneshjou, R.; Yuksekgonul, M.; Cai, Z.R.; Novoa, R.; Zou, J.Y. Skincon: A skin disease dataset densely annotated by domain experts for fine-grained debugging and analysis. Advances in Neural Information Processing Systems 2022, 35, 18157–18167. [Google Scholar]
- Krenzer, A.; Makowski, K.; Hekalo, A.; Fitting, D.; Troya, J.; Zoller, W.G.; Hann, A.; Puppe, F. Fast machine learning annotation in the medical domain: a semi-automated video annotation tool for gastroenterologists. BioMedical Engineering OnLine 2022, 21, 33. [Google Scholar] [CrossRef]
- Chen, H.; Gouin-Vallerand, C.; Bouchard, K.; Gaboury, S.; Couture, M.; Bier, N.; Giroux, S. Contrastive Self-Supervised Learning for Sensor-Based Human Activity Recognition: A Review. IEEE Access 2024. [Google Scholar] [CrossRef]
- Liu, S.; Zhao, L.; Chen, D.; Song, Z. Contrastive learning for image complexity representation. arXiv 2024, arXiv:2408.03230. [Google Scholar] [CrossRef]
- Ren, X.; Wei, W.; Xia, L.; Huang, C. A comprehensive survey on self-supervised learning for recommendation. ACM Computing Surveys 2025, 58, 1–38. [Google Scholar] [CrossRef]
- Prince, J.S.; Alvarez, G.A.; Konkle, T. Contrastive learning explains the emergence and function of visual category-selective regions. Science Advances 2024, 10, eadl1776. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.; Asmatullah, L.; Malik, A.; Khan, S.; Asif, H. A Survey on Self-supervised Contrastive Learning for Multimodal Text-Image Analysis. arXiv 2025, arXiv:2503.11101. [Google Scholar]
- Zeng, D.; Wu, Y.; Hu, X.; Xu, X.; Shi, Y. Contrastive learning with synthetic positives. In Proceedings of the European Conference on Computer Vision; Springer, 2024; pp. 430–447. [Google Scholar]
- Xu, Z.; Dai, Y.; Liu, F.; Wu, B.; Chen, W.; Shi, L. Swin MoCo: Improving parotid gland MRI segmentation using contrastive learning. Medical Physics 2024, 51, 5295–5307. [Google Scholar] [CrossRef] [PubMed]
- Jaiswal, A.; Babu, A.R.; Zadeh, M.Z.; Banerjee, D.; Makedon, F. A survey on contrastive self-supervised learning. Technologies 2020, 9, 2. [Google Scholar] [CrossRef]
- Gui, J.; Chen, T.; Zhang, J.; Cao, Q.; Sun, Z.; Luo, H.; Tao, D. A Survey on Self-supervised Learning: Algorithms, Applications, and Future Trends. IEEE Transactions on Pattern Analysis and Machine Intelligence 2024. [Google Scholar] [CrossRef]
- Wang, W.C.; Ahn, E.; Feng, D.; Kim, J. A review of predictive and contrastive self-supervised learning for medical images. Machine Intelligence Research 2023, 20, 483–513. [Google Scholar] [CrossRef]
- Shurrab, S.; Duwairi, R. Self-supervised learning methods and applications in medical imaging analysis: A survey. PeerJ Computer Science 2022, 8, e1045. [Google Scholar] [CrossRef]
- Khosla, P.; Teterwak, P.; Wang, C.; Sarna, A.; Tian, Y.; Isola, P.; Maschinot, A.; Liu, C.; Krishnan, D. Supervised contrastive learning. Advances in neural information processing systems 2020, 33, 18661–18673. [Google Scholar]
- Hu, H.; Wang, X.; Zhang, Y.; Chen, Q.; Guan, Q. A comprehensive survey on contrastive learning. Neurocomputing 2024, 610, 128645. [Google Scholar] [CrossRef]
- Wu, J.; Chen, J.; Wu, J.; Shi, W.; Wang, X.; He, X. Understanding contrastive learning via distributionally robust optimization. Advances in Neural Information Processing Systems 2024, 36. [Google Scholar]
- Le-Khac, P.H.; Healy, G.; Smeaton, A.F. Contrastive representation learning: A framework and review. Ieee Access 2020, 8, 193907–193934. [Google Scholar] [CrossRef]
- Kundu, R. The Beginner’s Guide to Contrastive Learning. https://www.v7labs.com/blog/contrastivelearning- guide, 2022. Accessed: 2025-10-10.
- Falcon, W.; Cho, K. A framework for contrastive self-supervised learning and designing a new approach. arXiv 2020, arXiv:2009.00104. [Google Scholar] [CrossRef]
- Wu, L.; Zhuang, J.; Chen, H. Voco: A simple-yet-effective volume contrastive learning framework for 3d medical image analysis. In Proceedings of the Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2024; pp. 22873–22882. [Google Scholar]
- Tang, C.; Zeng, X.; Zhou, L.; Zhou, Q.; Wang, P.; Wu, X.; Ren, H.; Zhou, J.; Wang, Y. Semi-supervised medical image segmentation via hard positives oriented contrastive learning. Pattern Recognition 2024, 146, 110020. [Google Scholar] [CrossRef]
- Zhang, C.; Zhang, K.; Pham, T.X.; Niu, A.; Qiao, Z.; Yoo, C.D.; Kweon, I.S. Dual temperature helps contrastive learning without many negative samples: Towards understanding and simplifying moco. In Proceedings of the Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 2022; pp. 14441–14450. [Google Scholar]
- Hoffmann, D.T.; Behrmann, N.; Gall, J.; Brox, T.; Noroozi, M. Ranking info noise contrastive estimation: Boosting contrastive learning via ranked positives. Proceedings of the Proceedings of the AAAI Conference on Artificial Intelligence 2022, 36, 897–905. [Google Scholar] [CrossRef]
- Xu, L.; Xie, H.; Li, Z.; Wang, F.L.; Wang, W.; Li, Q. Contrastive learning models for sentence representations. ACM Transactions on Intelligent Systems and Technology 2023, 14, 1–34. [Google Scholar] [CrossRef]
- Zheng, M.; Wang, F.; You, S.; Qian, C.; Zhang, C.; Wang, X.; Xu, C. Weakly supervised contrastive learning. In Proceedings of the Proceedings of the IEEE/CVF International Conference on Computer Vision, 2021; pp. 10042–10051. [Google Scholar]
- Robinson, J.; Chuang, C.Y.; Sra, S.; Jegelka, S. Contrastive learning with hard negative samples. arXiv 2020, arXiv:2010.04592. [Google Scholar]
- Liu, A.H.; Chang, H.J.; Auli, M.; Hsu, W.N.; Glass, J. Dinosr: Self-distillation and online clustering for self-supervised speech representation learning. Advances in Neural Information Processing Systems 2024, 36. [Google Scholar]
- Caron, M.; Touvron, H.; Misra, I.; Jégou, H.; Mairal, J.; Bojanowski, P.; Joulin, A. Emerging properties in self-supervised vision transformers. In Proceedings of the Proceedings of the IEEE/CVF international conference on computer vision, 2021; pp. 9650–9660. [Google Scholar]
- Chen, X.; Fan, H.; Girshick, R.; He, K. Improved baselines with momentum contrastive learning. arXiv 2020, arXiv:2003.04297. [Google Scholar] [CrossRef]
- Chen, T.; Kornblith, S.; Norouzi, M.; Hinton, G. A simple framework for contrastive learning of visual representations. In Proceedings of the International conference on machine learning. PMLR, 2020; pp. 1597–1607. [Google Scholar]
- Azizi, S.; Mustafa, B.; Ryan, F.; Beaver, Z.; Freyberg, J.; Deaton, J.; Loh, A.; Karthikesalingam, A.; Kornblith, S.; Chen, T.; et al. Big self-supervised models advance medical image classification. In Proceedings of the Proceedings of the IEEE/CVF international conference on computer vision, 2021; pp. 3478–3488. [Google Scholar]
- Chaitanya, K.; Erdil, E.; Karani, N.; Konukoglu, E. Contrastive learning of global and local features for medical image segmentation with limited annotations. Advances in neural information processing systems 2020, 33, 12546–12558. [Google Scholar]
- Ciga, O.; Xu, T.; Martel, A.L. Self supervised contrastive learning for digital histopathology. Machine Learning with Applications 2022, 7, 100198. [Google Scholar] [CrossRef]
- Guo, Z.; Zhang, Y.; Qiu, Z.; Dong, S.; He, S.; Gao, H.; Zhang, J.; Chen, Y.; He, B.; Kong, Z.; et al. An improved contrastive learning network for semi-supervised multi-structure segmentation in echocardiography. Frontiers in cardiovascular medicine 2023, 10, 1266260. [Google Scholar] [CrossRef] [PubMed]
- Luo, G.; Xie, W.; Gao, R.; Zheng, T.; Chen, L.; Sun, H. Unsupervised anomaly detection in brain MRI: Learning abstract distribution from massive healthy brains. Computers in biology and medicine 2023, 154, 106610. [Google Scholar] [CrossRef] [PubMed]
- Krishnan, R.; Rajpurkar, P.; Topol, E.J. Self-supervised learning in medicine and healthcare. Nature Biomedical Engineering 2022, 6, 1346–1352. [Google Scholar] [CrossRef] [PubMed]
- Pick, F.; Xie, X.; Wu, L.Y. Contrastive Multitask Transformer for Hospital Mortality and Length-of-Stay Prediction. In Proceedings of the International Conference on AI in Healthcare; Springer, 2024; pp. 134–145. [Google Scholar]
- Sun, M.; Yang, X.; Niu, J.; Gu, Y.; Wang, C.; Zhang, W. A cross-modal clinical prediction system for intensive care unit patient outcome. Knowledge-Based Systems 2024, 283, 111160. [Google Scholar] [CrossRef]
- Cai, T.; Huang, F.; Nakada, R.; Zhang, L.; Zhou, D. Contrastive Learning on Multimodal Analysis of Electronic Health Records. arXiv 2024, arXiv:2403.14926. [Google Scholar] [CrossRef]
- Zhong, X.; Batmanghelich, K.; Sun, L. Enhancing Biomedical Multi-modal Representation Learning with Multi-scale Pre-training and Perturbed Report Discrimination. In Proceedings of the 2024 IEEE Conference on Artificial Intelligence (CAI); IEEE, 2024; pp. 480–485. [Google Scholar]
- Bepler, T.; Berger, B. Learning the protein language: Evolution, structure, and function. Cell systems 2021, 12, 654–669. [Google Scholar] [CrossRef]
- Liu, X.; Xu, X.; Xu, X.; Li, X.; Xie, G. Representation Learning for Multi-omics Data with Heterogeneous Gene Regulatory Network. In Proceedings of the 2021 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 2021; pp. 702–705. [Google Scholar] [CrossRef]
- Li, S.; Ma, J.; Zhao, T.; Jia, Y.; Liu, B.; Luo, R.; Huang, Y. CellContrast: Reconstructing spatial relationships in single-cell RNA sequencing data via deep contrastive learning. Patterns 2024, 5. [Google Scholar] [CrossRef]
- Zhang, R.; Wu, H.; Liu, C.; Li, H.; Wu, Y.; Li, K.; Wang, Y.; Deng, Y.; Chen, J.; Zhou, F.; et al. Pepharmony: A multi-view contrastive learning framework for integrated sequence and structure-based peptide encoding. arXiv 2024, arXiv:2401.11360. [Google Scholar] [CrossRef]
- Zhang, Y.; Jiang, H.; Miura, Y.; Manning, C.D.; Langlotz, C.P. Contrastive learning of medical visual representations from paired images and text. In Proceedings of the Machine learning for healthcare conference. PMLR, 2022; pp. 2–25. [Google Scholar]
- Huang, S.C.; Shen, L.; Lungren, M.P.; Yeung, S. Gloria: A multimodal global-local representation learning framework for label-efficient medical image recognition. In Proceedings of the Proceedings of the IEEE/CVF international conference on computer vision, 2021; pp. 3942–3951. [Google Scholar]
- Boecking, B.; Usuyama, N.; Bannur, S.; Castro, D.C.; Schwaighofer, A.; Hyland, S.; Wetscherek, M.; Naumann, T.; Nori, A.; Alvarez-Valle, J.; et al. Making the most of text semantics to improve biomedical vision–language processing. In Proceedings of the European conference on computer vision; Springer, 2022; pp. 1–21. [Google Scholar]
- Bannur, S.; Hyland, S.; Liu, Q.; Perez-Garcia, F.; Ilse, M.; Castro, D.C.; Boecking, B.; Sharma, H.; Bouzid, K.; Thieme, A.; et al. Learning to exploit temporal structure for biomedical vision-language processing. In Proceedings of the Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2023; pp. 15016–15027. [Google Scholar]
- You, K.; Gu, J.; Ham, J.; Park, B.; Kim, J.; Hong, E.K.; Baek, W.; Roh, B. Cxr-clip: Toward large scale chest x-ray language-image pre-training. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer, 2023; pp. 101–111. [Google Scholar]
- Tiu, E.; Talius, E.; Patel, P.; Langlotz, C.P.; Ng, A.Y.; Rajpurkar, P. Expert-level detection of pathologies from unannotated chest X-ray images via self-supervised learning. Nature biomedical engineering 2022, 6, 1399–1406. [Google Scholar] [CrossRef]
- Wang, Z.; Wu, Z.; Agarwal, D.; Sun, J. Medclip: Contrastive learning from unpaired medical images and text. In Proceedings of the Proceedings of the Conference on Empirical Methods in Natural Language Processing. Conference on Empirical Methods in Natural Language Processing, 2022, Vol. 2022, p. 3876.
- Huang, Z.; Bianchi, F.; Yuksekgonul, M.; Montine, T.J.; Zou, J. A visual–language foundation model for pathology image analysis using medical twitter. Nature medicine 2023, 29, 2307–2316. [Google Scholar] [CrossRef]
- Lu, M.Y.; Chen, B.; Williamson, D.F.; Chen, R.J.; Liang, I.; Ding, T.; Jaume, G.; Odintsov, I.; Le, L.P.; Gerber, G.; et al. A visual-language foundation model for computational pathology. Nature medicine 2024, 30, 863–874. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Xu, Y.; Usuyama, N.; Xu, H.; Bagga, J.; Tinn, R.; Preston, S.; Rao, R.; Wei, M.; Valluri, N.; et al. Biomedclip: a multimodal biomedical foundation model pretrained from fifteen million scientific image-text pairs. arXiv 2023, arXiv:2303.00915. [Google Scholar]
- Lin, W.; Zhao, Z.; Zhang, X.; Wu, C.; Zhang, Y.; Wang, Y.; Xie, W. Pmc-clip: Contrastive language-image pre-training using biomedical documents. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer, 2023; pp. 525–536. [Google Scholar]
- Huang, W.; Li, C.; Zhou, H.Y.; Yang, H.; Liu, J.; Liang, Y.; Zheng, H.; Zhang, S.; Wang, S. Enhancing representation in radiography-reports foundation model: A granular alignment algorithm using masked contrastive learning. Nature Communications 2024, 15, 7620. [Google Scholar] [CrossRef] [PubMed]
- Koleilat, T.; Asgariandehkordi, H.; Rivaz, H.; Xiao, Y. Medclip-sam: Bridging text and image towards universal medical image segmentation. In Proceedings of the International conference on medical image computing and computer-assisted intervention; Springer, 2024; pp. 643–653. [Google Scholar]
- Liu, Z.; Alavi, A.; Li, M.; Zhang, X. Self-supervised contrastive learning for medical time series: A systematic review. Sensors 2023, 23, 4221. [Google Scholar] [CrossRef] [PubMed]
- Diamant, N.; Reinertsen, E.; Song, S.; Aguirre, A.D.; Stultz, C.M.; Batra, P. Patient contrastive learning: A performant, expressive, and practical approach to electrocardiogram modeling. PLoS computational biology 2022, 18, e1009862. [Google Scholar] [CrossRef]
- Yuan, Y.; Van Duyn, J.; Yan, R.; Huang, Z.; Vesal, S.; Plis, S.; Hu, X.; Kwak, G.H.; Xiao, R.; Fedorov, A. Learning ECG Representations via Poly-Window Contrastive Learning. arXiv 2025, arXiv:2508.15225. [Google Scholar] [CrossRef]
- Wang, Y.; Han, Y.; Wang, H.; Zhang, X. Contrast everything: A hierarchical contrastive framework for medical time-series. Advances in Neural Information Processing Systems 2023, 36, 55694–55717. [Google Scholar]
- Chen, W.; Wang, H.; Zhang, L.; Zhang, M. Temporal and spatial self supervised learning methods for electrocardiograms. Scientific Reports 2025, 15, 6029. [Google Scholar] [CrossRef]
- Raghu, A.; Chandak, P.; Alam, R.; Guttag, J.; Stultz, C. Contrastive pre-training for multimodal medical time series. In Proceedings of the NeurIPS 2022 Workshop on Learning from Time Series for Health, 2022. [Google Scholar]
- Guo, H.; Xu, X.; Wu, H.; Liu, B.; Xia, J.; Cheng, Y.; Guo, Q.; Chen, Y.; Xu, T.; Wang, J.; et al. Multi-scale and multi-modal contrastive learning network for biomedical time series. Biomedical Signal Processing and Control 2025, 106, 107697. [Google Scholar] [CrossRef]
- Sun, X.; Yang, Y.; Dong, X. Enhancing Contrastive Learning-based Electrocardiogram Pretrained Model with Patient Memory Queue. arXiv 2025, arXiv:2506.06310. [Google Scholar]
| Application Domain | Author(s) | Year | Method | Application |
| Medical Imaging | Azizi et al. [36] | 2021 | SimCLR-based pretraining on unlabeled chest X-rays | Learned transferable visual representations for medical image classification and segmentation using unlabeled data. |
| Chaitanya et al. [37] | 2020 | Semi-supervised contrastive framework for MRI | Improved segmentation and classification in MRI with limited labels using augmented positive and negative pairs. | |
| Ciga et al. [38] | 2022 | Self-supervised contrastive learning for histopathology | Enhanced cancer detection and tissue differentiation in biopsy samples using augmentation-invariant representations. | |
| Guo et al. [39] | 2023 | Multi-scale contrastive loss for cardiac MRI segmentation | Captured both global and local structures, improving accuracy of myocardium and ventricle segmentation. | |
| Luo et al. [40] | 2023 | Self-supervised anomaly detection using contrastive loss | Detected abnormal regions in brain MRI scans by distinguishing normal and pathological patches. | |
| Electronic Health Records | Krishnan et al. [41] | 2022 | Self-supervised contrastive learning on augmented EHR views | Modeled temporal and clinical correlations for mortality and heart failure prediction. |
| Pick et al. [42] | 2024 | Contrastive patient representation learning | Improved prediction of hospital mortality and length-of-stay through patient-level embeddings. | |
| Sun et al. [43] | 2024 | Cross-modal contrastive framework for EHR integration | Aligned structured and unstructured EHR data to predict disease progression and complications. | |
| Cai et al. [44] | 2024 | Distributed large-scale contrastive learning | Scalable training on large EHR datasets for improved generalization across patient populations. | |
| Genomics and Proteomics | Zhong et al. [45] | 2024 | Multi-scale contrastive learning (MSCL) for genomics | Identified disease-associated genetic markers by modeling gene and pathway-level interactions. |
| Bepler and Berger [46] | 2021 | Contrastive protein sequence representation learning | Learned structural and functional protein embeddings for improved function prediction and drug discovery. | |
| Liu et al. [47] | 2022 | Multi-omics contrastive learning (MoHeG / GenCL) | Integrated genomics, transcriptomics, and epigenomics to predict disease susceptibility and treatment outcomes. | |
| Li et al. [48] | 2024 | CellContrast for single-cell RNA sequencing | Enhanced clustering and identification of rare cell types in scRNA-seq data. | |
| Zhang et al. [49] | 2024 | Pepharmony: sequence–structure contrastive learning | Predicted protein–protein interactions with improved accuracy using multimodal peptide representations. | |
| Multimodal and Cross-Domain Learning | Zhang et al. [50] | 2022 | ConVIRT (image–text alignment) | Learned chest X-ray representations by aligning images and radiology reports with bidirectional contrastive loss. |
| Huang et al. [51] | 2021 | GLoRIA (global–local image–text alignment) | Improved retrieval and classification on MIMIC-CXR through local region–phrase alignment. | |
| Boecking et al. [52] | 2022 | BioViL (biomedical vision–language model) | Enhanced zero-shot radiology performance using domain-specific text pretraining. | |
| Bannur et al. [53] | 2023 | BioViL-T (temporal alignment) | Improved disease progression tracking in chest X-rays via temporal contrastive learning. | |
| You et al. [54] | 2023 | CXR-CLIP (prompt-based multimodal CL) | Combined image–label and image–text supervision for robust chest X-ray recognition. | |
| Tiu et al. [55] | 2022 | CheXzero (CLIP-style vision–language model) | Achieved radiologist-level zero-shot classification on the CheXpert benchmark. | |
| Wang et al. [56] | 2022 | MedCLIP (knowledge-aware matching loss) | Reduced false negatives in radiology by decoupling image–text corpora for efficient pretraining. | |
| Huang et al. [57] | 2023 | PLIP (pathology vision–language foundation model) | Achieved state-of-the-art performance in pathology classification and zero-shot transfer. | |
| Lu et al. [58] | 2024 | CONCH (large-scale histopathology pretraining) | Trained on 1.17M image–caption pairs for generalizable pathology retrieval and segmentation. | |
| Zhang et al. [59] | 2023 | BiomedCLIP (PubMed multimodal foundation model) | Pretrained on 15M image–text pairs for broad biomedical zero/few-shot applications. | |
| Lin et al. [60] | 2023 | PMC-CLIP (literature-derived pretraining) | Improved biomedical VQA and retrieval from 1.6M figure–caption pairs. | |
| Huang et al. [61] | 2024 | MaCo (masked contrastive learning) | Applied to chest X-rays for enhanced zero-shot and localized recognition. | |
| Koleilat et al. [62] | 2024 | MedCLIP + SAM (text-driven segmentation) | Enabled multimodal segmentation across ultrasound, MRI, and CT without explicit labels. | |
| Time-Series and Physiological Signals | Liu et al. [63] | 2023 | Systematic review of contrastive time-series methods | Identified key design trends in self-supervised ECG/EEG contrastive learning. |
| Diamant et al. [64] | 2022 | PCLR (patient-level contrastive learning) | Leveraged same-patient ECGs to improve cardiac disease prediction tasks. | |
| Yuan et al. [65] | 2025 | Poly-window contrastive learning | Modeled temporal overlap in ECGs to enhance representation efficiency. | |
| Wang et al. [66] | 2023 | COMET (hierarchical contrastive framework) | Applied multi-level contrastive learning for ECG and EEG classification with few labels. | |
| Chen et al. [67] | 2025 | CLOCS (spatiotemporal contrastive model) | Improved robustness in cardiac signals under lead and time variation. | |
| Raghu et al. [68] | 2022 | Multimodal temporal contrastive pretraining | Integrated physiological signals with lab and vitals data for outcome prediction. | |
| Guo et al. [69] | 2025 | MBSL (multi-scale multimodal contrastive learning) | Combined respiration, heart rate, and motion signals for multi-task biomedical inference. | |
| Sun et al. [70] | 2025 | PMQ (patient memory queue) | Mitigated false negatives in ECG pretraining by leveraging intra-patient memory banks. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




