Submitted:
16 December 2025
Posted:
17 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Traditional Approaches
1.2. Machine Learning Methods
1.3. Deep Learning Advances
1.4. Transformer-Based Models
1.5. Research Gap and Motivation
1.6. Contributions of This Study
- (a)
- We introduce a novel hybrid architecture that effectively combines the global contextual modeling capabilities of DNABERT-2 with the local motif detection strengths of convolutional networks, enabling comprehensive analysis of both long-range dependencies and precise local patterns in promoter sequences.
- (b)
- Our model achieves state-of-the-art performance with a ROC AUC of 97.28% and accuracy of 93.10% in 5-fold cross-validation, significantly outperforming existing methods, while maintaining robust generalization on independent test sets.
- (c)
- We develop an innovative feature fusion strategy that integrates global sequence embeddings with locally refined features, creating a biologically grounded representation that effectively handles both typical 70 promoters and challenging non-typical promoters for practical applications in synthetic biology.
2. Literature Review
2.1. Early Rule-Based and Consensus-Driven Methods
2.2. Feature-Engineered Machine Learning Approaches
2.3. Deep Learning for Automated Feature Learning
2.4. The Rise of Pre-trained Transformer Models in Genomics
2.5. Hybrid Architectures and Remaining Challenges
3. Materials and Methods
3.1. Dataset Construction
3.1.1. Data Sources and Processing
3.2. Sequence Encoding
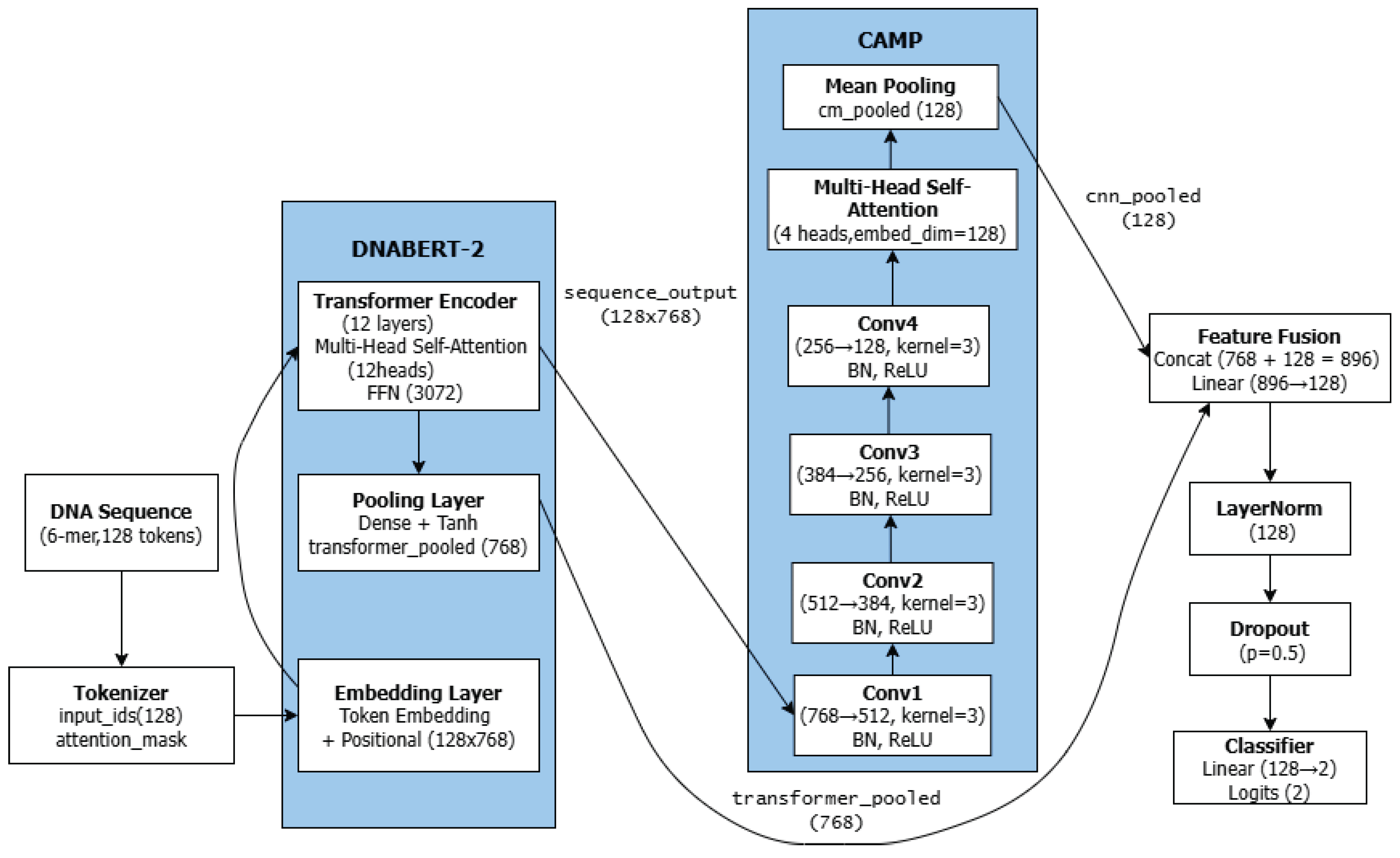
3.3. Model Architecture
3.4. DNABERT-2 Feature Extractor
- A sequence-level pooled embedding (transformer_pooled, 768-dimensional) that encapsulates global sequence semantics.
- Token-wise contextual embeddings (sequence_output, dimensions: [batch_size, 128, 768]), which retain positional information, are subsequently passed to the CNN module for local feature refinement.
3.5. CAMP Module: CNN, Multi-Head Attention, and Mean Pooling
3.5.1. Convolutional Neural Network
- Conv1: 768 → 512 channels, kernel size = 3, padding = "same"
- Conv2: 512 → 384 channels, kernel size = 3, padding = "same"
- Conv3: 384 → 256 channels, kernel size = 3, padding = "same"
- Conv4: 256 → 128 channels, kernel size = 3, padding = "same"
3.5.2. Multi-Head Self-Attention
3.5.3. Mean Pooling
3.6. Output Layer
3.7. Model Training and Evaluation
3.7.1. Training Strategy
3.7.2. Hyperparameter Optimization
3.8. Statistical Analysis
3.9. Code Availability
4. Results and Discussion
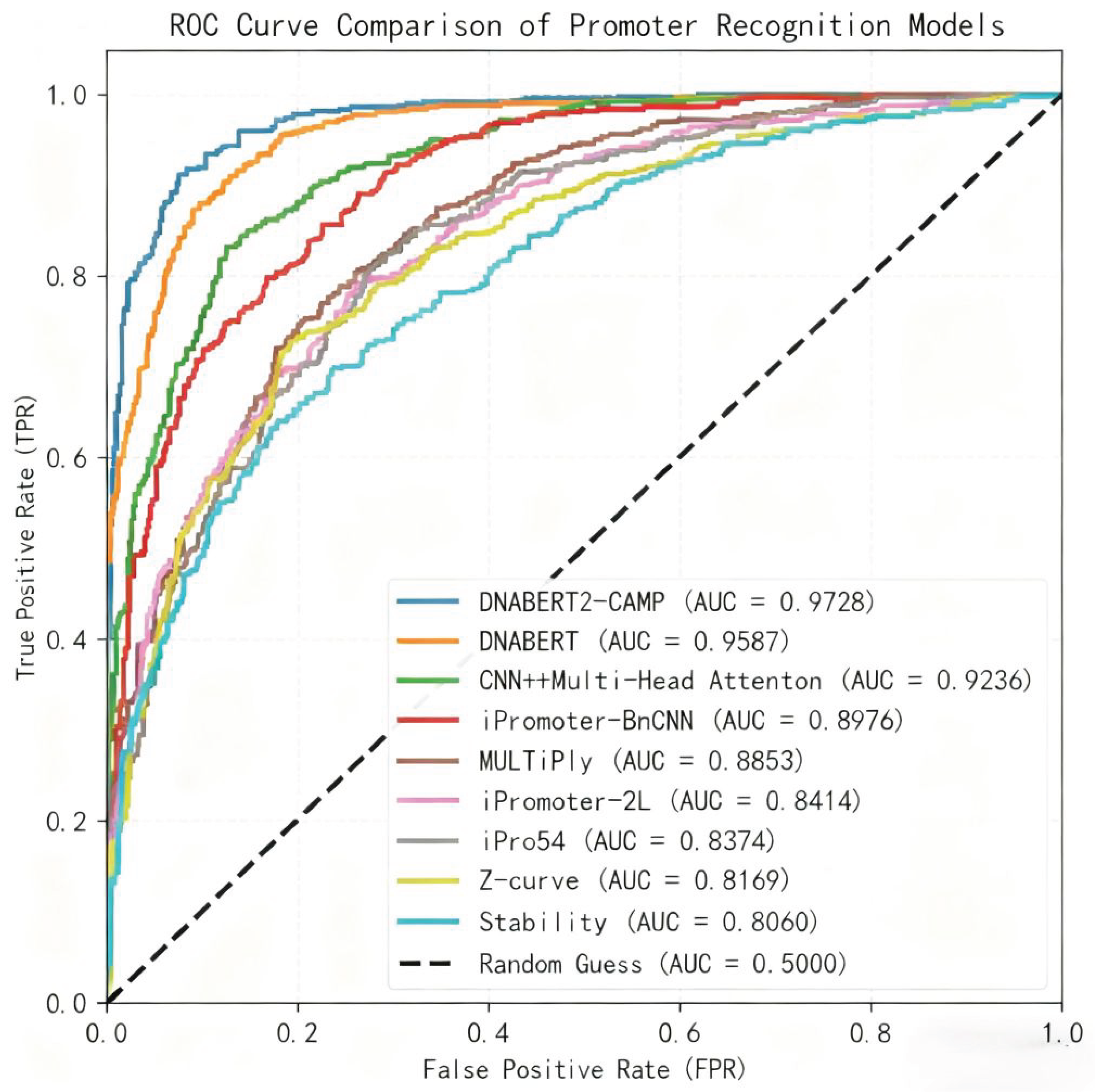
4.1. Model Performance and Comparative Analysis
4.2. Statistical Validation of Performance Improvements
4.3. Impact of Optimization and Comparative Analysis with Existing Methods
4.4. Analysis of Model Decisions and Biological Basis
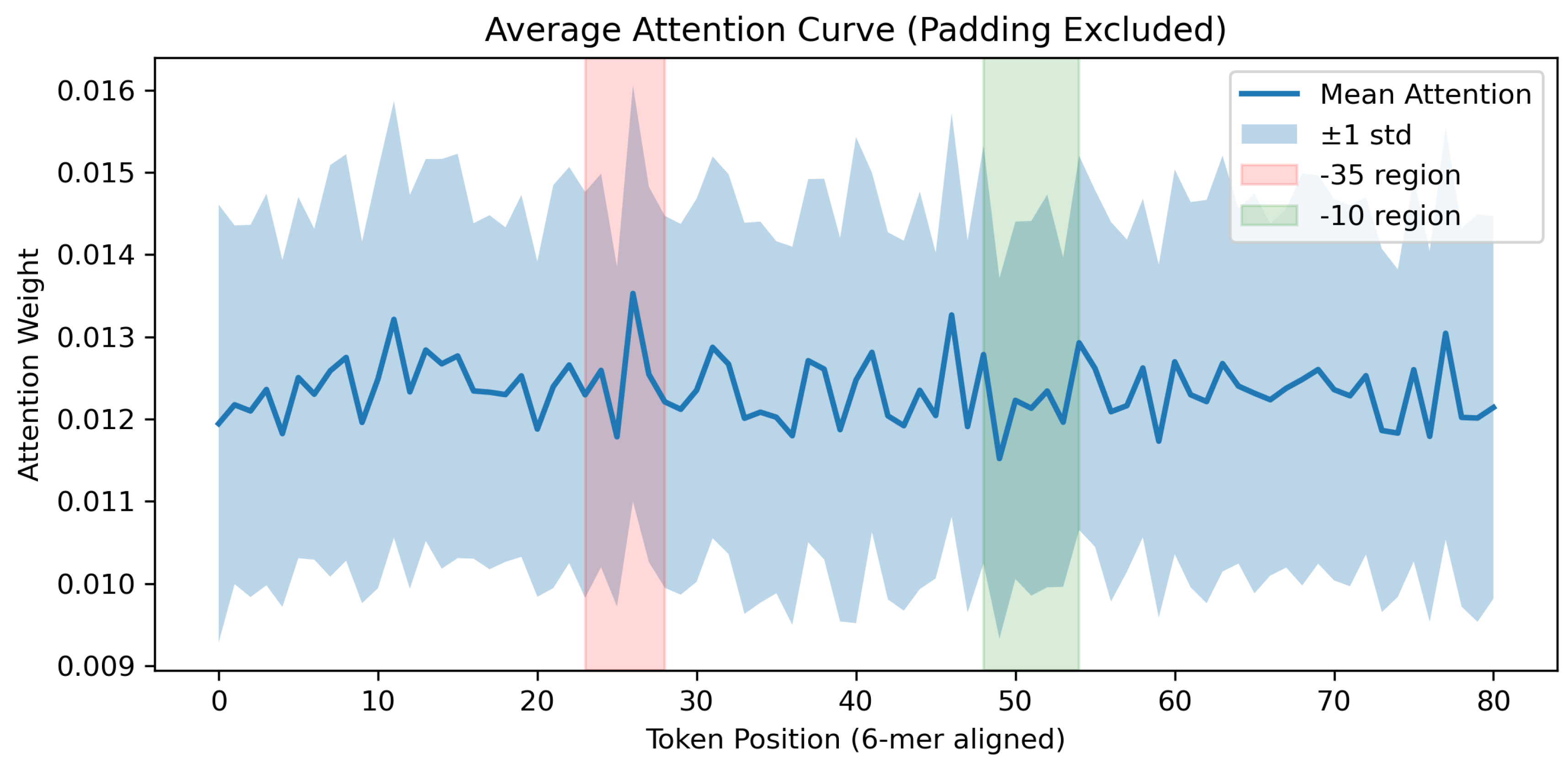
4.4.1. Attention-based Interpretability Analysis
4.4.2. CNN Motif Activation and Peak Analysis
4.5. Computational Considerations, Limitations, and Future Directions
- (a)
- Cross-species application: Leveraging the "species-agnostic" 6-mer tokenization and pre-trained embeddings to extend the model’s application to cross-species promoter prediction.
- (b)
- Enhanced interpretability: Utilizing techniques such as SHAP value analysis and saliency maps to uncover novel biological insights into promoter architecture[37].
- (c)
- Addressing data scarcity: Exploring data augmentation, transfer learning, and few-shot learning strategies for rare promoter classes.
- (d)
- Model efficiency: Investigating model compression techniques to facilitate deployment in high-throughput synthetic biology pipelines.
- (e)
- Multi-task learning: Developing frameworks to simultaneously predict promoter strength, transcription factor binding sites, and other regulatory elements for a more comprehensive view of gene regulation [38].
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Purnick, P.E.M.; Weiss, R. The second wave of synthetic biology: from modules to systems. Nature Reviews Molecular Cell Biology 2009, 10, 410–422. [Google Scholar] [CrossRef] [PubMed]
- Gruber, T.M.; Gross, C.A. The sigma70 promoter recognition network of Escherichia coli. Nucleic Acids Research 2014, 42, 7470–7485. [Google Scholar]
- Harley, C.B.; Reynolds, R.P. Sequence analysis of E. coli promoter DNA. Nucleic Acids Research 1987, 15, 2343–2361. [Google Scholar] [CrossRef]
- Faith, J.J.; Hayete, B.; Thaden, J.T.; Mogno, I.; Wierzbowski, J.; Cottarel, G.; Kasif, S.; Collins, J.J.; Gardner, T.S. Large-scale mapping and validation of Escherichia coli transcriptional regulation from a compendium of expression profiles. PLoS Biology 2007, 5, e8. [Google Scholar] [CrossRef]
- Alper, H.; Fischer, C.; Nevoigt, E.; Stephanopoulos, G. Tuning genetic control through promoter engineering. Proc. Natl. Acad. Sci. USA 2005, 102, 12678–12683. [Google Scholar] [CrossRef]
- Cox, R.S., III; Surette, M.G.; Elowitz, M.B. Programming gene expression with combinatorial promoters. Molecular Systems Biology 2007, 3, 145. [Google Scholar] [CrossRef]
- Towsey, M.; Timms, P.; Hogan, J.; et al. Promoter classification using DNA structural profiles. Bioinformatics 2008, 24, 23–29. [Google Scholar]
- Durbin, R.; Eddy, S.R.; Krogh, A.; Mitchison, G. Biological sequence analysis: probabilistic models of proteins and nucleic acids; Cambridge University Press, 1998. [Google Scholar]
- Umarov, R.; Solovyev, V. Promoter recognition using convolutional deep learning neural networks. PLOS ONE 2017, 12, e0171410. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Troyanskaya, O.G. Predicting effects of noncoding variants with deep learning-based sequence model. Nature Methods 2015, 12, 931–934. [Google Scholar] [CrossRef]
- Pascanu, R.; Mikolov, T.; Bengio, Y. On the Difficulty of Training Recurrent Neural Networks. Int. Conf. Mach. Learn. 2013, 28, 1310–1318. [Google Scholar]
- Ji, Y.; Zhou, Z.; Liu, H.; Davuluri, R.V. DNABERT: pre-trained Bidirectional Encoder Representations from Transformers model for DNA-language in genome. Bioinformatics 2021, 37, 2112–2120. [Google Scholar] [CrossRef] [PubMed]
- Hawley, D.K.; McClure, W.R. Compilation and Analysis of Escherichia coli Promoter Sequences. Nucleic Acids Res. 1983, 11, 2237–2240. [Google Scholar] [CrossRef]
- Abbas, M.M.; Mohie-Eldin, M.M.; El-Manzalawy, Y. Assessing the Effects of Data Selection and Representation on the Development of Reliable E. coli Sigma-70 Promoter Region Predictors. PLOS ONE 2015, 10, e0119721. [Google Scholar] [CrossRef]
- Lin, H.; Deng, E.Z.; Ding, H.; Chen, W.; Chou, K.C. iPro54-PseKNC: A Sequence-Based Predictor for Identifying Sigma-54 Promoters in Prokaryote with Pseudo k-Tuple Nucleotide Composition. Nucleic Acids Res. 2014, 42, 12961–12972. [Google Scholar] [CrossRef]
- Amin, R.; Rahman, C.R.; Ahmed, S.; et al. iPromoter-BnCNN: A Novel Branched CNN-Based Predictor for Identifying and Classifying Sigma Promoters. Bioinformatics 2020, 36, 4869–4875. [Google Scholar] [CrossRef]
- Shujaat, M.; Wahab, A.; Tayara, H.; et al. pcPromoter-CNN: A CNN-Based Prediction and Classification of Promoters. Genes 2020, 11, 1529. [Google Scholar] [CrossRef]
- Li, F.; Chen, J.; Ge, Z.; et al. Computational Prediction and Interpretation of Both General and Specific Types of Promoters in Escherichia coli by Exploiting a Stacked Ensemble-Learning Framework. Brief. Bioinform. 2021, 22, 2126–2140. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Yuan, Y.; Li, Y. Recognition of Escherichia coli Promoters Based on Attention Mechanisms. In Proceedings of the Proc. 7th Int. Conf. Comput. Biol. Bioinform., Kuala Lumpur, Malaysia, 12 2023; pp. 6–15. [Google Scholar] [CrossRef]
- Gama-Castro, S.; Salgado, H.; Santos-Zavaleta, A.; et al. RegulonDB Version 10.5: Tackling Challenges to Unify Classic and High Throughput Knowledge of Gene Regulation in E. coli K-12. Nucleic Acids Res. 2020, 48, D212–D220. [Google Scholar]
- Wheeler, D.L.; Barrett, T.; Benson, D.A.; Bryant, S.H.; Canese, K.; Chetvernin, V.; Church, D.M.; DiCuccio, M.; Edgar, R.; Federhen, S.; et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Research 2007, 35, D5–D12. [Google Scholar] [CrossRef]
- Li, W.; Godzik, A. CD-HIT: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 2006, 22, 1658–1659. [Google Scholar] [CrossRef] [PubMed]
- Kohavi, R. A Study of Cross-Validation and Bootstrap for Accuracy Estimation and Model Selection. In Proceedings of the Proc. 14th Int. Joint Conf. Artif. Intell., 1995; Vol. 2, pp. 1137–1145. [Google Scholar]
- Rovira, J.; O’Connor, K.; Bileschi, M.; et al. BERTology meets Biology: Interpreting Attention in Protein Language Models. EMBL Bioarchive 2020. Preprint: MSB-2020-00012. [Google Scholar]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention Is All You Need. In Proceedings of the Advances in Neural Information Processing Systems, 2017; Vol. 30. [Google Scholar]
- Goodfellow, I.; Bengio, Y.; Courville, A. Deep Learning; MIT Press, 2016. [Google Scholar]
- Ba, J.L.; Kiros, J.R.; Hinton, G.E. Layer Normalization. arXiv preprint arXiv:1607.06450 2016. arXiv:1607.06450.
- Srivastava, N.; Hinton, G.; Krizhevsky, A.; Sutskever, I.; Salakhutdinov, R. Dropout: A Simple Way to Prevent Neural Networks from Overfitting. Journal of Machine Learning Research 2014, 15, 1929–1958. [Google Scholar]
- Chen, X.; Zhang, L. Deep Learning for Imbalanced Genomic Data Analysis. Acta Biochim. Biophys. Sin. 2020, 52, 512–520. [Google Scholar]
- Chawla, N.V.; Bowyer, K.W.; Hall, L.O.; Kegelmeyer, W.P. SMOTE: Synthetic Minority Over-sampling Technique. J. Artif. Intell. Res. (JAIR) 2002, 16, 321–357. [Google Scholar] [CrossRef]
- Loshchilov, I.; Hutter, F. Decoupled Weight Decay Regularization. In Proceedings of the Int. Conf. Learn. Represent., 2019. [Google Scholar]
- Prechelt, L. Early Stopping-But When? In Neural Networks: Tricks of the Trade; Springer, 1998; pp. 55–69. [Google Scholar]
- Matthews, B.W. Comparison of the predicted and observed secondary structure of T4 phage lysozyme. Biochimica et Biophysica Acta (BBA) - Protein Structure 1975, 405, 442–451. [Google Scholar] [CrossRef]
- Bradley, A.P. The Use of the Area Under the ROC Curve in the Evaluation of Machine Learning Algorithms. Pattern Recognit. 1997, 30, 1145–1159. [Google Scholar] [CrossRef]
- DeLong, E.R.; DeLong, D.M.; Clarke-Pearson, D.L. Comparing the areas under two or more correlated receiver operating characteristic curves. Biometrics 1988, 44, 837–845. [Google Scholar] [CrossRef]
- Towsey, M.; Hogan, J.M. The Prediction of Bacterial Transcription Start Sites Using SVMs. Int. J. Neural Syst. 2006, 16, 363–370. [Google Scholar] [CrossRef]
- Lundberg, S.M.; Lee, S.I. A Unified Approach to Interpreting Model Predictions. In Proceedings of the NeurIPS, 2017. [Google Scholar]
- Powers, D.M.W. Evaluation: From Precision, Recall and F-Measure to ROC, Informedness, Markedness and Correlation. J. Mach. Learn. Technol. 2020, 2, 37–63. [Google Scholar]
| Data Source | 70 | Non-70 | Negative | Total |
|---|---|---|---|---|
| Literature [8] | 1,500 | - | 2,860 | 4,360 |
| RegulonDB | 1,860 | 1,000 | - | 2,860 |
| NCBI | - | - | 1,500 | 1,500 |
| Total | 3,360 | 1,000 | 4,360 | 8,720 |
| Hyperparameter | Tuning Range | Final Value |
|---|---|---|
| Learning Rate | - | |
| Batch Size (Training) | 4, 8, 16 | 8 |
| Batch Size (Evaluation) | 4, 8, 16 | 4 |
| Gradient Accumulation Steps | 2, 4, 8 | 4 |
| Weight Decay | 0.005 - 0.1 | 0.05 |
| Warmup Steps | 100 - 500 | 300 |
| Learning Rate Scheduler | cosine, linear | cosine |
| Max Gradient Norm | 0.6 - 1.0 | 0.8 |
| Training Epochs | 5 - 20 | 15 |
| Early Stopping Patience | 3 - 15 | 10 |
| Early Stopping Threshold | 0.1 - 0.3 | 0.1 |
| Random Seed | 42, 123 | 42 |
| Evaluation Metric | Value |
|---|---|
| Accuracy | 0.9310 |
| Precision | 0.9439 |
| Recall | 0.9242 |
| F1 Score | 0.9288 |
| ROC AUC | 0.9728 |
| MCC | 0.8604 |
| Specificity | 0.9300 |
| Evaluation Metric | Internal Test Set | Independent Test Set |
|---|---|---|
| Accuracy | 0.9190 | 0.8983 |
| Precision | 0.9253 | 0.9078 |
| Recall | 0.9146 | 0.8903 |
| F1 Score | 0.9162 | 0.8962 |
| ROC AUC | 0.9572 | 0.9279 |
| MCC | 0.8359 | 0.8014 |
| Specificity | 0.9183 | 0.8958 |
| Model | ACC (%) | Sn (%) | Sp (%) | ROC AUC (%) | MCC |
|---|---|---|---|---|---|
| Z-curve [15] | 80.33 | 76.68 | 82.50 | 81.69 | 0.5980 |
| Stability [14] | 77.96 | 75.95 | 78.64 | 80.60 | 0.5613 |
| iPro54 [15] | 80.47 | 76.83 | 82.71 | 83.74 | 0.5983 |
| iPromoter-2L [17] | 81.74 | 79.23 | 83.58 | 84.14 | 0.6327 |
| MULTiPly [18] | 86.93 | 87.26 | 86.20 | 88.53 | 0.7368 |
| iPromoter-BnCNN [16] | 88.16 | 88.73 | 87.24 | 89.76 | 0.7549 |
| CNN + Multi-Head Attention [19] | 90.24 | 90.35 | 90.13 | 92.36 | 0.8016 |
| DNABERT [12] | 90.99 | 88.97 | 89.12 | 95.87 | 0.8255 |
| DNABERT2-CAMP | 93.10 | 92.42 | 93.00 | 97.28 | 0.8604 |
| Dataset / Metric | DNABERT2-CAMP | DNABERT | Test | p-value |
|---|---|---|---|---|
| Cross-validation (5-fold AUC) | 0.9728 | 0.9587 | Wilcoxon signed-rank | 0.0625(two-sided), 0.0313(one-sided) |
| Independent test set(AUC) | 0.9279 | 0.9150 | DeLong test | 0.004 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




