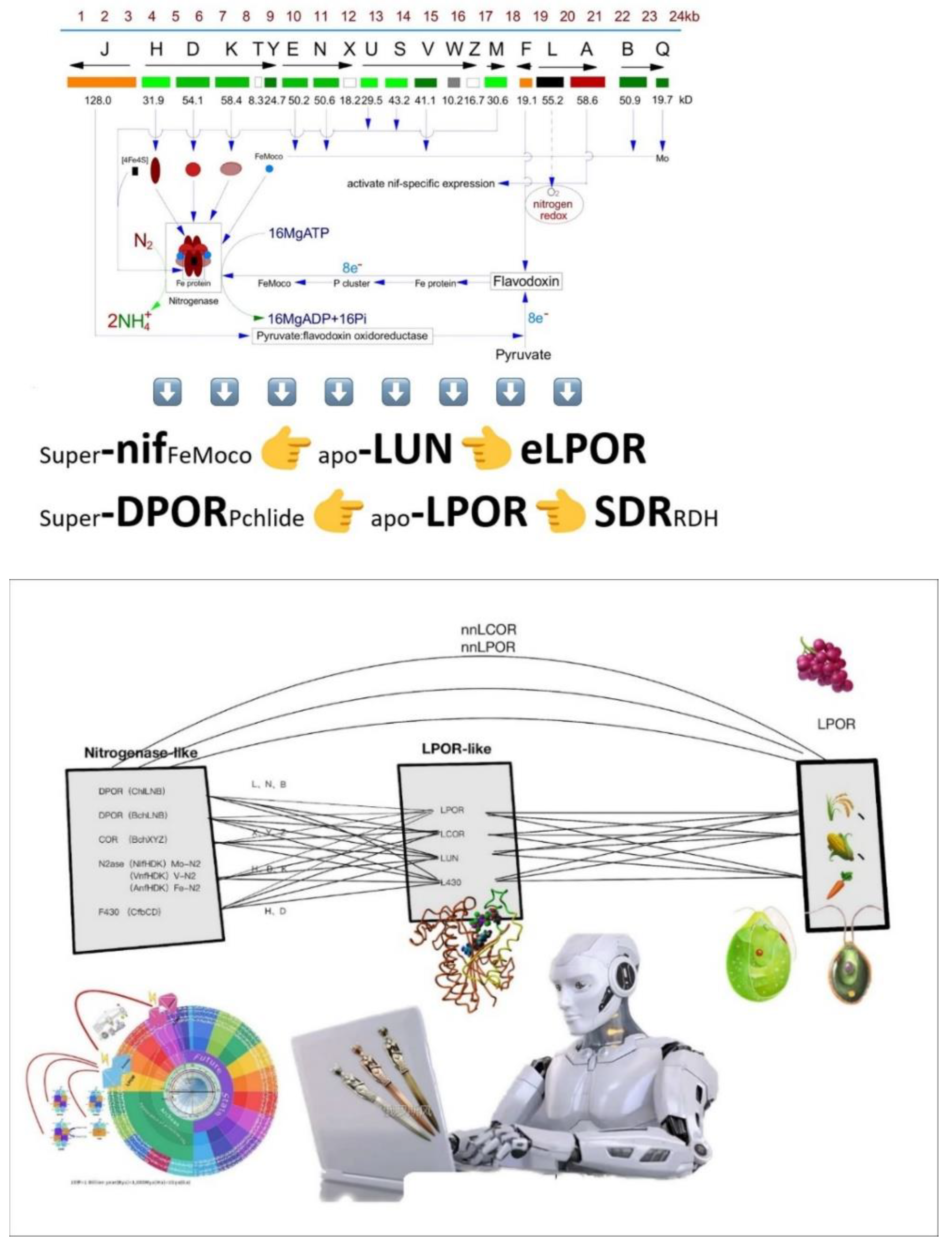
2. Evolutionary relationships among archael Coenzyme F430 reductase, DPOR, COR, nitrogenase and LPOR
- 1)
Common Origin: Evolutionary Basis of a Single Subdomain Precursor
Light-dependent protochlorophyllide oxidoreductase (LPOR), dark-operative protochlorophyllide oxidoreductase (DPOR), chlorophyll oxidoreductase (COR), nitrogenase, and coenzyme F430 reductase (F430 enzyme) exhibit significant structural and functional connections, suggesting they may have originated from a common single subdomain precursor. The F430 enzyme (CfbCD), found only in archaea, is the most ancient nitrogenase-like protein—considered the common ancestor of nitrogenase (NifHDK), DPOR (BchLNB/ChlLNB), and COR (BchXYZ). Structural analysis reveals high similarity between LPOR and the BchL subunit of DPOR (DALI Z-score 5.5), with slightly lower similarity to BchN and BchB (DALI Z-scores of 2.6 and 4.5, respectively), with all values exceeding the structural relatedness threshold (DALI Z-score 2). The Cbc subunit of the F430 enzyme (C-terminal and M domains) shows structural homology with subunits of DPOR, COR, and nitrogenase (DALI Z-scores 3.5–7.1), suggesting that the subunits of all these enzymes likely evolved through gene duplication from a single subdomain precursor.
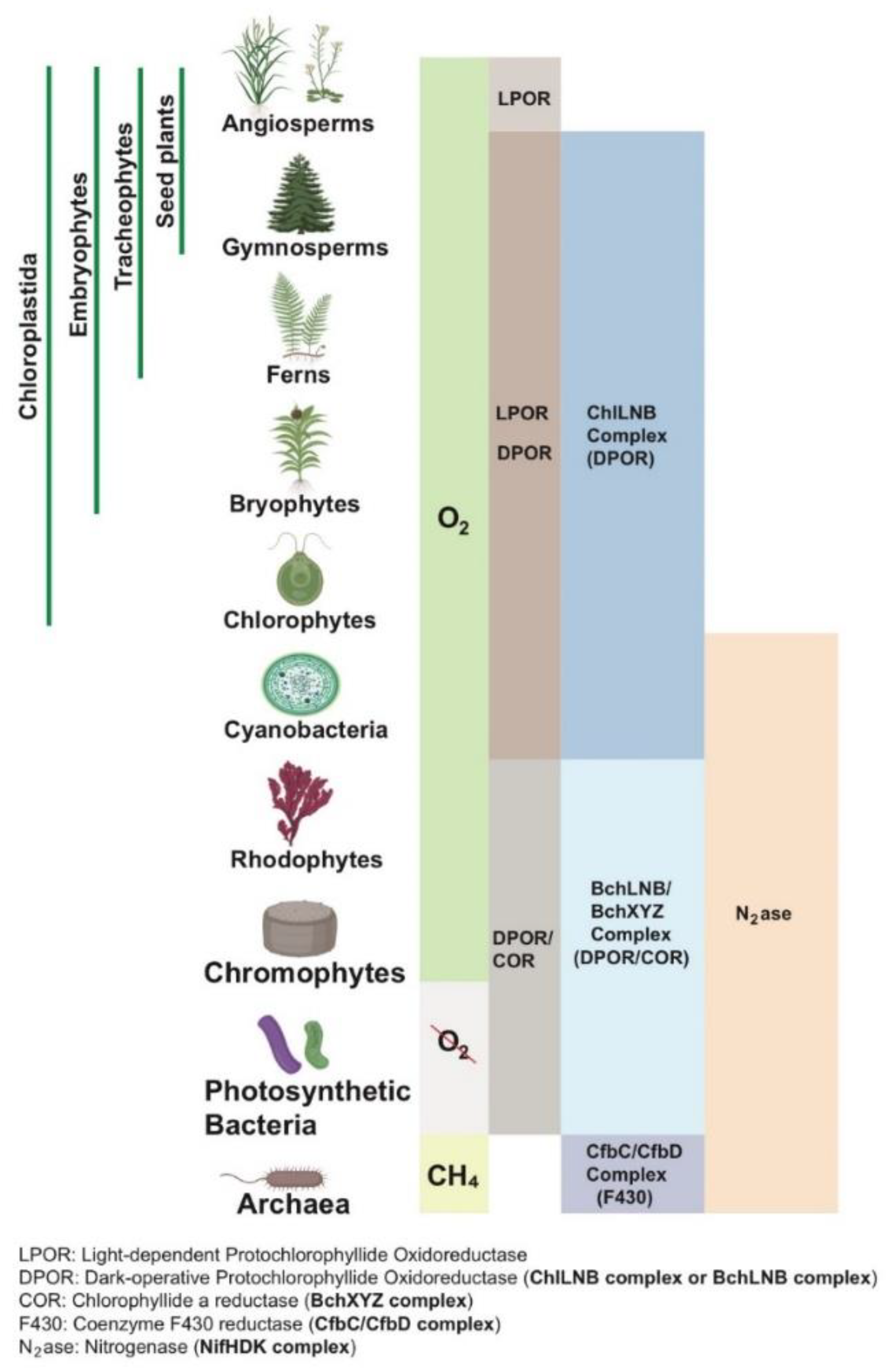
Approximately 71% of LPOR’s structure (92/128 amino acids) closely matches nitrogenase-like proteins, differing primarily at its C-terminus and an additional region (approximately 45 amino acids) absent in nitrogenase-like proteins. This region may represent a unique evolutionary adaptation for LPOR’s photocatalytic function. The hypothesis of a single subdomain precursor as the common origin is supported by principles of structural continuity, similarity scores, amino acid substitution rates, and the presence of the additional region. Species diagrams created using BioRender.com clearly illustrate the distribution and evolutionary sequence of these enzymes across microbial and plant kingdoms (
Figure 4).
- 2)
Evolutionary Pathway of LPOR: From COR via RDH to a Single-Subunit Enzyme
The evolution of LPOR from a multi-subunit nitrogenase-like protein, chlorophyll oxidoreductase (COR), to a single-subunit photoenzyme involved retinol dehydrogenase (RDH) as a critical intermediate. Structural comparisons using FoldSeek against the AlphaFold Swiss-Pro database revealed high similarity between RDH and LPOR, suggesting RDH mediated the evolutionary transition from COR to LPOR (bioRxiv preprint). LPOR likely originated from the BchY subunit of COR through two gene recombination events that integrated RDH structural modules. The N-terminus of BchY shares structural homology with the C-terminus of RDH, while the N-terminus of RDH is similar to the C-terminus of LPOR. This suggests that LPOR simplified the multi-subunit complex through gene fusion.
COR, found in anaerobic photosynthetic bacteria, resembles the Fe protein (NifH) and MoFe protein (NifDK) of nitrogenase. In contrast, LPOR’s single-subunit structure—NADPH-dependent and originating in cyanobacteria—has become ubiquitous across the plant kingdom, including chlorophytes, land plants, vascular plants, and seed plants. This reflects adaptation from anaerobic to aerobic environments.
RDH, which catalyzes retinal metabolism in organisms lacking COR and LPOR, laid the foundation for LPOR’s photocatalytic function. LPOR’s 'additional region' was likely acquired from RDH through gene recombination, facilitating adaptation to oxygen-rich environments and the development of a light-driven mechanism. In cyanobacteria, green algae, mosses, ferns, and gymnosperms, DPOR and LPOR coexist, whereas LPOR dominates in angiosperms, underscoring its critical role in aerobic photosynthesis (
Figure 4a–4c and Supplementary
Figure 2).
- 3)
Structural Conservation and Functional Divergence of Nitrogenase-Like Proteins
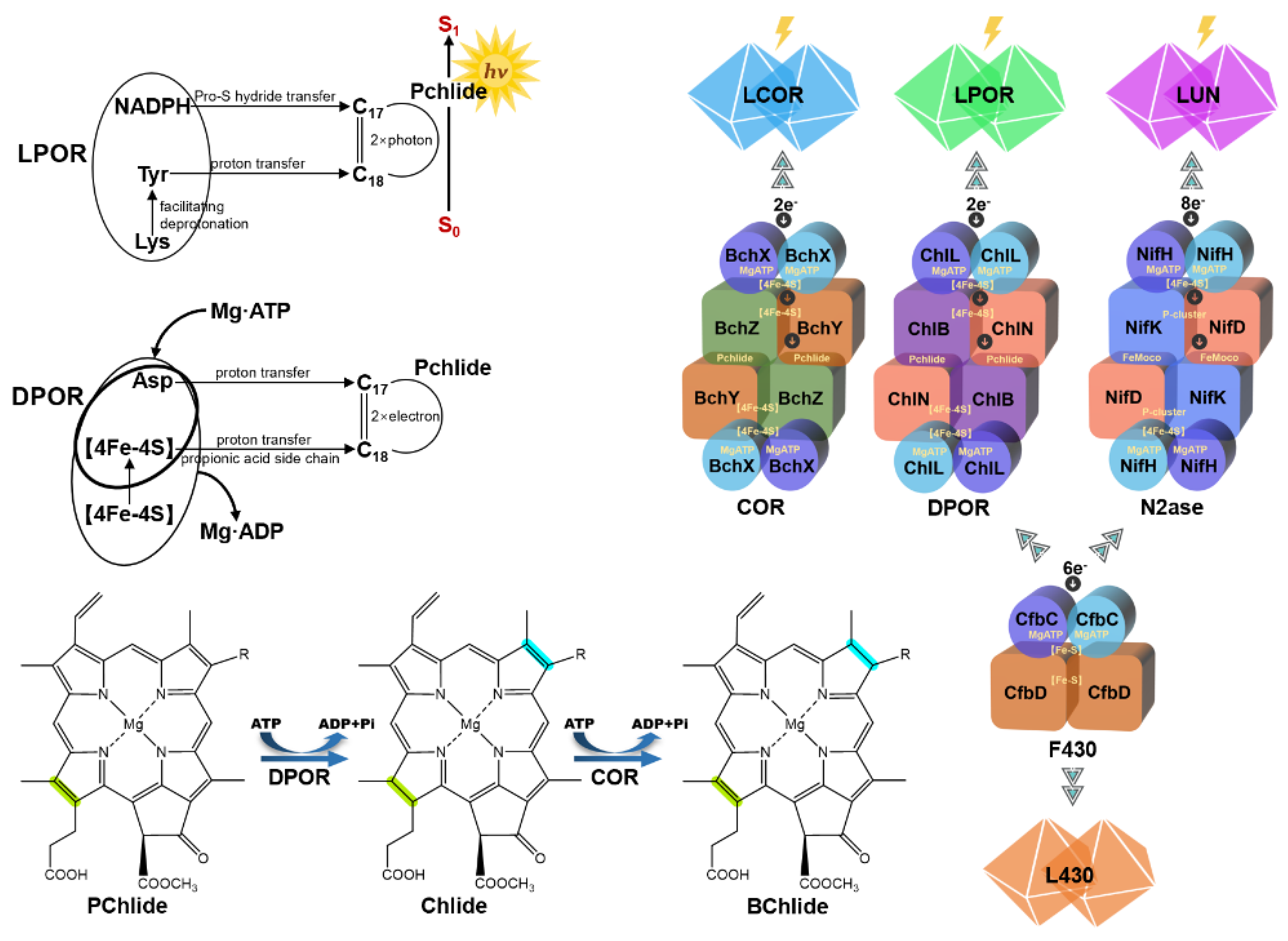
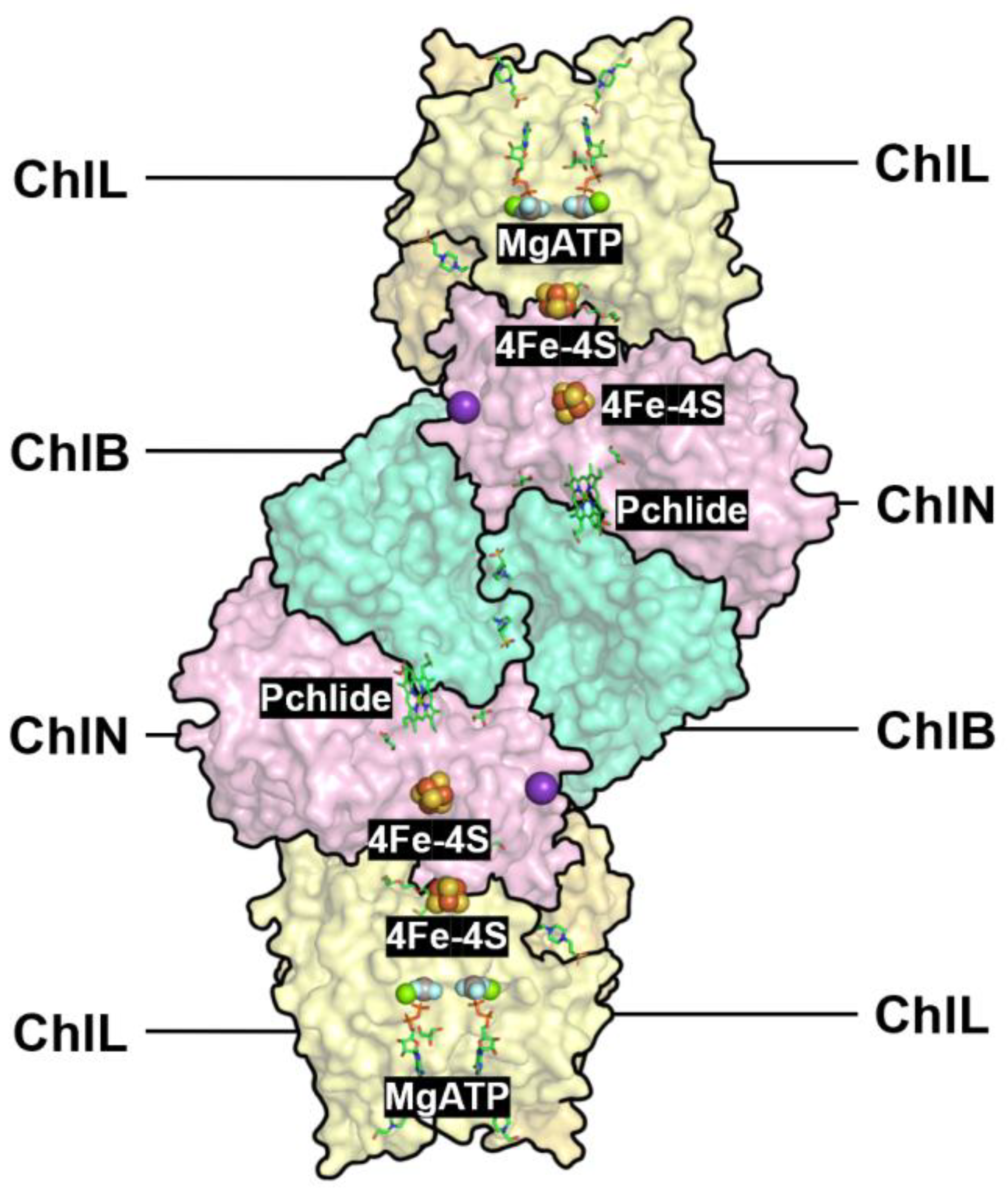
Nitrogenase-like proteins (nitrogenase, DPOR, COR, and F430 enzyme) exhibit high structural conservation but have diverged functionally to meet distinct metabolic demands. Nitrogenase catalyzes the reduction of nitrogen gas to ammonia, DPOR and COR are involved in chlorophyll and bacteriochlorophyll biosynthesis, and the F430 enzyme facilitates coenzyme F430 synthesis for archaeal methane metabolism. Using the iron-based nitrogenase from Rhodobacter capsulatus (PDB ID: 8OEF) as an example, literature shows that its AnfH, AnfD, and AnfK subunits are structurally similar to DPOR’s BchL, BchN, and BchB subunits (DALI Z-scores 2.6–5.5). The Pchlide-binding site in DPOR, spatially overlapping with the FeMo-cofactor site in nitrogenase, indicates that DPOR adapted its structure to accommodate Pchlide, while nitrogenase optimized the FeMo-cofactor for nitrogen reduction. COR (in anaerobic photosynthetic bacteria) and the F430 enzyme (in archaea) evolved specific catalytic capabilities for bacteriochlorophyll precursors and the F430 cofactor, respectively. DPOR is present in both anaerobic and aerobic photosynthetic bacteria, as well as plants (except angiosperms), making it the only nitrogenase-like protein spanning both prokaryotes and eukaryotes.
In contrast, LPOR, a single-subunit enzyme, exhibits a highly evolved catalytic mechanism, with precise structures observed in cyanobacteria and plants (
Figure 4a–4c). The key to functional divergence lies in adaptive changes in substrate-binding sites. Meanwhile, structural conservation reflects the evolutionary trajectory of these enzymes from a single precursor. The Cbc subunit of the F430 enzyme is considered the ancestral template for nitrogenase-like proteins, providing the structural foundation for the diversification of DPOR, COR, and nitrogenase.
- 4)
Evolutionary Environmental Context: Transition from Anaerobic to Aerobic Metabolism
The evolution of nitrogenase-like proteins and LPOR is closely tied to changes in Earth’s atmospheric oxygen levels. In the early anaerobic Earth environment, nitrogenase, DPOR, COR, and the F430 enzyme served as key enzymes for anaerobic metabolism, catalyzing nitrogen fixation, pigment synthesis, and methane metabolism, respectively. The F430 enzyme is exclusive to archaea, while COR is limited to anaerobic photosynthetic bacteria. DPOR is found in both anaerobic and aerobic photosynthetic bacteria and plants (except angiosperms). Nitrogenase is restricted to prokaryotes, reflecting its ancient origin (
Figure 4). With the Great Oxidation Event (GOE), cyanobacteria increased atmospheric oxygen levels through LPOR-mediated oxygenic photosynthesis. LPOR’s oxygen tolerance allowed it to supplant the oxygen-sensitive DPOR, becoming the core enzyme for chlorophyll synthesis in cyanobacteria and plants. This includes chlorophytes, land plants, vascular plants, and seed plants (bioRxiv preprint).
LPOR and DPOR share functional similarity in cyanobacteria, as both catalyze Pchlide reduction to Chlide. However, structural analysis indicates a shared origin. The TFT motif in DPOR is present in some LPORs and is absent in cyanobacterial LPOR, suggesting convergent evolution. Through the inheritance of LPOR and DPOR, the cyanobacterial photosynthetic system evolved into chloroplasts, laying the foundation for eukaryotic photosynthesis.
The evolutionary branch of RDH shifted from energy metabolism in archaea and bacteria to retinal metabolism in animal visual systems. This indicates that LPOR and RDH together drove the evolutionary transition from microbes to macroscopic organisms.
2.1. Binding Mode of Undefined LPOR with NADPH and Pchlide
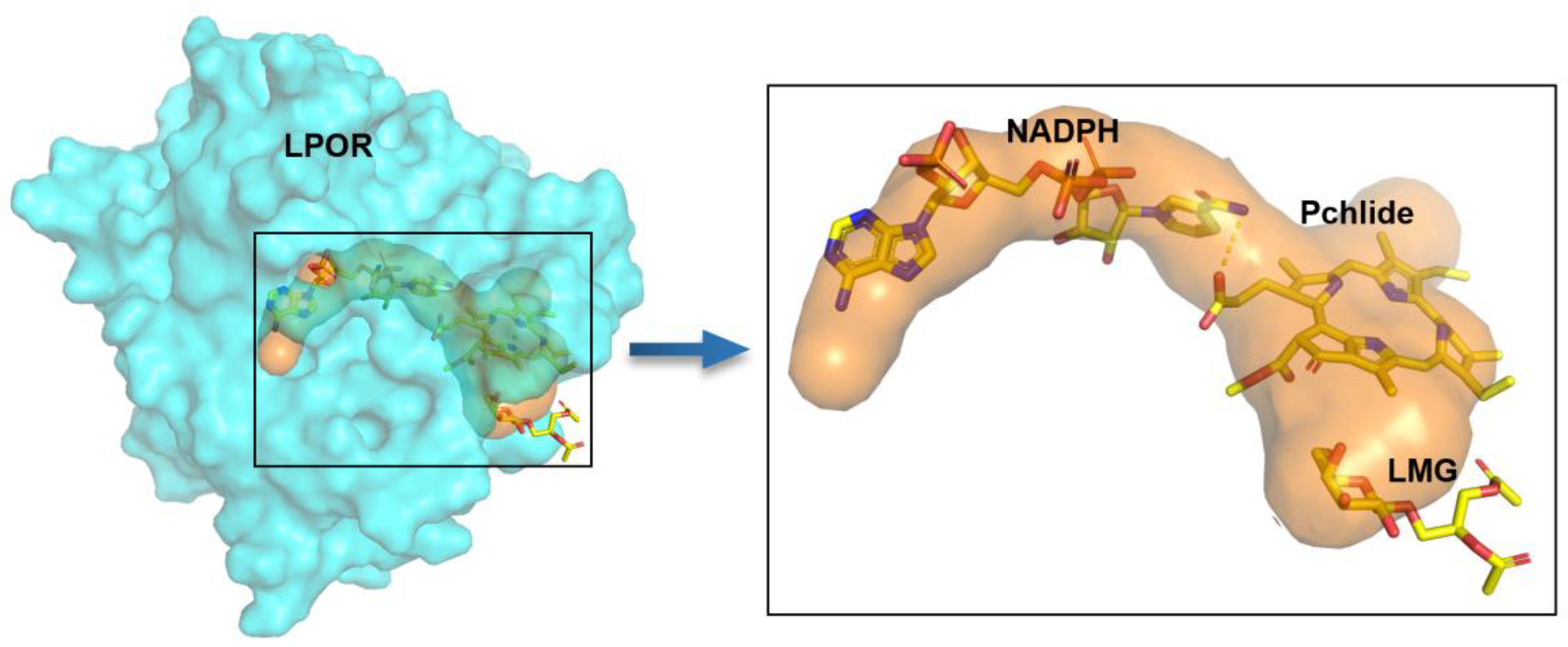
To further understand the binding mode of uncharacterized LPOR with NADPH and Pchlide, we predicted the three-dimensional structure of the uncharacterized LPOR using AlphaFold. In a previous paper, we compared the modelled LPOR structures with the LPOR crystal structure (PDB: 7JK9) [Sun XZ et al. (2025)]. In this article, we further elaborate the binding mode of Pchlide-NADPH with the uncharacterized LPOR, using the predicted three-dimensional structure of the uncharacterized LPOR and molecular docking. As shown in
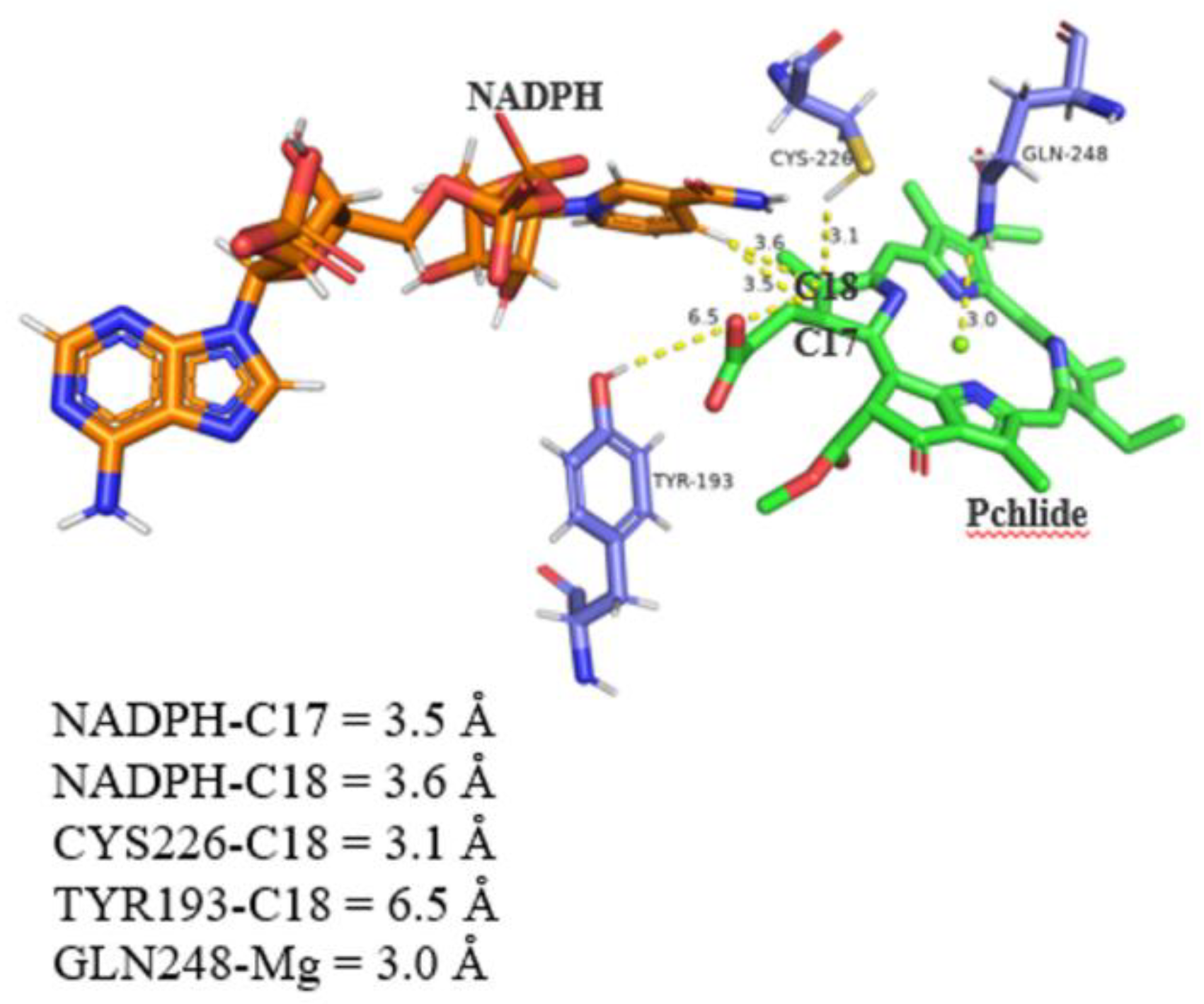
Figure 5, the distance between NADPH and C17 and C18 demonstrated a favorable transfer distance of <4 Å: 3.5 Å and 3.6 Å, respectively. In addition, we analyzed and visualized the NADPH and Pchlide ion channels of LPOR using the CAVER 3.0.3 plugin in PyMOL. The results further illustrated the potential pathways through which substrates may access the active site (
Figure 6).
The conserved, catalytically important CYS226 residue appears optimally positioned to participate in the reaction chemistry because it is 3.1 Å from the C18 position of the Pchlide substrate. Besides, GLN248 was found facing inwards to the active site, and the distance was close to the central Mg atom (3.0 Å) of Pchlide, potentially forming an adduct. However, the distance between Tyr193 and the C18 carbon is less favorable for proton transfer, at 6.5 Å. In all, the interaction mode is similar to light-dependent protochlorophyllide oxidoreductase (LPOR) bound to NADPH-Pchlide (PDB:7JK9) [Taylor et al. (2022)]. To explain the interaction, we also analyzed the active site of other uncharacterized LPOR structures (angiosperm LPOR, monocot LPOR, fern LPOR, gymnosperm LPOR, bryophyte LPOR, and algae LPOR) with the Pchlide–NADPH model, including TYR, CYS, GLN, and their respective distances to the C17/C18 region and central Mg atom of Pchlide (S-
Figure 2A-2F).
2.2. Final structural milestone for nitrogenase-like protein complex
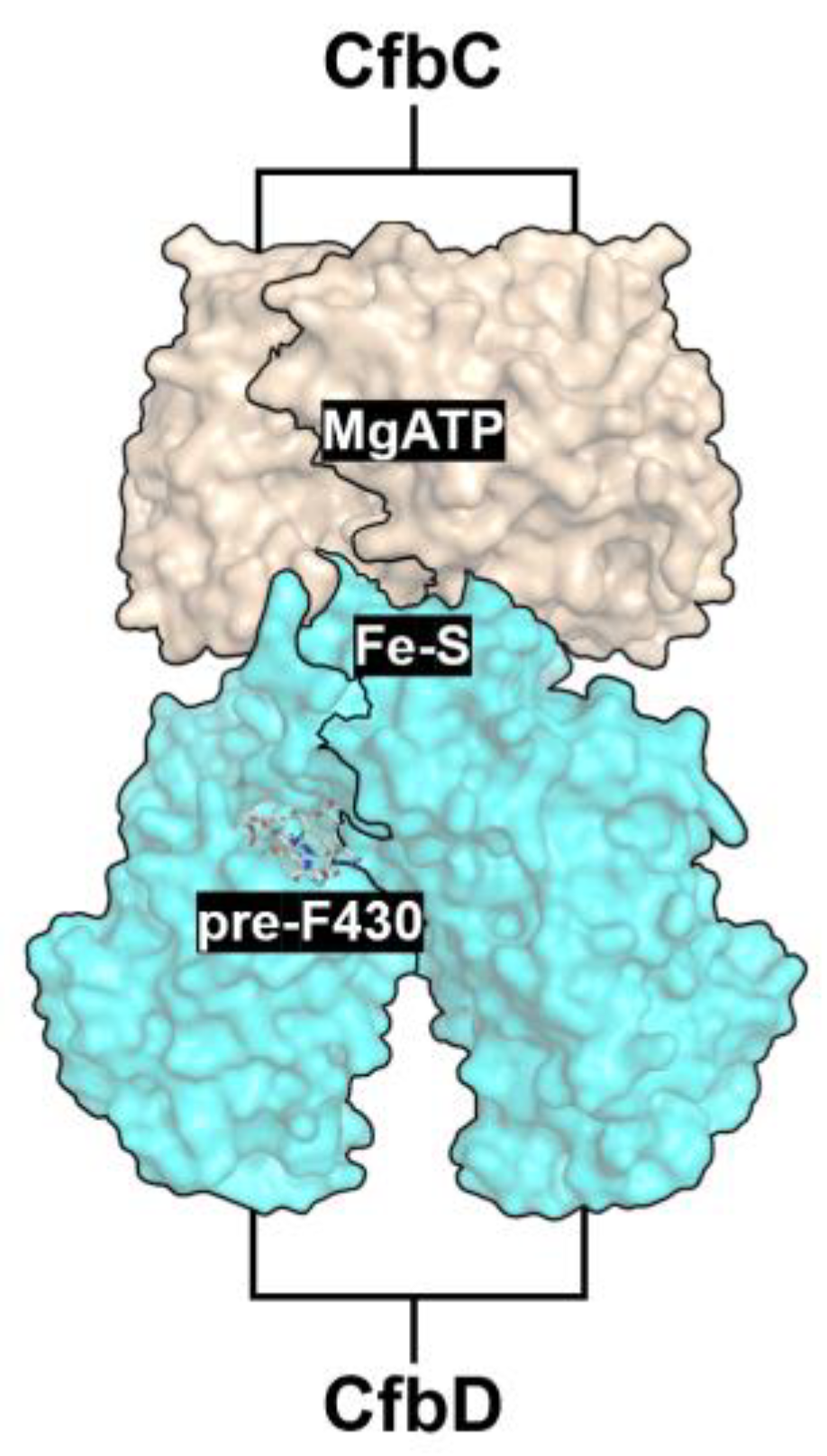
Protein complex CfbC/D, Coenzyme F430 reductase (hereafter referred to as F430,
Figure 7), is an ancient nitrogenase-like catalyst consisting of the CfbC Fe protein (equivalent to NifH) and CfbD (equivalent to NifD, ChlN, BchN). It is involved in the biosynthesis of Coenzyme F430, a redox-active nickel tetrahydrocorphin-containing prosthetic group. This complex could represent a simplified and ancestral version of N2ase, DPOR, or COR.
Coenzyme F430 is a redox-active nickel tetrahydrocorphin-containing prosthetic group in the active site of methyl-coenzyme M reductase (MCR), which catalyzes the final step in methane biosynthesis by methanogenic archaea.
The CfbC (UniProt ID: Q46FL1) and CfbD (UniProt ID: Q46FL2) proteins from Methanosarcina barkeri were subjected to AlphaFold-based structure prediction. Using the AlphaFold online platform, two sequences each of CfbC and CfbD were uploaded to simulate the quaternary structure of the F430 reductase enzyme complex (
Figure 8b), with the CfbC subunits shown in dark orange and the CfbD subunits in cyan. Representative CfbD sequences from various species were collected for sequence alignment (
Figure 8d), and the strictly conserved residues were identified and recorded. The predicted structure of F430 reductase was then annotated in PyMOL, with strictly conserved residues highlighted in red and moderately conserved residues in orange (
Figure 7,
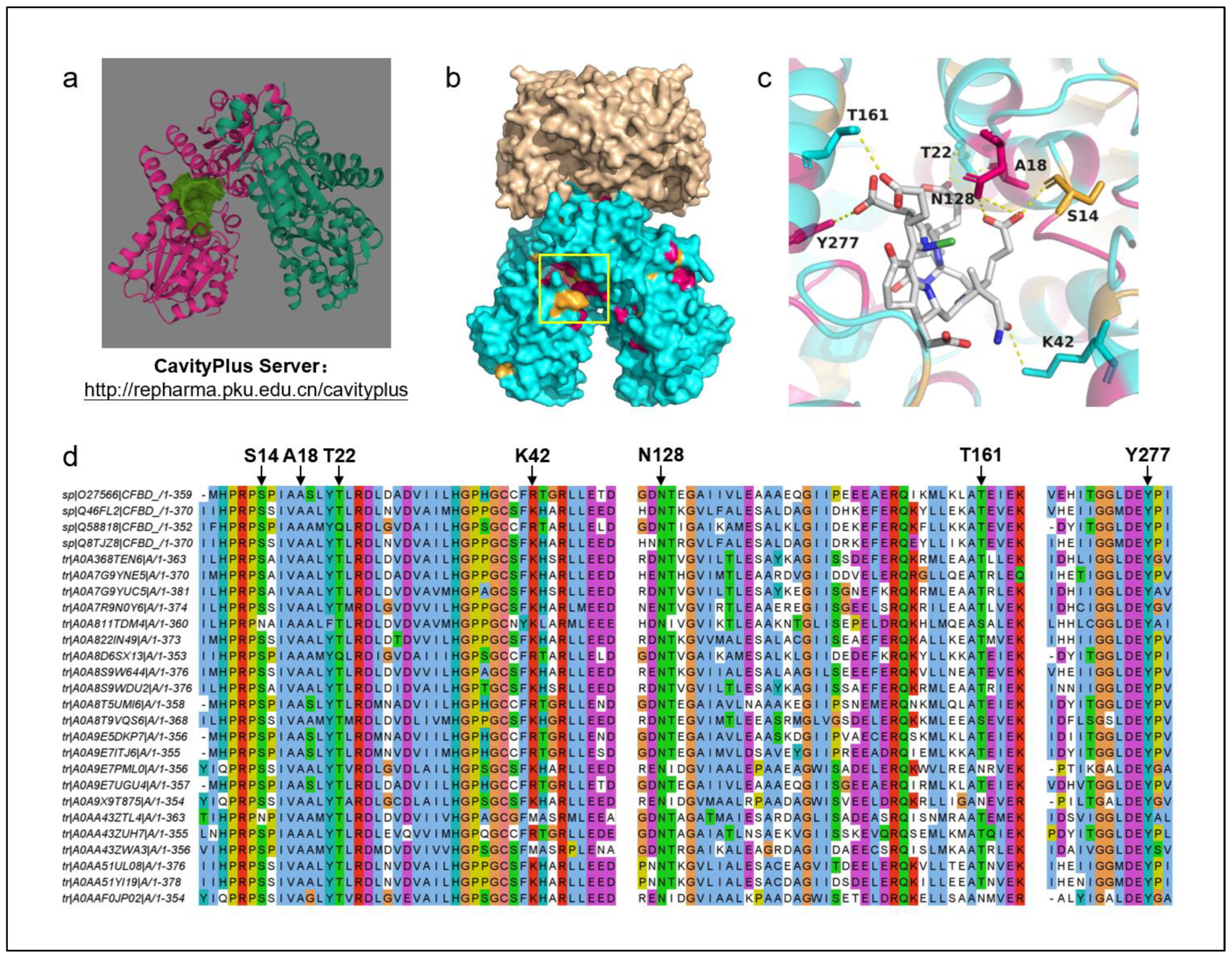
Figure 8b). A cluster of conserved residues was observed within the yellow-boxed pocket region.
Additionally, the predicted structure of F430 reductase and the F430 substrate molecule were uploaded to the CavityPlus server platform to predict the active pockets (
Figure 8a). The predicted active pocket is shown in dark green, with the results indicating a high correspondence between the predicted pocket region and the conserved regions identified from sequence alignment.
Molecular docking analysis of the predicted F430 reductase structure with the F430 substrate was performed using AutoDock Vina 1.1.2. The results revealed a binding energy of -8.1 kcal/mol, indicating a strong binding affinity. Interacting amino acid residues were annotated (
Figure 8c) and marked on the sequence alignment diagram. The results show that most of the interacting residues are conserved.
Based on our current results and global efforts working on Coenzyme F430 reductase, we expect that the X-ray crystal structure of either CfbC or CfbD, as well as the CfbC/D complex, will be achieved soon.
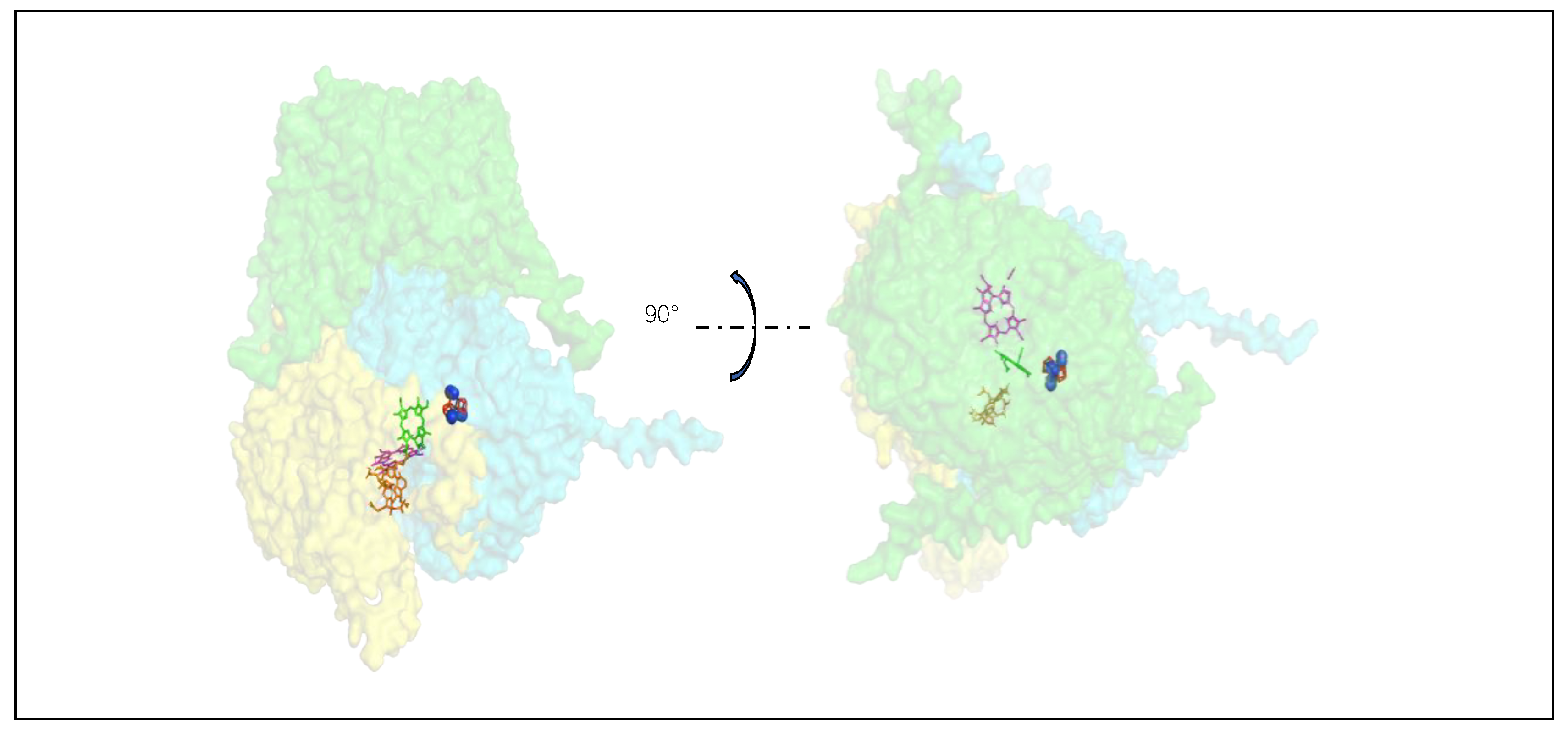
Based on real structures of N2ase, DPOR, and AlphaFold structures of COR and F430, we provide an overview of the substrate-channeling pathways (
Figure 9).
Green subunit: NifH, ChlL, BchX, CfbC; Cyan subunit: NifD, ChlN, BchY, CfbD; Yellow subunit: NifK, ChlB, BchZ, CfbD; Red ligand with blue balls: FeMo-cofactor with N2; Blue ligand: Pchlide; Magenta ligand: Chlide; Orange ligand: F430 cofactor.
According to our analysis, L contributes β1, α5, α11; N/B contributes α3; N contributes α9; and B contributes β7, β8. The majority of the remaining structures are similar among the three. Except for α6 and α7, which are unique to LPOR, corresponding structures can be found for the other regions.
In the article referenced [Buhr et al. (2008)], α6 and α7 belong to the extra loop of LPOR, which is an additional loop compared with the barley LPOR and Escherichia coli AHI genes. However, that article was from 2008, and at that time, relying on homologous modeling, it was impossible to identify spiral structures on the loop.
The three loops between β4-α5-α6-α7 and the two loops between β6-α9-α10 surround the substrate, which may be related to pocket formation. The extra loop formed between α6 and α7 is closest to the substrate.
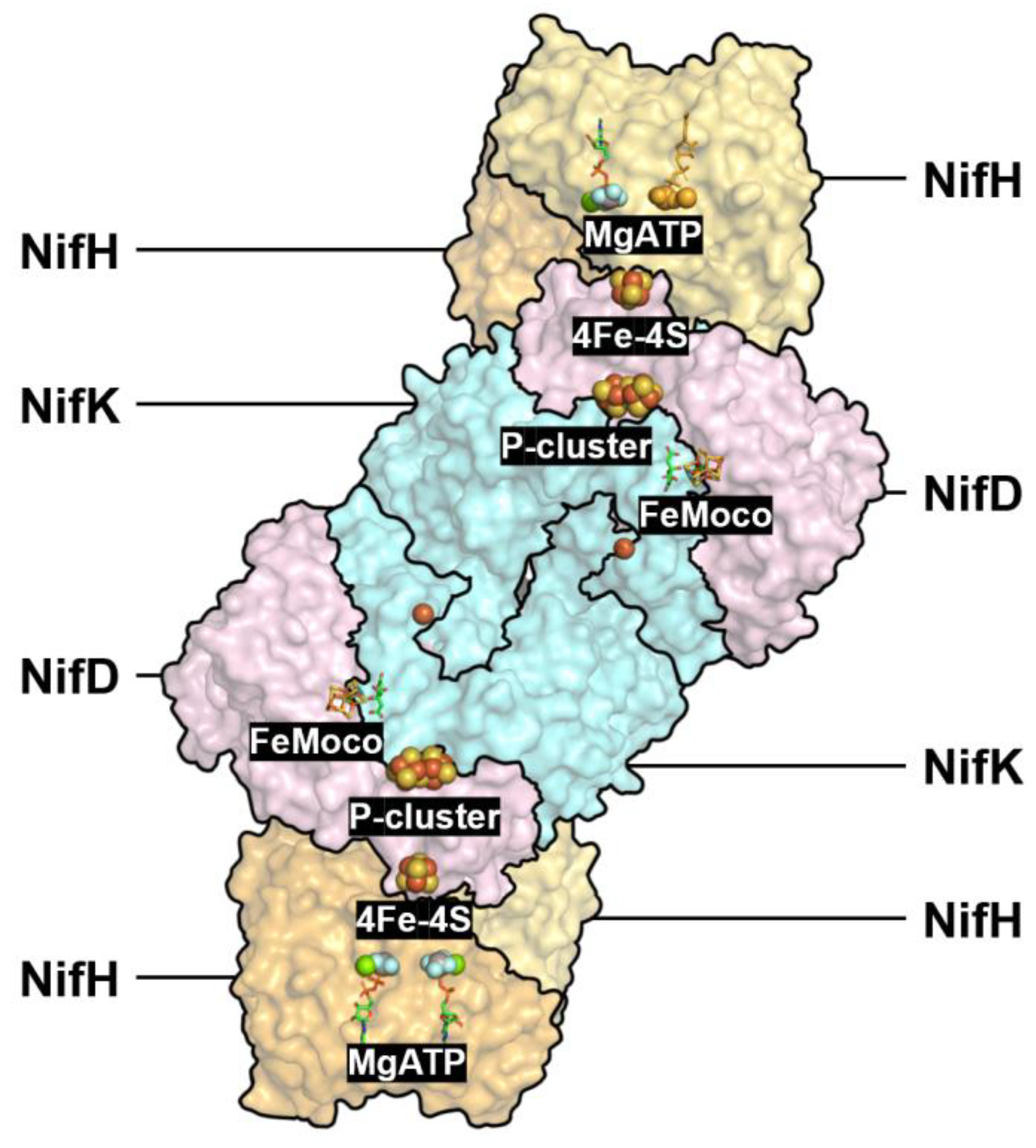
2.3. FeMoco docking with three-subunit N2ase
The nitrogenase enzyme, responsible for biological nitrogen fixation, relies on the intricate coordination and stabilization of its iron-molybdenum cofactor (FeMoco), a unique [MoFe₇S₉C] cluster. This cluster is embedded within the MoFe protein and serves as the active site for N₂ reduction [Spatzal, T., et al. (2011)]. Structural and spectroscopic studies have provided significant insights into its binding interactions and electron transfer pathways.
(1) Direct Coordination by Cysteine and Histidine: Crystallographic and spectroscopic analyses have revealed that FeMoco is coordinated within the α-subunit of the MoFe protein via interactions with amino acid residues, including homocitrate and histidine ligands [Einsle, O., et al. (2002); Lancaster, K. M., et al. (2011)]. The Mo atom is coordinated by a histidine residue, while Fe-S interactions stabilize the cluster within the protein scaffold [Rees, D. C., et al. (2005)]. Mutagenesis studies have demonstrated that specific residues such as α-His442 and α-Cys275 play a critical role in FeMoco binding and activity [Kaiser, J. T., et al. (2011)].
Cysteine (Cys275) directly coordinates the molybdenum atom in FeMoco, as demonstrated by Einsle et al. (2002) through high-resolution structural analysis. Similarly, Lancaster et al. (2011) identified histidine (His442) as crucial for stabilizing FeMoco, with its coordination role further supported by spectroscopic evidence. These residues ensure the proper positioning and electronic configuration of FeMoco.
(2) Stabilization via Hydrogen Bonding: Serine and threonine (Thr252) contribute to the stabilization of FeMoco through hydrogen bonding, as highlighted by Spatzal et al. (2016). Asparagine (Asn335) also plays a significant role by forming hydrogen bonds that stabilize the cofactor environment, as discussed by Howard and Rees (1994).
(3) Hydrophobic Environment: Hydrophobic residues such as leucine, isoleucine, and valine create a protective environment around FeMoco, shielding it from solvent interactions. Rees and Howard (2000) emphasized the importance of these residues in preserving the cofactor's reactivity and stability.
(4) Electron Transfer Modulation: Glutamate (Glu240) is critical for electron transfer within nitrogenase, as described by Danyal et al. (2011). This residue modulates electron density, facilitating efficient electron transfer from the Fe protein to FeMoco, a key step in nitrogen fixation.
(5) Indirect Support by Lysine and Arginine: Lysine and arginine residues indirectly support FeMoco by stabilizing the surrounding hydrogen bond network, as noted by Seefeldt et al. (2009). These residues contribute to the overall stability and functionality of the cofactor.
(6) FeMoco Assembly: FeMoco assembly occurs through a multi-step biosynthetic pathway involving chaperone proteins that mediate cluster maturation and insertion into the MoFe protein [Rubio, L. M., & Ludden, P. W. (2005)]. Studies using electron paramagnetic resonance (EPR) and X-ray absorption spectroscopy (XAS) have provided insights into the electronic structure of FeMoco and its redox properties [Bjornsson, R., et al. (2017)]. Additionally, computational modeling has identified potential conformational changes that regulate FeMoco insertion and catalytic efficiency [Dance, I. (2013)].
(7) Engineering Nitrogenase Variants: Understanding FeMoco binding and stability opens avenues for engineering nitrogenase variants with improved efficiency and alternative metal cofactors. This intricate network underscores the evolutionary sophistication of nitrogenase. Recent advancements in artificial nitrogenase design, including single-subunit light-driven nitrogenase, highlight the potential for synthetic biology approaches to enhance biological nitrogen fixation [Hu, Y., et al. (2019); Raugei, S., et al. (2018)].
2.4. Pchlide docking with single-subunit LPOR
Although various tetrapyrrole cofactors (e.g., bilin, coenzyme B12) can act as photoreceptive pigments in light sensing, only chlorophyll (and certain related derivatives) have been found to participate in biological photocatalysis [Taylor et al., 2022].
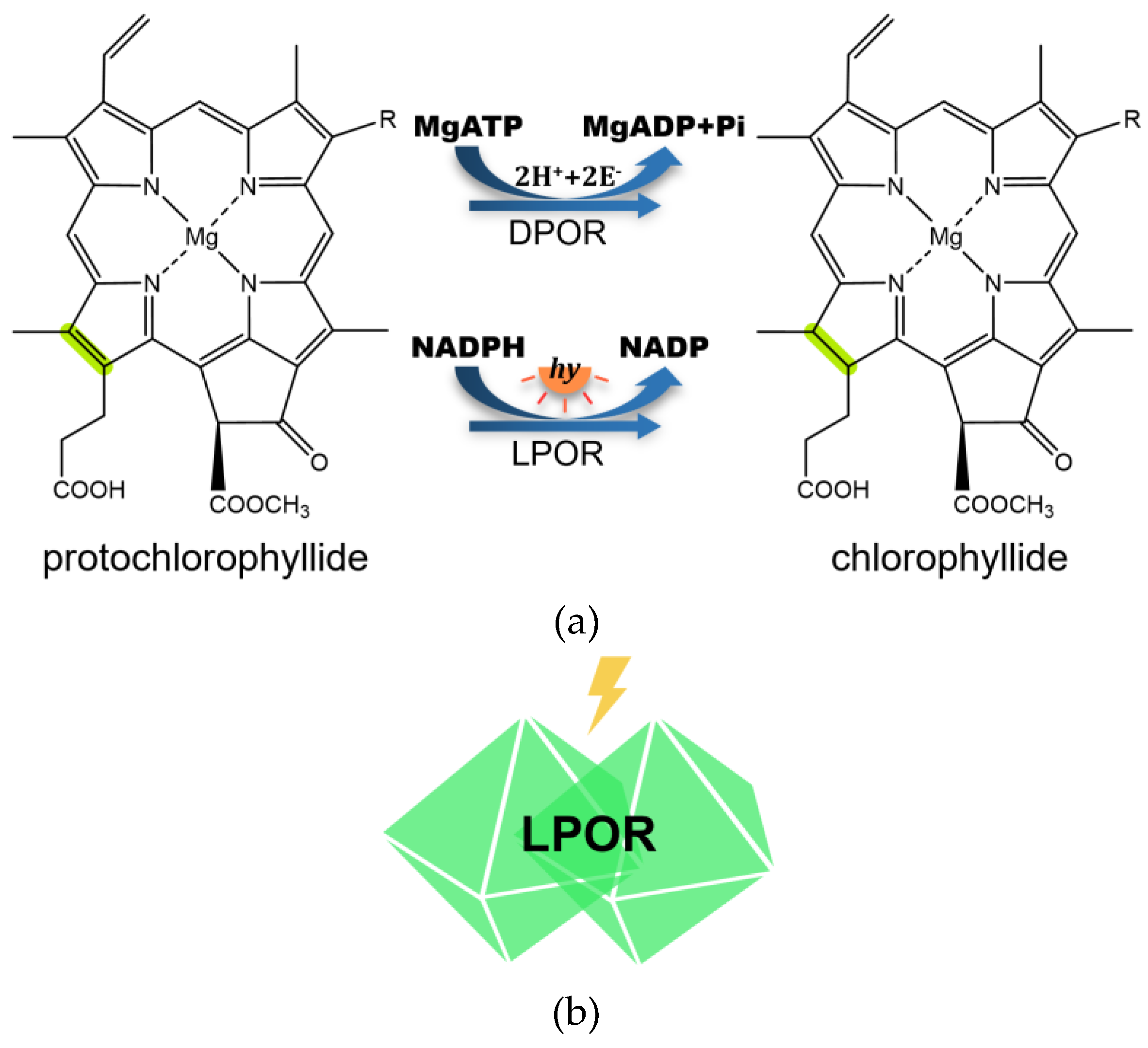
Chlorophyll is the primary pigment responsible for capturing light energy in photosynthetic organisms, with the reduction of the C17=C18 double bond in protochlorophyllide (Pchlide) being a critical step in forming chlorophyllide (Chlide). This reaction is catalyzed by light-dependent protochlorophyllide oxidoreductase (LPOR), which utilizes light energy to drive the reduction process.
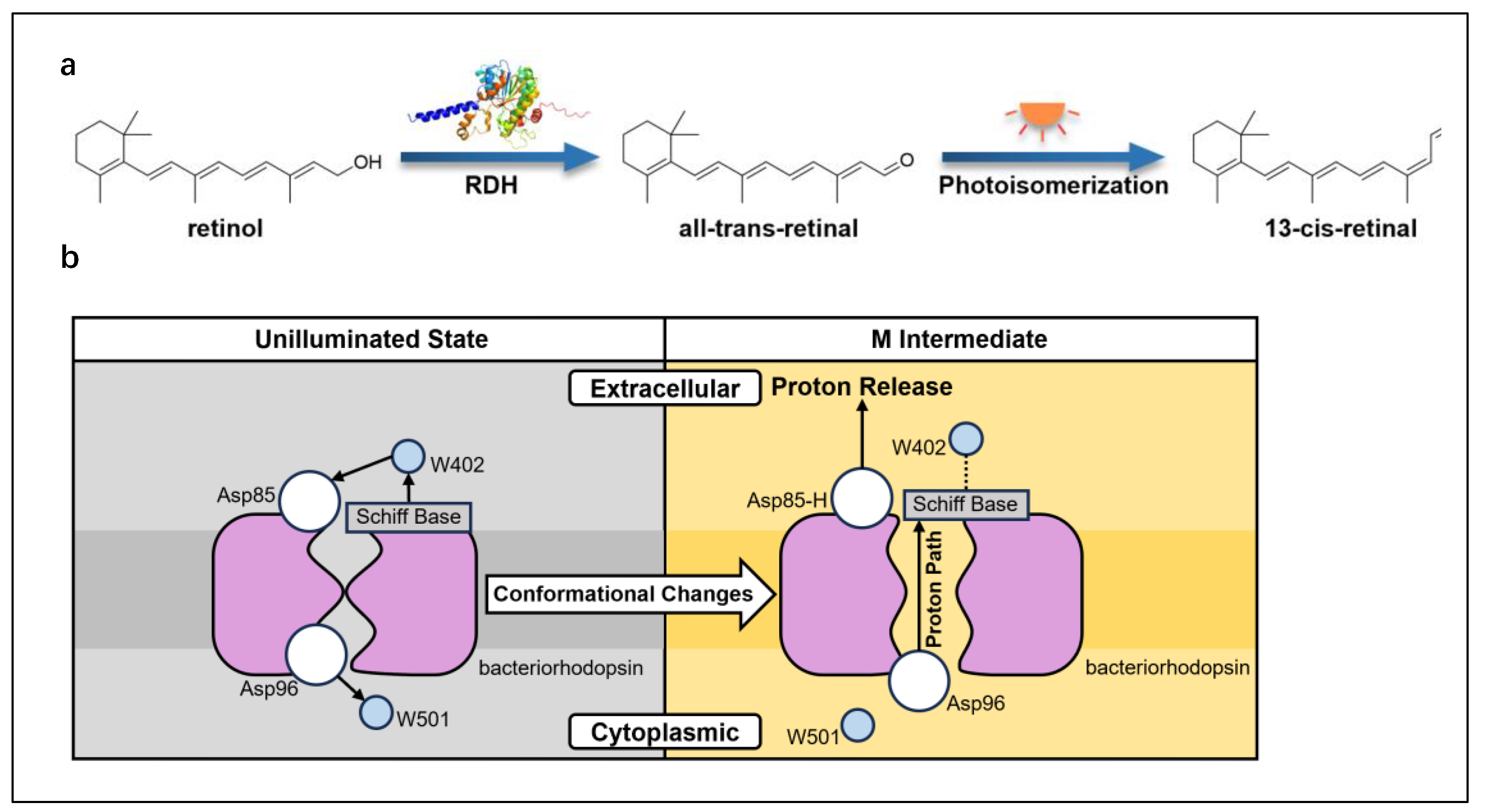
Additionally, RDH catalyzes the conversion of retinol to retinal, which in bacteriorhodopsin undergoes photoisomerization (
Figure 10a) to induce conformational changes in the protein, driving proton translocation across the membrane (
Figure 10b) and generating a proton gradient to support energy conversion [Lanyi and Luecke, 2001].
Understanding the photochemical mechanisms of LPOR and the role of retinal in proton pumping is essential for optimizing photosynthetic efficiency and advancing sustainable agricultural practices.
The reaction mechanism of LPOR involves a series of light-dependent steps, including the excitation of Pchlide, hydrogen transfer, and proton transfer.
(1) Light Absorption and Excitation: LPOR absorbs light, exciting Pchlide from its ground state (S0) to an excited state (S1 or S2). This excitation is crucial for initiating the reduction process. The excited Pchlide undergoes internal conversion (IC) and internal conversion fluorescence (ICF) to dissipate excess energy and stabilize in the S1 state [Heyes et al., 2009].
(2) Hydrogen Transfer (HYT): The excited Pchlide accepts a hydride (H⁻) from NADPH, transferring it to the C17 position of Pchlide. This step is rapid, with a rate constant of approximately 2.0 × 10⁶ s⁻¹. This hydrogen transfer results in the formation of a Pchlide-H⁻ intermediate, with NADP+ as a byproduct.
(3) Proton Transfer (PT): A proton (H⁺) is transferred from a tyrosine residue (usually Tyr189 or Tyr193) to the C18 position of Pchlide. This step is slower, with a rate constant of approximately 2.9 × 10⁴ s⁻¹. The proton transfer completes the reduction of Pchlide to Chlide, which is the precursor to chlorophyll [Taylor et al., 2024].
(4) Product Formation and Enzyme Regeneration: The reduced Pchlide (Chlide) is released from the enzyme, and the enzyme returns to its initial state, ready for another catalytic cycle. This process is essential for the continuous functioning of LPOR [Silva & Cheng, 2022].
Recent structural studies, including X-ray crystallography and cryo-electron microscopy (cryo-EM), have provided detailed views of the LPOR active site and the binding of Pchlide and NADPH. These studies have revealed the importance of specific amino acid residues in the catalytic process. Tyr189 and Tyr193 are crucial for proton transfer, although recent studies suggest that they may not directly donate the proton but rather stabilize the Pchlide intermediate [Taylor et al., 2024]. Cys226 has been proposed as a potential proton donor, with recent studies indicating its involvement in both hydrogen and proton transfer steps [Heyes et al., 2009]. Gln248 interacts with the central Mg atom of Pchlide, tuning its electronic properties and facilitating efficient photochemistry [Silva & Cheng, 2022; Pesara, et al. (2024)].
Computational methods, such as molecular dynamics (MD) simulations and quantum mechanics/molecular mechanics (QM/MM) calculations, have been instrumental in understanding the dynamics and energetics of the LPOR reaction mechanism. These studies have provided insights into the binding modes of Pchlide and the roles of key residues in the catalytic process [Menon et al., 2010; Pesara, et al. (2024)].
The detailed understanding of the LPOR reaction mechanism has significant implications for chlorophyll biosynthesis and photosynthesis. Efficient reduction of Pchlide to Chlide is essential for the formation of chlorophyll, which in turn affects the efficiency of light capture and energy conversion in photosynthetic organisms. Enhancing the activity of LPOR could lead to improved photosynthetic efficiency and increased crop yields, particularly under suboptimal light conditions [Heyes et al., 2009].
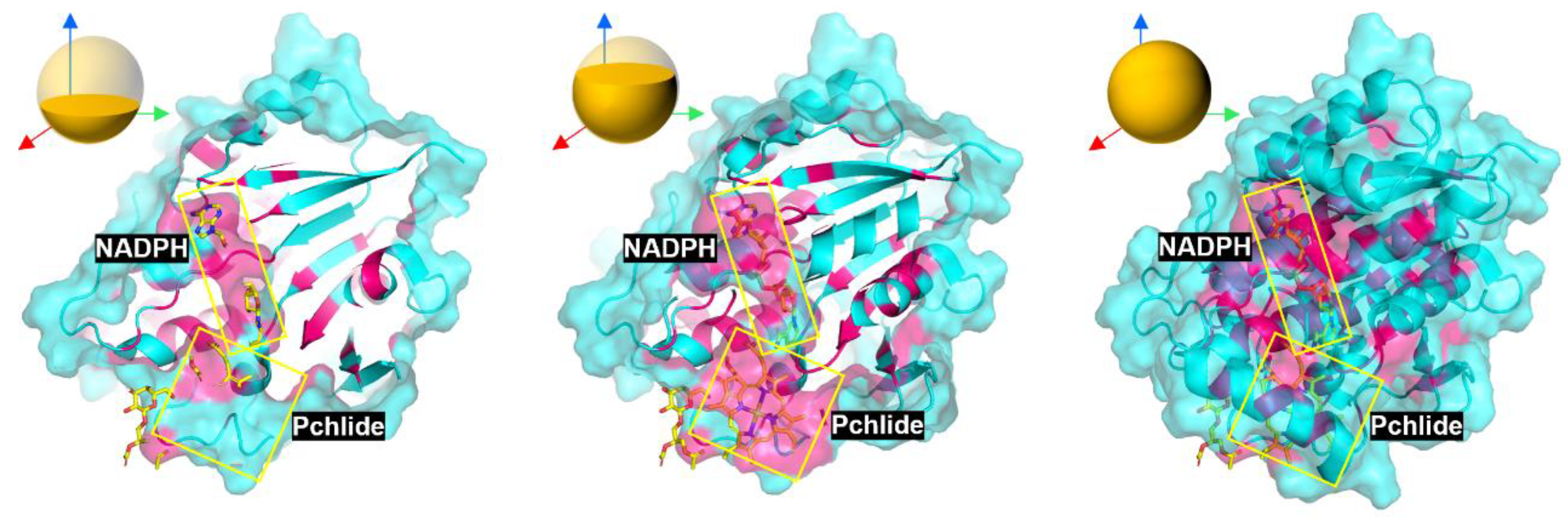
To further elucidate the roles of these residues in the LPOR reaction mechanism and address controversies regarding their contributions, we performed a multiple sequence alignment of LPOR orthologs from various species. Highly conserved residues identified in the alignment were mapped onto the LPOR structure (PDB ID: 7JK9) for visualization. In this representation, strictly conserved residues were highlighted in bright red, while non-conserved or variable residues were depicted in cyan by default. Given that the substrate-binding pocket is located within the protein interior, a layered scanning approach was employed to analyze the spatial distribution of these residues (
Figure 11).
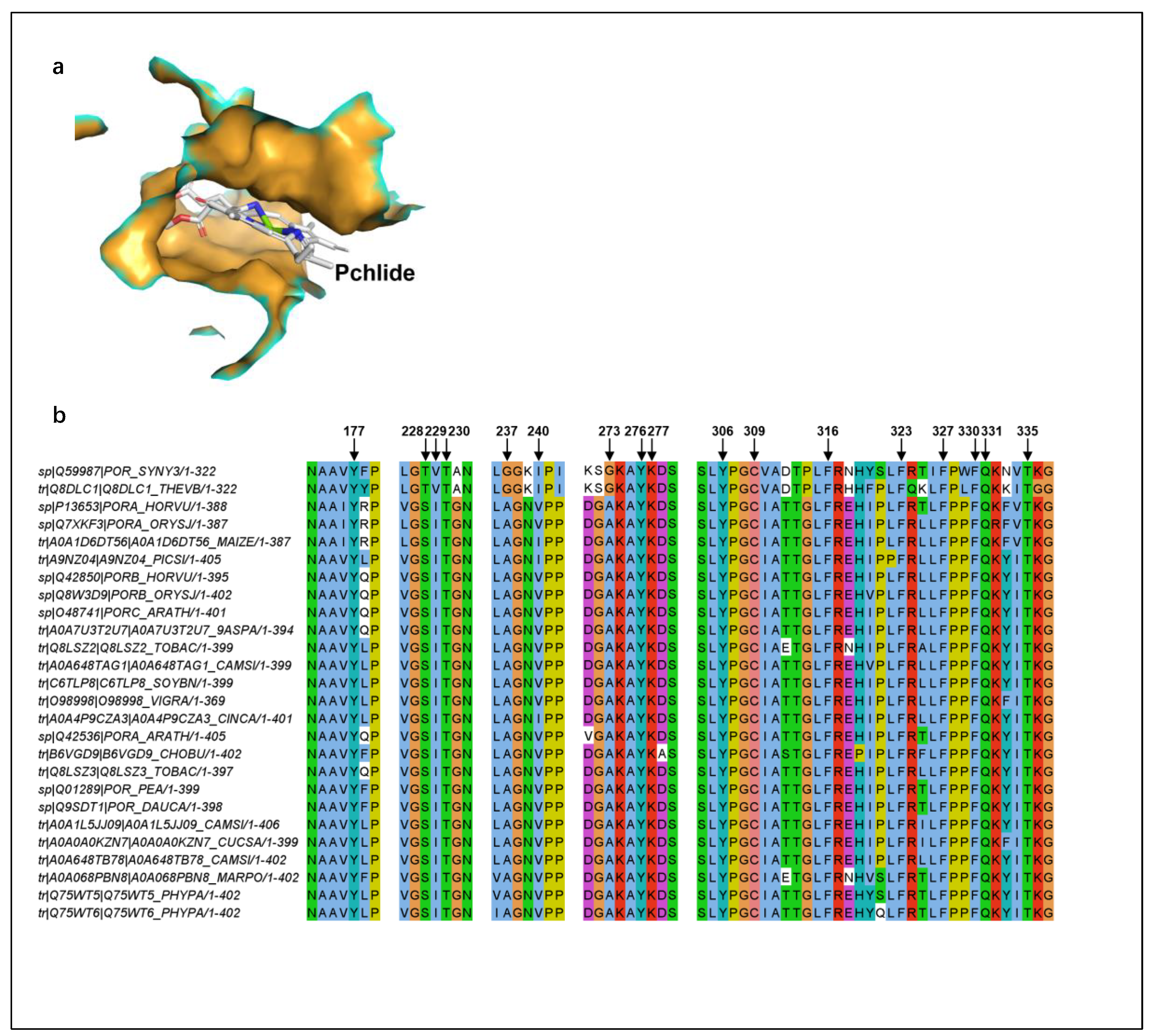
The first layer scan focused on the NADPH-binding region, revealing a cluster of conserved residues likely stabilizing cofactor binding and facilitating hydride transfer. The second layer scan targeted the Pchlide-binding pocket, highlighting conserved motifs around the porphyrin ring that may modulate substrate excitation and protonation. The third layer scan provided an overview of the entire structure, demonstrating the overall conservation pattern and its correlation with functional domains. Additionally, residues surrounding Pchlide were highlighted in PyMOL (
Figure 12a, orange region), all of which directly contact the substrate Pchlide. The corresponding positions of these residues in the sequence alignment have been indicated (
Figure 12b). This analysis underscores the evolutionary conservation of key active-site residues, supporting their critical role in catalysis.
Future research on LPOR should focus on: (1) Structural Dynamics – Investigating the dynamic changes in the LPOR active site upon light excitation and during the catalytic cycle; (2) Enzyme Engineering – Developing LPOR variants with enhanced activity and stability through rational design and directed evolution; (3) In Vivo Studies – Conducting in vivo studies to understand the physiological context of LPOR activity and its impact on plant growth and development.
2.5. FeMoco docking with engineered LPOR
Based on structural insights from nitrogenase-like proteins (DPOR and COR) and SDR-like proteins (RDH and LPOR), we designed a hypothetical, new light-driven electron transfer protein by incorporating FeMoco into the LPOR scaffold. Previous studies identified the substrate binding mode in the Pchlide-NADPH-LPOR complex and explored the reaction mechanism using quantum-chemical methods. These studies revealed how the LPOR active site promotes photo-driven reduction through initial electron transfer from Tyr189 to the substrate, then hydride transfer from NADPH and internal electron transfer from Cys222 to the deprotonated Tyr189 radical. Given the structural and functional similarities between nitrogenase-like proteins and LPOR, we hypothesized that FeMoco could be accommodated within the LPOR scaffold, enabling light-driven electron transfer to FeMoco, potentially eliminating the need for ATP-driven electron transfer observed in natural nitrogenases.
The key electron-transfer event in LPOR is facilitated by active-site amino acids rather than just the interplay of NADPH and Pchlide molecular orbitals. Considering the structural relationship between nitrogenase and DPOR, and the alignment of substrate binding sites, we hypothesized that the LPOR scaffold could accommodate FeMoco and catalyze light-driven electron transfer from Tyr189 to FeMoco.
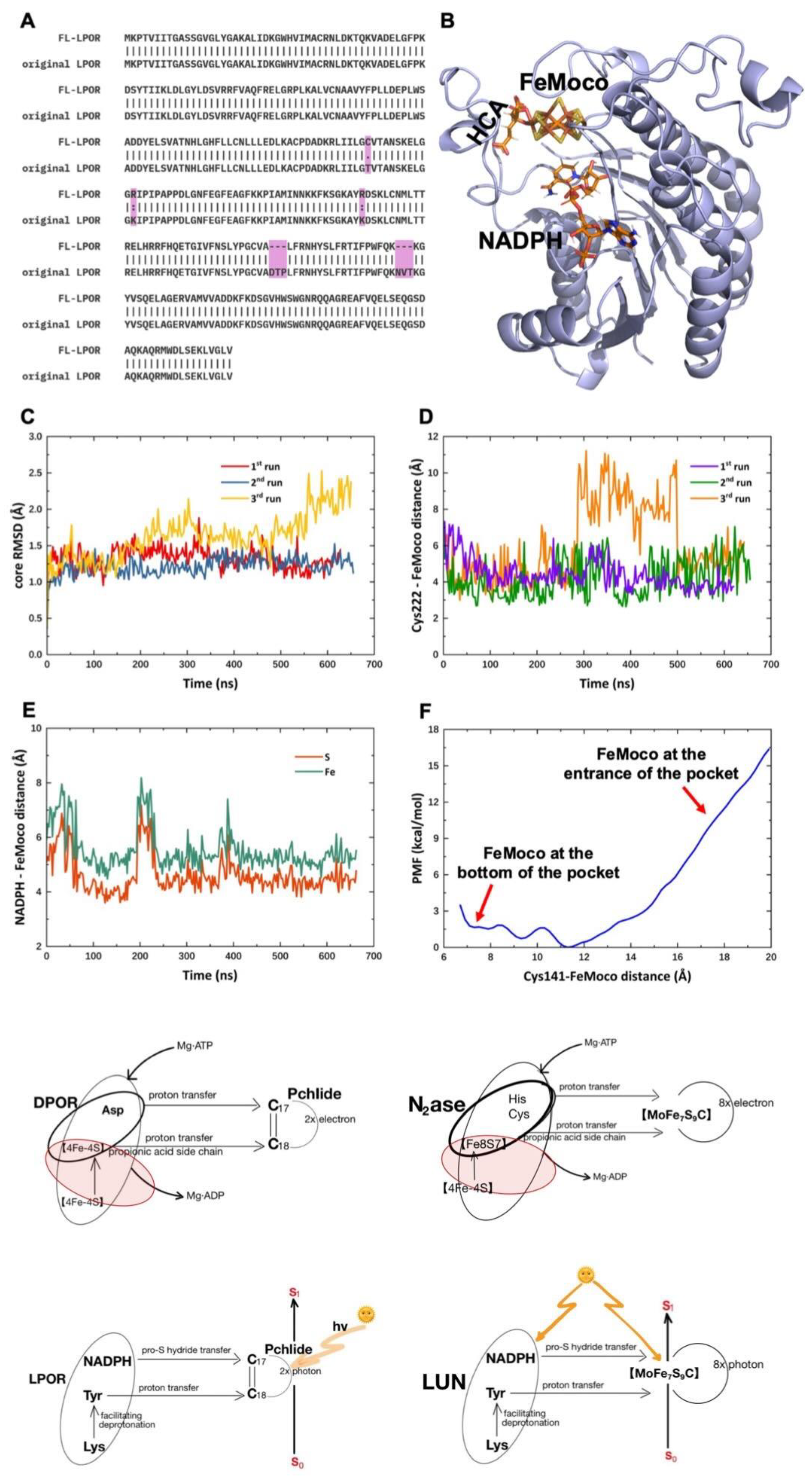
FeMoco is covalently linked to nitrogenase through a His-Molybdenum bond and a Cys-Fe bond. In LPOR, the substrate binding cavity is partially occluded by a highly mobile loop containing a His residue, which we repurposed for FeMoco binding. Since no Cys exists at an appropriate position to bind the opposite end of FeMoco, we mutated Thr141 to Cys. By removing amino acids adjacent to the candidate His in the mobile loop, we obtained a modified LPOR model, designated FeMoco-loaded engineered LPOR (FL-LPOR), with His and Cys residues positioned to bind FeMoco [
Figure 13].
To better mimic the Arg-rich environment of natural nitrogenase, two Lys residues were replaced with Arg in FL-LPOR. The modeled structure of FL-LPOR is similar to LPOR, allowing for molecular docking with FeMoco at comparable positions. Molecular dynamics simulations (>500 ns each) tested the stability of FL-LPOR, showing that the protein remains stable with high mobility in the mobile loop and C-terminal portions. The distance between FeMoco and Cys222 remains short, suggesting a reliable electron transfer pathway. However, the distance between NADPH and FeMoco is more variable, indicating that the interaction with NADPH might need further optimization for consistent hydride donation to FeMoco.
Potential of mean force (PMF) graphs show that FeMoco entry into the LPOR cavity is thermodynamically favorable, with a negative energy change indicating spontaneous binding. This suggests that engineered LPOR can readily accommodate FeMoco, crucial for enabling subsequent catalytic reactions.
The minimum complement of
nif genes required for FeMoco biosynthesis and assembly varies [
Figure 14] [Cheng (1998)] [Cheng (2005)]. The NifEN protein complex is usually considered essential for FeMoco synthesis. However, Payá-Tormo et al. (2024) showed that a thermophilic Roseiflexus bacterium could assemble FeMoco without the scaffold NifEN. The study demonstrated that NifH, NifB, and apo-NifDK proteins produced in
E. coli were sufficient for FeMoco maturation and insertion into the NifDK protein. This finding challenges the conventional view and suggests that alternative pathways for FeMoco biosynthesis may exist [Xie et al., 2014; Shang et al., 2024; He et al., 2022; Payá-Tormo et al., 2024].
2.6. Predictions and Propelling Engineering
- 1)
N2ase-born FeMoco may shift from multi-subunit N2ase to single-subunit engineered LPOR in nitrogen fixation cyanobacteria, such as Anabaena;
- 2)
FeMoco-fitted LPOR-like LUN may have evolved in some cyanobacteria with superior nitrogen fixation ability, which requires further investigation;
- 3)
FeMoco-like LPOR-like LUN may have evolved in some C4 plants with superior growth rates, which requires further investigation.
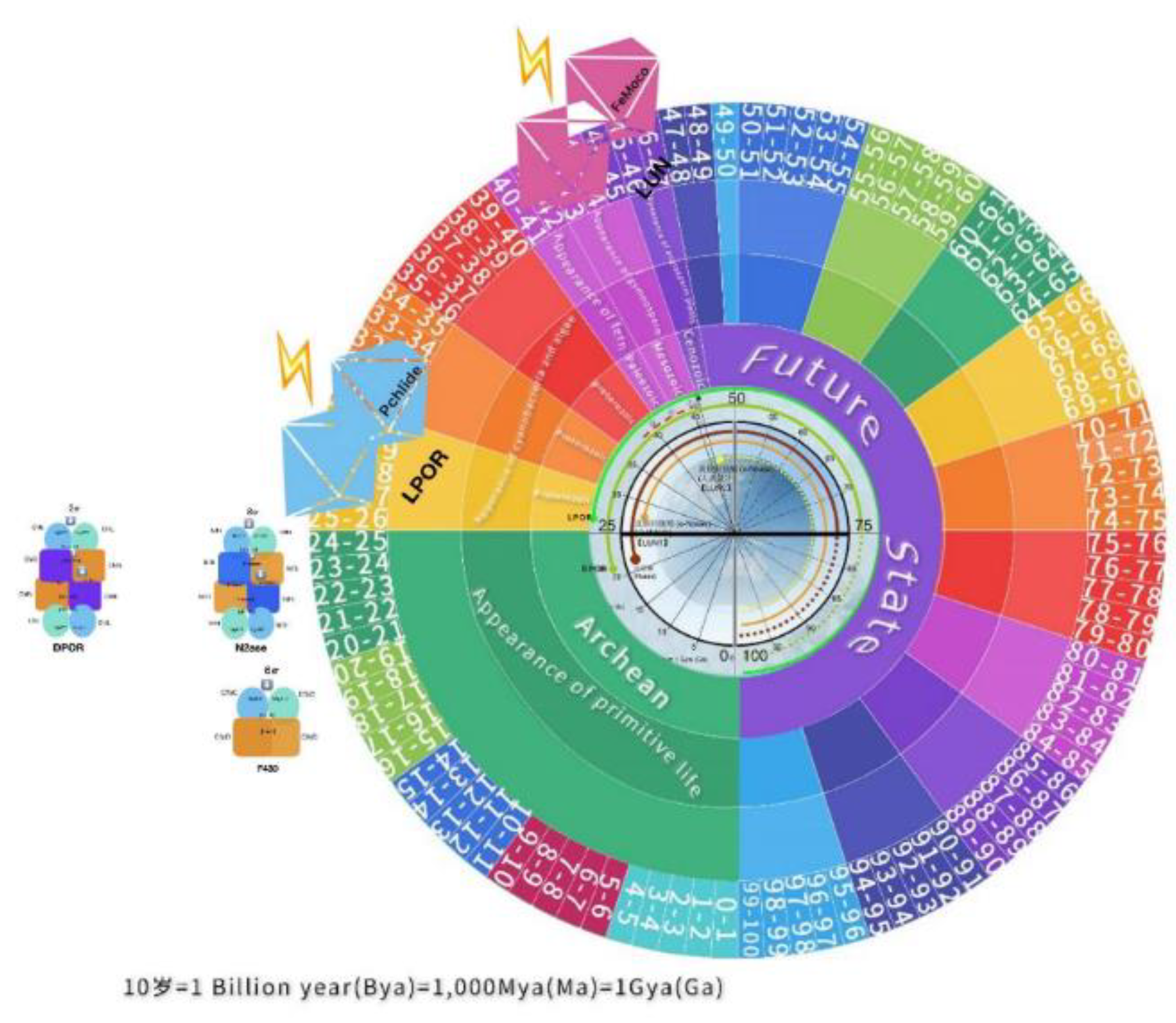
On the other hand, positioning evolutionary events, organisms, and molecules in geological time, in an Earth Clock manner, may be helpful for understanding the visible past or somehow predictable future. However, while noting for certain that our planet Earth could have "full life" for another 100 years (
Figure 15), it may encounter unexpected events, such as natural disasters or existential threats from outer space.
As far as we can see, the visible images of the nitrogenase-related catalytic protein complex structures are nearly complete. A precise version, where scientists may be able to embark on achieving comparative results [Cheng (2008)] and more effectively [Taylor et al., 2022], is forthcoming.
2.7. AI-aid designing for non-natural LPOR
In the first strategy plan for reinventing a possible ancestral non-natural Light-Dependent Protochlorophyllide Oxidoreductase (LPOR), we adopted a homology-guided design approach to construct the substrate-binding pocket—a critical domain for LPOR’s core function of catalyzing protochlorophyllide reduction. This design was rooted in comparative analysis of multi-subunit sequences from four evolutionarily related enzymes: Dark-Operative Protochlorophyllide Oxidoreductase (DPOR), Chlorophyllide Oxidoreductase (COR), Nitrogenase (N₂ase), and F₄₃₀-dependent Methyl-Coenzyme M Reductase. These enzymes share conserved structural folds and redox-active cofactor-binding regions, making their sequences a reliable template to infer the ancestral LPOR’s substrate-interacting residues. By aligning conserved motifs across these homologs and mapping residue positions critical for substrate coordination (e.g., hydrophobic pockets and polar interaction sites), we generated a preliminary substrate-binding pocket model that balances ancestral sequence features with LPOR’s specific substrate preferences.
Complementing this, we incorporated two key elements to establish a functional electron transfer chain—essential for LPOR’s redox catalysis. First, we integrated the NADPH-binding motif, a universally conserved domain in reductases that enables binding of the NADPH cofactor (the primary electron donor for LPOR). Second, we included the YKDSK motif, a sequence identified in homologous redox enzymes (e.g., DPOR and N₂ase) that mediates electron transfer between the cofactor and the active site via conserved lysine and aspartate residues. Together, these motifs formed a predicted electron transfer pathway, linking NADPH oxidation to substrate reduction in the ancestral non-natural LPOR.
To validate and illustrate this design process,
Figure 13 and
Figure 14 provide visualizations of the reconstructed substrate-binding pocket—showing residue alignments with homologs, predicted substrate (protochlorophyllide) docking, and key interaction sites (e.g., hydrogen bonds between pocket residues and substrate functional groups). Meanwhile,
Supplementary Table 1 (S-table 1) compiles quantitative data supporting the design: including sequence conservation scores of pocket residues across DPOR/COR/N₂ase/F₄₃₀ homologs, predicted binding affinities of the reconstructed pocket for protochlorophyllide (calculated via molecular docking simulations), and alignment statistics for the NADPH-binding and YKDSK motifs. Collectively, these figures and the table serve as concrete examples of how we translated evolutionary sequence data into a testable model of an ancestral non-natural LPOR, bridging structural homology with functional enzyme design.
2.8. de novo Light-driven catalysts: LUN、LCOR、L430
This paper introduces the entire history of thirty years of effort in the structural elucidation of the ancient anaerobic enzyme nitrogenase, the nitrogenase-like DPOR protein complex, and their ancestral F430 reductase. The comparison of these nitrogenase-like catalysts has been demonstrated in terms of structural complexes, mechanistic illustration, and substrates and products (
Figure 6). Evolutionary relationships with nitrogenase-like catalysts and the structural implications of light-utilizing nitrogenase (LUN) are shown in
Figure 16.In turn, focusing on the evolutionary invention of the remarkable enzyme LPOR and its position in the geological age on the 'Earth clock' (
Figure S3), we look forward to the discovery or invention of a greater masterpiece, the LUN. Additionally, several other higher-version catalysts (LCOR, L430) could potentially be designed from the lower-version ones (COR, F430) (
Figure 16,
Table 1).Key catalysts for nitrogen reduction and porphyrin reduction have been summarized in
Table 1.
Based on the successful evolutionary story of LPOR from DPOR, and the preliminary evidence we provide, one could postulate that the so-far theoretical higher versions of catalysts may be able to be designed from the lower version ones (
Figure 15). At present, it still seems very ambitious and bold, but no less scientific, proposing that there may be a completely undiscovered novel nitrogenase or a designable LUN that could appear somewhere on the Earth Clock [Cheng (2013)]. However, the central aim of this paper is to ask whether the evolutionary and structural legacy of the nitrogenase-like protochlorophyllide oxidoreductase (POR) could guide the future of biological nitrogen fixation with a light-utilizing nitrogenase (LUN). This could revolutionize biological nitrogen fixation engineering.
In this aspect, there may be two opposite opinions. One may be scientifically seeking more support: a) comparative implications; b) chemical engineering; c) synthetic biology; d) In the near future, molecular design work may be carried out using gigantic quantum assimilation technologies or AI-aided and structural mimicking approaches (in fact, we have already developed a set of highly evolved de novo nitrogenase-related catalysts [LUN, LCOR, L430] as ancestral versions) (S-table 1).
On the other hand, particularly for LUN, a philosophical objection may be: a) chemically impossible; b) why hasn’t it dominated the plant kingdom so far; c) wait for another billion years according to the Earth Clock [Cheng (2013)].
This paper uses AlphaFold to predict the structure of the representative LPORs from the LPOR phylogenetic tree and analyzes the factors affecting the efficiency of protein activity based on the structure. At the same time, based on comparative analysis, we propose relevant design schemes for LUN and look forward to its continuous improvements. Preliminary results on AI-designed non-natural LCOR can be seen [Sun XZ et al. (2025)].
In the near future, AI technology may have great potential to cope with gigantic structural optimization for specific substrate targets, such as Chlide a or a FeMoco-like cage for N₂, etc. After all, there may still be a long way down this road and technical difficulties in designing such de novo enzymes biochemically and biologically, but as the pioneer Professor Ray Dixon once said, we cannot afford not to try it for the cause of agricultural food production, ultimately sustaining burgeoning world populations.


 Minimum nif genes
required for FeMoco biosynthesis and assembly.
Minimum nif genes
required for FeMoco biosynthesis and assembly.
 Minimum nif genes
required for FeMoco biosynthesis and assembly.
Minimum nif genes
required for FeMoco biosynthesis and assembly.