Submitted:
17 November 2025
Posted:
18 November 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Related Works
- Direct regression prediction of AST level, considered as an independent predictor of cardiovascular risk and not as a marker of hepatological disorders;
- Integration of routine biochemical, anthropometric, and behavioral parameters, including inflammation, body weight, and lifestyle indicators, enhances the clinical relevance of the model;
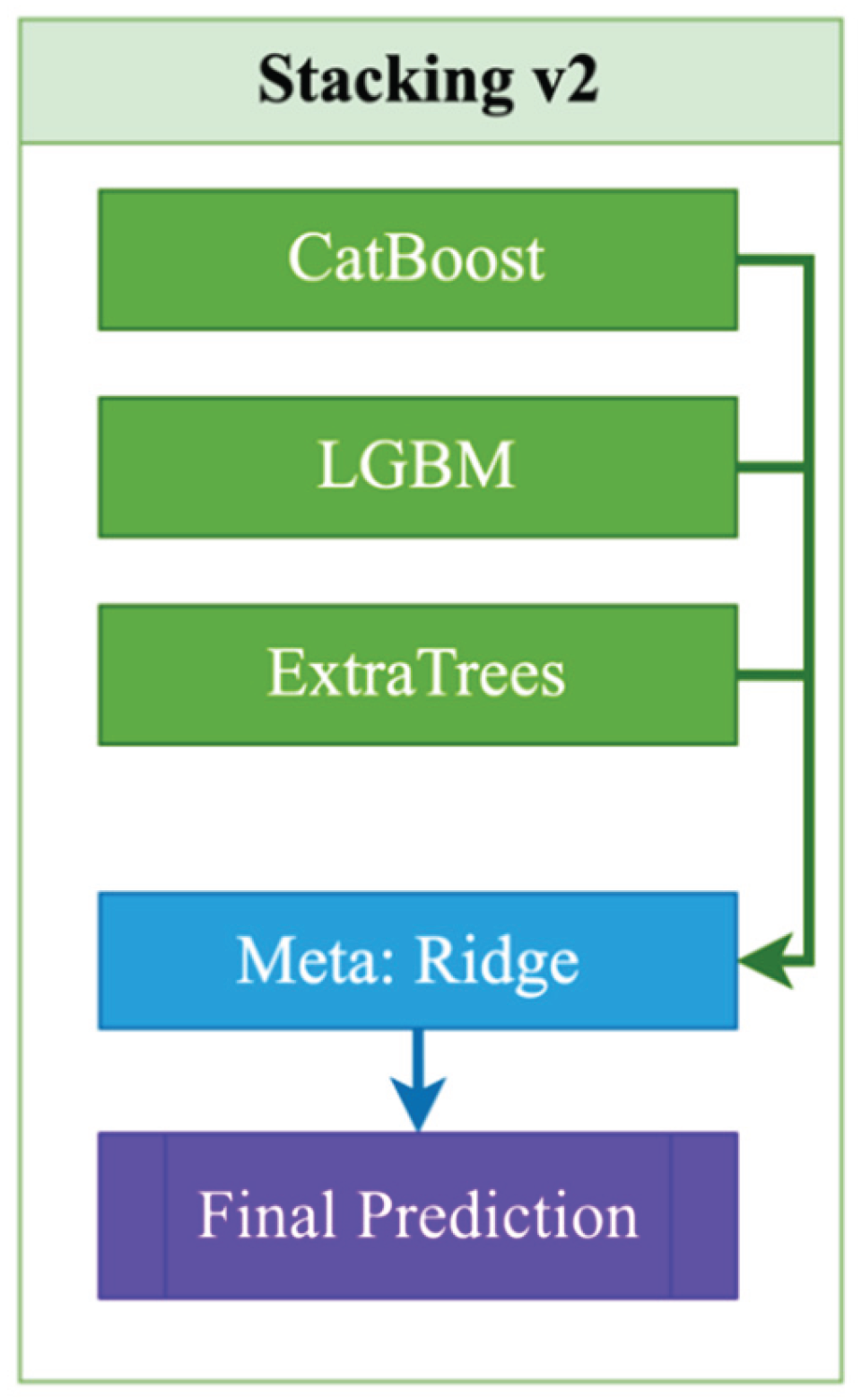
- Use of a stacking ensemble (Stacking v2), which combines the capabilities of modern algorithms and an interpretable meta-model to improve accuracy and stability;
- The use of SHAP and mutual information to analyze the significance of features ensures the interpretability of the model and its applicability in the clinical environment;
- Validation on a large and representative NHANES dataset (1988–2018) covering a wide range of health data from the US population.
3. Materials and Methods
3.1. Dataset Collection
3.2. Rationale for a Method Selection
- Removing emissions. In the first step, we excluded observations with suspiciously high values (s ≤ 200) from the dataset. This helps reduce the influence of anomalies and noise on model training. It's especially important when dealing with biomarkers, as technical or clinical errors can lead to outliers.
- Removing gaps. Removing rows with missing values in the target variable (AST) and critical predictors ensures the correctness of the training process. This step is necessary to maintain the quality of predictions and prevent distortions.
- Transformation of categorical variables. One-hot encoding of categorical features (e.g., demographic and questionnaire data) is applied, which allows them to be efficiently included in machine learning models without violating assumptions about the numerical nature of the input data.
- Scaling of Numerical Features. Numerical features are normalized (z-transformed) to equalize scales and prevent features with high variance from dominating the analysis. This is especially important for linear and gradient-boosted models that are sensitive to scale.
- Split into training and validation samples. The standard split of the sample (train/test split) is used to assess the quality of the model objectively. This allows you to control overfitting and tune hyperparameters.
- Base models (Base regressors). The following algorithms were selected to build a forecast of the AST level:
- Linear Regression — a basic benchmark for estimating linear relationships.
- Random Forest — a stochastic model that is robust to outliers and works well with small samples.
- XGBoost — a powerful gradient boosting that provides high accuracy and control over overfitting.
- CatBoost — an optimized boosting algorithm that works efficiently with categorical features without the need for manual coding.
- LightGBM — a fast and scalable boosting algorithm, especially effective on large and sparse data.
- Extra Trees — an improved version of Random Forest that uses additional stochasticity to improve generalization.
- 7.
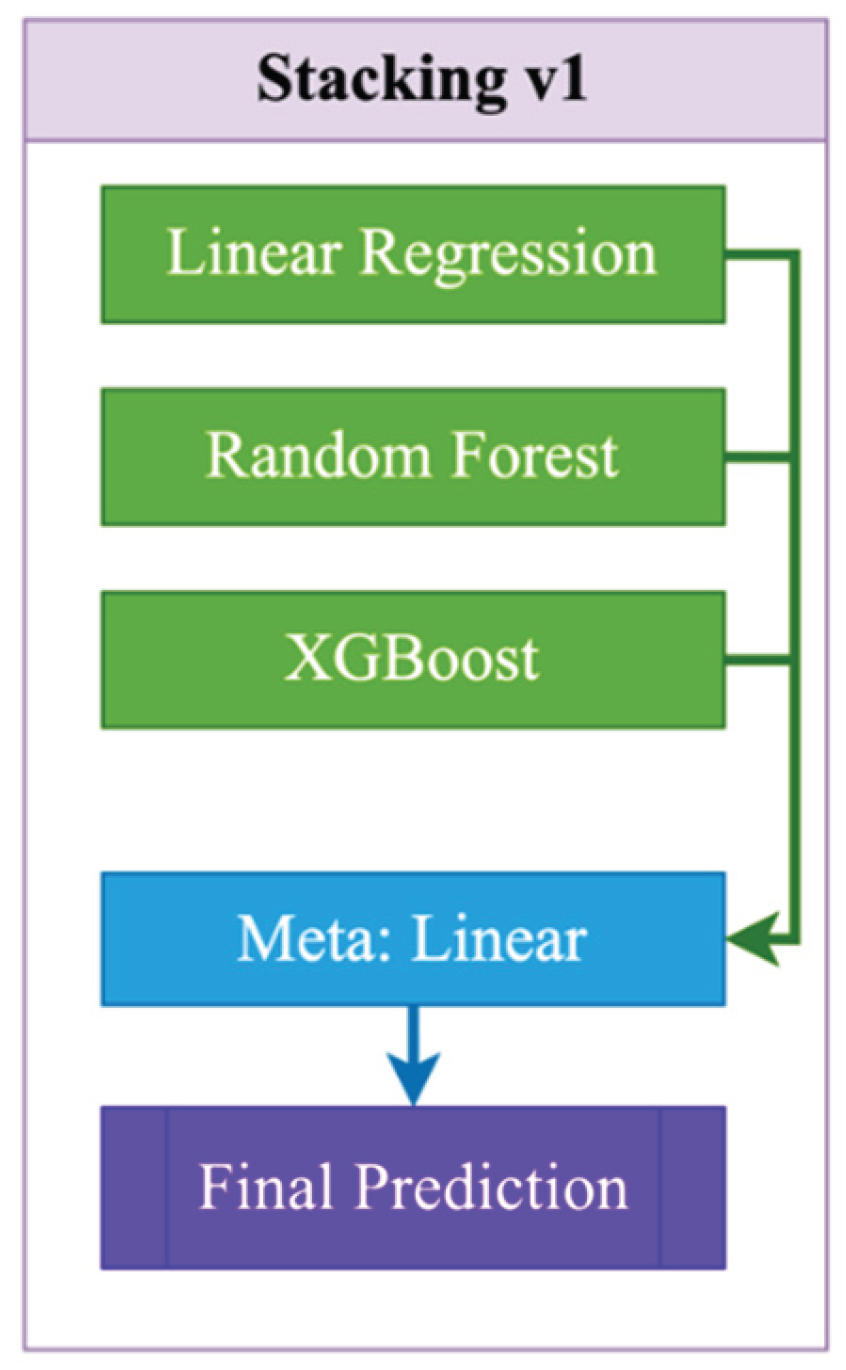
- Stacking. As shown in Figure 1, Stacking v1 is a simple two-level ensemble scheme in which base models (Linear Regression, Random Forest, and XGBoost) are independently trained on the original features (1):
3.3. Stages of Model Implementation
- n — number of observations (patients);
- d — number of initial features;
- feature matrix;
- feature vector for the i-th patient;
- target variable vector (ACT level, LBXSASSI).
- ▪
- — age;
- ▪
- — gender;
- ▪
- — weight;
- ▪
- — height;
- ▪
- — ferritin;
- ▪
- — homocysteine;
- ▪
- — total cholesterol;
- ▪
- — LDL-C;
- ▪
- — glucose;
- ▪
- — hemoglobin;
- ▪
- — creatinine;
- ▪
- — hs-CRP;
- ▪
- — alkaline phosphatase (ALP);
- ▪
- — gamma-GT;
- ▪
- — LDH;
- ▪
- — urea;
- ▪
- — uric acid;
- ▪
- — leukocytes;
- ▪
- average number of alcoholic drinks per day;
- ▪
- — physical activity.
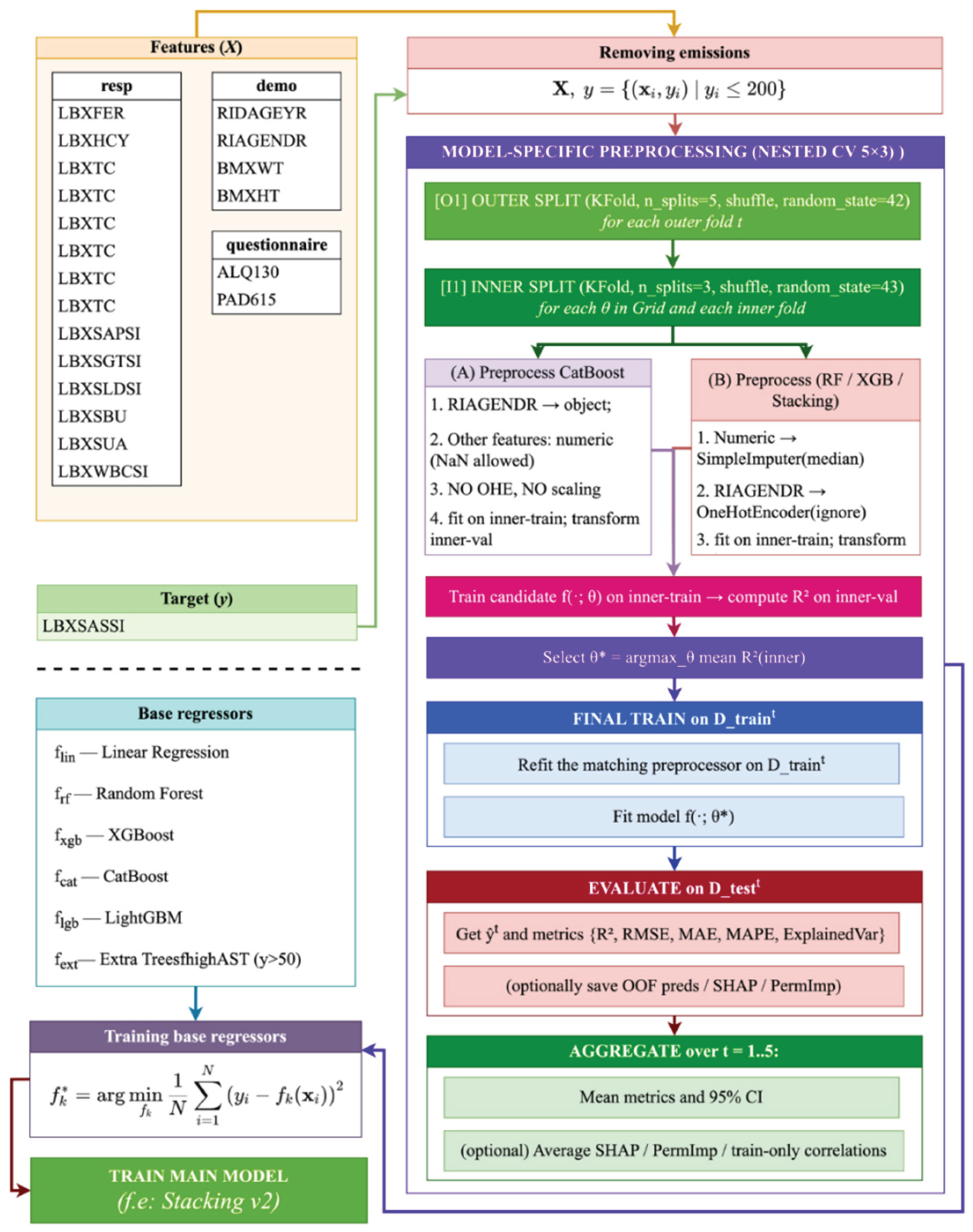
- For CatBoost: the gender feature (RIAGENDR) was converted to a categorical type (object), with missing values explicitly marked as "__MISSING__". All other features were retained in their numeric form. No one-hot encoding (OHE) or feature scaling was applied, as CatBoost natively handles both categorical variables and missing values.
- For Random Forest, XGBoost, and Stacking models: numerical features with missing values were imputed using SimpleImputer(strategy="median") to support tree-based models and meta-learners. The gender feature (RIAGENDR) was encoded using OneHotEncoder(handle_unknown="ignore"). Feature scaling was selectively applied only where necessary (e.g., for linear models or the meta-level in stacking).
- Loading and merging data. The first stage involves loading three tables containing clinical, demographic, and questionnaire data, after which they are combined using a unique patient identifier (SEQN), allowing for the formation of a single data structure for subsequent analysis and model building (4).
- Remove outliers. Removes records where the target variable (5)
- Removing gaps. Only those records are left where there are no gaps for the selected features and target variable (6):
- Transformation of categorical variables. Gender indicator is encoded using the one-hot method (7):
- Scaling of numerical features. For each numerical feature ()), except for categorical ones, standardization is applied (8):
- Training base regressors. For each base algorithm we train a regression function (9):
- — Linear Regression
- — Random Forest
- — XGBoost
- — CatBoost
- — LightGBM
- — Extra Trees
- local regressor for high AST values (trains only on cases with y>50)
- Linear correlation (10):
- pearman/Kendall: uses nonparametric measures for robustness.
- Mutual Information (11):
- SHAP values (12):
4. Results
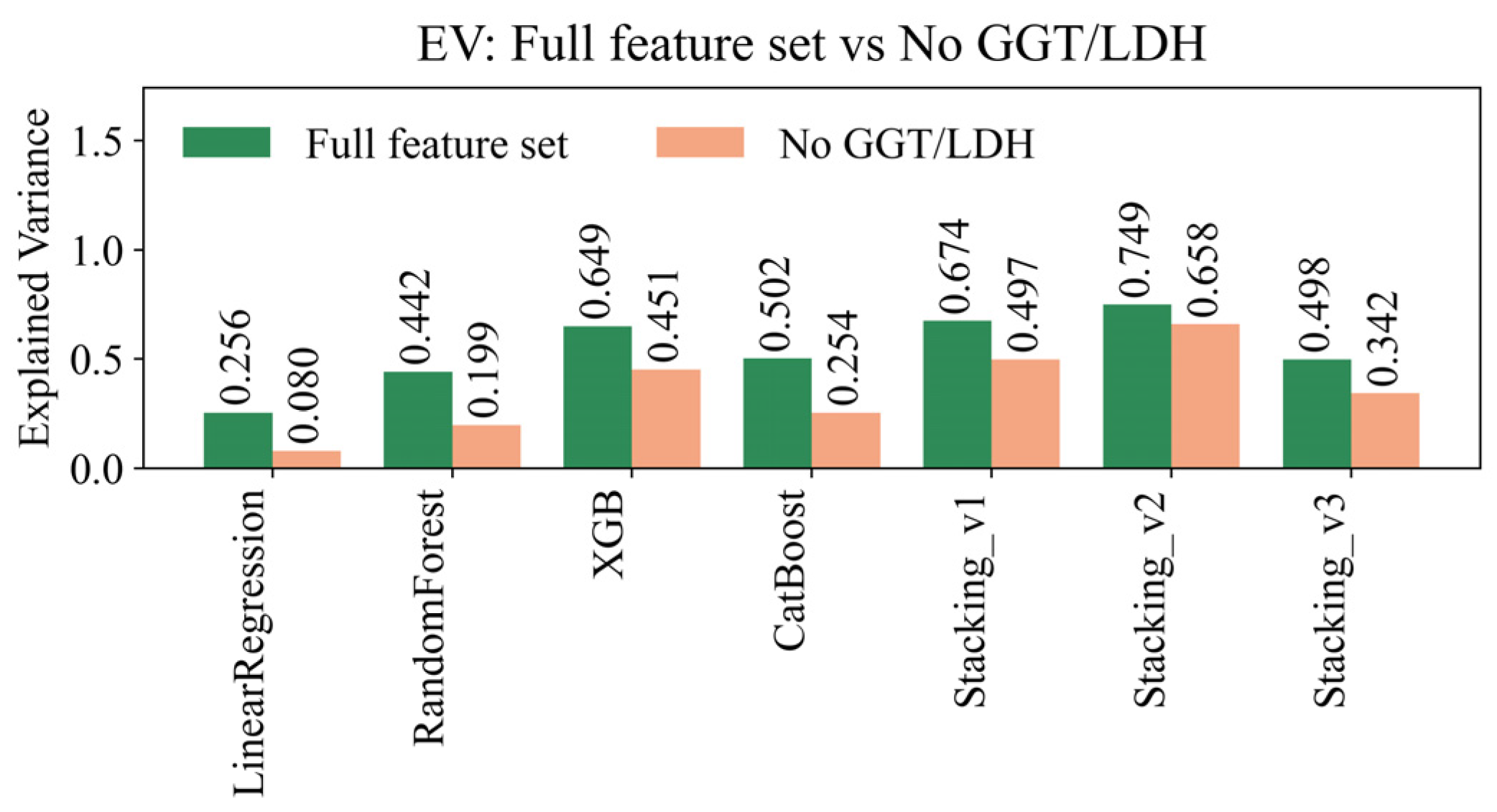
4.1. Assessment of Forecast Quality
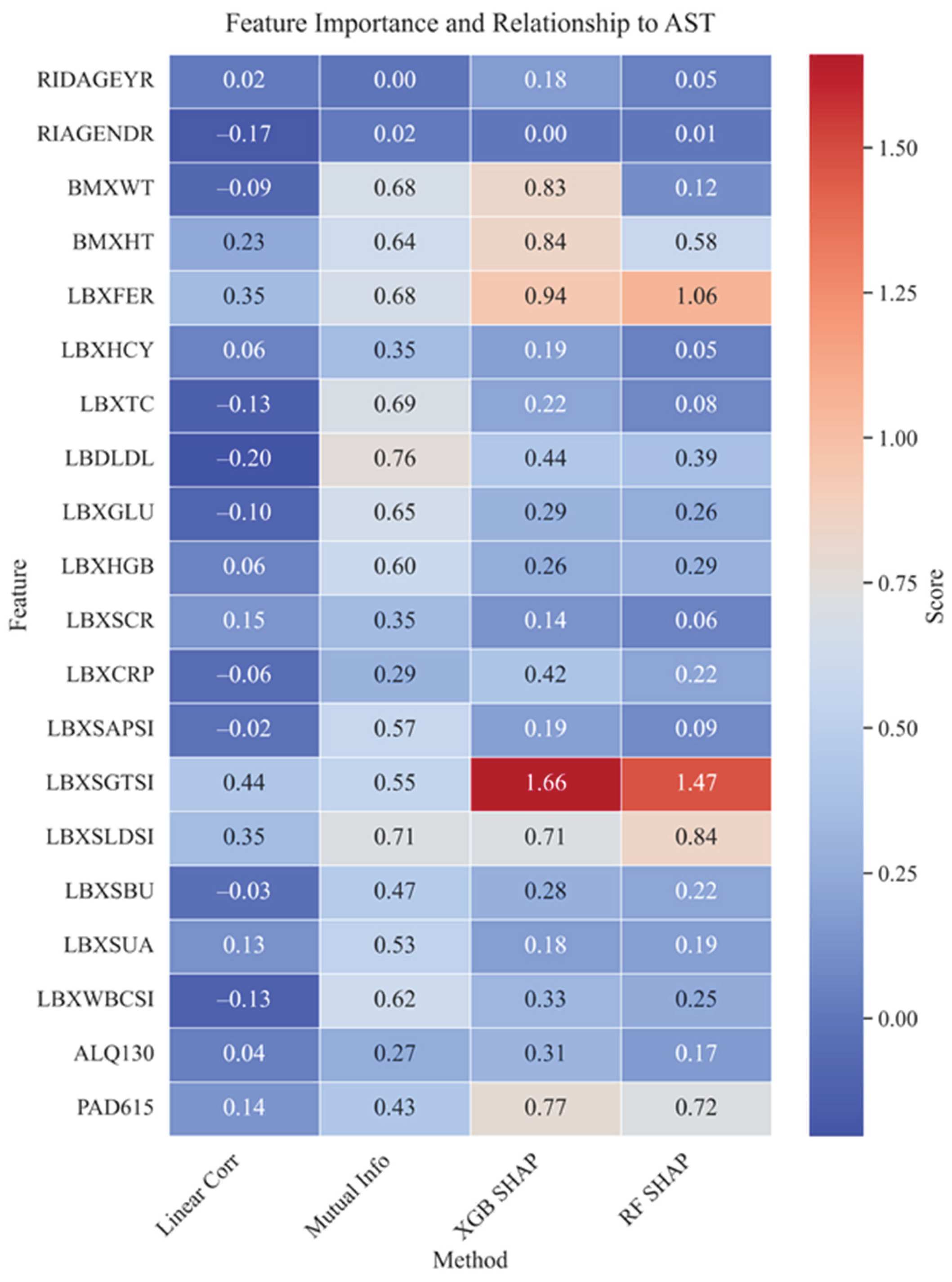
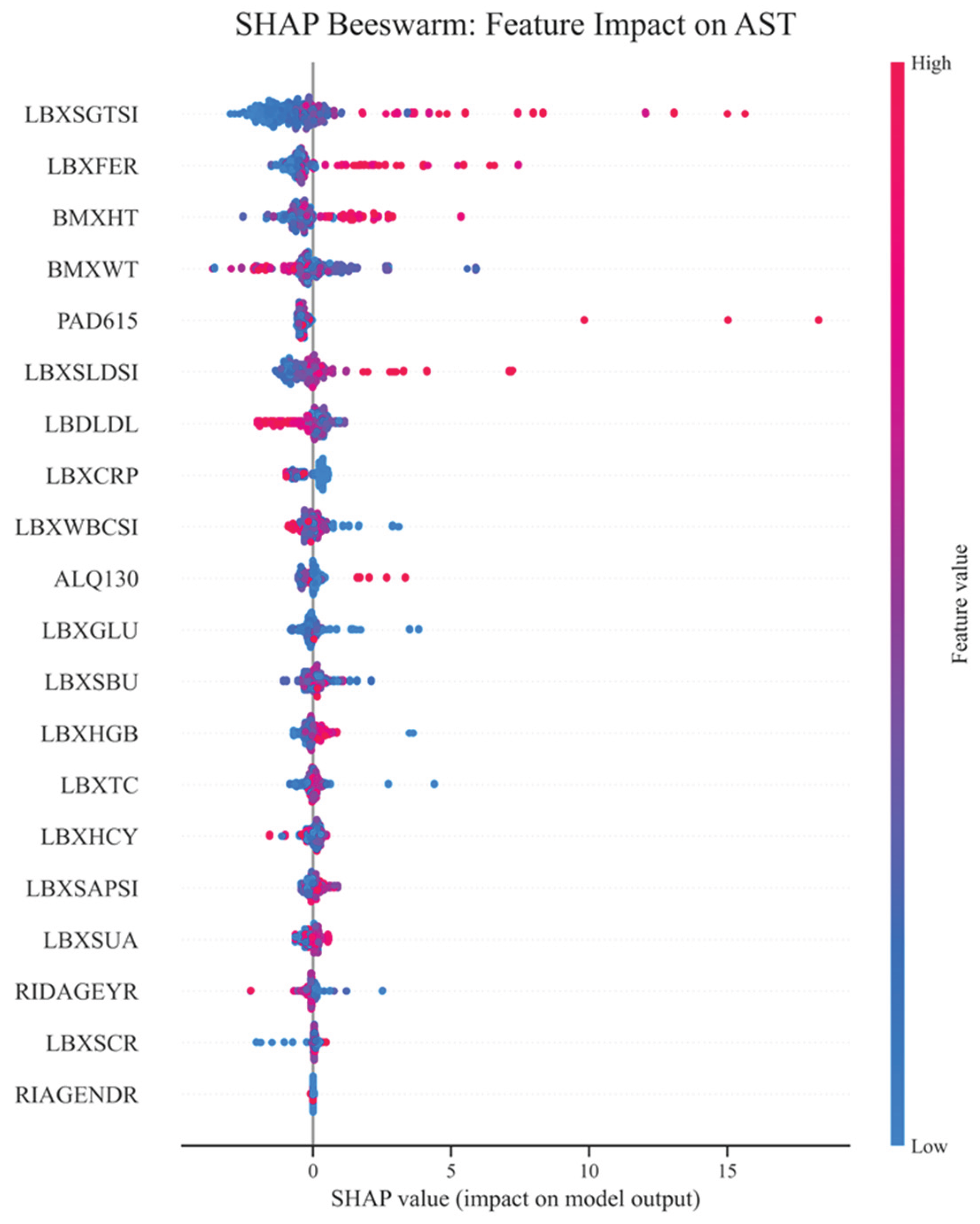
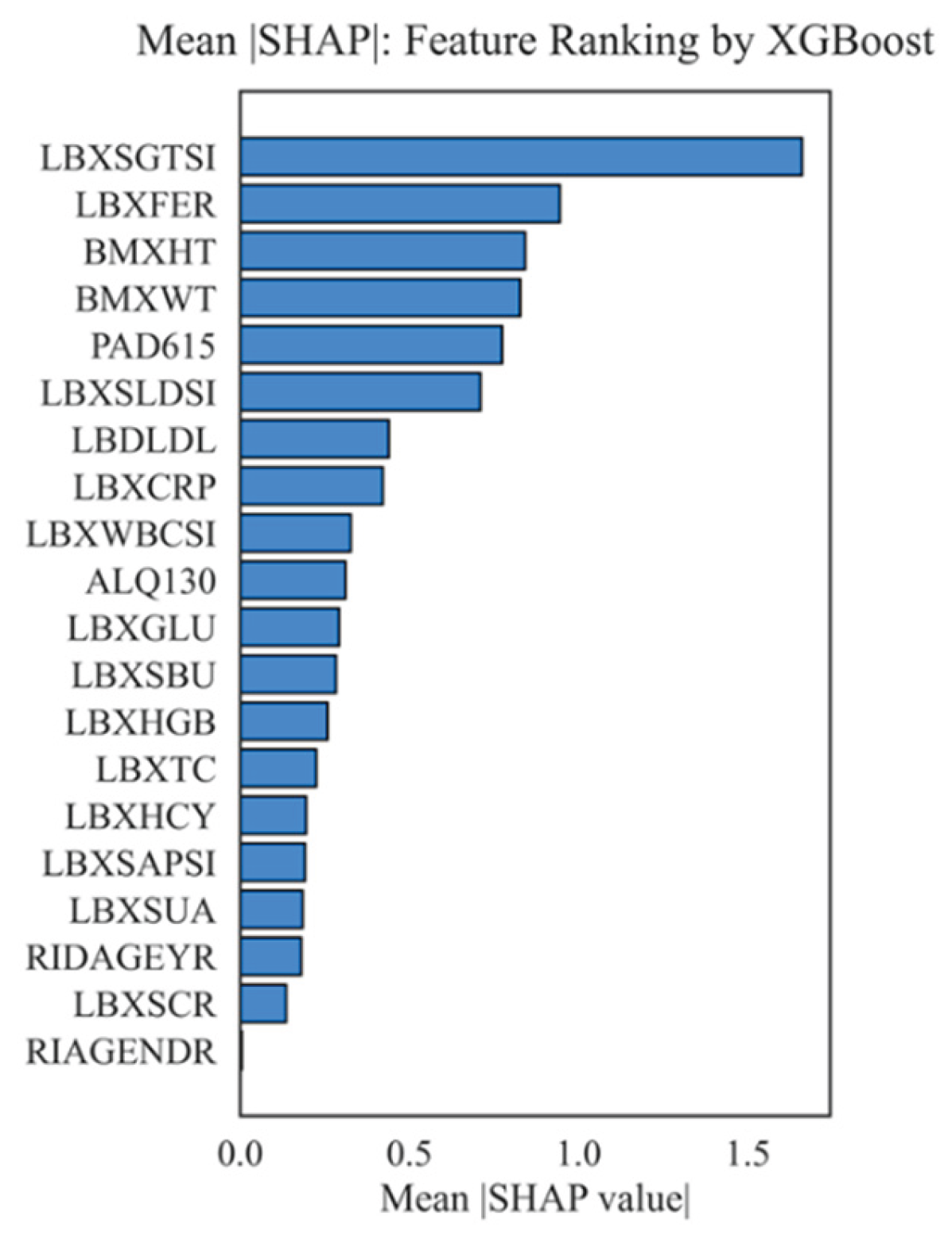
4.2. Importance of Features, Their Interactions, and Correlations
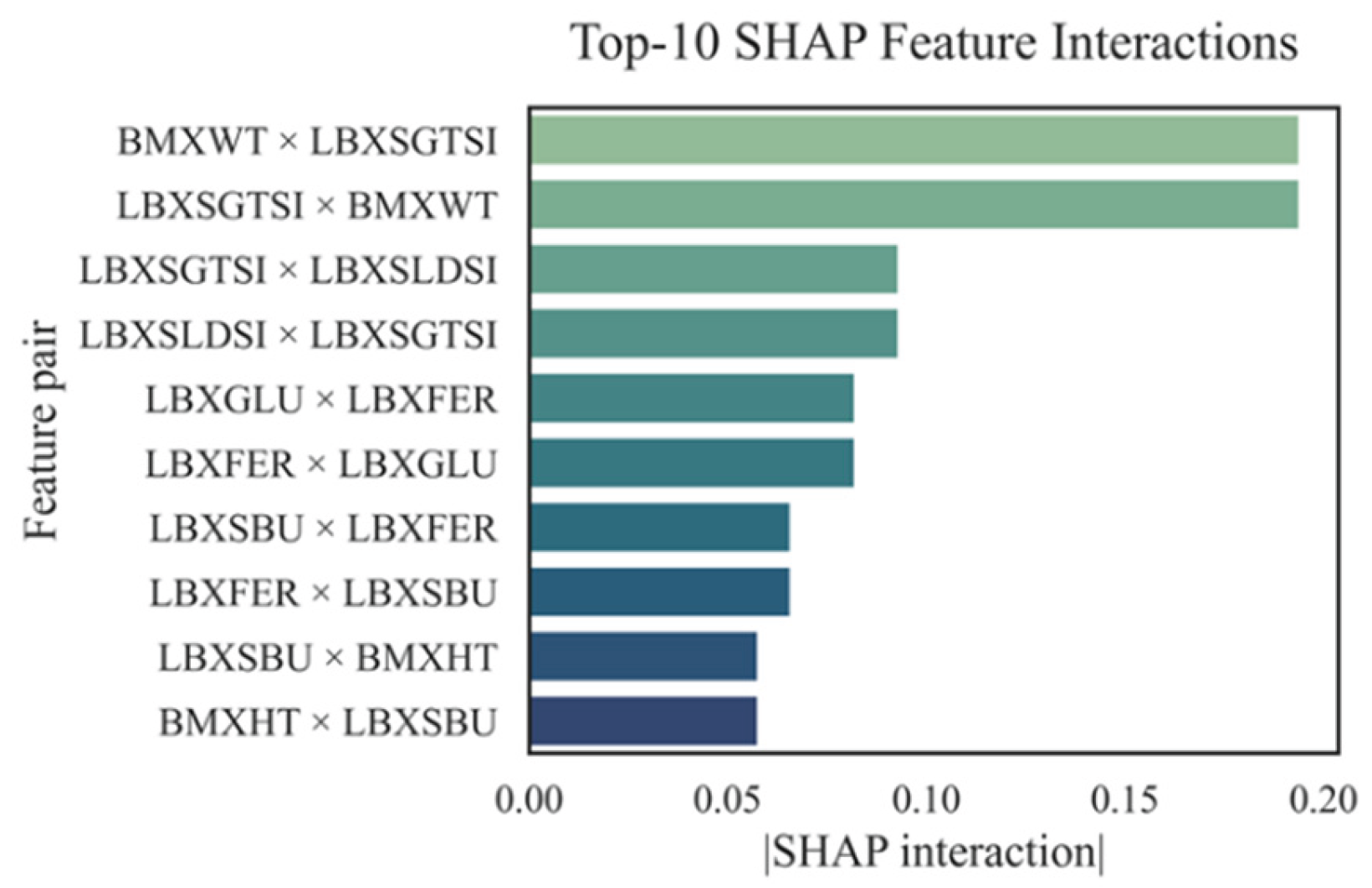
4.3. SHAP Interactions
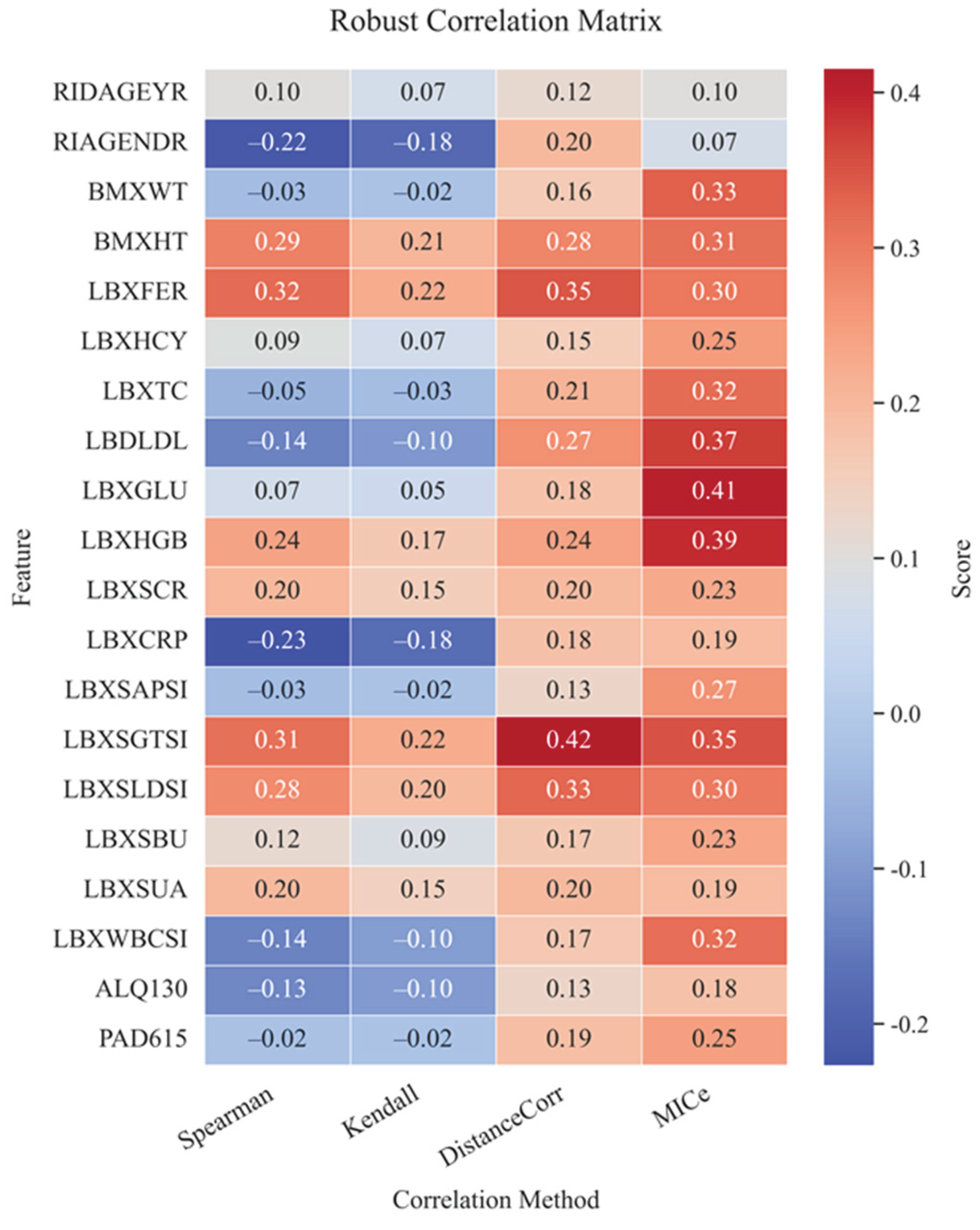
4.4. Correlation Analysis
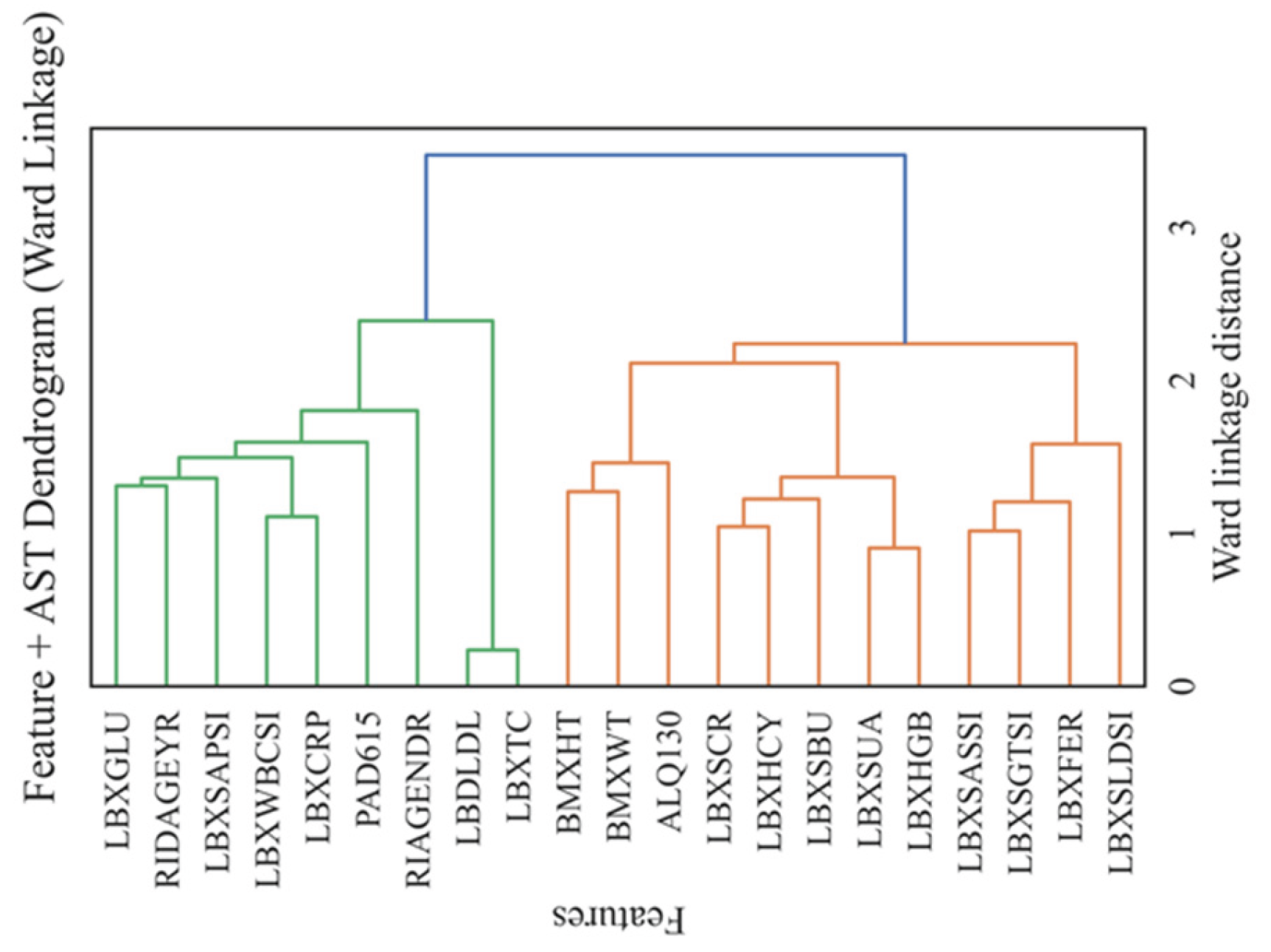
4.5. Clustering and Dendrogram
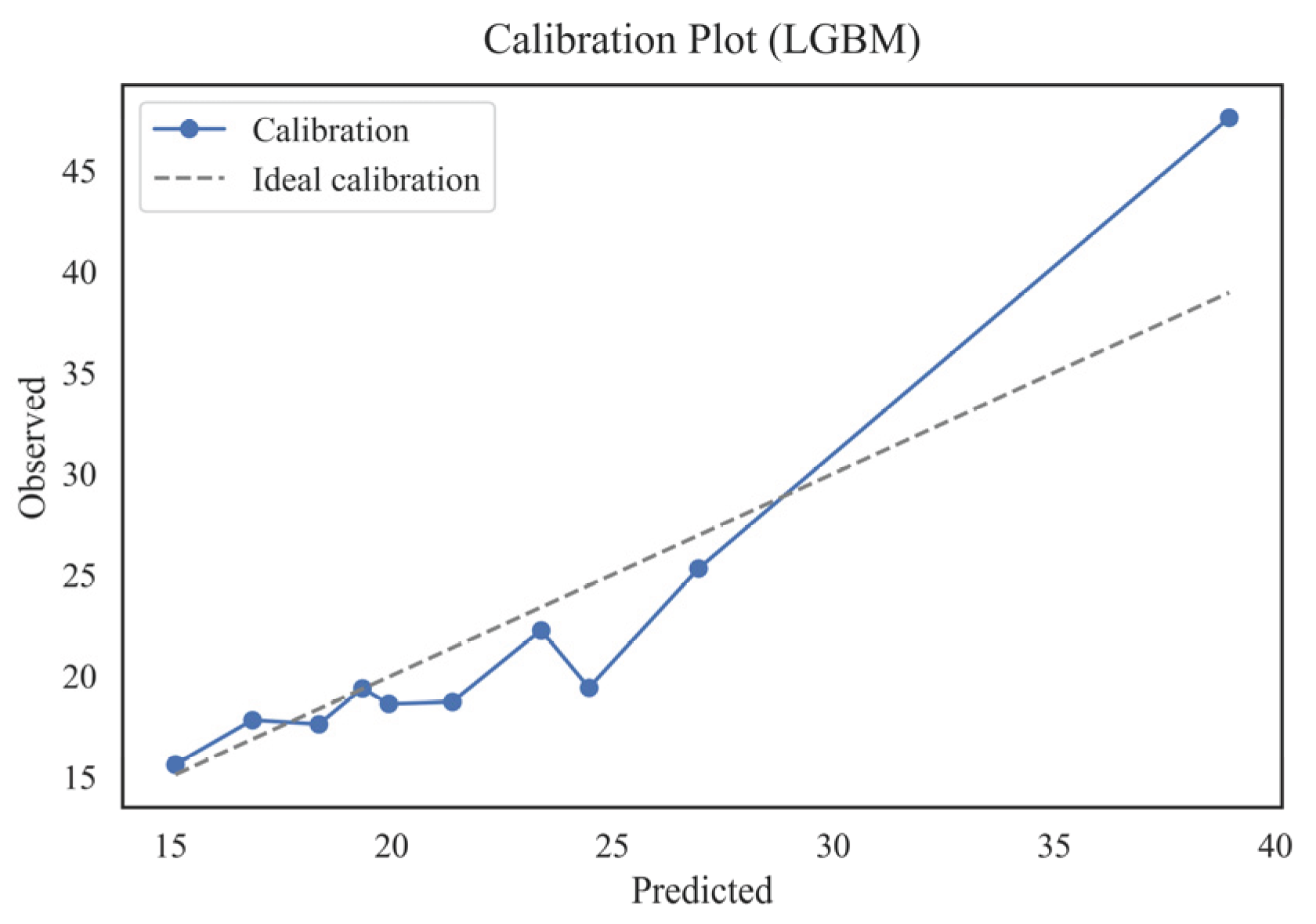
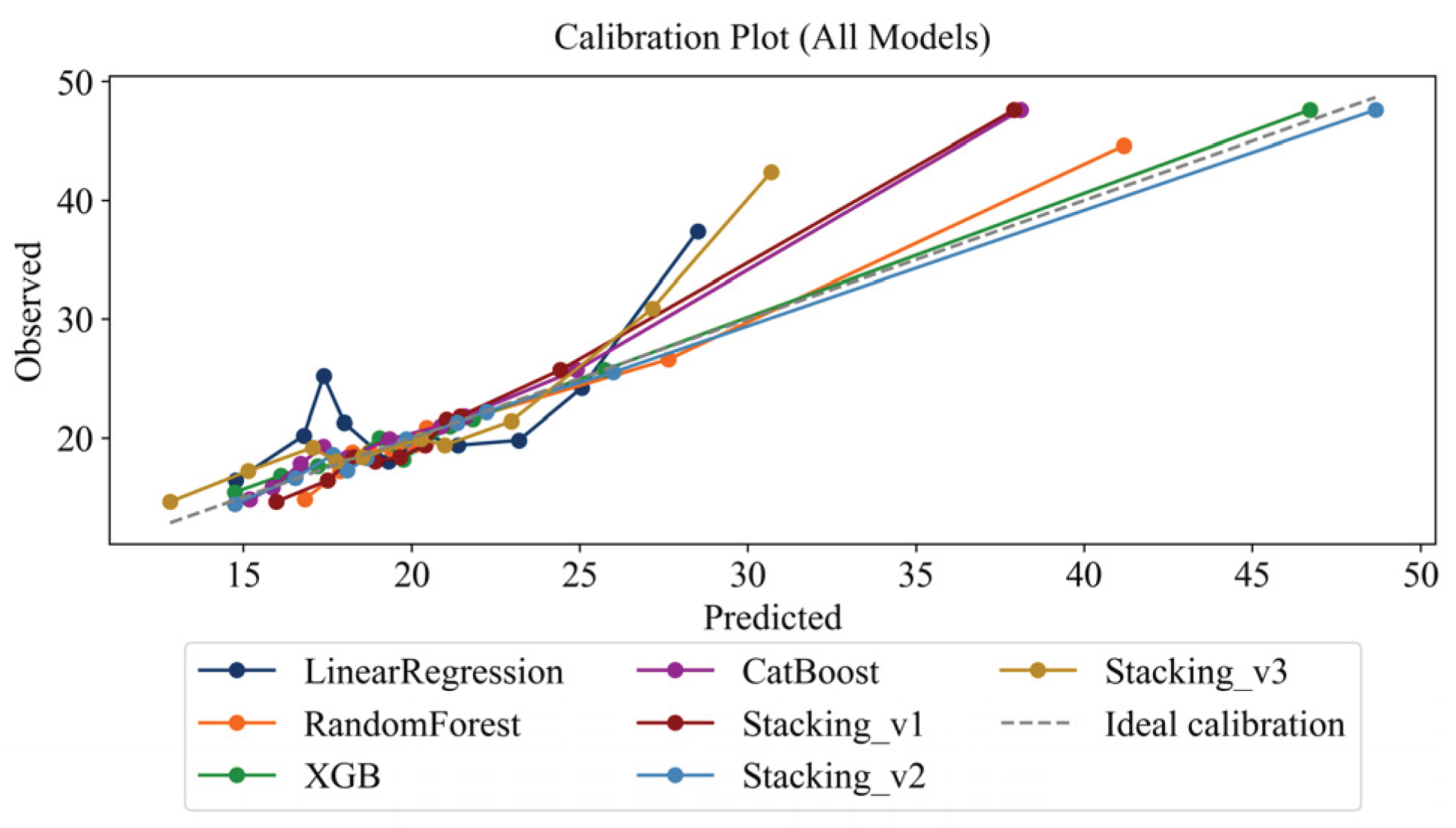
4.6. Assessing Interpretability and Calibration
- Stacking_v2 was the most stable and accurate model in terms of calibration;
- Deviations below 5% across all quantiles were considered clinically acceptable, given that the typical physiological variability of AST measurements is approximately 3–4 U/L.
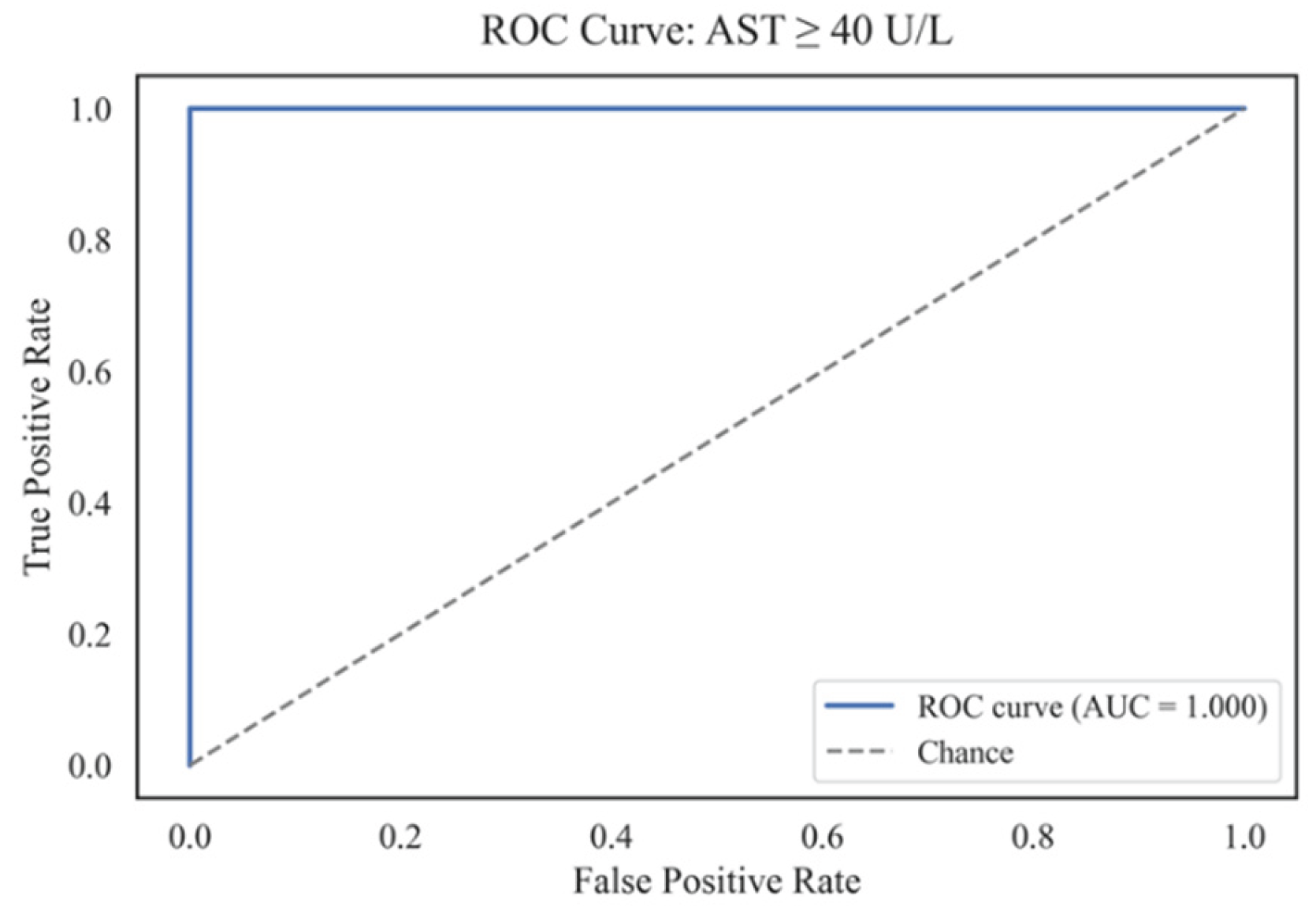
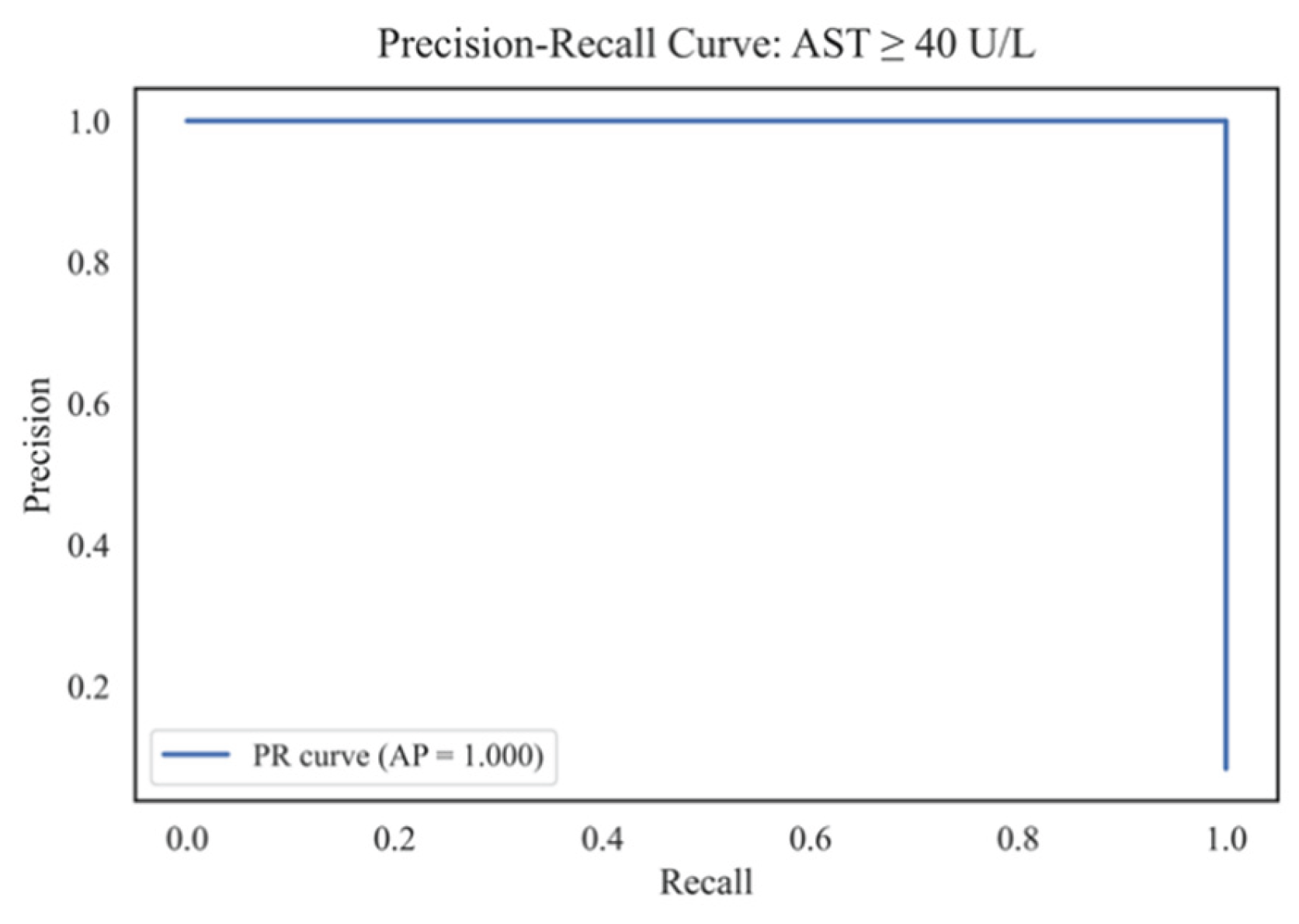
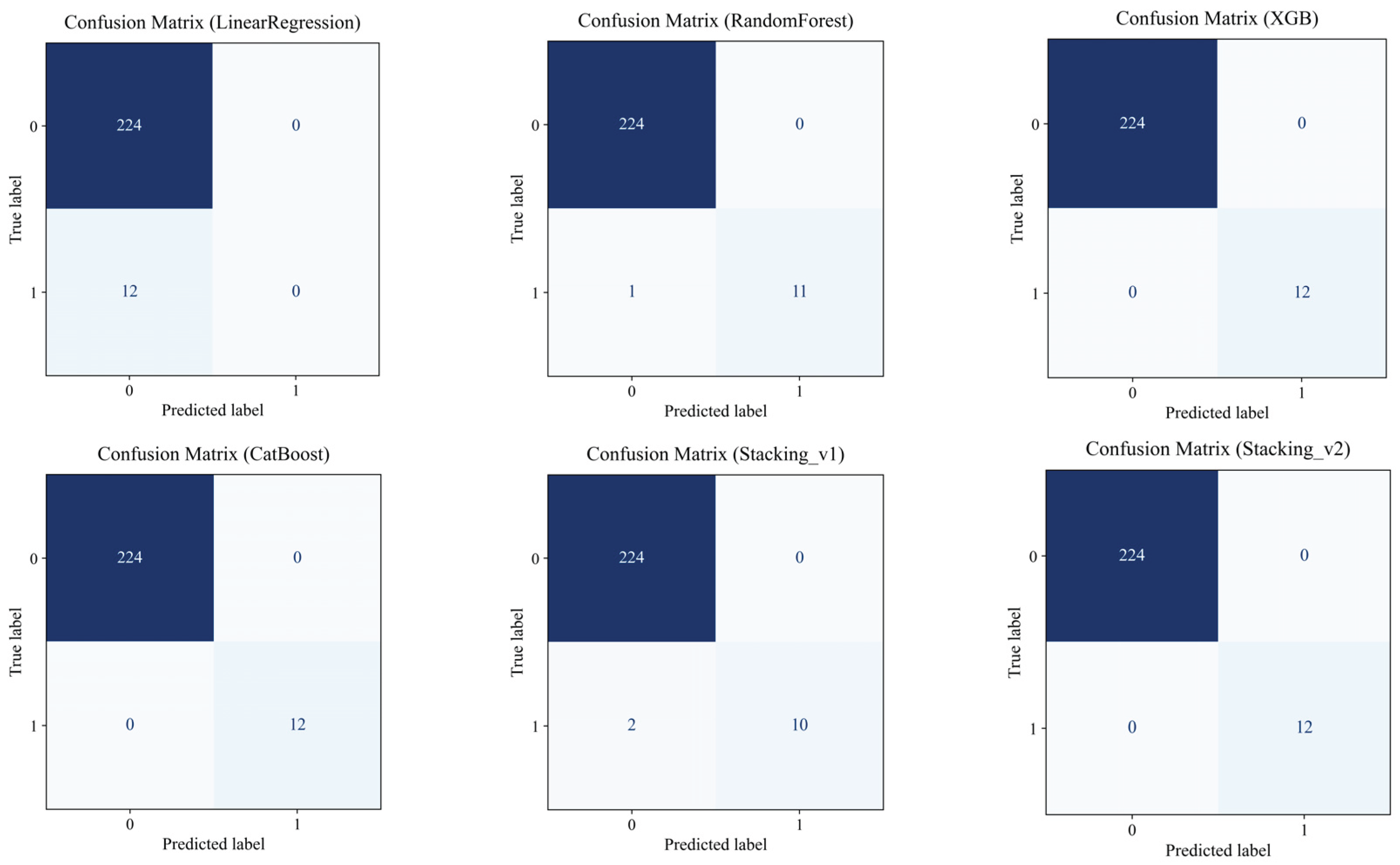
4.7. Predicting the Risk of Exceeding the AST Threshold
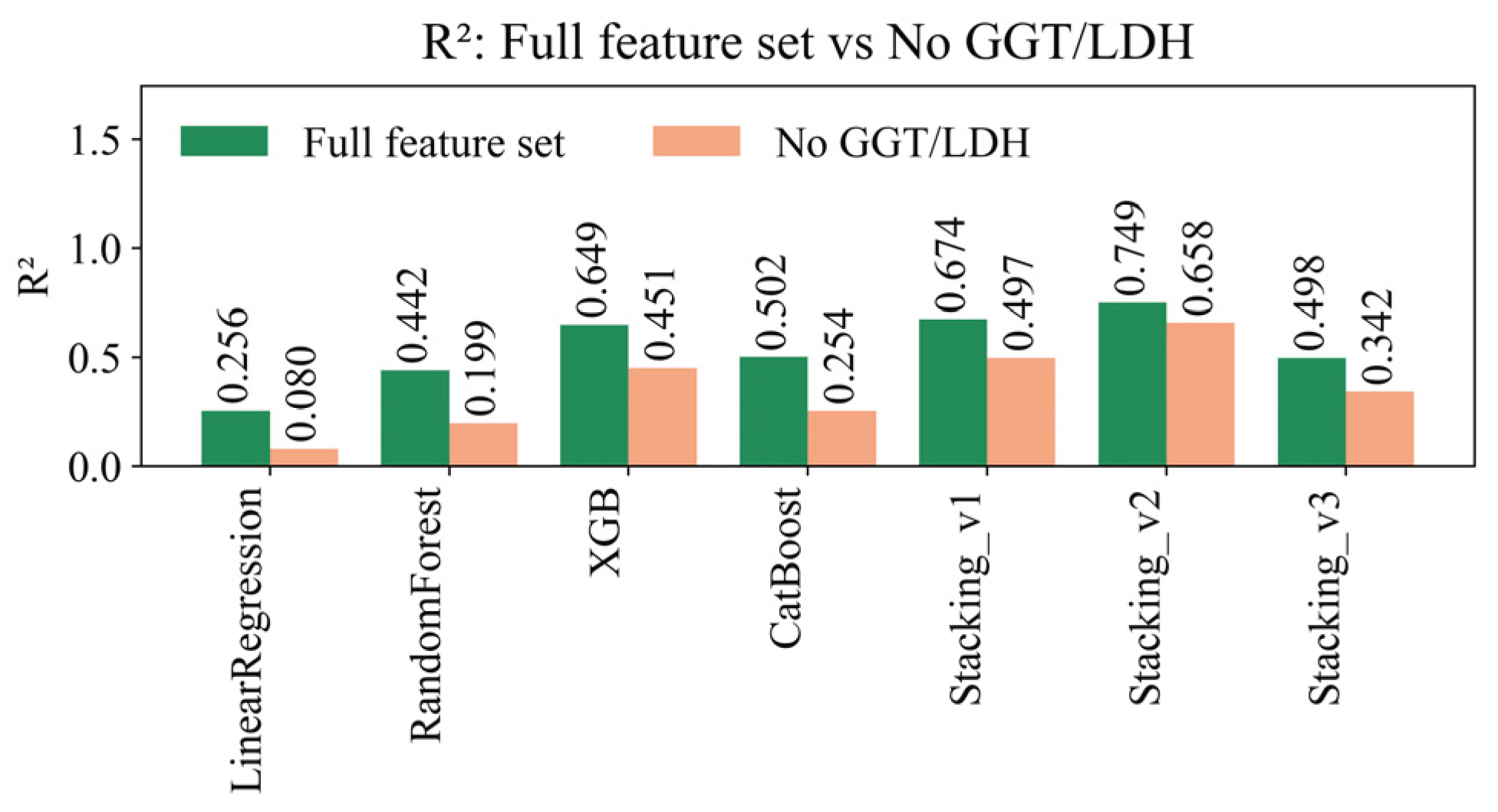
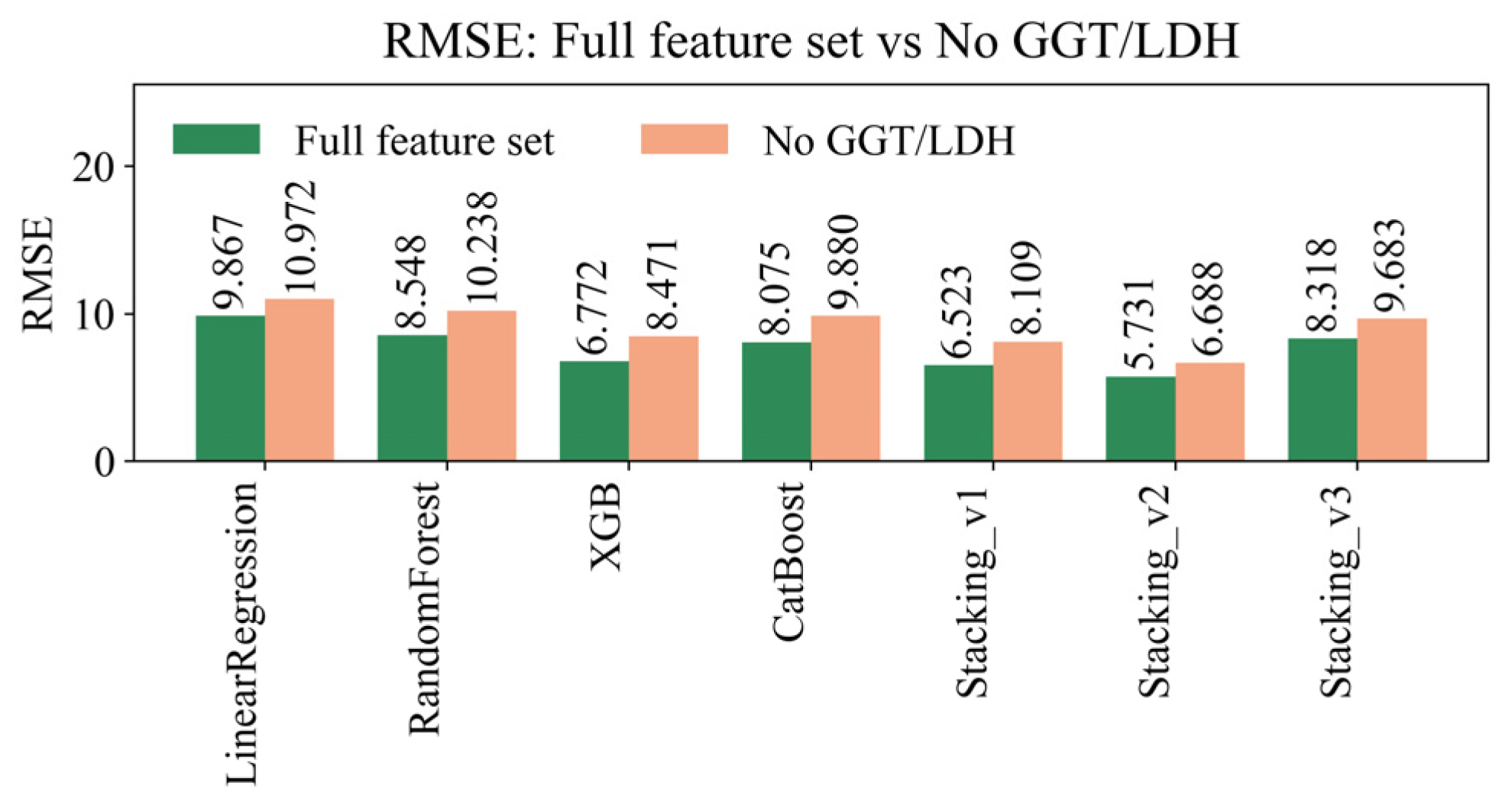
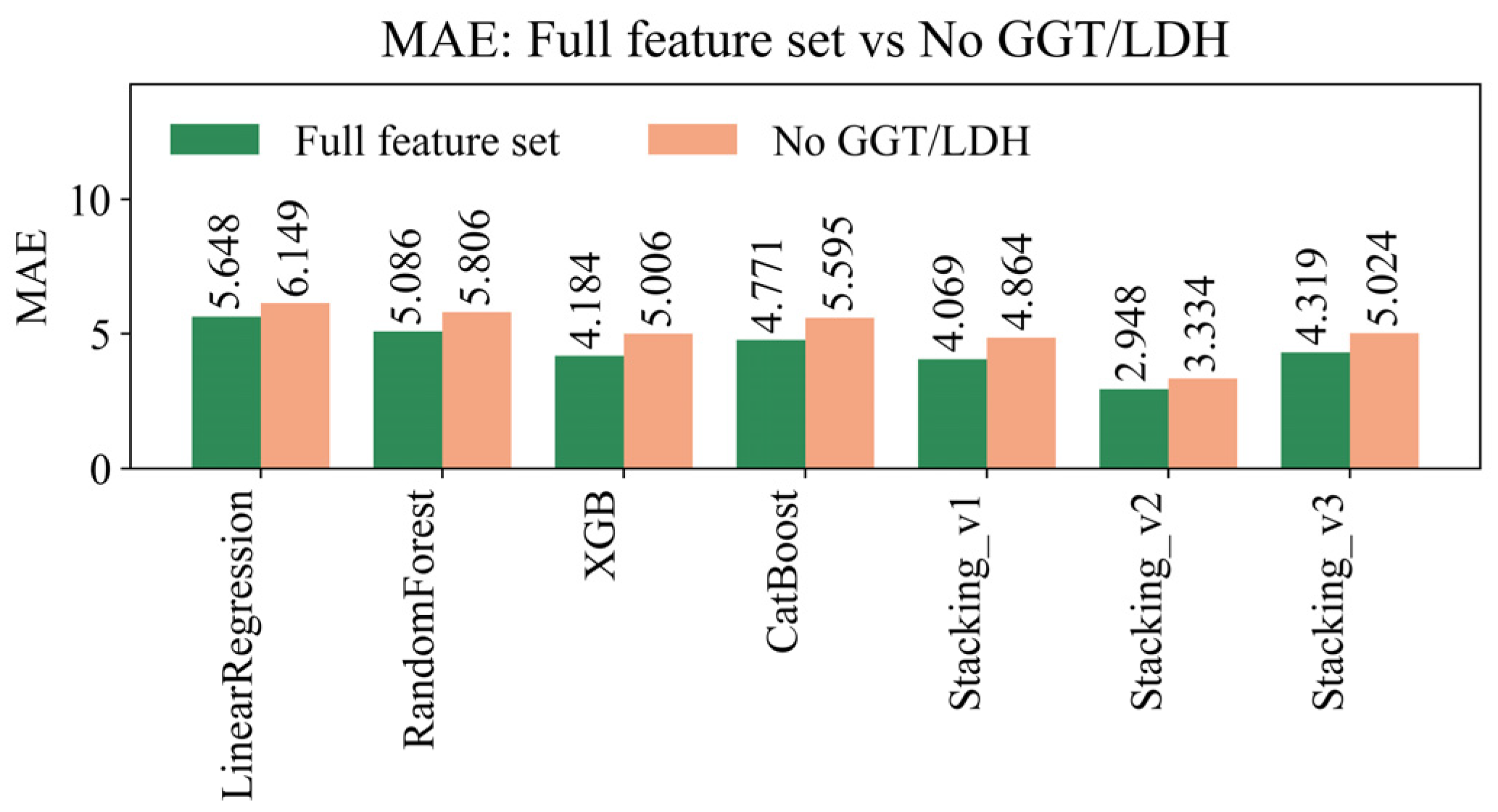
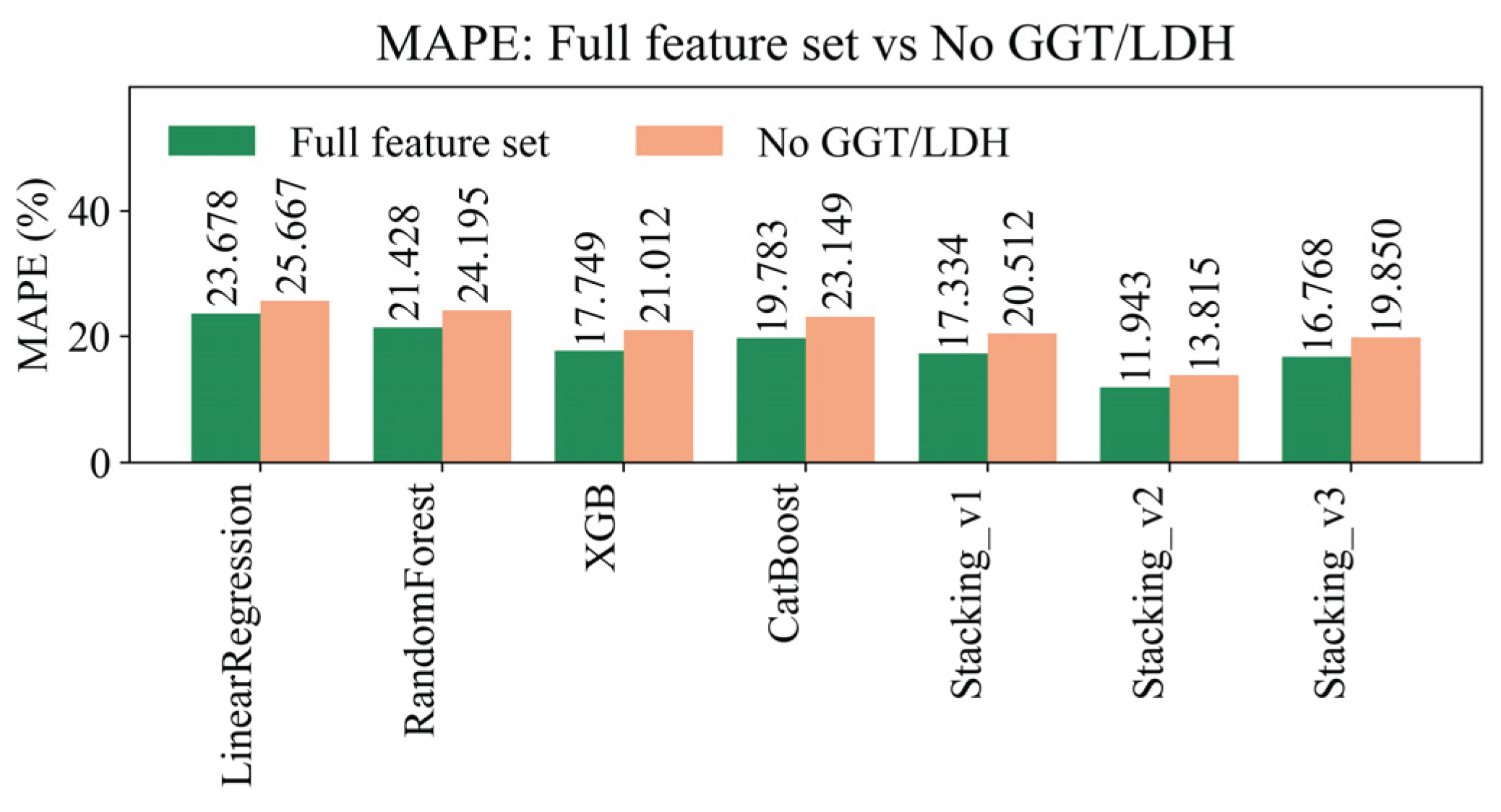
4.8. Sensitivity Analysis of Missing Data Handling Methods
| Model | R²_mean | RMSE_mean | MAE_mean | MAPE_mean |
|---|---|---|---|---|
| LinearRegression | 0.215 | 6.813 | 4.923 | 22.96 |
| RandomForest | 0.629 | 4.389 | 3.198 | 15.14 |
| XGB | 0.728 | 3.234 | 1.369 | 6.47 |
| CatBoost | 0.759 | 3.648 | 1.916 | 8.26 |
| Stacking_v1 | 0.690 | 3.668 | 2.241 | 10.72 |
| Stacking_v2 | 0.879 | 2.204 | 1.272 | 6.42 |
| Stacking_v3 | 0.615 | 5.130 | 2.766 | 10.82 |
4.9. Mediator Analysis
5. Discussion
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AST | Aspartate Aminotransferase |
| ALP | Alkaline Phosphatase |
| γ-GT | Gamma-Glutamyl Transferase |
| LDH | Lactate Dehydrogenase |
| hs-CRP | High-sensitivity C-Reactive Protein |
| BMI | Body Mass Index |
| MAE | Mean Absolute Error |
| RMSE | Root Mean Square Error |
| R² | Coefficient of Determination |
| MAPE | Mean Absolute Percentage Error |
| SHAP | SHapley Additive exPlanations |
| NHANES | National Health and Nutrition Examination Survey |
| RF | Random Forest |
| XGBoost | Extreme Gradient Boosting |
| CatBoost | Categorical Boosting |
| LGBM | Light Gradient Boosting Machine |
| MICe | Maximal Information Coefficient (enhanced version) |
| SEQN | Sequence Number (unique identifier in NHANES) |
| AUC | Area Under Curve |
| ROC | Receiver Operating Characteristic |
| PR-AUC | Precision-Recall Area Under Curve |
References
- Zhou, W.; Wang, Y.; Yu, H.; et al. Machine Learning Model Identifies Circulating Biomarkers Associated with Cardiovascular Disease. Sci. Rep. 2024, 14, 77352. [Google Scholar] [CrossRef]
- Roseiro, M.; Henriques, J.; Paredes, S.; Rocha, T.; Sousa, J. An Interpretable Machine Learning Approach to Estimate the Influence of Inflammation Biomarkers on Cardiovascular Risk Assessment. Comput. Methods Programs Biomed. 2023, 230, 107347. [Google Scholar] [CrossRef] [PubMed]
- Lüscher, T.F.; Wenzl, F.A.; D’Ascenzo, F.; Friedman, P.A.; Antoniades, C. Artificial Intelligence in Cardiovascular Medicine: Clinical Applications. Eur. Heart J. 2024, 45, 4291–4304. [Google Scholar] [CrossRef]
- da Costa, C.A.; Zeiser, F.A.; da Rosa Righi, R.; Antunes, R.S.; Alegretti, A.P.; Bertoni, A.P.; Rigo, S.J. Internet of Things and Machine Learning for Smart Healthcare. In IoT and ML for Information Management: A Smart Healthcare Perspective; Springer: Singapore, 2024; pp. 95–133. [Google Scholar] [CrossRef]
- Kwak, S.; Lee, H.J.; Kim, S.; Park, J.B.; Lee, S.P.; Kim, H.K.; Kim, Y.J. Machine Learning Reveals Sex-Specific Associations Between Cardiovascular Risk Factors and Incident Atherosclerotic Cardiovascular Disease. Sci. Rep. 2023, 13, 9364. [Google Scholar] [CrossRef]
- Ben-Assuli, O.; Ramon-Gonen, R.; Heart, T.; Jacobi, A.; Klempfner, R. Utilizing Shared Frailty with the Cox Proportional Hazards Regression: Post Discharge Survival Analysis of CHF Patients. J. Biomed. Inform. 2023, 140, 104340. [Google Scholar] [CrossRef]
- Guo, X.; Ma, M.; Zhao, L.; Wu, J.; Lin, Y.; Fei, F.; Ye, B. The Association of Lifestyle with Cardiovascular and All-Cause Mortality Based on Machine Learning: A Prospective Study from the NHANES. BMC Public Health 2025, 25, 319. [Google Scholar] [CrossRef]
- Miyachi, Y.; Ishii, O.; Torigoe, K. Design, Implementation, and Evaluation of the Computer-Aided Clinical Decision Support System Based on Learning-to-Rank: Collaboration Between Physicians and Machine Learning in the Differential Diagnosis Process. BMC Med. Inform. Decis. Mak. 2023, 23, 26. [Google Scholar] [CrossRef]
- Guo, L.; Tahir, A.M.; Zhang, D.; Wang, Z.J.; Ward, R.K. Automatic Medical Report Generation: Methods and Applications. APSIPA Trans. Signal Inf. Process. 2024, 13, e7. [Google Scholar] [CrossRef]
- Mesinovic, M.; Watkinson, P.; Zhu, T. Explainable AI for Clinical Risk Prediction: A Survey of Concepts, Methods, and Modalities. arXiv 2023, arXiv:2308.08407. [Google Scholar] [CrossRef]
- Sharma, P.; Sharma, P.; Sharma, K.; Varma, V.; Patel, V.; Sarvaiya, J.; Shah, K. Revolutionizing Utility of Big Data Analytics in Personalized Cardiovascular Healthcare. Bioengineering 2025, 12, 463. [Google Scholar] [CrossRef]
- Lai, T. Interpretable Medical Imagery Diagnosis with Self-Attentive Transformers: A Review of Explainable AI for Health Care. BioMedInformatics 2024, 4, 113–126. [Google Scholar] [CrossRef]
- Wang, Y.; Ni, B.; Xiao, Y.; Lin, Y.; Jiang, Y.; Zhang, Y. Application of Machine Learning Algorithms to Construct and Validate a Prediction Model for Coronary Heart Disease Risk in Patients with Periodontitis: A Population-Based Study. Front. Cardiovasc. Med. 2023, 10, 1296405. [Google Scholar] [CrossRef] [PubMed]
- Liao, W.; Voldman, J. A Multidatabase ExTRaction PipEline (METRE) for Facile Cross Validation in Critical Care Research. J. Biomed. Inform. 2023, 141, 104356. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Sun, X.; Zhao, H.; Xie, F.; Li, B.; Zhang, J. Biomarker Identification and Risk Assessment of Cardiovascular Disease Based on Untargeted Metabolomics and Machine Learning. Sci. Rep. 2024, 14, 25755. [Google Scholar] [CrossRef]
- Hu, Q.; Chen, Y.; Zou, D.; He, Z.; Xu, T. Predicting Adverse Drug Event Using Machine Learning Based on Electronic Health Records: A Systematic Review and Meta-Analysis. Front. Pharmacol. 2024, 15, 1497397. [Google Scholar] [CrossRef]
- Zhu, G.; Song, Y.; Lu, Z.; Yi, Q.; Xu, R.; Xie, Y.; Xiang, Y. Machine Learning Models for Predicting Metabolic Dysfunction-Associated Steatotic Liver Disease Prevalence Using Basic Demographic and Clinical Characteristics. J. Transl. Med. 2025, 23, 381. [Google Scholar] [CrossRef]
- Yang, B.; Lu, H.; Ran, Y. Advancing Non-Alcoholic Fatty Liver Disease Prediction: A Comprehensive Machine Learning Approach Integrating SHAP Interpretability and Multi-Cohort Validation. Front. Endocrinol. 2024, 15, 1450317. [Google Scholar] [CrossRef]
- Wang, Y.; Liu, L.; Wang, C. Trends in Using Deep Learning Algorithms in Biomedical Prediction Systems. Front. Neurosci. 2023, 17, 1256351. [Google Scholar] [CrossRef]
- Ali, G.; Mijwil, M.M.; Adamopoulos, I.; Buruga, B.A.; Gök, M.; Sallam, M. Harnessing the Potential of Artificial Intelligence in Managing Viral Hepatitis. Mesopotamian J. Big Data 2024, 2024, 128–163. [Google Scholar] [CrossRef]
- Yang, Y.; Liu, J.; Sun, C.; Shi, Y.; Hsing, J.C.; Kamya, A.; Zhu, S. Nonalcoholic Fatty Liver Disease (NAFLD) Detection and Deep Learning in a Chinese Community-Based Population. Eur. Radiol. 2023, 33, 5894–5906. [Google Scholar] [CrossRef]
- Khaled, O.M.; Elsherif, A.Z.; Salama, A.; Herajy, M.; Elsedimy, E. Evaluating Machine Learning Models for Predictive Analytics of Liver Disease Detection Using Healthcare Big Data. Int. J. Electr. Comput. Eng. 2025, 15, 1162–1174. [Google Scholar] [CrossRef]
- McGettigan, B.M.; Shah, V.H. Every Sheriff Needs a Deputy: Targeting Non-Parenchymal Cells to Treat Hepatic Fibrosis. J. Hepatol. 2024, 81, 20–22. [Google Scholar] [CrossRef] [PubMed]
- Farhadi, S.; Tatullo, S.; Ferrian, F. Comparative Analysis of Ensemble Learning Techniques for Enhanced Fatigue Life Prediction. Sci. Rep. 2025, 15, 11136. [Google Scholar] [CrossRef]
- Bing, Z. , Lemke C., Cheng L., et al. "Energy-efficient and damage-recovery slithering gait design for a snake-like robot based on reinforcement learning and inverse reinforcement learning," Neural Networks, vol. 138, pp. 212–222, 2021.
- Hao, X. , Wang R., Guo Y., et al. "Multi-modal Self-paced Locality Preserving Learning for Diagnosis of Alzheimer’s Disease," IEEE Transactions on Cognitive and Developmental Systems, vol. 14, no. 2, pp. 423–434, 2022.
- Deng, R. , Chen Z.M., Chen H., et al. "Learning to refine object boundaries," Neurocomputing, vol. 417, pp. 142–153, 2020.
- Xie, Z. , Chen X. "Subsampling for partial least-squares regression via an influence function," Knowledge-Based Systems, vol. 238, 107803, 2022.
- Liu, S.; Zhang, J.; Xiang, Y.; Zhou, W.; Xiang, D. A Study of Data Pre-Processing Techniques for Imbalanced Biomedical Data Classification. Int. J. Bioinform. Res. Appl. 2020, 16, 290–318. [Google Scholar] [CrossRef]
- Bumbu, M.G.; Niculae, M.; Ielciu, I.; Hanganu, D.; Oniga, I.; Benedec, D.; Marcus, I. Comprehensive Review of Functional and Nutraceutical Properties of Craterellus cornucopioides (L.) Pers. Nutrients 2024, 16, 831. [Google Scholar] [CrossRef]
- Wang, Z.; Gu, Y.; Huang, L.; Liu, S.; Chen, Q.; Yang, Y.; Ning, W. Construction of Machine Learning Diagnostic Models for Cardiovascular Pan-Disease Based on Blood Routine and Biochemical Detection Data. Cardiovasc. Diabetol. 2024, 23, 351. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The views, opinions, and data presented in this publication are solely those of the individual author(s) and contributors. They do not necessarily reflect the views of the publisher and/or the editorial board. The publisher and editors disclaim any responsibility for any harm, damage, or injury to individuals or property arising from the application of any ideas, methods, instructions, or products discussed in this content. |
| Ref. | Study Focus | Methods | Key Findings | Identified Gaps |
|---|---|---|---|---|
| [16] | Predicting liver enzyme elevation in RA patients on methotrexate | Random Forest classifier on EHR data | The ML model accurately predicts transaminase elevation | Specific to RA patients, limited generalizability |
| [17] | ML models for MASLD prediction using demographic and clinical data | Comparison of 10 ML algorithms, including XGBoost and Random Forest | High accuracy in MASLD screening; accessible features | Did not focus on AST-specific prediction |
| [18] | ML with SHAP for NAFLD prediction | ML models with SHAP interpretability | Robust predictive tool for NAFLD; high accuracy and generalizability | Lacks longitudinal data and lifestyle factors |
| [19] | ML models for cardiovascular disease diagnosis using routine blood tests | Logistic Regression, Random Forest, SVM, XGBoost, DNN | Effective diagnosis using accessible blood data; SHAP for interpretation | Focused on cardiovascular diseases, not liver-specific |
| [20] | ML models in liver transplantation prognostication | A systematic review of ML applications | ML models outperform traditional scoring systems in predicting post-transplant complications. | Emphasis on transplantation, not general AST prediction |
| [21] | Early liver disease prediction using deep learning | Deep learning algorithms | A promising approach for rapid and accurate liver disease diagnosis | Requires further validation and integration into clinical practice |
| [22] | Comparison of ML models for liver disease detection using big data | Evaluation of three ML models on 32,000 records | Enhanced prediction and management of liver diseases | Specific models and features are not detailed |
| [23] | ML model to predict liver-related outcomes post-hepatitis B cure | ML-based risk prediction model | Accurate forecasting of liver-related outcomes after functional cure | Focused on hepatitis B, not general AST prediction |
| [24] | Comparative analysis of ensemble learning techniques for fatigue life prediction | Boosting, stacking, bagging vs. linear regression and KNN | Ensemble models outperform traditional methods in prediction tasks | Application in fatigue life; relevance to AST prediction, indirect |
| Model | Type/Role | Key Settings |
|---|---|---|
| Linear Regression | Basic reference point | fit_intercept=True |
| Random Forest | Ensemble of trees | n_estimators=300, max_depth=8, random_state=42 |
| XGBoost | Gradient Boosting | n_estimators=300, max_depth=8, learning_rate=0.05, objective="reg:squarederror", random_state=42 |
| CatBoost | Robust Boosting (Native Categories) | iterations=1200, depth=6, learning_rate=0.03, loss_function ∈ {Huber:δ=2.0, RMSE} — is selected in the internal CV |
| Stacking_v1 | Base: Linear + RF + XGB; meta: LinearRegression | passthrough=False |
| Stacking_v2 | Base: CatBoost + LGBM + ExtraTrees; Meta: Ridge | LGBM(n_estimators=2000, lr=0.015, subsample=0.8, colsample_bytree=0.7, reg_lambda=0.8); ExtraTrees(n_estimators=700, max_features="sqrt", min_samples_leaf=2); Ridge(alpha=1.0) |
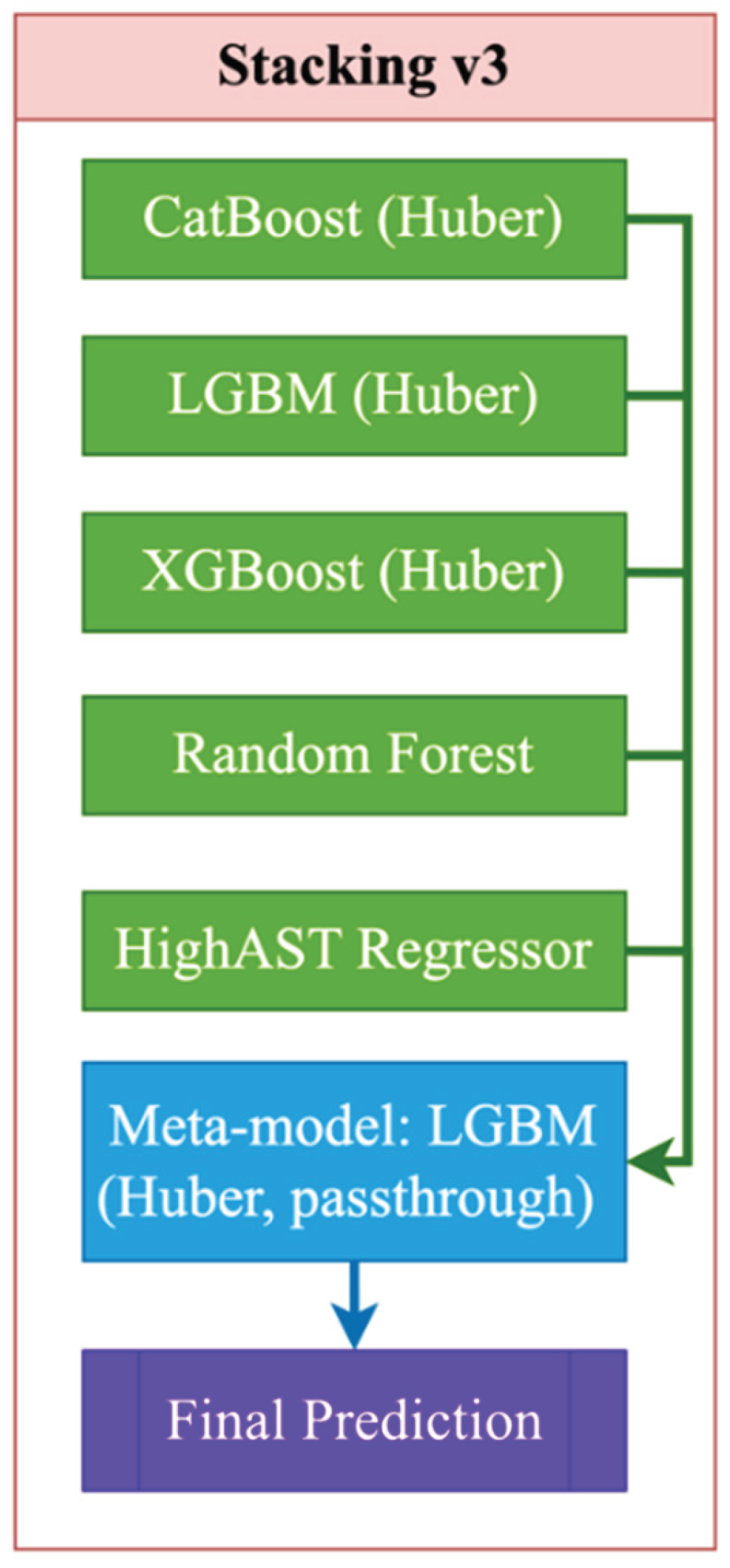
| Stacking_v3 | Base: CatBoost(Huber) + LGBM(Huber) + XGB + RF + HighASTRegressor; Meta: LGBM(Huber) | HighASTRegressor: y>50, LGBM(objective="huber", n_estimators=400, num_leaves=32); meta: LGBM(objective="huber", n_estimators=400, num_leaves=32), passthrough=True |
| Variable | N | Min | 25% | Median | 75% | Max | IQR |
|---|---|---|---|---|---|---|---|
| RIDAGEYR | 201 850 | 0.00 | 17.00 | 34.00 | 57.00 | 90.00 | 40.00 |
| RIAGENDR | 201 850 | 1.00 | 1.00 | 2.00 | 2.00 | 2.00 | 1.00 |
| BMXWT | 199 811 | 21.80 | 61.20 | 72.90 | 86.50 | 371.00 | 25.30 |
| BMXHT | 200 050 | 118.50 | 159.00 | 166.00 | 173.50 | 206.50 | 14.50 |
| LBXFER | 145 110 | 1.04 | 28.40 | 61.00 | 133.00 | 3 234.00 | 104.60 |
| LBXHCY | 99 679 | 1.65 | 6.00 | 7.67 | 9.87 | 156.30 | 3.87 |
| LBXTC | 201 601 | 59.00 | 161.00 | 188.00 | 220.00 | 813.00 | 59.00 |
| LBDLDL | 89 515 | 9.00 | 88.00 | 111.00 | 138.00 | 629.00 | 50.00 |
| LBXGLU | 123 350 | 21.00 | 89.30 | 97.00 | 109.30 | 686.20 | 20.00 |
| LBXHGB | 200 863 | 4.95 | 13.00 | 14.02 | 15.10 | 19.90 | 2.10 |
| LBXSCR | 201 844 | 0.10 | 0.70 | 0.90 | 1.10 | 17.80 | 0.40 |
| LBXCRP | 177 089 | 0.01 | 0.13 | 0.21 | 0.41 | 29.60 | 0.28 |
| LBXSAPSI | 201 841 | 7.00 | 62.00 | 78.00 | 101.00 | 1 378.00 | 39.00 |
| LBXSGTSI | 185 105 | 1.00 | 13.00 | 19.00 | 29.00 | 2 274.00 | 16.00 |
| LBXSLDSI | 201 729 | 4.00 | 122.00 | 141.00 | 166.00 | 1 274.00 | 44.00 |
| LBXSBU | 201 848 | 1.00 | 10.00 | 12.00 | 16.00 | 122.00 | 6.00 |
| LBXSUA | 201 847 | 0.20 | 4.30 | 5.20 | 6.20 | 18.00 | 1.90 |
| LBXWBCSI | 200 867 | 1.40 | 5.70 | 6.90 | 8.40 | 400.00 | 2.70 |
| ALQ130 | 50 467 | 1.00 | 1.00 | 2.00 | 3.00 | 999.00 | 2.00 |
| PAD615 | 9 190 | 10.00 | 60.00 | 120.00 | 240.00 | 9 999.00 | 180.00 |
| LBXSASSI | 201 850 | 6.00 | 18.00 | 21.00 | 26.00 | 200.00 | 8.00 |
| Model | Brief Description | Key parameters |
|---|---|---|
| Linear Regression | Basic Interpretable Model | default |
| Random Forest | Decision Tree Ensemble | n_estimators=300, max_depth=8, random_state=42 |
| XGBoost | Gradient Boosting | n_estimators=300, max_depth=8, learning_rate=0.05, objective='reg:squared error, random_state=42 |
| CatBoost | Robust Gradient Boosting | iterations=1200, depth=6, learning_rate=0.03, loss_function='Huber:delta=2.0', random_state=42 |
| Stacking_v1 | Linear + RF + XGBoost | meta=Linear Regression |
| Stacking_v2 | CatBoost + LGBM + ExtraTrees | meta=Ridge Regression (alpha=1.0) |
| Stacking_v3 | CatBoost + LGBM + XGB + RF + high_AST | meta=LGBM (huber), passthrough=True, регрессoр “high_AST”: LGBM (objective='huber', n_estimators=400) |
| Advanced Stacking | Specialized regressor for high_AST | LGBM (separately on the subsample with high AST) |
| Metric | LinearRegression | RandomForest | XGB | CatBoost | Stacking_v1 | Stacking_v2 | Stacking_v3 |
|---|---|---|---|---|---|---|---|
| R2 (mean) | 0.215 | 0.629 | 0.728 | 0.759 | 0.690 | 0.916 | 0.612 |
| R2 (std) | 0.285 | 0.253 | 0.360 | 0.135 | 0.333 | 0.143 | 0.087 |
| RMSE (mean) | 6.813 | 4.389 | 3.234 | 3.648 | 3.668 | 2.274 | 5.150 |
| RMSE (std) | 0.976 | 1.084 | 1.898 | 0.832 | 1.675 | 0.925 | 1.634 |
| MAE (mean) | 4.923 | 3.198 | 1.369 | 1.916 | 2.241 | 1.303 | 2.779 |
| MAE (std) | 0.562 | 0.673 | 0.807 | 0.381 | 0.881 | 0.507 | 0.601 |
| MAPE (mean) | 22.96 | 15.14 | 6.47 | 8.26 | 10.72 | 6.564 | 10.84 |
| MAPE (std) | 2.02 | 3.67 | 4.18 | 2.01 | 4.89 | 2.863 | 1.77 |
| ExplainedVar (mean) | 0.217 | 0.659 | 0.744 | 0.771 | 0.716 | 0.926 | 0.633 |
| ExplainedVar (std) | 0.285 | 0.233 | 0.331 | 0.136 | 0.290 | 0.097 | 0.087 |
| Model | R² (±95% CI) | RMSE (±95% CI) | MAE (±95% CI) | MAPE (±95% CI) | EV (±95% CI) |
|---|---|---|---|---|---|
| LinearRegression | 0.2558 ± 0.0128 | 9.8675 ± 0.2617 | 5.6477 ± 0.0506 | 23.6785 ± 0.0871 | 0.2559 ± 0.0128 |
| RandomForest | 0.4416 ± 0.0075 | 8.5477 ± 0.2190 | 5.0857 ± 0.0524 | 21.4277 ± 0.1158 | 0.4416 ± 0.0075 |
| XGBoost | 0.6492 ± 0.0143 | 6.7716 ± 0.1559 | 4.1842 ± 0.0451 | 17.7494 ± 0.1199 | 0.6492 ± 0.0143 |
| CatBoost | 0.5017 ± 0.0071 | 8.0747 ± 0.2146 | 4.7712 ± 0.0474 | 19.7826 ± 0.1306 | 0.5017 ± 0.0071 |
| Stacking_v1 | 0.6743 ± 0.0174 | 6.5227 ± 0.1365 | 4.0688 ± 0.0450 | 17.3341 ± 0.1131 | 0.6743 ± 0.0174 |
| Stacking_v2 | 0.7491 ± 0.0065 | 5.7307 ± 0.2088 | 2.9480 ± 0.0396 | 11.9428 ± 0.0857 | 0.7491 ± 0.0065 |
| Stacking_v3 | 0.4982 ± 0.0041 | 8.318 ± 0.0490 | 4.319 ± 0.0176 | 16.768 ± 0.0407 | 0.4982 ± 0.0041 |
| Mediator | Path | Coef. | SE | p-Value | 95% CI (Low) | 95% CI (Upper) | Significance | Description of the Effect |
|---|---|---|---|---|---|---|---|---|
| LBXFER Ferritin |
Direct | 0.0231 | 0.0040 | 1.20e-8 | 0.0155 | 0.0309 | Yes | Major contribution via direct path (significant) |
| Indirect | -0.0003 | 0.0010 | 0.672 | -0.0026 | 0.0013 | No | The indirect effect is not significant. | |
| Total | 0.0229 | 0.0040 | 3.58e-8 | 0.0150 | 0.0309 | Yes | The overall effect remains | |
| LBXSGTSI Gamma-GT |
Direct | 0.0138 | 0.0041 | 9.42e-4 | 0.0057 | 0.0218 | Yes | Significant direct influence |
| Indirect | 0.0092 | 0.0027 | 0.000 | 0.0047 | 0.0154 | Yes | The indirect effect is statistically significant. | |
| Total | 0.0229 | 0.0040 | 3.58e-8 | 0.0150 | 0.0309 | Yes | Overall mediation effect | |
| BMXHT Height (cm) |
Direct | 0.0201 | 0.0042 | 2.55e-6 | 0.0119 | 0.0284 | Yes | Significant direct path |
| Indirect | 0.0028 | 0.0019 | 0.008 | 0.0004 | 0.0078 | Yes | The indirect effect is significant. | |
| Total | 0.0229 | 0.0040 | 3.58e-8 | 0.0150 | 0.0309 | Yes | The overall effect is maintained | |
| LBXSBU Urea (BUN) |
Direct | 0.0262 | 0.0042 | 1.97e-9 | 0.0179 | 0.0345 | Yes | The main effect is direct. |
| Indirect | -0.0033 | 0.0015 | 0.000 | -0.0071 | -0.0013 | Yes | The indirect effect is statistically significant. | |
| Total | 0.0229 | 0.0040 | 3.58e-8 | 0.0150 | 0.0309 | Yes | The final effect is confirmed | |
| LBXGLU Fasting glucose |
Direct | 0.0241 | 0.0040 | 7.70e-9 | 0.0162 | 0.0320 | Yes | The direct effect is clearly expressed. |
| Indirect | -0.0011 | 0.0007 | 0.008 | -0.0035 | -0.0002 | Yes | The indirect effect is expressed. | |
| Total | 0.0229 | 0.0040 | 3.58e-8 | 0.0150 | 0.0309 | Yes | The overall effect is confirmed | |
| LBXSLDSI |
Direct | 0.0198 | 0.0039 | 6.32e-7 | 0.0122 | 0.0274 | Yes | Significant direct contribution |
| Indirect | 0.0031 | 0.0019 | 0.004 | 0.0009 | 0.0082 | Yes | The indirect effect is statistically significant. | |
| Total | 0.0229 | 0.0040 | 3.58e-8 | 0.0150 | 0.0309 | Yes | The overall effect is expressed | |
| PAD615 activity in min | Direct | 0.0238 | 0.0040 | 8.59e-9 | 0.0159 | 0.0316 | Yes | The main contribution is direct. |
| Indirect | -0.0008 | 0.0008 | 0.240 | -0.0030 | 0.0003 | No | The indirect effect is insignificant. | |
| Total | 0.0229 | 0.0040 | 3.58e-8 | 0.0150 | 0.0309 | Yes | The final effect is confirmed. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




