Submitted:
08 September 2025
Posted:
10 September 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
| Article | Pemphigus subtype 1 | Number of MicroRNAs studied | Internal control | Key microRNAs |
Material and internal control |
|---|---|---|---|---|---|
| Valentino A. et al. [43] | PV | 86 | U6 | hsa-miR-148a-3p hsa-miR-146b-5p hsa-miR-126 hsa-miR-139 |
Blood plasma derived exosomes |
| Khabou B. et al. [44] | PF | 6 | n/a 2 | hsa-miR-17-5p hsa-miR-21-5 hsa-miR-146a-5p hsa-miR-155-5p hsa-miR-338-3p hsa-miR-21 |
Peripheral blood mononuclear cells/ skin biopsy samples |
| He W. et al. [32] |
PV | 12 | 5S рРНК | hsa-miR-125b-5p hsa-miR-146a-5p hsa-miR-148a-3p hsa-miR-150-5p hsa-miR-155-5p hsa-miR-181a-5p hsa-miR181b-5p hsa-miR-326 hsa-miR-338-3p hsa-miR-423-5p hsa-miR-424-5p hsa-miR-584-5p |
Blood plasma |
| Xu M. et al. [45] |
PV | 1 | U6 | hsa-miR-338-3p | Peripheral blood mononuclear cells |
| Lin N. et al. [20] |
PV | 1 | U6 | hsa-miR-338-3p | Peripheral blood mononuclear cells |
| Liu Q. et al. [24] |
PV | 1 | 18srRNA | hsa-miR-338-3p | Peripheral blood mononuclear cells |
| Wang M. et al. [21] |
PV | 124 | U6 | hsa-miR-424-5p | Peripheral blood mononuclear cells |
2. Results
2.1. Patient Selection
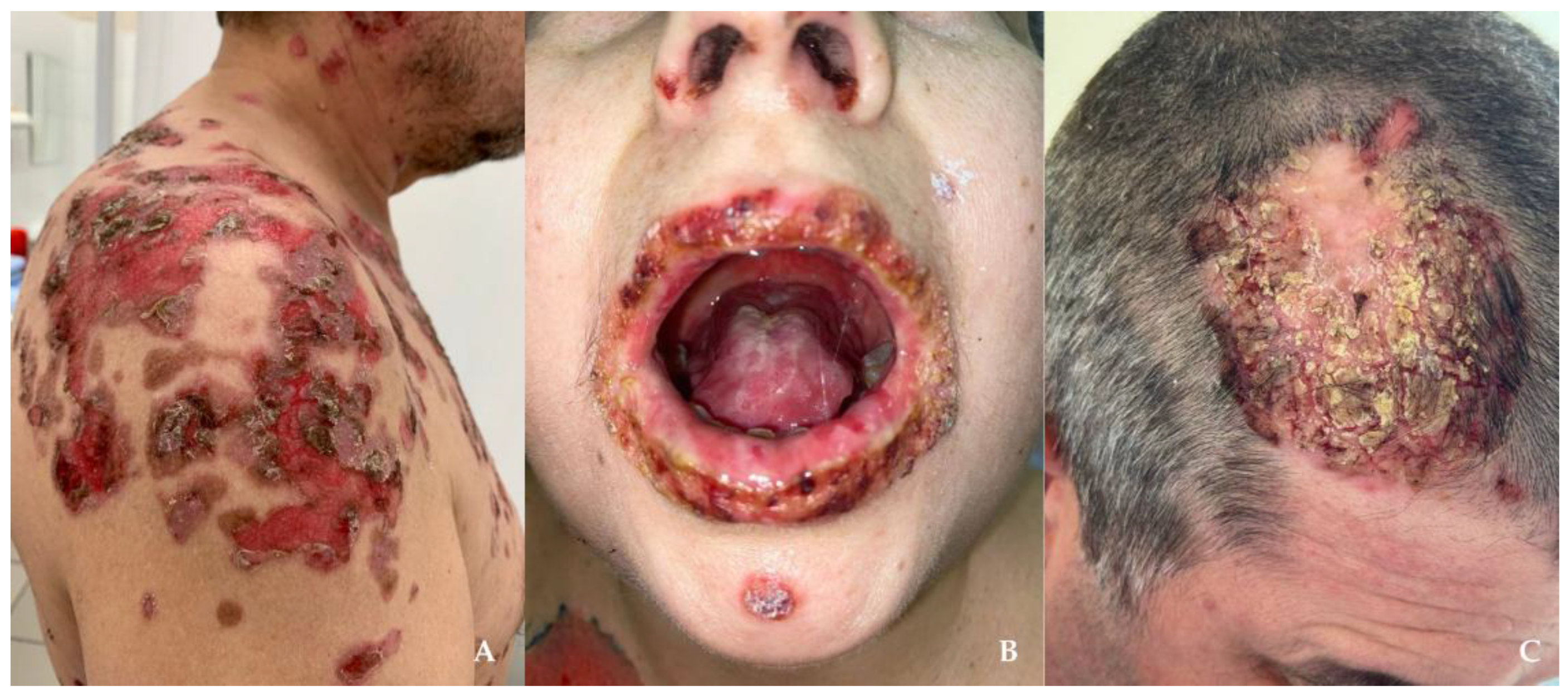
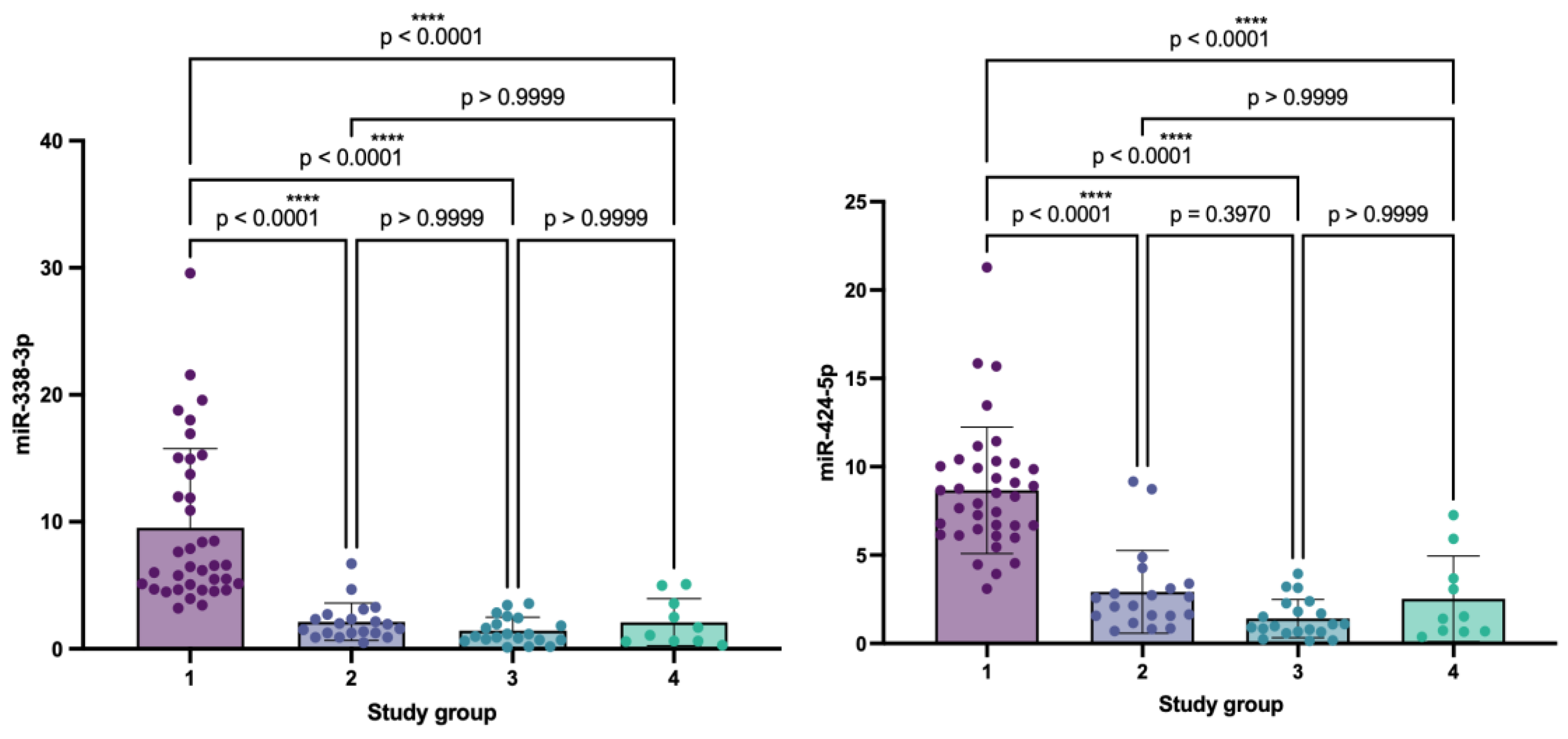
2.2. Analysis of Relative microRNA Expression Levels in Patients with Pemphigus and Control Groups
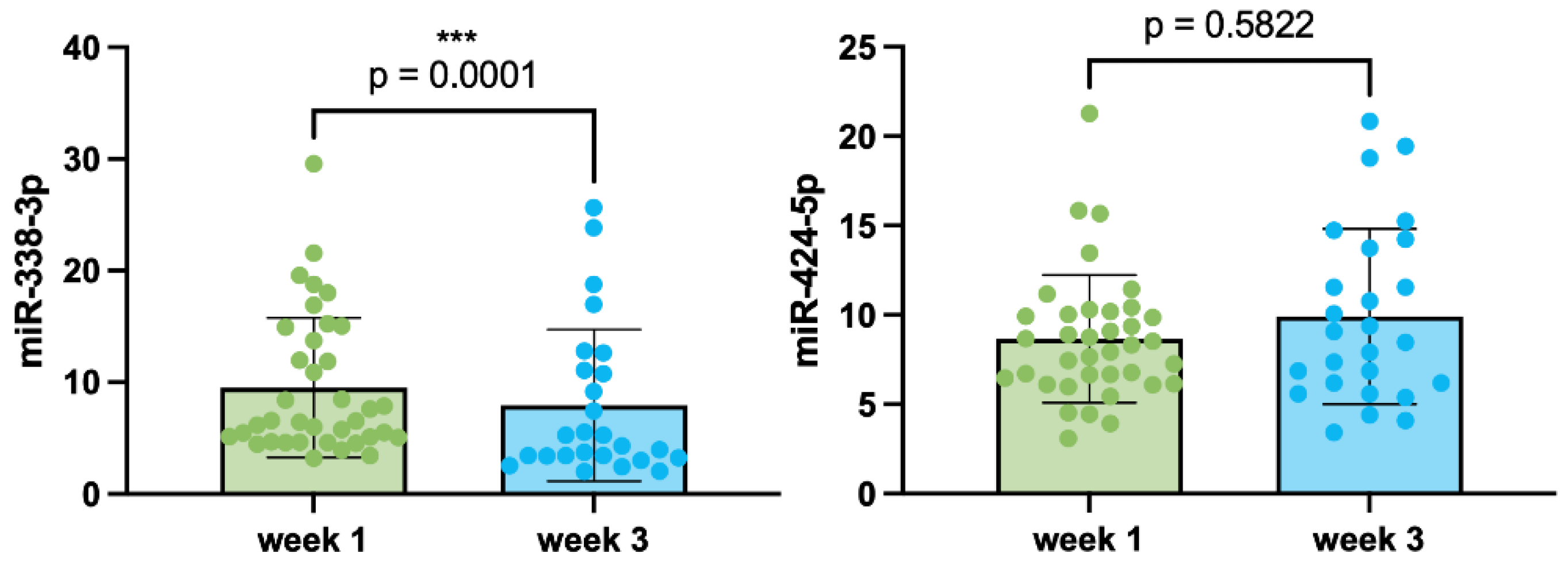
2.3. Relative Expression Levels of miR-338-3p and miR-424-5p in Patients with Pemphigus During Oral Systemic Glucocorticosteroid Therapy
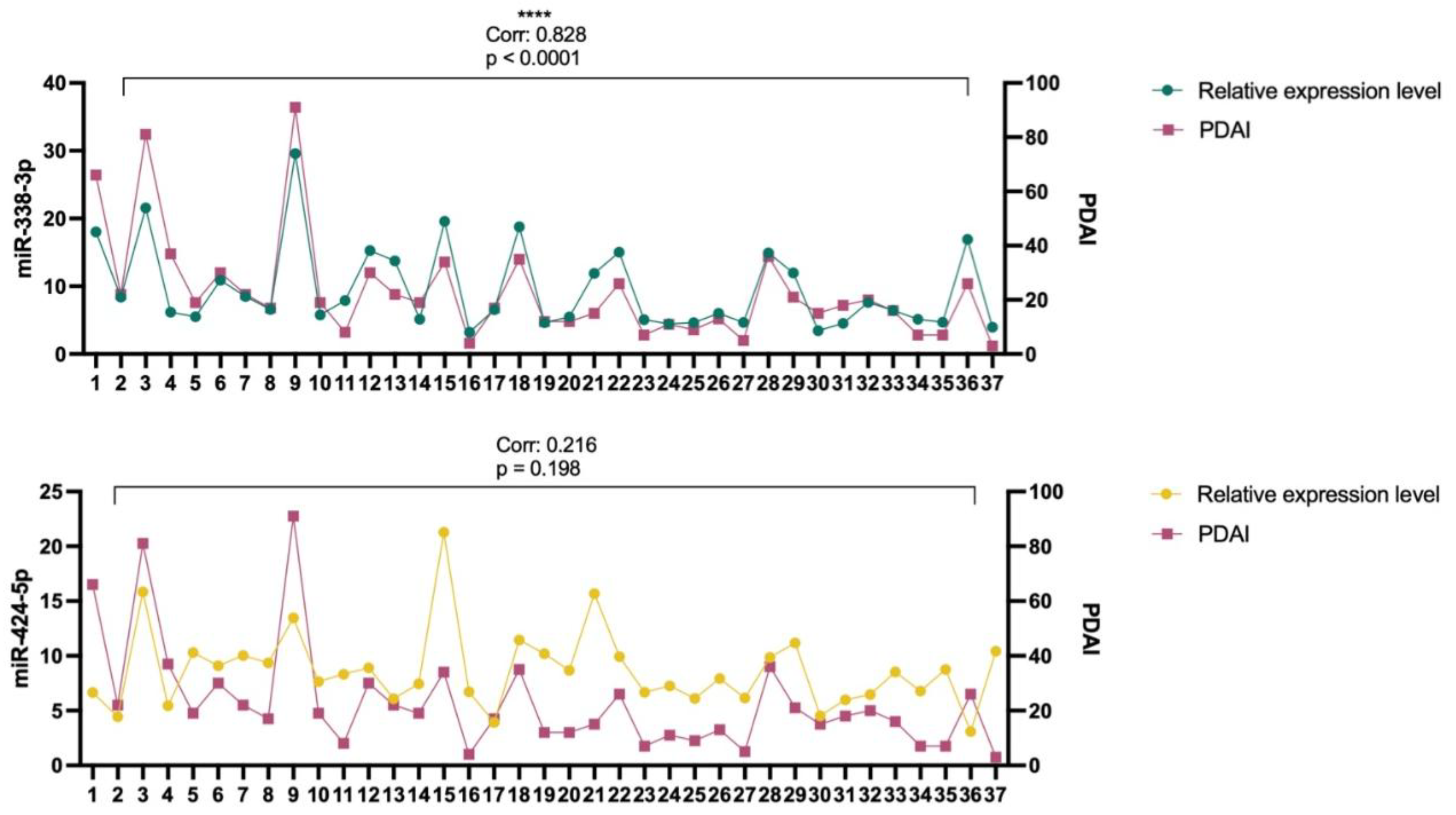
2.4. Correlation Analysis of miR-338-3p and miR-424-5p Expression Levels with Disease Severity Assessed by the Pemphigus Disease Area Index (PDAI)
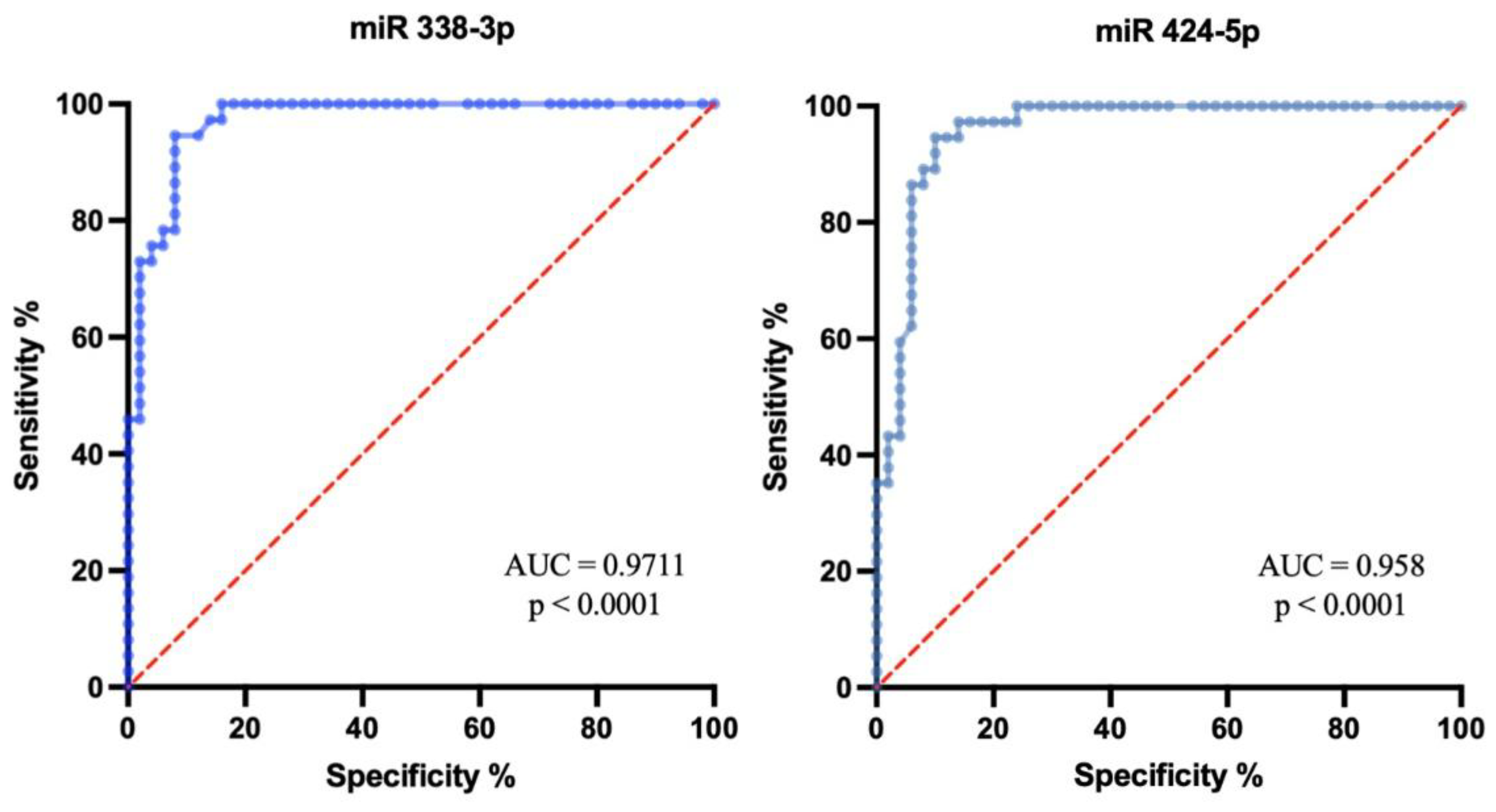
2.5. Diagnostic Value of Relative miR-338-3p and miR-424-5p Expression Levels in Pemphigus
3. Discussion
4. Materials and Methods
4.1. Patients, Study Design, and Approvals
4.2. Quantitative Real-Time PCR
4.3. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Olisova, O.Y.; Teplyuk, N.P. An Illustrated Guide to Dermatology. For Preparing Practitioners for Accreditation; GE-OTAR-Media: Moscow, Russia, 2023; p. 376. [Google Scholar]
- Malik, A.M.; Tupchong, S.; Huang, S.; Are, A.; Hsu, S.; Motaparthi, K. An Updated Review of Pemphigus Diseases. Medicina 2021, 57, 1080. [Google Scholar] [CrossRef]
- Schmidt, E.; Kasperkiewicz, M.; Joly, P. Pemphigus. Lancet 2019, 394, 882–894. [Google Scholar] [CrossRef]
- Amagai, M.; Tanikawa, A.; Committee for Guidelines for the Management of Pemphigus Disease. Japanese Guidelines for the Management of Pemphigus. J. Dermatol. 2014, 41, 471–486. [Google Scholar] [CrossRef]
- Kridin, K. Pemphigus Group: Overview, Epidemiology, Mortality, and Comorbidities. Immunol. Res. 2018, 66, 255–270. [Google Scholar] [CrossRef]
- Gonçalves, G.A.; Brito, M.M.; Salathiel, A.M.; Ferraz, T.S.; Alves, D.; Roselino, A.M. Incidence of Pemphigus Vulgaris Exceeds That of Pemphigus Foliaceus in a Region Where Pemphigus Foliaceus Is Endemic: Analysis of a 21-Year Historical Series. An. Bras. Dermatol. 2011, 86, 1109–1112. [Google Scholar] [CrossRef]
- Costan, V.V.; Popa, C.; Hâncu, M.F.; Porumb-Andrese, E.; Toader, M.P. Comprehensive Review on the Pathophysiology, Clinical Variants and Management of Pemphigus. Exp. Ther. Med. 2021, 22, 1335. [Google Scholar] [CrossRef] [PubMed]
- Harel-Raviv, M.; Srolovitz, H.; Gornitsky, M. Pemphigus Vulgaris: The Potential for Error. A Case Report. Spec. Care Dentist. 1995, 15, 61–64. [Google Scholar] [CrossRef] [PubMed]
- Morishima-Koyano, M.; Nobeyama, Y.; Fukasawa-Momose, M.; et al. Case of Pemphigus Foliaceus Misdiagnosed as a Single Condition of Erythrodermic Psoriasis and Modified by Brodalumab. J. Dermatol. 2020, 47, e201–e202. [Google Scholar] [CrossRef] [PubMed]
- Daltaban, Ö.; Özçentik, A.; Karakaş, A.; et al. Clinical Presentation and Diagnostic Delay in Pemphigus Vulgaris: A Prospective Study from Turkey. J. Oral Pathol. Med. 2020, 49, 681–686. [Google Scholar] [CrossRef]
- Khamaganova, I.V.; Malyarenko, E.N.; Denisova, E.V.; Vorontsova, I.V.; Plieva, K.T. Mistakes of Diagnostics in Pemphigus Vulgaris: Case Report. Russ. J. Skin Venereal Dis. 2017, 20, 30–33. [Google Scholar] [CrossRef]
- Teplyuk, N.P.; Kolesova, Y.V.; Mak, D.V.; Lepekhova, A.A.; Toshchakov, S.V.; Fedotcheva, T.A. Pemphigus: New Approaches to Diagnosis and Disease Severity Assessment. Russ. J. Skin Venereal Dis. 2023, 26, 515–526. [Google Scholar] [CrossRef]
- Kridin, K.; Bergman, R. The Usefulness of Indirect Immunofluorescence in Pemphigus and the Natural History of Patients with Initial False-Positive Results: A Retrospective Cohort Study. Front. Med. 2018, 5, 266. [Google Scholar] [CrossRef] [PubMed]
- Giurdanella, F.; Nijenhuis, A.M.; Diercks, G.F.H.; Jonkman, M.F.; Pas, H.H. Keratinocyte Binding Assay Identifies Anti-Desmosomal Pemphigus Antibodies Where Other Tests Are Negative. Front. Immunol. 2018, 9, 839. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, E.; Dähnrich, C.; Rosemann, A.; et al. Novel ELISA Systems for Antibodies to Desmoglein 1 and 3: Correlation of Disease Activity with Serum Autoantibody Levels in Individual Pemphigus Patients. Exp. Dermatol. 2010, 19, 458–463. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front. Endocrinol. 2018, 9, 402. [Google Scholar] [CrossRef]
- Sanz-Rubio, D.; Martin-Burriel, I.; Gil, A.; Cubero, P.; Forner, M.; Khalyfa, A.; Marin, J.M. Stability of Circulating Exosomal miRNAs in Healthy Subjects. Sci. Rep. 2018, 8, 10306. [Google Scholar] [CrossRef]
- Matias-Garcia, P.R.; Wilson, R.; Mussack, V.; Reischl, E.; Waldenberger, M.; Gieger, C.; Anton, G.; Peters, A.; Kuehn-Steven, A. Impact of Long-Term Storage and Freeze-Thawing on Eight Circulating microRNAs in Plasma Samples. PLoS ONE 2020, 15, e0227648. [Google Scholar] [CrossRef]
- Ward Gahlawat, A.; Lenhardt, J.; Witte, T.; Keitel, D.; Kaufhold, A.; Maass, K.K.; Pajtler, K.W.; Sohn, C.; Schott, S. Evaluation of Storage Tubes for Combined Analysis of Circulating Nucleic Acids in Liquid Biopsies. Int. J. Mol. Sci. 2019, 20, 704. [Google Scholar] [CrossRef]
- Lin, N.; Liu, Q.; Wang, M.; Wang, Q.; Zeng, K. Usefulness of miRNA-338-3p in the Diagnosis of Pemphigus and Its Correlation with Disease Severity. PeerJ 2018, 6, e5388. [Google Scholar] [CrossRef]
- Wang, M.; Liang, L.; Li, L.; et al. Increased miR-424-5p Expression in Peripheral Blood Mononuclear Cells from Patients with Pemphigus. Mol. Med. Rep. 2017, 15, 3479–3484. [Google Scholar] [CrossRef]
- Rodriguez, M.S.; Egaña, I.; Lopitz-Otsoa, F.; Aillet, F.; Lopez-Mato, M.P.; Dorronsoro, A.; Lobato-Gil, S.; Sutherland, J.D.; Barrio, R.; Trigueros, C.; Lang, V. The RING Ubiquitin E3 RNF114 Interacts with A20 and Modulates NF-κB Activity and T-Cell Activation. Cell Death Dis. 2014, 5, e1399. [Google Scholar] [CrossRef]
- Yang, P.; Lu, Y.; Li, M.; Zhang, K.; Li, C.; Chen, H.; Tao, D.; Zhang, S.; Ma, Y. Identification of RNF114 as a Novel Positive Regulatory Protein for T Cell Activation. Immunobiology 2014, 219, 432–439. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Cui, F.; Wang, M.; Xiong, H.; Peng, X.; Liang, L.; Li, L.; Zhang, J.; Peng, X.; Zeng, K. Increased Expression of microRNA-338-3p Contributes to Production of Dsg3 Antibody in Pemphigus Vulgaris Patients. Mol. Med. Rep. 2018, 18, 550–556. [Google Scholar] [CrossRef]
- Satyam, A.; Khandpur, S.; Sharma, V.K.; Sharma, A. Involvement of TH1/TH2 Cytokines in the Pathogenesis of Autoimmune Skin Disease—Pemphigus Vulgaris. Immunol. Investig. 2009, 38, 498–509. [Google Scholar] [CrossRef]
- Lee, S.H.; Hong, W.J.; Kim, S.C. Analysis of Serum Cytokine Profile in Pemphigus. Ann. Dermatol. 2017, 29, 438–445. [Google Scholar] [CrossRef]
- Rizzo, C.; Fotino, M.; Zhang, Y.; et al. Direct Characterization of Human T Cells in Pemphigus Vulgaris Reveals Elevated Autoantigen-Specific Th2 Activity in Association with Active Disease. Clin. Exp. Dermatol. 2005, 30, 535–540. [Google Scholar] [CrossRef]
- Teplyuk, N.P.; Mak, D.V.; Kolesova, Y.V.; Lepekhova, A.A.; Fedotcheva, T.A.; Ulchenko, D.N. The miR-338-3p Expression Level in Pemphigus Diagnosis. Russ. J. Skin Venereal Dis. 2024, 27, 448–462. [Google Scholar] [CrossRef]
- Li, X.; Ishii, N.; Ohata, C.; Furumura, M.; Hashimoto, T. Signalling Pathways in Pemphigus Vulgaris. Exp. Dermatol. 2014, 23, 155–156. [Google Scholar] [CrossRef]
- Chernyavsky, A.I.; Arredondo, J.; Kitajima, Y.; Sato-Nagai, M.; Grando, S.A. Desmoglein versus Non-Desmoglein Signaling in Pemphigus Acantholysis: Characterization of Novel Signaling Pathways Downstream of Pemphigus Vulgaris Antigens. J. Biol. Chem. 2007, 282, 13804–13812. [Google Scholar] [CrossRef] [PubMed]
- Berkowitz, P.; Hu, P.; Liu, Z.; Diaz, L.A.; Enghild, J.J.; Chua, M.P.; Rubenstein, D.S. Desmosome Signaling. Inhibition of p38MAPK Prevents Pemphigus Vulgaris IgG-Induced Cytoskeleton Reorganization. J. Biol. Chem. 2005, 280, 23778–23784. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Xing, Y.; Li, C.; et al. Identification of Six microRNAs as Potential Biomarkers for Pemphigus Vulgaris: From Diagnosis to Pathogenesis. Diagnostics 2022, 12, 3058. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, Z.; Gemeinhart, R.A. Progress in microRNA Delivery. J. Control. Release 2013, 172, 962–974. [Google Scholar] [CrossRef] [PubMed]
- Slack, F.J.; Chinnaiyan, A.M. The Role of Non-Coding RNAs in Oncology. Cell 2019, 179, 1033–1055. [Google Scholar] [CrossRef]
- Pozniak, T.; Shcharbin, D.; Bryszewska, M. Circulating microRNAs in Medicine. Int. J. Mol. Sci. 2022, 23, 3996. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Cannavicci, A.; Dai, S.-C.; Wang, C.; Kutryk, M.J.B. MicroRNA Signature of Human Blood Mononuclear Cells. Mol. Cell. Biochem. [CrossRef]
- Papara, C.; Zillikens, D.; Sadik, C.D.; Baican, A. MicroRNAs in Pemphigus and Pemphigoid Diseases. Autoimmun. Rev. 2021, 20, 102852. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Xu, M.; Tian, X.; et al. Research Advances in the Detection of miRNA. J. Pharm. Anal. 2019, 9, 217–226. [Google Scholar] [CrossRef]
- Kramer, M.F. Stem-Loop RT-qPCR for miRNAs. Curr. Protoc. Mol. Biol. 2011. Chapter 15, Unit15.10. [Google Scholar] [CrossRef]
- Chen, C.; Ridzon, D.A.; Broomer, A.J.; et al. Real-Time Quantification of MicroRNAs by Stem–Loop RT–PCR. Nucleic Acids Res. 2005, 33, e179. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2(−ΔΔCT) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Valentino, A.; Leuci, S.; Galderisi, U.; et al. Plasma Exosomal microRNA Profile Reveals miRNA 148a-3p Downregulation in the Mucosal-Dominant Variant of Pemphigus Vulgaris. Int. J. Mol. Sci. 2023, 24, 11493. [Google Scholar] [CrossRef] [PubMed]
- Khabou, B.; Fakhfakh, R.; Tahri, S.; et al. miRNA Implication in the Pathogenesis and the Outcome of Tunisian Endemic Pemphigus Foliaceus. Exp. Dermatol. 2023, 32, 1132–1142. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Liu, Q.; Li, S.; et al. Increased Expression of miR-338-3p Impairs Treg-Mediated Immunosuppression in Pemphigus Vulgaris by Targeting RUNX1. Exp. Dermatol. 2020, 29, 623–629. [Google Scholar] [CrossRef] [PubMed]
| Sample | Gender 1 /Age |
Diagnosis 2 | Disease duration (years) |
Lesion localization |
GCS 3 dose (mg) at admission/week 3 | PDAI 4 at admission/week 3 |
| 1 | M/53 | PV | 1.1 | Skin/mucosa | 0 / 80 | 66 / 38 |
| 2 | M/52 | PF | 3.0 | Skin | 0 / 80 | 22 / 16 |
| 3 | M/36 | PV | 1.0 | Skin/mucosa | 0 / 100 | 81 / 64 |
| 4 | F/45 | PV | 3.4 | Skin/mucosa | 15 / 80 | 37 / 27 |
| 5 | F/64 | PF | 0.1 | Skin | 0 / 80 | 19 / 16 |
| 6 | F/56 | PV | 0.8 | Skin/mucosa | 0 / 80 | 30 / 22 |
| 7 | F/30 | PF | 1.0 | Skin | 0 / 80 | 22 / 11 |
| 8 | M/38 | PV | 0.1 | Skin/mucosa | 0 / 80 | 17 / 10 |
| 9 | F/52 | PV | 0.2 | Skin/mucosa | 0 / 100 | 91 / 67 |
| 10 | M/38 | PF | 0.4 | Skin | 0 / 80 | 19 / 10 |
| 11 | F/55 | PV | 1.2 | Mucosa | 17.5 / 80 | 8 / 8 |
| 12 | F/55 | PV | 1.1 | Mucosa | 17.5 / 80 | 30 / 28 |
| 13 | F/36 | PV | 0.8 | Mucosa | 17.5 / 80 | 22 / 15 |
| 14 | F/37 | PV | 0.7 | Skin/mucosa | 0 / 80 | 19 / 14 |
| 15 | F/77 | PV | 0.7 | Skin/mucosa | 0 / 80 | 34 / 24 |
| 16 | F/50 | PV | 0.9 | Skin/mucosa | 20 / 60 | 4 / 2 |
| 17 | F/60 | PV | 1.0 | Mucosa | 25 / – * | 17 / – * |
| 18 | F/56 | PV | 5.0 | Skin/mucosa | 15 / 100 | 35 / 54 |
| 19 | F/83 | PV | 0.2 | Mucosa | 0 / – * | 12 / – * |
| 20 | F/60 | PV | 0.1 | Skin/mucosa | 0 / 80 | 12 / 9 |
| 21 | F/44 | PV | 6.0 | Skin/mucosa | 12.5 / 70 | 15 / 12 |
| 22 | F/58 | PV | 4.0 | Skin/mucosa | 0 / 80 | 26 / 16 |
| 23 | F/44 | PV | 4.0 | Skin | 10 / 80 | 7 / 6 |
| 24 | F/80 | PV | 16.0 | Skin | 5 / – * | 11 / – * |
| 25 | F/50 | PV | 0.5 | Mucosa | 0 / 80 | 9 / 7 |
| 26 | M/61 | PF | 0.5 | Skin | 15 / – * | 13 / – * |
| 27 | M/33 | PF | 0.7 | Skin | 60 / 80 | 5 / 3 |
| 28 | M/61 | PV | 2.0 | Skin/mucosa | 0 / 80 | 36 / 26 |
| 29 | F/62 | PV | 0.4 | Skin/mucosa | 0 / – * | 21 / – * |
| 30 | F/59 | PV | 0.7 | Skin/mucosa | 45 / – * | 15 / – * |
| 31 | F/69 | PF | 0.1 | Skin | 0 / – * | 18 / – * |
| 32 | F/52 | PV | 0.6 | Skin | 60 / 80 | 20 / 15 |
| 33 | M/47 | PF | 0.9 | Skin | 30 / 80 | 16 / 10 |
| 34 | F/76 | PV | 9.0 | Skin/mucosa | 12.5 / – * | 7 / – * |
| 35 | F/40 | PF | 3.0 | Skin | 50 / – * | 7 / – * |
| 36 | F/82 | PV | 0.5 | Skin/mucosa | 25 / – * | 26 / – * |
| 37 | F/49 | PV | 0.7 | Mucosa | 15 / – * | 3 / – * |
| Group | Number of patients |
Age (mean ± SD) | Male, n (%) |
Female, n (%) |
miR-338-3p relative expression (2−ΔΔCt method) |
miR-424-5p relative expression (2−ΔΔCt method) |
|---|---|---|---|---|---|---|
| 1. Active pemphigus | 37 | 54 ± 13.93 | 9 (24.3%) | 28 (75.7%) | 9.53 (SD ± 6.24) | 8.67 (SD ± 3.58) |
| 2. Remission pemphigus | 20 | 56 ± 12.81 | 7 (35%) | 13 (65%) | 2.13 (SD ± 1.47) | 2.92 (SD ± 2.34) |
| 3. Healthy controls | 20 | 34 ± 11.5 | 5 (25%) | 15 (75%) | 1.42 (SD ± 1.06) | 1.41 (SD ± 1.08) |
| 4. Other bullous dermatoses | 10 | 65 ± 18.58 | 3 (30%) | 7 (70%) | 2.10 (SD ± 1.85) | 2.53 (SD ± 2.42) |
| Primer | Nucleotide sequence |
|---|---|
| Stem-loop reverse transcription primers | |
| RT-oligo-miR-338-3p | 5’-GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACAGGTCG-3’ |
| RT-oligo-miR-424-5p | 5’-GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACGTCGTC-3’ |
| RT-oligo-U6 | 5’-GTCGTGTCTGAGGCTGACTGAGACCTATTCGCACCTGACACGACGGCCATGC-3’ |
| Amplification primers | |
| Forward - miR-338-3p | 5’-TACGTTGTTTTAGTGACTACGACCT-3’ |
| Reverse - miR-338-3p | 5’-GTGCAGGGTCCGAGGTATTC-3’ |
| Forward - miR-424-5p | 5’-CCAAGTTTTGTACTTAACGACGAC-3’ |
| Reverse - miR-424-5p | 5’-GTGCAGGGTCCGAGGTATTC-3’ |
| Forward - U6 | 5’-GGCCGCATACAGAGAAGATTA-3’ |
| Reverse - U6 | 5’-CTGAGGCTGACTGAGACCT-3’ |
| Forward - miR-338-3p | 5’-TACGTTGTTTTAGTGACTACGACCT-3’ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




