Submitted:
11 July 2025
Posted:
15 July 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
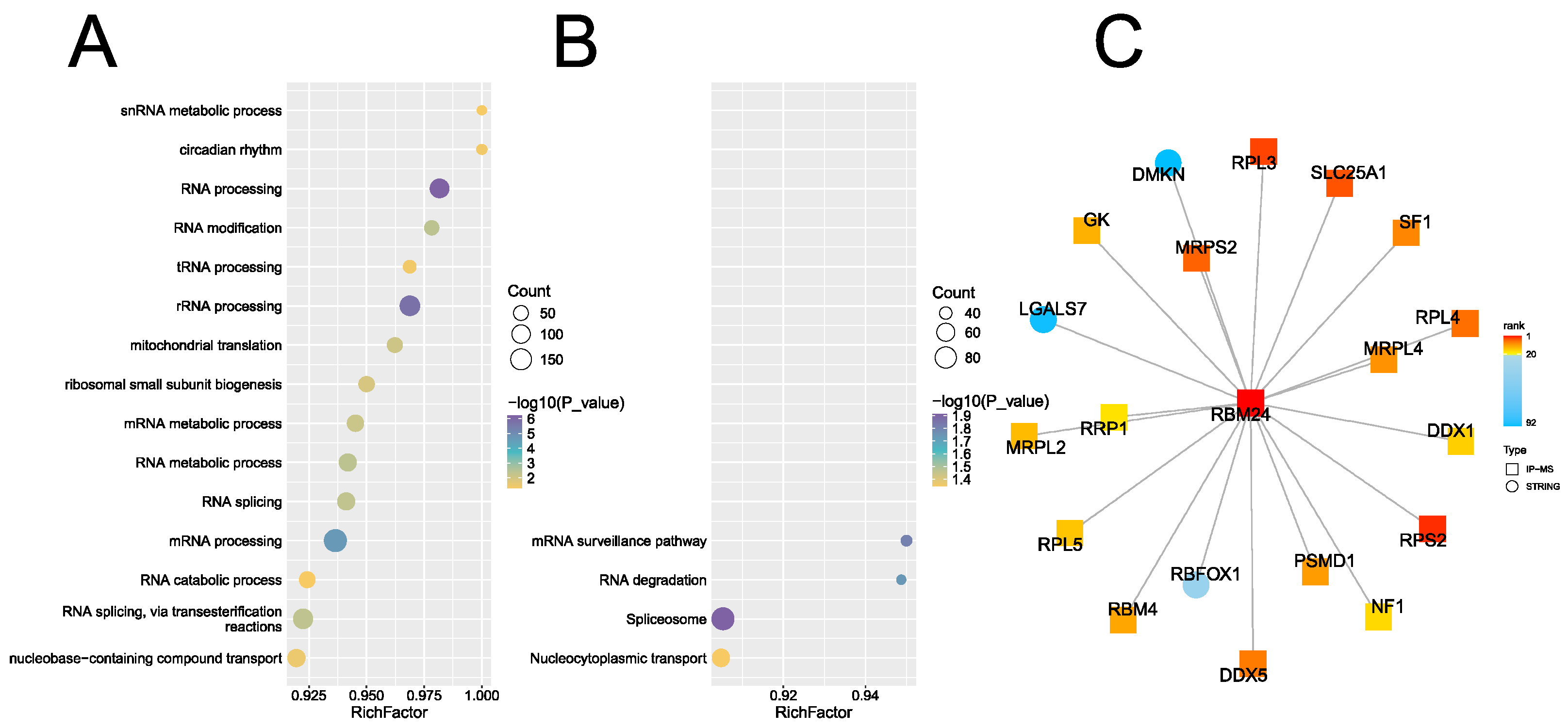
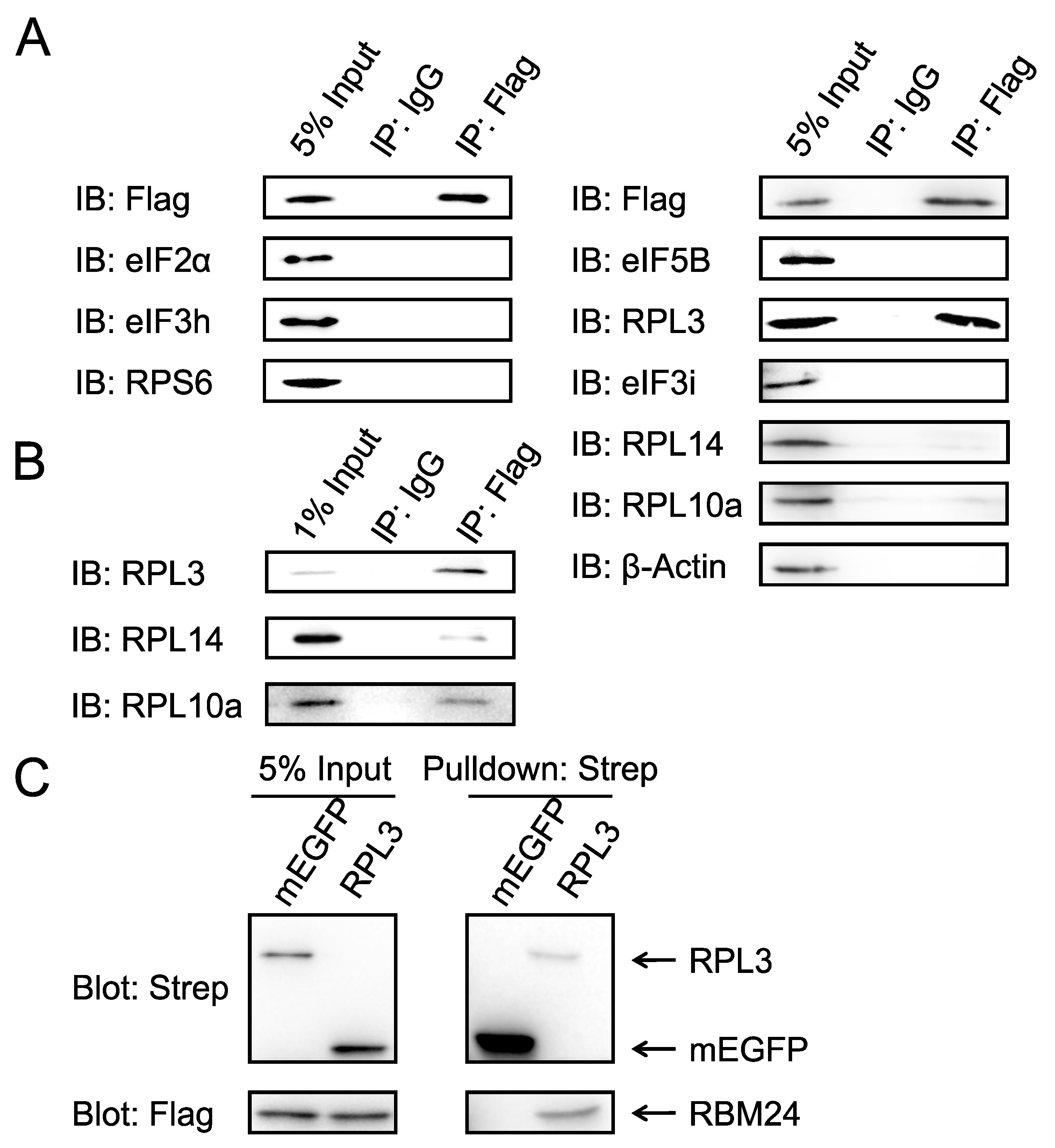
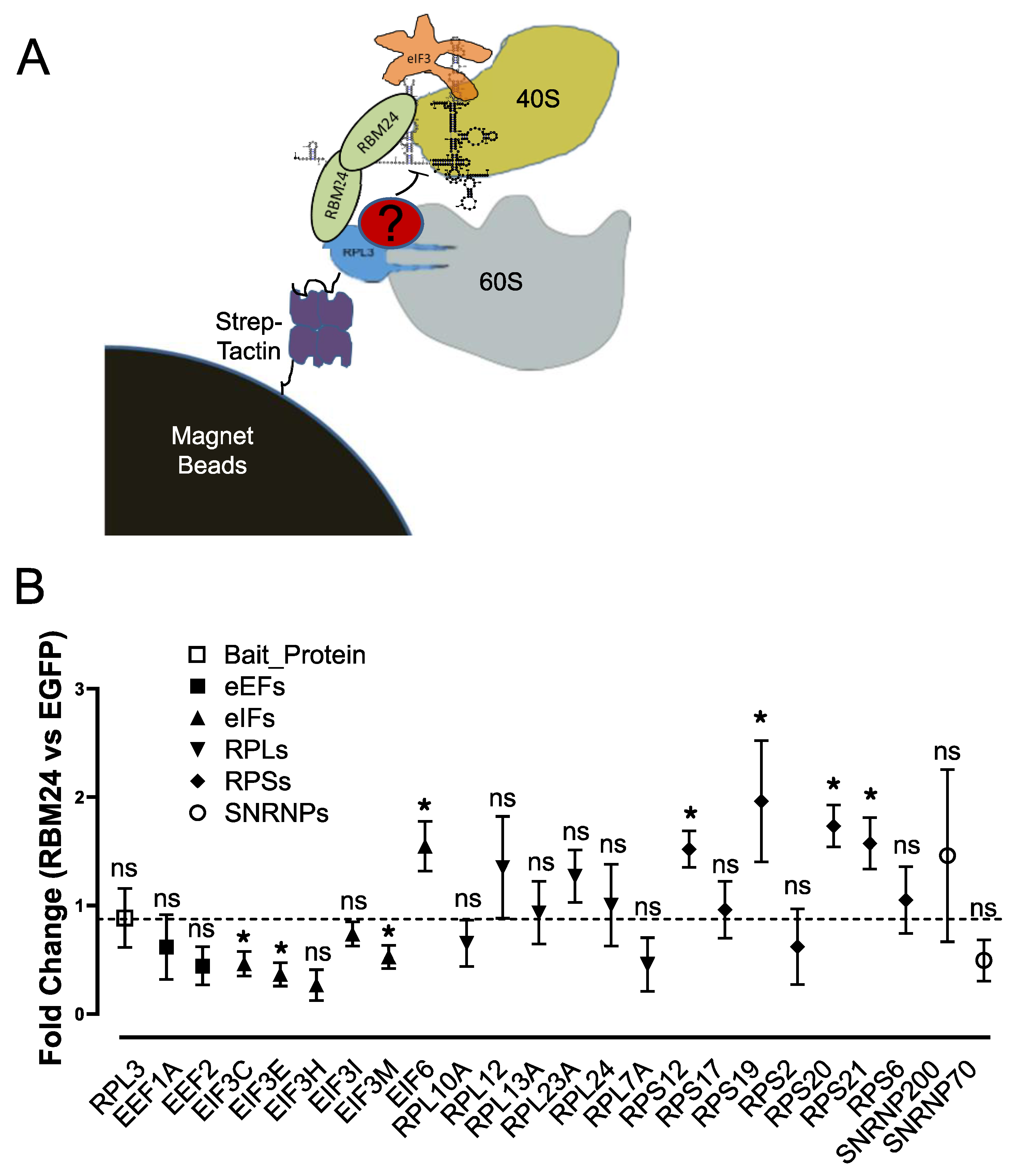
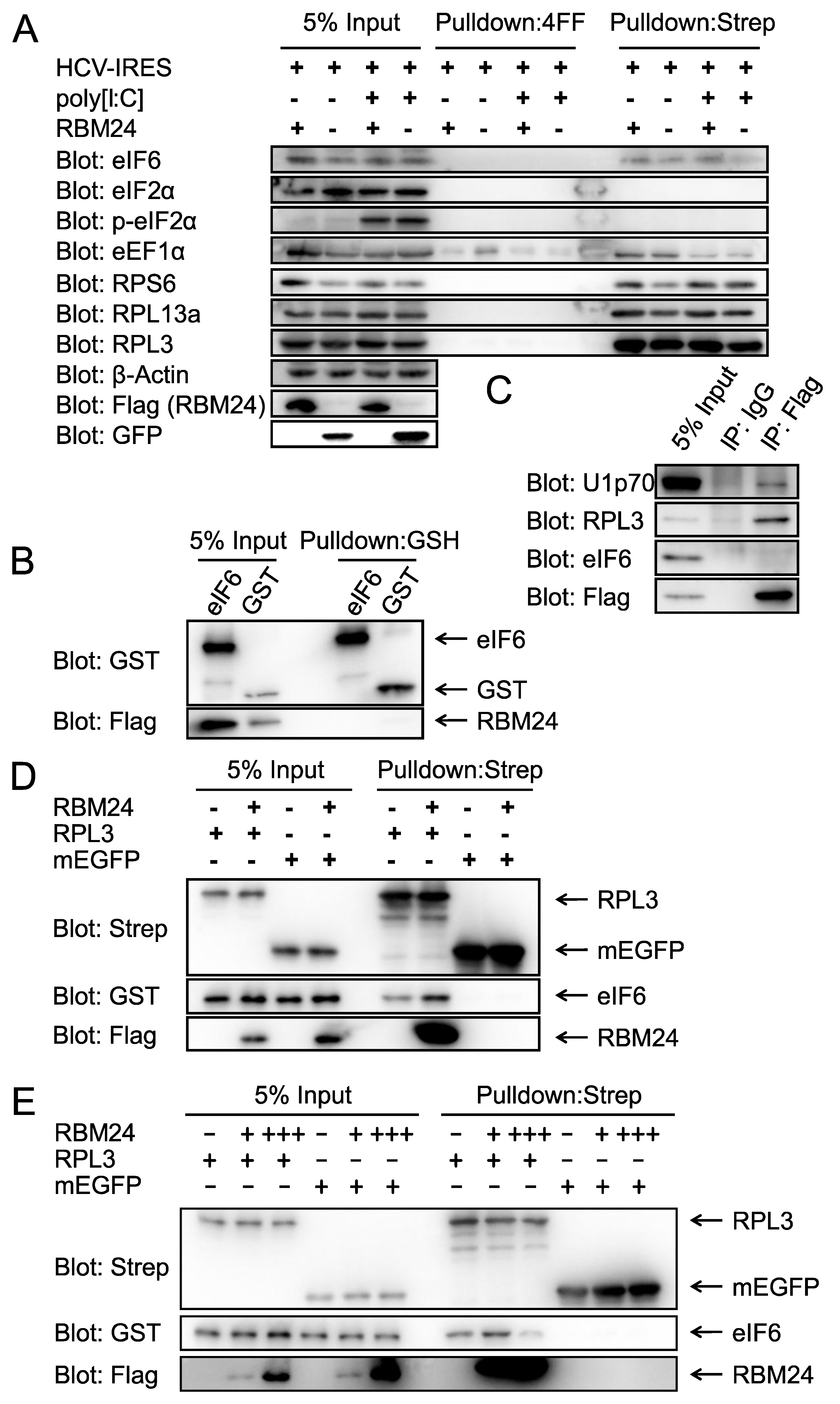
2.1. RBM24 Interacts With RPL3 in Both IP and Pulldown Assays
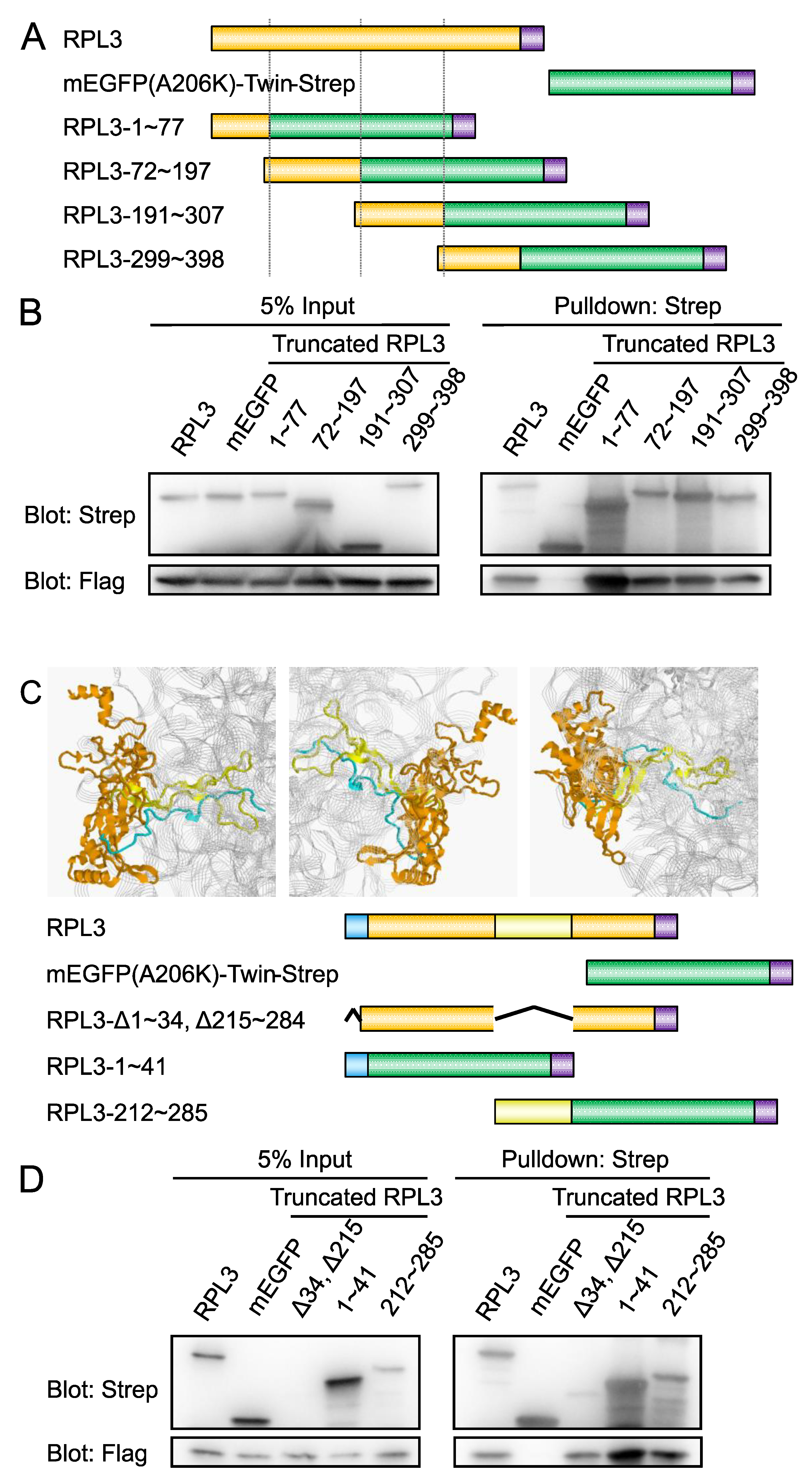
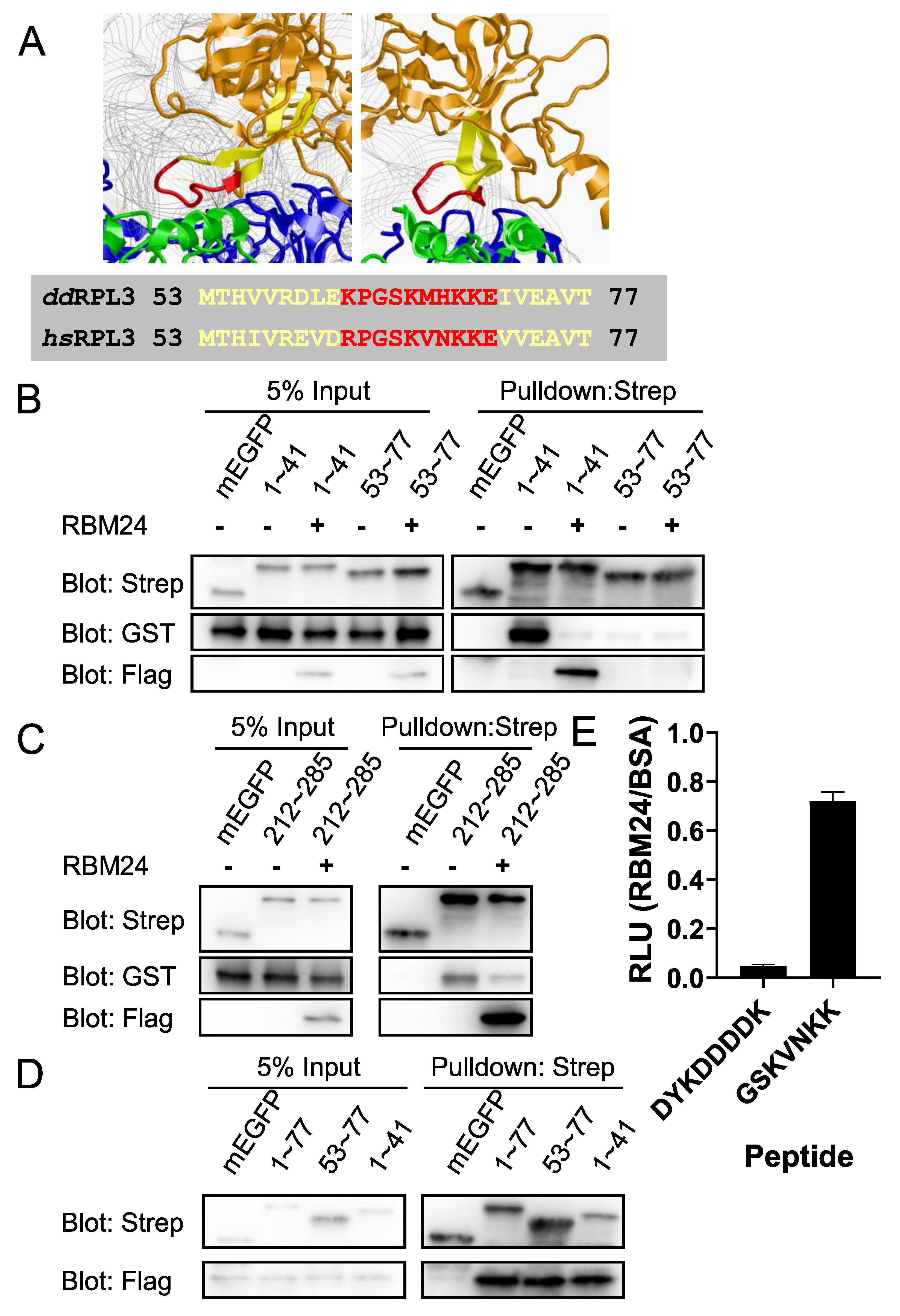
2.2. RBM24 Binds Both Exposed and 28S-embeded Region of RPL3
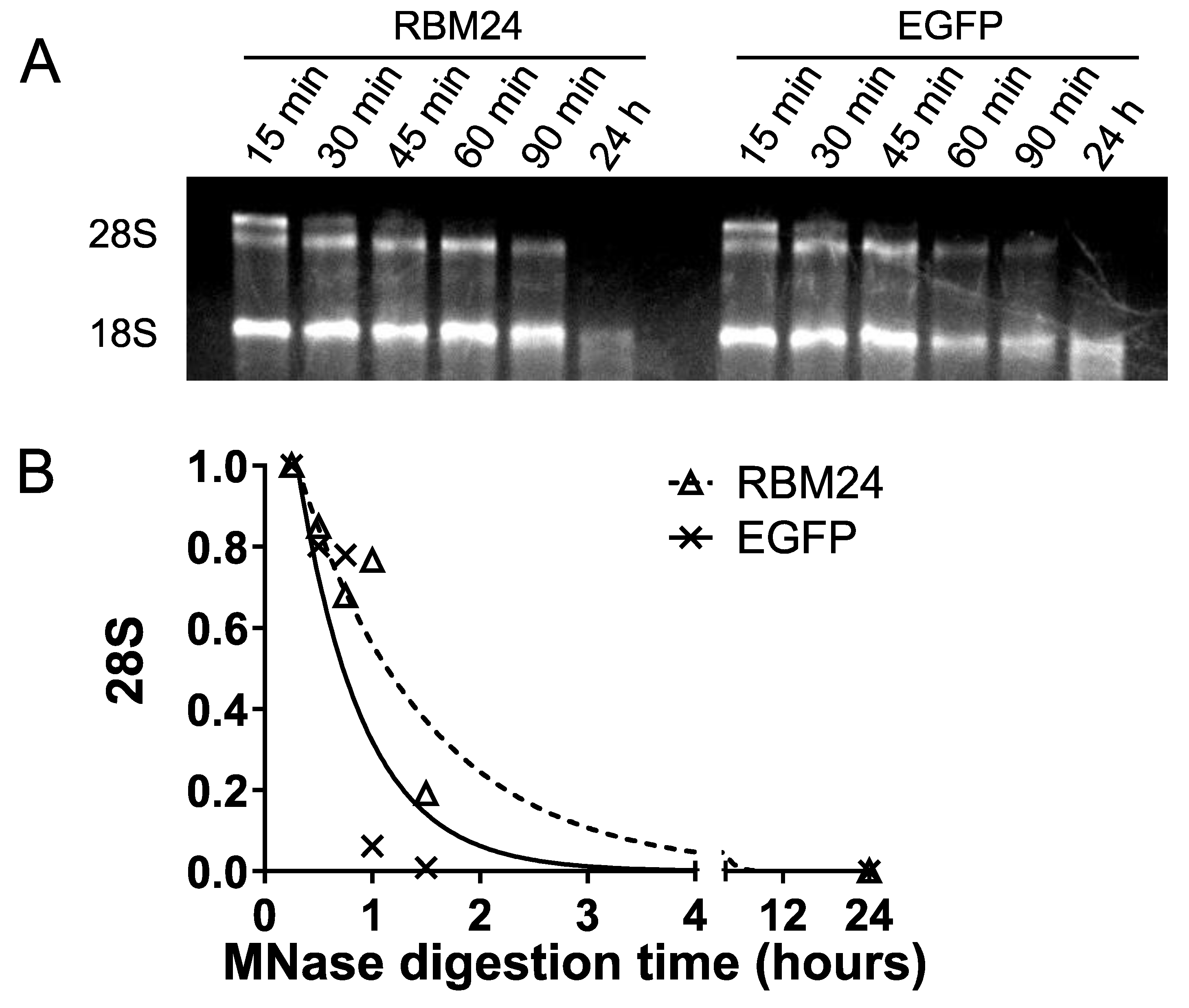
2.3. RBM24 does not Vulnerate 28S rRNA to Micrococcal Nuclease
2.4. RBM24 Enhances the Interaction Between the RPL3 and eIF6
2.5. RBM24 Blocks the Binding of eIF6 to 28S-embeded Regions in RPL3
2.6. RPL3 Derived Peptide Rescues RBM24 Mediated Translation Inhibition
2.7. Cellular IRES Candidates Potentially Targeted by RBM24
3. Discussion
4. Materials and Methods
4.1. Cell Culture and Transfection
4.2. Plasmids
4.3. Recombinant Proteins
4.4. 28.S Degradation Assay and Urea-Agarose Gel Electrophoresis
4.5. In vitro Transcription
4.6. Western Blot, Co-Immunoprecipitation and IP-MS Analyses
4.7. Twin Strep Tag Pulldown and GST Pulldown
4.8. Endogenous Tagging of RPL3
4.9. Ribosome Affinity Capture and Quantification Mass Spectrometry
4.10. PDB Files Visualizing
4.11. Peptide Competition Assay
4.12. RNA Immunoprecipitation Followed by High Throughput Sequencing
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lin, Y.; May, G.E.; Kready, H.; Nazzaro, L.; Mao, M.; Spealman, P.; Creeger, Y.; McManus, C.J. Impacts of uORF codon identity and position on translation regulation. Nucleic Acids Res 2019, 47, 9358-9367. https://doi.org/10.1093/nar/gkz681. [CrossRef]
- Sonenberg, N.; Hinnebusch, A.G. Regulation of translation initiation in eukaryotes: mechanisms and biological targets. Cell 2009, 136, 731-745. https://doi.org/10.1016/j.cell.2009.01.042. [CrossRef]
- Ryoo, H.D.; Vasudevan, D. Two distinct nodes of translational inhibition in the Integrated Stress Response. BMB Rep 2017, 50, 539-545. https://doi.org/10.5483/bmbrep.2017.50.11.157. [CrossRef]
- Hershey, J.W.B.; Sonenberg, N.; Mathews, M.B. Principles of Translational Control. Cold Spring Harb Perspect Biol 2019, 11. https://doi.org/10.1101/cshperspect.a032607. [CrossRef]
- Jiang, Y.; Zhang, M.; Qian, Y.; Xu, E.; Zhang, J.; Chen, X. Rbm24, an RNA-binding protein and a target of p53, regulates p21 expression via mRNA stability. J Biol Chem 2014, 289, 3164-3175. https://doi.org/10.1074/jbc.M113.524413. [CrossRef]
- Xu, E.; Zhang, J.; Zhang, M.; Jiang, Y.; Cho, S.J.; Chen, X. RNA-binding protein RBM24 regulates p63 expression via mRNA stability. Mol Cancer Res 2014, 12, 359-369. https://doi.org/10.1158/1541-7786.MCR-13-0526. [CrossRef]
- Shao, M.; Lu, T.; Zhang, C.; Zhang, Y.Z.; Kong, S.H.; Shi, D.L. Rbm24 controls poly(A) tail length and translation efficiency of crystallin mRNAs in the lens via cytoplasmic polyadenylation. Proc Natl Acad Sci U S A 2020, 117, 7245-7254. https://doi.org/10.1073/pnas.1917922117. [CrossRef]
- Yang, J.; Hung, L.H.; Licht, T.; Kostin, S.; Looso, M.; Khrameeva, E.; Bindereif, A.; Schneider, A.; Braun, T. RBM24 is a major regulator of muscle-specific alternative splicing. Dev Cell 2014, 31, 87-99. https://doi.org/10.1016/j.devcel.2014.08.025. [CrossRef]
- Ohe, K.; Yoshida, M.; Nakano-Kobayashi, A.; Hosokawa, M.; Sako, Y.; Sakuma, M.; Okuno, Y.; Usui, T.; Ninomiya, K.; Nojima, T., et al. RBM24 promotes U1 snRNP recognition of the mutated 5' splice site in the IKBKAP gene of familial dysautonomia. RNA 2017, 23, 1393-1403. https://doi.org/10.1261/rna.059428.116. [CrossRef]
- Zhang, T.; Lin, Y.; Liu, J.; Zhang, Z.G.; Fu, W.; Guo, L.Y.; Pan, L.; Kong, X.; Zhang, M.K.; Lu, Y.H., et al. Rbm24 Regulates Alternative Splicing Switch in Embryonic Stem Cell Cardiac Lineage Differentiation. Stem Cells 2016, 34, 1776-1789. https://doi.org/10.1002/stem.2366. [CrossRef]
- Liu, J.; Kong, X.; Zhang, M.; Yang, X.; Xu, X. RNA binding protein 24 deletion disrupts global alternative splicing and causes dilated cardiomyopathy. Protein Cell 2019, 10, 405-416. https://doi.org/10.1007/s13238-018-0578-8. [CrossRef]
- Zhang, M.; Zhang, Y.; Xu, E.; Mohibi, S.; de Anda, D.M.; Jiang, Y.; Zhang, J.; Chen, X. Rbm24, a target of p53, is necessary for proper expression of p53 and heart development. Cell Death Differ 2018, 25, 1118-1130. https://doi.org/10.1038/s41418-017-0029-8. [CrossRef]
- Cao, H.; Zhao, K.; Yao, Y.; Guo, J.; Gao, X.; Yang, Q.; Guo, M.; Zhu, W.; Wang, Y.; Wu, C., et al. RNA binding protein 24 regulates the translation and replication of hepatitis C virus. Protein Cell 2018, 9, 930-944. https://doi.org/10.1007/s13238-018-0507-x. [CrossRef]
- Yao, Y.; Yang, B.; Cao, H.; Zhao, K.; Yuan, Y.; Chen, Y.; Zhang, Z.; Wang, Y.; Pei, R.; Chen, J., et al. RBM24 stabilizes hepatitis B virus pregenomic RNA but inhibits core protein translation by targeting the terminal redundancy sequence. Emerg Microbes Infect 2018, 7, 86. https://doi.org/10.1038/s41426-018-0091-4. [CrossRef]
- Yao, Y.; Sun, H.; Chen, Y.; Tian, L.; Huang, D.; Liu, C.; Zhou, Y.; Wang, Y.; Wen, Z.; Yang, B., et al. RBM24 inhibits the translation of SARS-CoV-2 polyproteins by targeting the 5'-untranslated region. Antiviral Res 2023, 209, 105478. https://doi.org/10.1016/j.antiviral.2022.105478. [CrossRef]
- Godet, A.C.; David, F.; Hantelys, F.; Tatin, F.; Lacazette, E.; Garmy-Susini, B.; Prats, A.C. IRES Trans-Acting Factors, Key Actors of the Stress Response. Int J Mol Sci 2019, 20. https://doi.org/10.3390/ijms20040924. [CrossRef]
- Kieft, J.S. Viral IRES RNA structures and ribosome interactions. Trends Biochem Sci 2008, 33, 274-283. https://doi.org/10.1016/j.tibs.2008.04.007. [CrossRef]
- Sun, C.; Querol-Audi, J.; Mortimer, S.A.; Arias-Palomo, E.; Doudna, J.A.; Nogales, E.; Cate, J.H. Two RNA-binding motifs in eIF3 direct HCV IRES-dependent translation. Nucleic Acids Res 2013, 41, 7512-7521. https://doi.org/10.1093/nar/gkt510. [CrossRef]
- Fraser, C.S.; Doudna, J.A. Structural and mechanistic insights into hepatitis C viral translation initiation. Nat Rev Microbiol 2007, 5, 29-38. https://doi.org/10.1038/nrmicro1558. [CrossRef]
- Lukavsky, P.J. Structure and function of HCV IRES domains. Virus Res 2009, 139, 166-171. https://doi.org/10.1016/j.virusres.2008.06.004. [CrossRef]
- Yamamoto, H.; Unbehaun, A.; Loerke, J.; Behrmann, E.; Collier, M.; Burger, J.; Mielke, T.; Spahn, C.M. Structure of the mammalian 80S initiation complex with initiation factor 5B on HCV-IRES RNA. Nat Struct Mol Biol 2014, 21, 721-727. https://doi.org/10.1038/nsmb.2859. [CrossRef]
- Dmitriev, S.E.; Terenin, I.M.; Andreev, D.E.; Ivanov, P.A.; Dunaevsky, J.E.; Merrick, W.C.; Shatsky, I.N. GTP-independent tRNA delivery to the ribosomal P-site by a novel eukaryotic translation factor. J Biol Chem 2010, 285, 26779-26787. https://doi.org/10.1074/jbc.M110.119693. [CrossRef]
- Jaafar, Z.A.; Oguro, A.; Nakamura, Y.; Kieft, J.S. Translation initiation by the hepatitis C virus IRES requires eIF1A and ribosomal complex remodeling. Elife 2016, 5. https://doi.org/10.7554/eLife.21198. [CrossRef]
- Gonzalez-Almela, E.; Williams, H.; Sanz, M.A.; Carrasco, L. The Initiation Factors eIF2, eIF2A, eIF2D, eIF4A, and eIF4G Are Not Involved in Translation Driven by Hepatitis C Virus IRES in Human Cells. Front Microbiol 2018, 9, 207. https://doi.org/10.3389/fmicb.2018.00207. [CrossRef]
- Terenin, I.M.; Dmitriev, S.E.; Andreev, D.E.; Shatsky, I.N. Eukaryotic translation initiation machinery can operate in a bacterial-like mode without eIF2. Nat Struct Mol Biol 2008, 15, 836-841. https://doi.org/10.1038/nsmb.1445. [CrossRef]
- Niepmann, M.; Gerresheim, G.K. Hepatitis C Virus Translation Regulation. Int J Mol Sci 2020, 21. https://doi.org/10.3390/ijms21072328. [CrossRef]
- Meskauskas, A.; Dinman, J.D. Ribosomal protein L3: gatekeeper to the A site. Mol Cell 2007, 25, 877-888. https://doi.org/10.1016/j.molcel.2007.02.015. [CrossRef]
- Ceci, M.; Gaviraghi, C.; Gorrini, C.; Sala, L.A.; Offenhauser, N.; Marchisio, P.C.; Biffo, S. Release of eIF6 (p27BBP) from the 60S subunit allows 80S ribosome assembly. Nature 2003, 426, 579-584. https://doi.org/10.1038/nature02160. [CrossRef]
- Jaako, P.; Faille, A.; Tan, S.; Wong, C.C.; Escudero-Urquijo, N.; Castro-Hartmann, P.; Wright, P.; Hilcenko, C.; Adams, D.J.; Warren, A.J. eIF6 rebinding dynamically couples ribosome maturation and translation. Nat Commun 2022, 13, 1562. https://doi.org/10.1038/s41467-022-29214-7. [CrossRef]
- Zacharias, D.A.; Violin, J.D.; Newton, A.C.; Tsien, R.Y. Partitioning of lipid-modified monomeric GFPs into membrane microdomains of live cells. Science 2002, 296, 913-916. https://doi.org/10.1126/science.1068539. [CrossRef]
- consortium, P.D.-K. PDBe-KB: collaboratively defining the biological context of structural data. Nucleic Acids Res 2022, 50, D534-D542. https://doi.org/10.1093/nar/gkab988. [CrossRef]
- Thoms, M.; Buschauer, R.; Ameismeier, M.; Koepke, L.; Denk, T.; Hirschenberger, M.; Kratzat, H.; Hayn, M.; Mackens-Kiani, T.; Cheng, J., et al. Structural basis for translational shutdown and immune evasion by the Nsp1 protein of SARS-CoV-2. Science 2020, 369, 1249-1255. https://doi.org/10.1126/science.abc8665. [CrossRef]
- Valbuena, F.M.; Fitzgerald, I.; Strack, R.L.; Andruska, N.; Smith, L.; Glick, B.S. A photostable monomeric superfolder green fluorescent protein. Traffic 2020, 21, 534-544. https://doi.org/10.1111/tra.12737. [CrossRef]
- Russo, A.; Pagliara, V.; Albano, F.; Esposito, D.; Sagar, V.; Loreni, F.; Irace, C.; Santamaria, R.; Russo, G. Regulatory role of rpL3 in cell response to nucleolar stress induced by Act D in tumor cells lacking functional p53. Cell Cycle 2016, 15, 41-51. https://doi.org/10.1080/15384101.2015.1120926. [CrossRef]
- Russo, A.; Russo, G. Ribosomal Proteins Control or Bypass p53 during Nucleolar Stress. Int J Mol Sci 2017, 18. https://doi.org/10.3390/ijms18010140. [CrossRef]
- Weis, F.; Giudice, E.; Churcher, M.; Jin, L.; Hilcenko, C.; Wong, C.C.; Traynor, D.; Kay, R.R.; Warren, A.J. Mechanism of eIF6 release from the nascent 60S ribosomal subunit. Nat Struct Mol Biol 2015, 22, 914-919. https://doi.org/10.1038/nsmb.3112. [CrossRef]
- Uhlen, M.; Fagerberg, L.; Hallstrom, B.M.; Lindskog, C.; Oksvold, P.; Mardinoglu, A.; Sivertsson, A.; Kampf, C.; Sjostedt, E.; Asplund, A., et al. Proteomics. Tissue-based map of the human proteome. Science 2015, 347, 1260419. https://doi.org/10.1126/science.1260419. [CrossRef]
- Milenkovic, I.; Santos Vieira, H.G.; Lucas, M.C.; Ruiz-Orera, J.; Patone, G.; Kesteven, S.; Wu, J.; Feneley, M.; Espadas, G.; Sabido, E., et al. Dynamic interplay between RPL3- and RPL3L-containing ribosomes modulates mitochondrial activity in the mammalian heart. Nucleic Acids Res 2023, 51, 5301-5324. https://doi.org/10.1093/nar/gkad121. [CrossRef]
- Yang, T.H.; Wang, C.Y.; Tsai, H.C.; Liu, C.T. Human IRES Atlas: an integrative platform for studying IRES-driven translational regulation in humans. Database (Oxford) 2021, 2021. https://doi.org/10.1093/database/baab025. [CrossRef]
- Papadopoulos, J.S.; Agarwala, R. COBALT: constraint-based alignment tool for multiple protein sequences. Bioinformatics 2007, 23, 1073-1079. https://doi.org/10.1093/bioinformatics/btm076. [CrossRef]
- Gandin, V.; Miluzio, A.; Barbieri, A.M.; Beugnet, A.; Kiyokawa, H.; Marchisio, P.C.; Biffo, S. Eukaryotic initiation factor 6 is rate-limiting in translation, growth and transformation. Nature 2008, 455, 684-688. https://doi.org/10.1038/nature07267. [CrossRef]
- Sanjana, N.E.; Shalem, O.; Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat Methods 2014, 11, 783-784. https://doi.org/10.1038/nmeth.3047. [CrossRef]
- Ku, H.K.; Lim, H.M.; Oh, K.H.; Yang, H.J.; Jeong, J.S.; Kim, S.K. Interpretation of protein quantitation using the Bradford assay: comparison with two calculation models. Anal Biochem 2013, 434, 178-180. https://doi.org/10.1016/j.ab.2012.10.045. [CrossRef]
- Rabilloud, T. Optimization of the cydex blue assay: A one-step colorimetric protein assay using cyclodextrins and compatible with detergents and reducers. PLoS One 2018, 13, e0195755. https://doi.org/10.1371/journal.pone.0195755. [CrossRef]
- Cui, W.; Xue, H.; Cheng, H.; Zhang, H.; Jin, J.; Wang, Q. Increasing the amount of phosphoric acid enhances the suitability of Bradford assay for proteomic research. Electrophoresis 2019, 40, 1107-1112. https://doi.org/10.1002/elps.201800430. [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat Methods 2012, 9, 671-675. https://doi.org/10.1038/nmeth.2089. [CrossRef]
- Xu, S.; Pei, R.; Guo, M.; Han, Q.; Lai, J.; Wang, Y.; Wu, C.; Zhou, Y.; Lu, M.; Chen, X. Cytosolic phospholipase A2 gamma is involved in hepatitis C virus replication and assembly. J Virol 2012, 86, 13025-13037. https://doi.org/10.1128/JVI.01785-12. [CrossRef]
- Bolt, M.W.; Mahoney, P.A. High-efficiency blotting of proteins of diverse sizes following sodium dodecyl sulfate-polyacrylamide gel electrophoresis. Anal Biochem 1997, 247, 185-192. https://doi.org/10.1006/abio.1997.2061. [CrossRef]
- Villanueva, M.A. Electrotransfer of proteins in an environmentally friendly methanol-free transfer buffer. Anal Biochem 2008, 373, 377-379. https://doi.org/10.1016/j.ab.2007.08.007. [CrossRef]
- Yeung, Y.G.; Stanley, E.R. A solution for stripping antibodies from polyvinylidene fluoride immunoblots for multiple reprobing. Anal Biochem 2009, 389, 89-91. https://doi.org/10.1016/j.ab.2009.03.017. [CrossRef]
- Han, S.; Cui, Y.; Helbing, D.L. Inactivation of Horseradish Peroxidase by Acid for Sequential Chemiluminescent Western Blot. Biotechnol J 2020, 15, e1900397. https://doi.org/10.1002/biot.201900397. [CrossRef]
- Yoshimi, K.; Kunihiro, Y.; Kaneko, T.; Nagahora, H.; Voigt, B.; Mashimo, T. ssODN-mediated knock-in with CRISPR-Cas for large genomic regions in zygotes. Nat Commun 2016, 7, 10431. https://doi.org/10.1038/ncomms10431. [CrossRef]
- Nagahora, H.T., JP) METHOD FOR PREPARING LONG-CHAIN SINGLE-STRANDED DNA. 2019.
- Heigwer, F.; Kerr, G.; Boutros, M. E-CRISP: fast CRISPR target site identification. Nat Methods 2014, 11, 122-123. https://doi.org/10.1038/nmeth.2812. [CrossRef]
- Pasloske, B.L.A., TX), Wu, William (Austin, TX) Method and reagents for inactivating ribonucleases RNase A, RNase I and RNase T1. 2004.
- Schmidt, T.G.M.; Eichinger, A.; Schneider, M.; Bonet, L.; Carl, U.; Karthaus, D.; Theobald, I.; Skerra, A. The Role of Changing Loop Conformations in Streptavidin Versions Engineered for High-affinity Binding of the Strep-tag II Peptide. J Mol Biol 2021, 433, 166893. https://doi.org/10.1016/j.jmb.2021.166893. [CrossRef]
- Burley, S.K.; Bhikadiya, C.; Bi, C.; Bittrich, S.; Chao, H.; Chen, L.; Craig, P.A.; Crichlow, G.V.; Dalenberg, K.; Duarte, J.M., et al. RCSB Protein Data Bank (RCSB.org): delivery of experimentally-determined PDB structures alongside one million computed structure models of proteins from artificial intelligence/machine learning. Nucleic Acids Res 2023, 51, D488-D508. https://doi.org/10.1093/nar/gkac1077. [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res 2000, 28, 235-242. https://doi.org/10.1093/nar/28.1.235. [CrossRef]
- Guex, N.; Peitsch, M.C. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 1997, 18, 2714-2723. https://doi.org/10.1002/elps.1150181505. [CrossRef]
- Bernstein, H.J. Recent changes to RasMol, recombining the variants. Trends Biochem Sci 2000, 25, 453-455. https://doi.org/10.1016/s0968-0004(00)01606-6. [CrossRef]
- Ma, J.; Chen, T.; Wu, S.; Yang, C.; Bai, M.; Shu, K.; Li, K.; Zhang, G.; Jin, Z.; He, F., et al. iProX: an integrated proteome resource. Nucleic Acids Res 2019, 47, D1211-D1217. https://doi.org/10.1093/nar/gky869. [CrossRef]
- Chen, T.; Ma, J.; Liu, Y.; Chen, Z.; Xiao, N.; Lu, Y.; Fu, Y.; Yang, C.; Li, M.; Wu, S., et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res 2022, 50, D1522-D1527. https://doi.org/10.1093/nar/gkab1081. [CrossRef]
- Sayers, E.W.; Bolton, E.E.; Brister, J.R.; Canese, K.; Chan, J.; Comeau, D.C.; Connor, R.; Funk, K.; Kelly, C.; Kim, S., et al. Database resources of the national center for biotechnology information. Nucleic Acids Res 2022, 50, D20-D26. https://doi.org/10.1093/nar/gkab1112. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



